Epidemiology and Genetic Diversity of Chlamydia pecorum in Cattle and Sheep from Western China
Abstract
1. Introduction
2. Method
2.1. Sample Collection
2.2. Nucleic Acid Extraction and Concentration
2.3. C. pecorum ompA Genotyping
2.4. Multilocus Sequence Typing
2.5. Comparison of MLST and ompA Phylogenetic Analyses in C. pecorum
3. Results
3.1. Epidemiological Characteristics of C. pecorum Infections
3.2. Molecular Typing and Evolutionary Characteristics of C. pecorum ompA
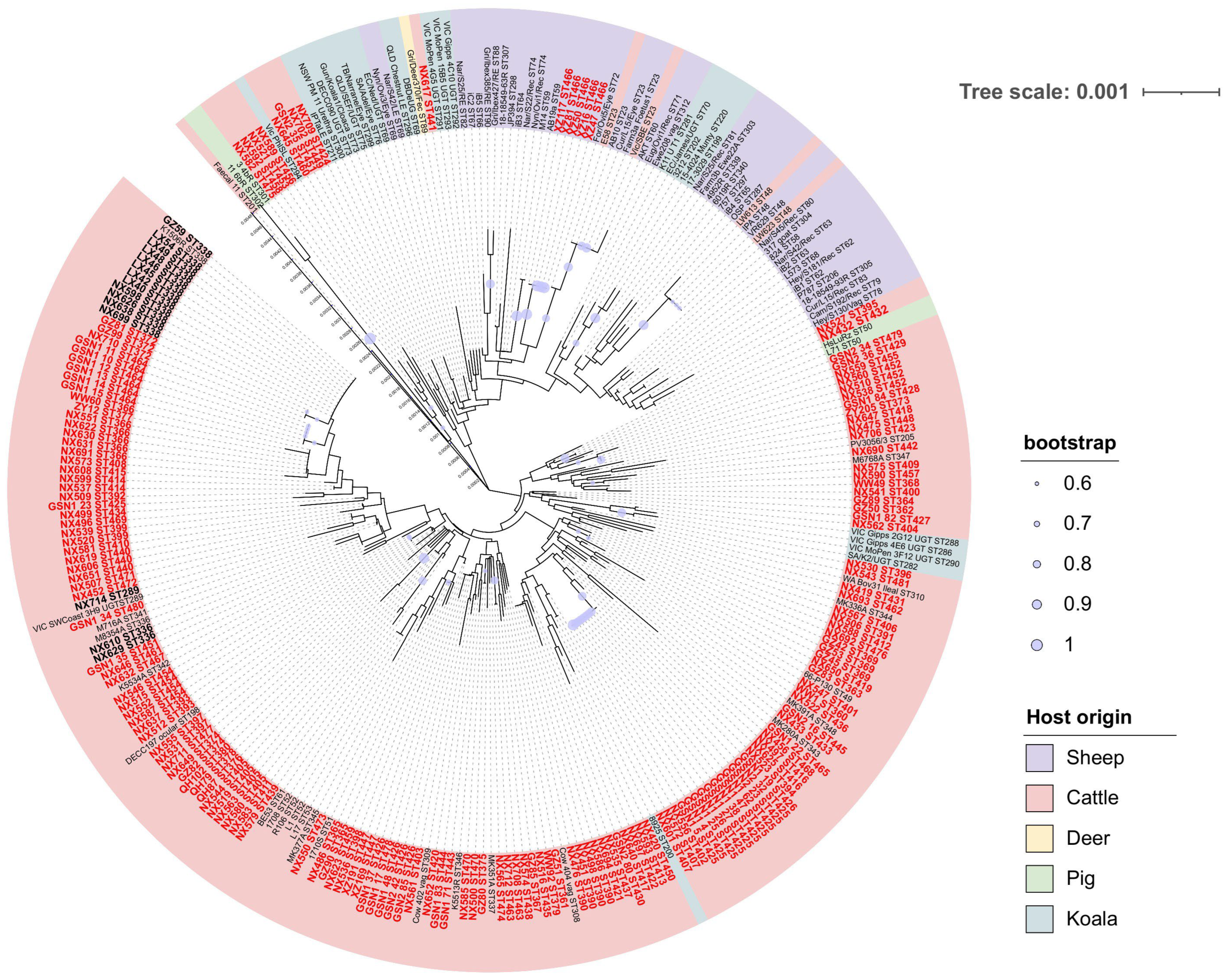
3.3. C. pecorum MLST Analyses
3.4. MLST and ompA Phylogenetic Tanglegram
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Borel, N.; Polkinghorne, A.; Pospischil, A. A Review on Chlamydial Diseases in Animals: Still a Challenge for Pathologists? Vet. Pathol. 2018, 55, 374–390. [Google Scholar] [CrossRef] [PubMed]
- Burnard, D.; Polkinghorne, A. Chlamydial infections in wildlife-conservation threats and/or reservoirs of ‘spill-over’ infections? Vet. Microbiol. 2016, 196, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Jelocnik, M.; Taylor-Brown, A.; O’Dea, C.; Anstey, S.; Bommana, S.; Masters, N.; Katouli, M.; Jenkins, C.; Polkinghorne, A. Detection of a range of genetically diverse chlamydiae in Australian domesticated and wild ungulates. Transbound. Emerg. Dis. 2019, 66, 1132–1137. [Google Scholar] [CrossRef] [PubMed]
- Marti, H.; Shima, K.; Boutin, S.; Rupp, J.; Clarke, I.N.; Laroucau, K.; Borel, N. Zoonotic and other veterinary chlamydiae—An update, the role of the plasmid and plasmid-mediated transformation. Pathog. Dis. 2024, 82, ftae030. [Google Scholar] [CrossRef]
- Fukushi, H.; Hirai, K. Proposal of Chlamydia pecorum sp. nov. for Chlamydia strains derived from ruminants. Int. J. Syst. Bacteriol. 1992, 42, 306–308. [Google Scholar] [CrossRef]
- Hagenbuch, F.; Loehrer, S.; Marti, H.; Kasimov, V.; Jelocnik, M.; Borel, N. Investigation of Chlamydia pecorum in livestock from Switzerland reveals a high degree of diversity in bovine strains. Vet. Microbiol. 2024, 292, 110057. [Google Scholar] [CrossRef]
- Walker, E.; Lee, E.J.; Timms, P.; Polkinghorne, A. Chlamydia pecorum infections in sheep and cattle: A common and under-recognised infectious disease with significant impact on animal health. Vet. J. 2015, 206, 252–260. [Google Scholar] [CrossRef]
- Cao, L.; He, L.; Wang, S.; Xu, L.; Zhuang, S. Severe community-acquired pneumonia caused by Chlamydia pecorum. Int. J. Infect. Dis. 2022, 124, 171–173. [Google Scholar] [CrossRef]
- Zhang, D.; Liu, D.; An, Y.; Li, W. Chlamydia pecorum pneumonia complicated with multiple organ failure detected by metagenomic next-generation sequencing: A case report. Chin. J. Infect. Control 2023, 22, 597–600. [Google Scholar]
- Sarovich, D.S.; Jelocnik, M.; Stewart, P.; Pollak, N.M.; Wong, J.S.F.; Kunesh, C.; Ojiako, C.; Hanger, J.; Gillett, A.; Valenza, L.; et al. Dominance of Vaccine-Specific Chlamydia pecorum ompA Genotypes in Koalas From North-Eastern Australia. Ecol. Evol. 2025, 15, e70973. [Google Scholar] [CrossRef]
- Kaltenboeck, B.; Heinen, E.; Schneider, R.; Wittenbrink, M.M.; Schmeer, N. OmpA and antigenic diversity of bovine Chlamydophila pecorum strains. Vet. Microbiol. 2009, 135, 175–180. [Google Scholar] [CrossRef]
- Jelocnik, M.; Walker, E.; Pannekoek, Y.; Ellem, J.; Timms, P.; Polkinghorne, A. Evaluation of the relationship between Chlamydia pecorum sequence types and disease using a species-specific multi-locus sequence typing scheme (MLST). Vet. Microbiol. 2014, 174, 214–222. [Google Scholar] [CrossRef]
- Jelocnik, M.; White, R.T.; Clune, T.; O’Connell, J.; Foxwell, J.; Hair, S.; Besier, S.; Tom, L.; Phillips, N.; Robbins, A.; et al. Molecular characterisation of the Australian and New Zealand livestock Chlamydia pecorum strains confirms novel but clonal ST23 in association with ovine foetal loss. Vet. Microbiol. 2023, 283, 109774. [Google Scholar] [CrossRef]
- Wang, Q.; Tian, T.; Yao, N.Q.; Chen, Z.Y.; Yang, Y.; Sun, Y.H.; Shi, J.F.; Li, J.M.; Shi, K.; Du, R. A Meta-Analysis of Bovine Chlamydia Prevalence in Cattle in China from 1989 to 2020. Vector Borne Zoonotic Dis. 2022, 22, 76–92. [Google Scholar] [CrossRef]
- Guo, Y.; Ge, S.; Luo, H.; Rehman, A.; Li, Y.; He, S. Occurrence of Trichophyton verrucosum in cattle in the Ningxia Hui autonomous region, China. BMC Vet. Res. 2020, 16, 187. [Google Scholar] [CrossRef]
- Szabo, K.V.; O’Neill, C.E.; Clarke, I.N. Diversity in Chlamydial plasmids. PLoS ONE 2020, 15, e0233298. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Jelocnik, M.; Polkinghorne, A.; Pannekoek, Y. Multilocus Sequence Typing (MLST) of Chlamydiales. Methods Mol. Biol. 2019, 2042, 69–86. [Google Scholar] [CrossRef] [PubMed]
- Feil, E.J.; Li, B.C.; Aanensen, D.M.; Hanage, W.P.; Spratt, B.G. eBURST: Inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 2004, 186, 1518–1530. [Google Scholar] [CrossRef]
- Francisco, A.P.; Bugalho, M.; Ramirez, M.; Carriço, J.A. Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC Bioinform. 2009, 10, 152. [Google Scholar] [CrossRef]
- Li, M.; Jelocnik, M.; Yang, F.; Gong, J.; Kaltenboeck, B.; Polkinghorne, A.; Feng, Z.; Pannekoek, Y.; Borel, N.; Song, C.; et al. Asymptomatic infections with highly polymorphic Chlamydia suis are ubiquitous in pigs. BMC Vet. Res. 2017, 13, 370. [Google Scholar] [CrossRef]
- Huson, D.H.; Scornavacca, C. Dendroscope 3: An interactive tool for rooted phylogenetic trees and networks. Syst. Biol. 2012, 61, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Hayati, M.; Chindelevitch, L. Computing the distribution of the Robinson-Foulds distance. Comput. Biol. Chem. 2020, 87, 107284. [Google Scholar] [CrossRef] [PubMed]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 124. [Google Scholar] [CrossRef]
- Loehrer, S.; Hagenbuch, F.; Marti, H.; Pesch, T.; Hässig, M.; Borel, N. Longitudinal study of Chlamydia pecorum in a healthy Swiss cattle population. PLoS ONE 2023, 18, e0292509. [Google Scholar] [CrossRef]
- Jelocnik, M.; Bachmann, N.L.; Seth-Smith, H.; Thomson, N.R.; Timms, P.; Polkinghorne, A.M. Molecular characterisation of the Chlamydia pecorum plasmid from porcine, ovine, bovine, and koala strains indicates plasmid-strain co-evolution. PeerJ 2016, 4, e1661. [Google Scholar] [CrossRef]
- Pannekoek, Y.; Morelli, G.; Kusecek, B.; Morré, S.A.; Ossewaarde, J.M.; Langerak, A.A.; van der Ende, A. Multi locus sequence typing of Chlamydiales: Clonal groupings within the obligate intracellular bacteria Chlamydia trachomatis. BMC Microbiol. 2008, 8, 42. [Google Scholar] [CrossRef] [PubMed]
- Dixit, P.D.; Pang, T.Y.; Maslov, S. Recombination-Driven Genome Evolution and Stability of Bacterial Species. Genetics 2017, 207, 281–295. [Google Scholar] [CrossRef]
- White, N.J.; Beckerman, A.P.; Snook, R.R.; Brockhurst, M.A.; Butlin, R.K.; Eyres, I. Experimental evolution of local adaptation under unidimensional and multidimensional selection. Curr. Biol. 2022, 32, 1310–1318.e1314. [Google Scholar] [CrossRef]
- Seth-Smith, H.; Bommana, S.; Dean, D.; Read, T.D.; Marti, H. Chlamydia suis undergoes interclade recombination promoting Tet-island exchange. BMC Genom. 2024, 25, 724. [Google Scholar] [CrossRef] [PubMed]
- Polkinghorne, A.; Hanger, J.; Timms, P. Recent advances in understanding the biology, epidemiology and control of chlamydial infections in koalas. Vet. Microbiol. 2013, 165, 214–223. [Google Scholar] [CrossRef]
- Nogarol, C.; Marchino, M.; Scala, S.; Belvedere, M.; Renna, G.; Vitale, N.; Mandola, M.L. Seroprevalence and Risk Factors Associated with Chlamydia abortus Infection in Sheep and Goats in North-Western Italy. Animals 2024, 14, 291. [Google Scholar] [CrossRef]
- Marti, H.; Suchland, R.J.; Rockey, D.D. The Impact of Lateral Gene Transfer in Chlamydia. Front. Cell Infect. Microbiol. 2022, 12, 861899. [Google Scholar] [CrossRef]
- Shima, K.; Wanker, M.; Skilton, R.J.; Cutcliffe, L.T.; Schnee, C.; Kohl, T.A.; Niemann, S.; Geijo, J.; Klinger, M.; Timms, P.; et al. The Genetic Transformation of Chlamydia pneumoniae. mSphere 2018, 3, e00412-18. [Google Scholar] [CrossRef]
- Prieur, J. Critical warning! Preventing the multidimensional apocalypse on planet Earth. Ecosyst. Serv. 2020, 45, 101161. [Google Scholar] [CrossRef] [PubMed]
- Albini, S.; Marti, H.; Imkamp, F.; Borel, N. Update on the zoonotic potential of Chlamydia. Schweiz. Arch. Tierheilkd. 2023, 165, 165. [Google Scholar] [CrossRef]
- Sohaili, A.; Mogaka, F.; Vanrompay, D.; Omollo, V.O.; Morre, S.A.; Thomas, P.P.M. A review of zoonotic chlamydiae species in Africa: Assessing their burden and potential impact through One Health perspective. One Health Outlook 2025, 7, 40. [Google Scholar] [CrossRef]
- Ramakers, B.P.; Heijne, M.; Lie, N.; Le, T.N.; van Vliet, M.; Claessen, V.P.J.; Tolsma, P.J.P.; De Rosa, M.; Roest, H.I.J.; Vanrompay, D.; et al. Zoonotic Chlamydia caviae Presenting as Community-Acquired Pneumonia. N. Engl. J. Med. 2017, 377, 992–994. [Google Scholar] [CrossRef] [PubMed]
- Borel, N.; Longbottom, D.; Greub, G.; Albini, S.; Vanrompay, D.; Laroucau, K. Zoonotic infections due to avian Chlamydia abortus: What are we missing? Lancet Microbe 2025, 6, 101197. [Google Scholar] [CrossRef]
- Qin, T.; Han, L.; Sun, Y.; Qin, X.; Xu, J. Silent outbreaks of psittacosis revealed using targeted next-generation sequencing. Lancet Microbe 2025, 6, 101061. [Google Scholar] [CrossRef] [PubMed]
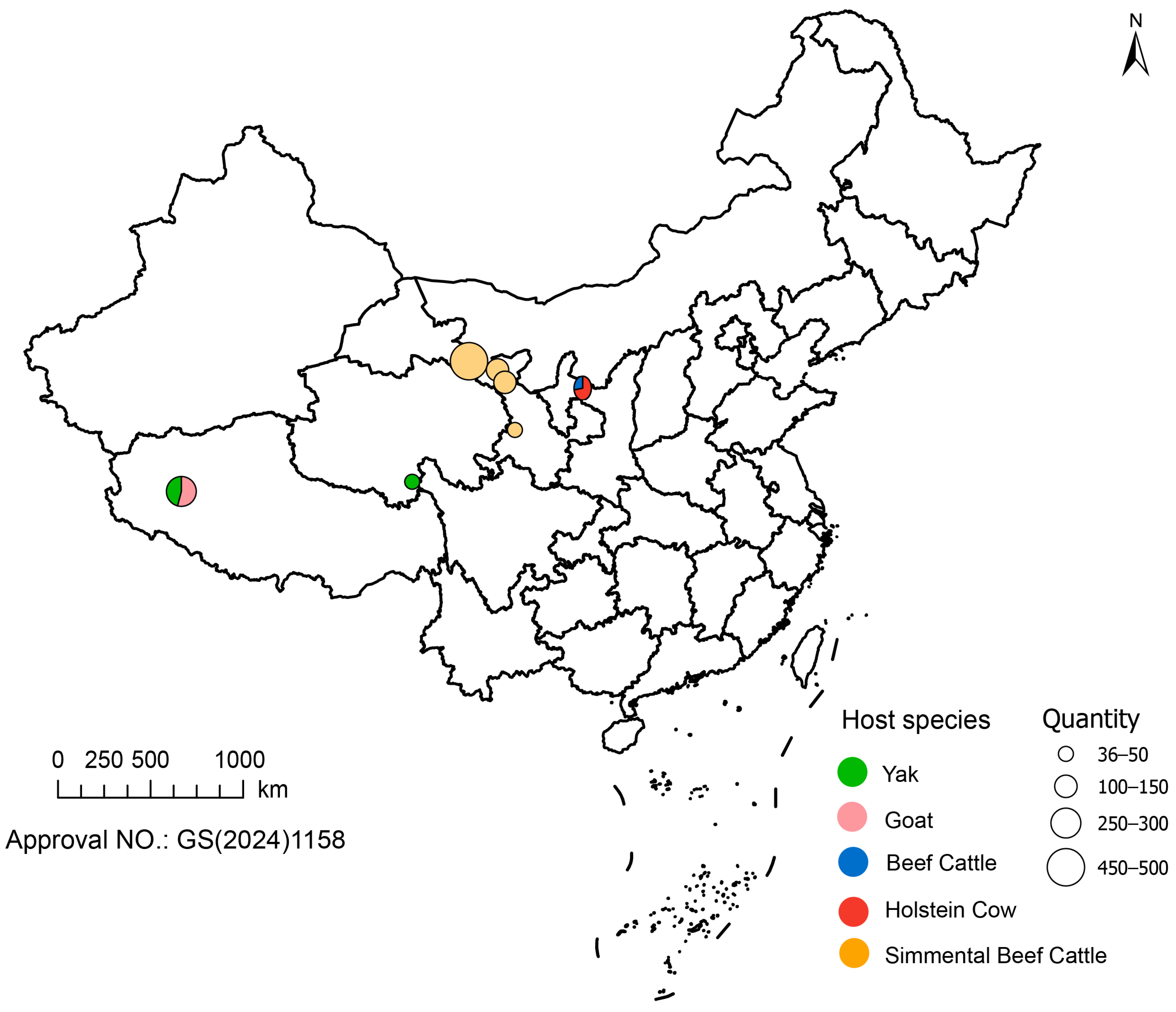



| Province | City/Prefecture | Host Species | Positivity |
|---|---|---|---|
| Gansu Province | Zhangye | Simmental Beef Cattle | 16.34% (75/459) |
| Wuwei | Simmental Beef Cattle | 6.80% (8/103) | |
| Linxia | Simmental Beef Cattle | 33.33% (12/36) | |
| Jinchang | Simmental Beef Cattle | 1.85% (2/108) | |
| Qinghai Province | Yushu | Yak | 12.00% (6/50) |
| Ningxia | Wuzhong | Holstein Cow (Adult) | 48.62% (88/181) |
| Wuzhong | Holstein Cow (Calf) | 0.00% (0/36) | |
| Wuzhong | Beef Cattle | 55.56% (45/81) | |
| Tibet | Ngari | Goat | 4.83% (7/145) |
| Ngari | Yak | 5.69% (7/123) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, M.; Wang, W.; Xu, D.; Qin, X.; Liang, J.; Guo, B.; Zhu, Z.; Teng, Z.; Bai, N.; Rong, B.; et al. Epidemiology and Genetic Diversity of Chlamydia pecorum in Cattle and Sheep from Western China. Pathogens 2025, 14, 1209. https://doi.org/10.3390/pathogens14121209
Zhang M, Wang W, Xu D, Qin X, Liang J, Guo B, Zhu Z, Teng Z, Bai N, Rong B, et al. Epidemiology and Genetic Diversity of Chlamydia pecorum in Cattle and Sheep from Western China. Pathogens. 2025; 14(12):1209. https://doi.org/10.3390/pathogens14121209
Chicago/Turabian StyleZhang, Mengtao, Wen Wang, Daqin Xu, Xincheng Qin, Junrong Liang, Bangcheng Guo, Zhen Zhu, Zhongqiu Teng, Nan Bai, Binguo Rong, and et al. 2025. "Epidemiology and Genetic Diversity of Chlamydia pecorum in Cattle and Sheep from Western China" Pathogens 14, no. 12: 1209. https://doi.org/10.3390/pathogens14121209
APA StyleZhang, M., Wang, W., Xu, D., Qin, X., Liang, J., Guo, B., Zhu, Z., Teng, Z., Bai, N., Rong, B., He, J., Dai, L., Zhang, X., & Qin, T. (2025). Epidemiology and Genetic Diversity of Chlamydia pecorum in Cattle and Sheep from Western China. Pathogens, 14(12), 1209. https://doi.org/10.3390/pathogens14121209






