Abstract
The Gram-negative bacterium Gallibacterium anatis is part of the normal avian respiratory, intestinal and reproductive tract microflora and can be transmitted horizontally and vertically. With the coexistence of other relevant factors, G. anatis becomes an opportunistic pathogen, economically damaging to the poultry industry. This bacterium’s prevalence and molecular epidemiology were investigated, and the antimicrobial treatment options for G. anatis infection in chicken flocks in Poland were assessed. Tracheal samples from 182 flocks were collected between April 2022 and March 2023. The bacterial prevalence was determined by PCR targeting the gyrB gene and 16–23S rRNA. Gallibacterium anatis was identified by matrix-assisted laser desorption/ionisation–time-of-flight mass spectrometry (MALDI-TOF) after culturing and PCR amplification. Isolates’ susceptibility to 11 antimicrobials was assessed with a disc diffusion test. Isolates were also tested for gyrB, GtxA and flfA virulence genes and blaROB, aphA, tetB and tetH antibiotic resistance genes by PCR. Forty-one flocks (22.5%) were positive through PCR. Antibiotic resistance was most frequently observed against tilmicosin, tylosin, enrofloxacin, amoxicillin, tetracycline and doxycycline. Multiple resistance to at least eight antibiotics occurred in 20% of isolates and to at least four in 100%. The occurrence of gyrB was noted in 100%, GtxA was detected in 89%, and flfA was found in 14% of positive samples. The tetB gene was present in 61.0% of positive samples, tetH was in 36.0%, aphA was in 16.7%, and blaROB was in 5.6%. Significant differences were found in G. anatis isolates related to the presence of the virulence genes GtxA and gyrB and the presence of resistance genes (p < 0.05) associated with resistance to tetracyclines, β-lactams and aminoglycosides. The continued rise in the resistance of G. anatis to a broadening range of antibiotics is a major problem for the poultry industry worldwide, as well as for public health. The findings of this study may expand the knowledge of the pathogenicity of G. anatis in poultry.
1. Introduction
Gallibacterium anatis, previously known as Pasteurella anatis, belongs to the phylum Proteobacteria, class Gammaproteobacteria and family Pasteurellaceae. It is a bacterium with pleomorphic cell morphology, and it is known as a commensal inhabitant of the respiratory, intestinal and reproductive tracts in poultry [1,2].
The pathogenicity of G. anatis depends on many factors related to the bacterial strain and the age, stress exposure and immune status of the bird [1,3,4]. In addition to these determinants, the severity of disease symptoms can be influenced by co-infection with other bacteria or viruses that damage the respiratory tract or cause immunosuppression [4,5]. Environmental factors can also exacerbate the disease. Mixed infection with G. anatis with viruses and bacteria such as Escherichia coli, Avibacterium paragallinarum and Mycoplasma gallisepticum may increase the rate of disease, resulting in increased morbidity and mortality [3,4,5,6]. The clinical form of the disease can lead to economic losses in the poultry industry by causing respiratory problems in birds and a wide range of pathological lesions in the ovaries and oviducts of laying hens. Infection of the reproductive tract of chickens can reduce egg production by 8–10%. In roosters, G. anatis is responsible for changes associated with reduced semen quality [4,5,7,8]. This opportunistic bacterium is responsible for the occurrence of many clinical signs, causing oophoritis, salpingitis, peritonitis and enteritis. Infections often remain undiagnosed [9].
Strains of G. anatis are widespread worldwide and have been isolated from various poultry species, such as chickens, turkeys, geese and ducks, but also partridges, guinea fowl and wild birds. It has been isolated in many countries in Europe, Asia and Africa, as well as the United States [5,10]. The widespread prevalence of G. anatis in poultry appears to be related to the route of its transmission in flocks. The main transmission route of G. anatis appears to be the respiratory system, but vertical transmission through the trans-ovarian, -oviduct and -eggshell routes is also possible, which was experimentally demonstrated in embryonated eggs. Gallibacterium anatis penetration into eggs is highly pathogenic to developing chicken embryos [9,10]
These bacteria can also instigate disease in non-avian species, in which they are involved in the pathogenesis of respiratory tract infections, such as tracheitis or aerosacculitis, and genital tract infections, such as salpingitis and oophoritis in cattle, horses, pigs, sheep and rabbits [1,9,11,12]. A recently described case of a woman who was in very bad health after renal transplantation was probably infected with G. anatis through food and developed bacteraemia and diarrhoea, which indicates that there are no species barriers to G. anatis [13].
Knowledge related to the virulence genes and antimicrobial susceptibility of G. anatis isolates in laying hens remains limited. The virulence factors of G. anatis involved in the colonisation and invasion of the tracheal epithelium, oropharyngeal tissues and oviducts, which include GtxA, bacterial fimbriae and tetracycline resistance determinants, are widespread and often found in multidrug-resistant bacterial species. Studies conducted around the world are revealing a large number of G. anatis strains showing multidrug resistance to three or more antibiotics. The growing multidrug resistance of G. anatis to antimicrobials is causing increased concern. Additionally, its range of host species being wide and not only avian makes it an important pathogen for its effect not only on livestock industries but also on public health. The purpose of this study was to present the prevalence of G. anatis in chicken flocks, as well as to assess the resistance of isolates to antibiotics. Additionally, we examined the presence of virulence and resistance genes for several classes of antibiotics.
2. Materials and Methods
2.1. Sample Collection
A total of 182 chicken flocks (162 layers and 20 broilers) from different parts of Poland were examined for G. anatis infection in 2022 and 2023. Tracheal swab samples were brought to the Department of Poultry Diseases at the National Veterinary Research Institute in Poland as part of a routine diagnostic test and monitoring programme. All examined birds were floor-reared, and most of them showed no respiratory or reproductive clinical signs. Some of the birds showed respiratory signs in the form of rales and coughing, and some of them had swollen heads.
2.2. DNA Extraction
Tracheal swabs were pooled separately into tubes containing Tris-EDTA buffer and processed for DNA extraction. Genomic DNA was extracted using a QIAamp DNA Mini kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations. The quantity and quality of the DNA were determined using the NanoDrop 1000 system (Thermo Scientific, Waltham, MA, USA). DNA extraction from the Tris-EDTA used for sample preparation was conducted as a negative control. Samples were frozen at −20 °C until further analysis.
2.3. Real-Time PCR
To detect G. anatis by real-time PCR, primers complementary to the gyrB gene were used as described by Wang et al., 2016 [14], with slight modifications. The reaction was carried out using a QuantiTect Probe PCR Kit (Qiagen, Hilden, Germany) in a total volume of 25 μL with 1.3 μL of each 10 μM primer, 0.5 μL of probe, 7.4 μL of distilled water and 2 μL of DNA in an ABI 7500 thermal cycler (Applied Biosystems, part of Thermo Fisher Scientific, Norwalk, CA, USA) under the following conditions: 95 °C for 3 min and 40 cycles of 95 °C for 3 s. The fluorescence data were collected during a 60 °C for 32 s annealing–extension step.
2.4. PCR
PCR was conducted using previously described specific G. anatis primers that amplify the 16S–23S rRNA gene [15]. The PCR assays were performed on positive samples obtained in real-time PCR. The reaction mixture contained Taq PCR Master Mix (Eurx, Gdańsk, Poland) in a volume of 12.5 μL, 1.5 μL of each 10 μM primer and 7.5 μL of distilled water with the addition of 2 μL of DNA to give a total reaction volume of 25 μL. The PCR procedure included an initial incubation for 1 min at 95 °C, 35 cycles for 40 s each at 95 °C, annealing at 50 °C for 40 s, and extension at 72 °C for 40 s, with a final extension at 72 °C for 2 min. The PCR amplicons were separated by electrophoresis on a 2% agarose E-gel plate (Invitrogen, part of Thermo Fisher Scientific, Waltham, MA, USA) containing ethidium bromide and were visualised by ultraviolet transillumination. The method was used to identify G. anatis from swab samples as well as isolates.
2.5. Isolation and Identification of Gallibacterium anatis
The supernatant of the swabs (10 μL) from samples positive through PCR was inoculated onto Columbia agar plates with 5% sheep’s blood and incubated at 37 °C under a 5% CO2 atmosphere for 24 h. The G. anatis colonies were verified by matrix-assisted laser desorption ionisation–time-of-flight mass spectrometry (MALDI-TOF MS). The bacterial colonies from the agar plate were transferred to the MALDI target plate and mixed with formic acid and α-cyano-4-hydroxycinnamic acid matrix solution. All mass spectra were analysed with Bruker Daltonics software (Bruker Corporation, Billerica, MA, USA).
2.6. Antibiotic Resistance
The antimicrobial susceptibility of G. anatis was determined by using the disc diffusion method with 11 different drugs (Oxoid discs, Basingstoke, UK). The test applied a bacteria volume of 100 uL of 1.5 × 107 CFU/mL (0.5 McFarland scale), distributed uniformly onto the Columbia agar with 5% sheep’s blood. Eleven antibiotics from nine classes were used: florfenicol (FFC, 30 μg)—quinolone class; doxycycline (DO, 30 μg) and tetracycline (TE, 30 μg)—tetracycline class; amoxicillin (AML, 25 μg)—β-lactam class; gentamicin (CN, 10 μg)—aminoglycoside class; enrofloxacin (ENR, 5 μg)—fluoroquinolone class; colistin (CT, 50 μg)—polymyxin class; ceftazidime (CAZ, 30 μg)—cephalosporin class; chloramphenicol (C, 30 μg)—chloramphenicol class; tilmicosin (TIL, 15 μg) and tylosin (TY, 30 μg)—macrolide class. The inhibition zones were interpreted visually. Eleven out of forty-one Gallibacterium isolates could not be recultivated after PCR identification; the DNA obtained from them was saved for further study.
2.7. Virulence and Resistance Genes
In addition, isolates of G. anatis were tested for the presence of the virulence genes gyrB, GtxA and flfA. All samples were also tested for the presence of the antibiotic resistance genes blaROB, aphA, tetB and tetH. A PCR method was used for both test steps with the starters described earlier [12] and DNA extracted from the G. anatis isolates using a QIAamp DNA Mini kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations.
2.8. Statistical Analysis
Venn diagrams were constructed showing the number of shared virulence and resistance genes. Statistical analysis was carried out using one-way ANOVA and the Mann–Whitney test to determine the statistical significance of the presence of virulence and resistance genes. The value of p < 0.05 was considered statistically significant. Statistical analyses were performed using the Social Science Statistics program (www.socscistatistics.com 3 April 2023).
3. Results
3.1. Isolation and Identification
All the swab samples were tested by real-time PCR, and confirmed isolates of G. anatis (n = 41) were obtained from layer chickens. Twenty-two of them dated from 2023 and nineteen were from 2022. All samples positive in the real-time PCR were subjected to a specific PCR and had an amplicon of approximately 1030 bp. The prevalence of G. anatis in chicken flocks was 22.5%. Colony growth was obtained for 30 samples that were confirmed by MALDI-TOF MS. Other samples showed contamination related to overgrowth bacteria such as Proteus sp. All positive samples had a colony suggestive of the haemolytic biovar of G. anatis.
3.2. Antibiotic Resistance
In this study, 11 different antibiotics from different classes were used. The percentages of the 30 isolates with antibiotic resistance are shown in Table 1. The most frequently resisted antibiotics were enrofloxacin (100%), tilmicosin (100%), tylosin (100%), amoxicillin (83%) and tetracycline (80%). Gallibacterium anatis isolates showed less resistance to doxycycline (39%), chloramphenicol (33%), florfenicol (22%), gentamicin (18%), colistin (17%) and ceftazidime (11%). Resistance to nine and eight antibiotics occurred in 10% of isolates, and resistance to seven and six antibiotics was noted in 6.7% and 13.3%, respectively. The largest proportion of isolates were resistant to at least five antibiotics, at 43.3%, and the next largest to four antibiotics, at 16.7%.

Table 1.
Antimicrobial susceptibility of the 30 Gallibacterium anatis isolates.
3.3. Presence of Virulence and Resistance Genes
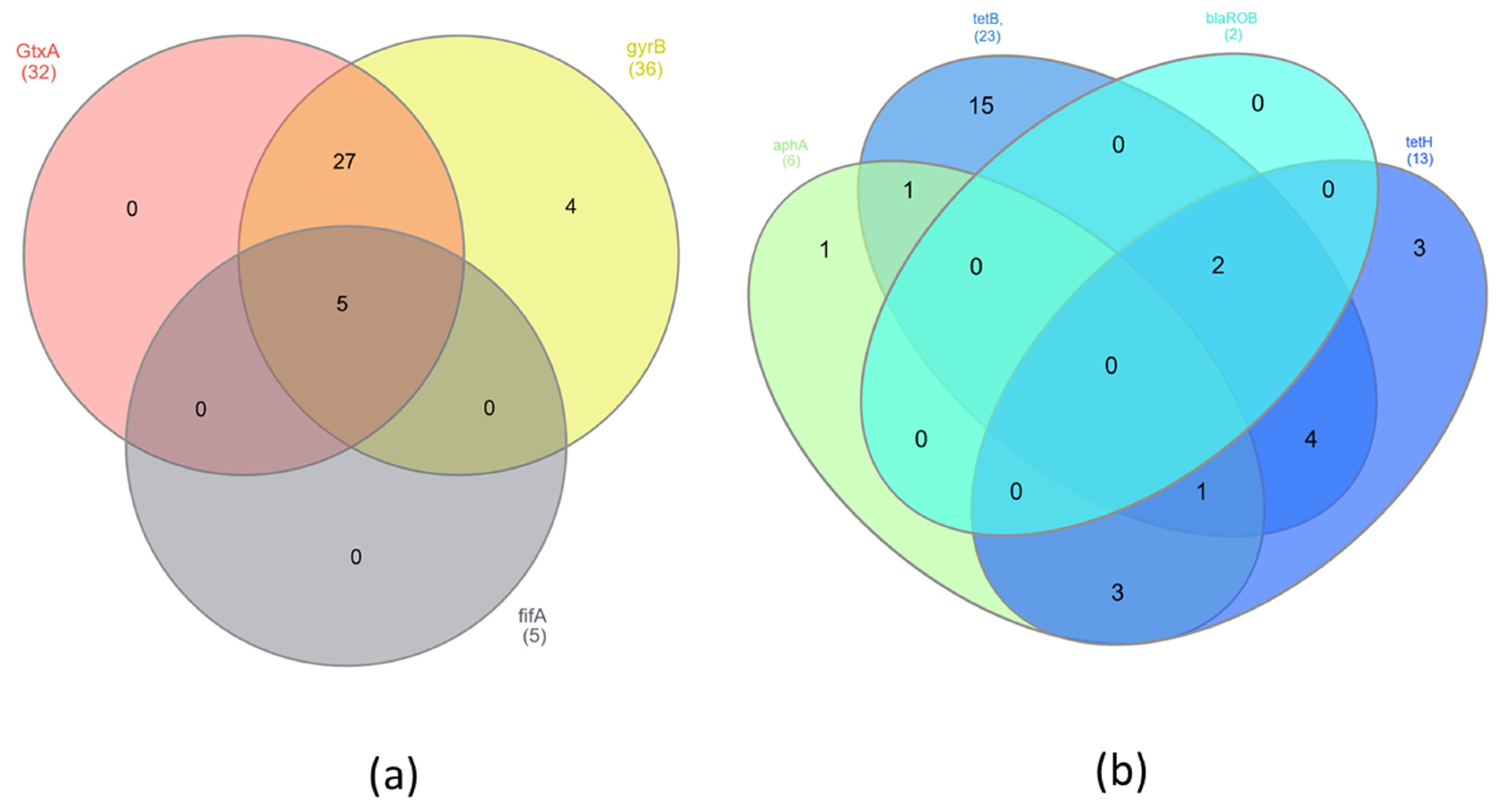
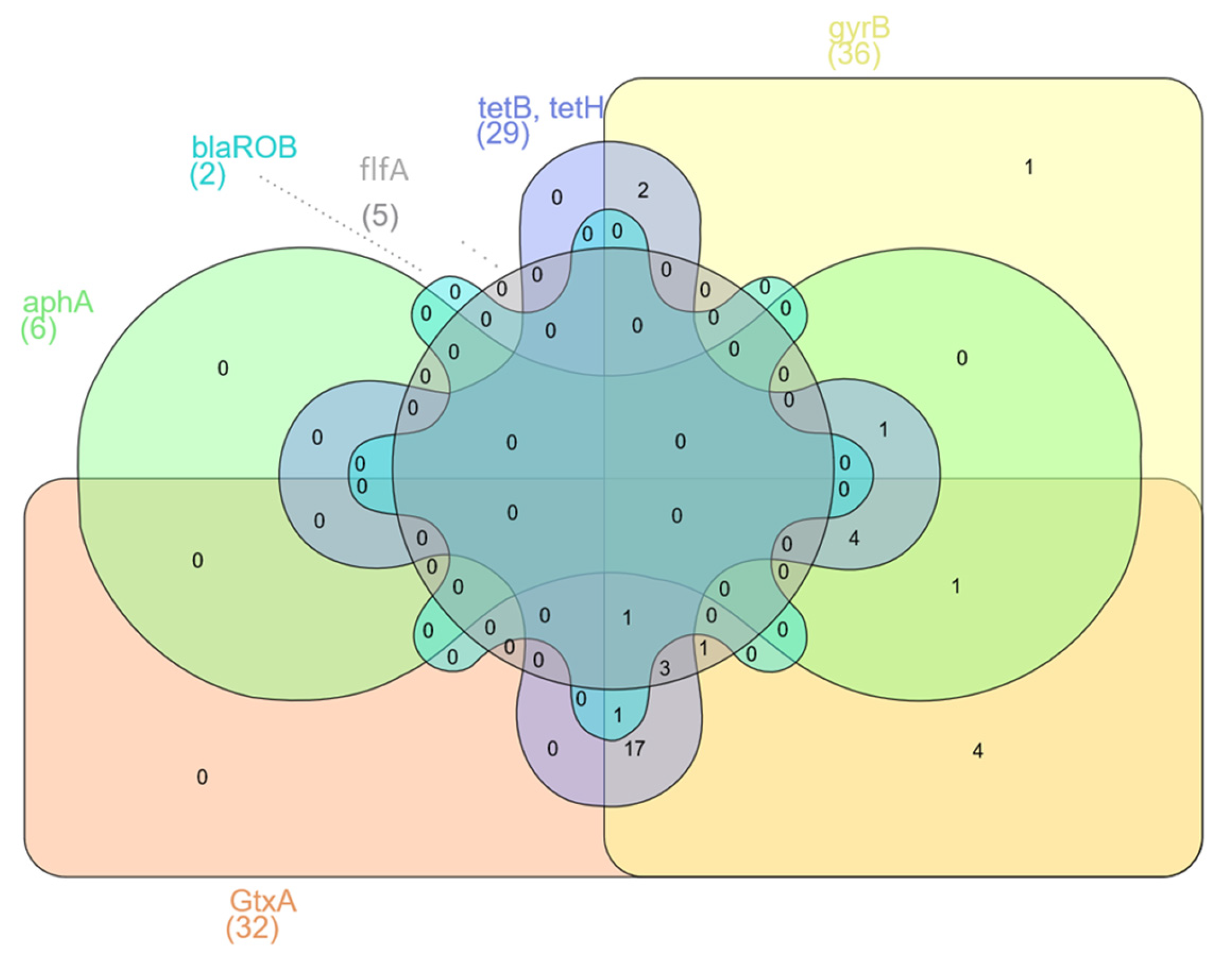
The presence of virulence and resistance genes was investigated in 36 samples by PCR. Five samples were excluded from the study because of poor DNA quality. The virulence gene GtxA was in 89% of isolates, gyrB was in 100%, and flfA was in 14% (Table 2). There were no statistically significant differences (p < 0.05) in the presence of GtxA and gyrB genes. However, statistically significant differences (p < 0.05) were found between occurrences of the virulence genes GtxA, gyrB and flfA. The antibiotic resistance gene tetB was found in 61.1% of isolates, tetH was in 36.1%, aphA was in 16.7%, and blaROB was in 5.6% (Table 2). The GtxA and gyrB genes were present in 27 (75%) isolates. However, all three tested virulence genes were identified in only five (13.2%) isolates (Figure 1a). The largest group of resistance genes present in G. anatis isolates were the tetB and tetH genes. The presence of both of these genes was found in seven (19.4%) isolates; additionally, the blaROB gene was also detected in two of them, and the aphA gene was identified in one of them (Figure 1b). The largest number of isolates (n = 17) contained both virulence genes GtxA and gyrB but also at least one of the resistance genes—tetB or tetH. Three isolates (8.3%) had all of the previously mentioned genes and additionally the flfA gene. In four (11.1%) isolates of G. anatis, besides two virulence genes (GtxA and gyrB), the resistance genes tetB, tetH and aphA were found (Figure 2). Significant differences were found associated with the presence of the virulence genes GtxA and gyrB and the presence of resistance genes (p < 0.05) associated with resistance to tetracyclines, β-lactams and aminoglycosides.

Table 2.
Presence of virulence and resistance genes in 36 G. anatis isolates.
Figure 1.
Venn diagrams showing presence of shared (a) virulence genes and (b) resistance genes in G. anatis isolates.
Figure 2.
Venn diagram showing presence in the G. anatis isolates of shared resistance and virulence genes.
4. Discussion
Gallibacterium anatis has been one of the pathogens causing reproductive problems in poultry in recent years. It is a component of the normal microbiota of chickens, but under certain circumstances, it can be a problem for bird health. There are many factors influencing the development of G. anatis infection. Some are host-related factors, such as age, stress or hormones, and others are environmental factors, such as seasonal changes or cold stress [1,16,17]. The epidemiological status of infections depends heavily upon the pathogenicity of the strain, the route of infection and the presence of a secondary bacterial agent, such as Mycoplasma gallisepticum, M. synoviae, Ornithobacterium rhinotracheale or E. coli [5,9,18,19,20,21]. Viral agents can also facilitate the development of the clinical form of the associated disease [22]. The presence of G. anatis in chicken flocks has been reported in Europe (in Germany, Denmark, Belgium and Austria), but also outside Europe (in Turkey, Morocco, Egypt and the USA) [9,11,21,22].
Two G. anatis biovars, one haemolytic and the other non-haemolytic, can be found in the respiratory tract and the lower genital tract of birds. In poultry, the haemolytic G. anatis biovar has been associated with systemic infections, including pericarditis, perihepatitis, peritonitis and septicaemia [3,5,23]. In this study, the presence of G. anatis was detected in the respiratory system of laying hens, in which, in some flocks, a decrease in laying was noted, as well as respiratory problems and swollen heads in parts of the flocks. The identification of G. anatis in chicken flocks in Poland was confirmed by real-time PCR and PCR methods, and its prevalence was 22.5%. The isolated species, including its biovar, was G. anatis biovar haemolytica. The range of tissue tropism of this G. anatis species is very wide and can include the respiratory, gastrointestinal and reproductive systems.
Many papers in recent years have reported that G. anatis is increasingly resistant to antibiotics. The frequency of multidrug resistance to different classes of antibiotics is also rising [1,24,25], which is demonstrated in this work. The current study determined the susceptibility of G. anatis isolates to eleven antimicrobial agents from the nine classes. Resistance to at least four antibiotics was shown in 100% of the tested strains. The highest percentage of isolates was the percentage resistant to five antibiotics—43.3%. Resistance to at least eight antibiotics was shown by 20% of field isolates. The highest rates of resistance were to macrolide and fluoroquinolone classes of antibiotics, which has been confirmed in several other studies in recent years [9,11,26]. In this study, G. anatis strains showed the greatest resistance equally to tylosin, tilmicosin and enrofloxacin—100%—which is similar to the results of other studies in the case of tylosin [1,24,25]. In the case of enrofloxacin, the rate is much higher than was observed among isolates in the USA (0%), Germany (33.3%) or Austria (58.2%) [21,22,27]. Our data on the susceptibility of G. anatis showed that 22% of isolates were resistant to florfenicol, which belongs to the quinolone class; this proportion was higher than in other reports from the EU [21,27]. However, other studies showed similar results to those of the present research for the antibiotics of the tetracycline class: doxycycline and tetracycline. In our study, the resistance levels for these antibiotics were 39% and 80%, respectively. Resistance to an antibiotic from the β-lactam class (amoxicillin) was 83%, much higher than the resistance of isolates to this antibiotic recorded by other authors [3,12,21,27]. It should be underlined that one isolate of G. anatis resistant to eight classes of antimicrobials and three isolates resistant to seven classes of antimicrobials were identified. Thirty-three percent of the isolates were resistant to four different classes of antibiotics, and this represented the largest group of G. anatis isolates. The results of antibiotic resistance and multidrug resistance obtained from G. anatis field isolates are very alarming and indicate the need to reduce the use of antibiotics in poultry production.
Several resistance genes in G. anatis isolates were identified by PCR. Statistical differences (p < 0.05) in the occurrence of antibiotic resistance genes were observed. All G. anatis isolates possessed at least one of the resistance genes tetB, tetH, aphA or blaROB. The tetracycline resistance gene tetB was the gene with the highest carriage rate, at 61%, and the rate for tetH was 36%, which is consistent with the results of Bojesen et al. (2011) and Abelazeem et al. (2022) [12,28]. Sixteen percent of G. anatis isolates had both the tetB and tetH genes. The detection of these genes in G. anatis isolates confirms their high antibiotic resistance to tetracycline and doxycycline. Aminoglycoside antibiotic resistance genes were investigated by attempting to detect the aphA resistance gene, which was identified and occurred in 17% of isolates in this study. Antibiotic resistance to gentamicin in the tested isolates was at similar levels. The presence of this gene was found to coincide with the tet genes in 14% of isolates. Statistical differences (p < 0.05) in the presence of antibiotic resistance genes tetB and aphA were observed. Among G. anatis isolates, low levels were found for the blaROB gene, which is a gene conferring resistance to antibiotics of the β-lactam class. Unfortunately, the low percentage of the gene detected does not coincide with the high antibiotic resistance of the isolates to amoxicillin. Samples in which the blaROB gene was found also contained tetB and tetH (Figure 1). The differences in the occurrence of these genes were statistically significant (p < 0.05). To expand knowledge in this area, future studies should be conducted on a higher number of resistance genes.
Additionally, we identified isolates containing virulence genes among G. anatis-positive chicken flocks. An 89% proportion of G. anatis isolates harboured the GtxA gene, which enables the production of RTX-like cytotoxins and accounts for the haemolytic and leukotoxic activities of G. anatis (Figure 1) [29]. Studies have shown that the GtxA protein has a significant inhibitory effect on lymphocyte growth. It is also considered an important pathogenic factor, affecting cell permeability and the expression of inflammatory factors, causing cell damage and apoptosis and significantly mediating the pathogenesis of salpingitis in poultry [29,30,31]. These results are comparable to data previously presented by other authors [30], which also relate to the presence of another virulence gene often used to identify G. anatis infections in chickens and turkeys. The gene gyrB encodes the ATPase domain of DNA gyrase, which is necessary for the replication of DNA in G. anatis, and it was detected in this study in all positive isolates. Gallibacterium anatis has the ability to adhere to glycoprotein receptors on the mucous membranes of birds. This is possible due to the presence of fimbriae, which have different sizes and shapes and are F17-like. Different types of fimbriae involved in the processes of colonisation and tissue adhesion were identified and grouped [10,12,32]. The flfA gene was another virulence gene detected in this study. The content of this gene in G. anatis isolates was much lower than those of GtxA or gyrB, which is consistent with another study (Figure 1) [12].
G. anatis infections can lead to a wide range of serious clinical signs in many parts of the avian body. The bacterium can colonise many important organs in the host organism. In addition, the alarming increase in its multidrug resistance to an ever-increasing range of antibiotics noted around the world can lead to infections and treatment failures in poultry flocks.
5. Conclusions
In this study, we showed the prevalence of G. anatis in laying chickens in Poland. The findings from the current study show the multidrug resistance of the isolated strains and the presence of genes related to the virulence and resistance of G. anatis bacteria isolated from laying chickens. The results of this study may expand the knowledge of the pathogenicity of G. anatis in poultry.
Author Contributions
O.K. processed samples, analysed data and wrote the manuscript; G.T. discussed the results and reviewed the manuscript; A.S. processed samples; and A.S.-D. reviewed and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
All samples were from commercial turkeys and chickens and were taken by veterinarians during routine diagnostic examinations. Formal ethical approval is not required for this kind of study (Directive 2010/63/EU Chapter I, Article 1, paragraph 5, points b, d, f).
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author. The data are not publicly available because of legislation protecting privacy.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Singh, S.V.; Singh, B.R.; Sinha, D.K.; Vinodh, K.O.R.; Prasanna, V.A.; Monika, B.; Sakshi, D. Gallibacterium Anatis: An Emerging Pathogen of Poultry Birds and Domiciled Birds. J. Vet. Sci. Technol. 2016, 7, 324. [Google Scholar] [CrossRef]
- Christensen, H.; Kuhnert, P.; Norskov-Lauritsen, N.; Planet, P.J.; Bisgaard, M. The Family Pasteurellaceae. In The Prokaryotes: Gammaproteobacteria; Springer: Berlin/Heidelberg, Germany, 2014; pp. 535–564. [Google Scholar]
- Elbestawy, A.R.; Ellakany, H.F.; Abd El-Hamid, H.S.; Bekheet, A.A.; Mataried, N.E.; Nasr, S.M.; Amarin, N.M. Immunology, Health, and Disease: Isolation, Characterization, and Antibiotic Sensitivity Assessment of Gallibacterium Anatis Biovar Haemolytica, from Diseased Egyptian Chicken Flocks during the Years 2013 and 2015. Poult. Sci. 2018, 97, 1519–1525. [Google Scholar] [CrossRef] [PubMed]
- Paudel, S.; Liebhart, D.; Aurich, C.; Hess, M.; Hess, C. Pathogenesis of Gallibacterium Anatis in a Natural Infection Model Fulfils Koch’s Postulates: 2. Epididymitis and Decreased Semen Quality Are the Predominant Effects in Specific Pathogen Free Cockerels. Avian Pathol. 2014, 43, 529–534. [Google Scholar] [CrossRef] [PubMed]
- Neubauer, C.; De Souza-Pilz, M.; Bojesen, A.M.; Bisgaard, M.; Hess, M. Tissue Distribution of Haemolytic Gallibacterium Anatis Isolates in Laying Birds with Reproductive Disorders. Avian Pathol. 2009, 38, 1–7. [Google Scholar] [CrossRef]
- Sid, H.; Benachour, K.; Rautenschlein, S. Co-Infection with Multiple Respiratory Pathogens Contributes to Increased Mortality Rates in Algerian Poultry Flocks. Avian Dis. 2015, 59, 440–446. [Google Scholar] [CrossRef]
- Wang, C.; Pors, S.E.; Olsen, R.H.; Bojesen, A.M. Transmission and Pathogenicity of Gallibacterium Anatis and Escherichia Coli in Embryonated Eggs. Vet. Microbiol. 2018, 217, 76–81. [Google Scholar] [CrossRef]
- Roberts, J.R. Factors Affecting Egg Internal Quality and Egg Shell Quality in Laying Hens. J. Poult. Sci. 2004, 41, 161–177. [Google Scholar] [CrossRef]
- Narasinakuppe Krishnegowda, D.; Dhama, K.; Kumar Mariappan, A.; Munuswamy, P.; Iqbal Yatoo, M.; Tiwari, R.; Karthik, K.; Bhatt, P.; Reddy, M.R. Etiology, Epidemiology, Pathology, and Advances in Diagnosis, Vaccine Development, and Treatment of Gallibacterium Anatis Infection in Poultry: A Review. Vet. Q. 2020, 40, 16–34. [Google Scholar]
- Persson, G.; Bojesen, A.M. Bacterial Determinants of Importance in the Virulence of Gallibacterium Anatis in Poultry. Vet. Res. 2015, 46, 57. [Google Scholar] [CrossRef]
- Van Driessche, L.; Vanneste, K.; Bogaerts, B.; De Keersmaecker, S.C.; Roosens, N.H.; Haesebrouck, F.; De Cremer, L.; Deprez, P.; Pardon, B.; Boyen, F. Isolation of Drug-Resistant Gallibacterium Anatis from Calves with Unresponsive Bronchopneumonia, Belgium. Emerg. Infect. Dis. 2020, 26, 721. [Google Scholar] [CrossRef]
- Algammal, A.M.; Hashem, M.E.A.; Alfifi, K.J.; Al-Otaibi, A.S.; Alatawy, M.; Eltarabili, R.M.; El-Ghany, W.A.A.; Hetta, H.F.; Hamouda, A.M.; Elewa, A.A.; et al. Sequence Analysis, Antibiogram Profile, Virulence and Antibiotic Resistance Genes of XDR and MDR Gallibacterium Anatis Isolated from Layer Chickens in Egypt. Infect. Drug Resist. 2022, 15, 4321–4334. [Google Scholar] [CrossRef]
- Ghislain Aubin, G.; Haloun, A.; Treilhaud, M.; Reynaud, A.; Corvec, S. Gallibacterium Anatis Bacteremia in a Human. J. Clin. Microbiol. 2013, 51, 3897–3899. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Robles, F.; Ramirez, S.; Riber, A.B.; Bojesen, A.M. Culture-Independent Identification and Quantification of Gallibacterium anatis (G. anatis) by Real-Time Quantitative PCR. Avian Pathol. 2016, 45, 538–544. [Google Scholar] [CrossRef] [PubMed]
- Bojesen, A.M.; Vazquez, M.E.; Robles, F.; Gonzalez, C.; Soriano, E.V.; Olsen, J.E.; Christensen, H. Specific Identification of Gallibacterium by a PCR Using Primers Targeting the 16S RRNA and 23S RRNA Genes. Vet. Microbiol. 2007, 123, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Bojesen, A.M.; Nielsen, O.L.; Christensen, J.P.; Bisgaard, M. In Vivo Studies of Gallibacterium Anatis Infection in Chickens. Avian Pathol. 2004, 33, 145–152. [Google Scholar] [CrossRef]
- Johnson, T.J.; Danzeisen, J.L.; Trampel, D.; Nolan, L.K.; Seemann, T.; Bager, R.J.; Bojesen, A.M. Genome Analysis and Phylogenetic Relatedness of Gallibacterium Anatis Strains from Poultry. PLoS ONE 2013, 8, e54844. [Google Scholar] [CrossRef]
- Castillo, G.; Koga, Y.; Alvarado, A.; Tinoco, R.; Fernández, D. Isolation and Biochemical Characterization of Pasteurella Multocida and Gallibacterium Anatis Strains in Poultry with Respiratory Signs. Rev. Investig. Vet. Peru 2014, 25, 516–522. [Google Scholar] [CrossRef]
- Kursa, O.; Tomczyk, G.; Sawicka-Durkalec, A.; Giza, A.; Słomiany-Szwarc, M. Bacterial Communities of the Upper Respiratory Tract of Turkeys. Sci. Rep. 2021, 11, 2544. [Google Scholar] [CrossRef]
- Kursa, O.; Tomczyk, G.; Adamska, K.; Chrzanowska, J.; Sawicka-Durkalec, A. The Microbial Community of the Respiratory Tract of Commercial Chickens and Turkeys. Microorganisms 2022, 10, 987. [Google Scholar] [CrossRef]
- Hess, C.; Grafl, B.; Bagheri, S.; Kaesbohrer, A.; Zloch, A.; Hess, M. Antimicrobial Resistance Profiling of Gallibacterium Anatis from Layers Reveals High Number of Multiresistant Strains and Substantial Variability Even between Isolates from the Same Organ. Microb. Drug Resist. 2020, 26, 169–177. [Google Scholar] [CrossRef]
- Shabbir, M.Z.; Kariyawasam, S.; Pierre, T.A.; Dunn, P.A.; Wallner-Pendleton, E.A.; Lu, H. Identification, 16S RRNA–Based Characterization, and Antimicrobial Profile of Gallibacterium Isolates from Broiler and Layer Chickens. J. Vet. Diagn. Investig. 2023, 35, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Christensen, H.; Bisgaard, M.; Bojesen, A.M.; Mutters, R.; Olsen, J.E. Genetic Relationship among Avian Isolates Classified as Pasteurella Haemolytica. “Actinobacillus Salpingitidis” or Pasteurella Anatis with Proposal of Gallibacterium Anatis Gen. Nov., Comb. Nov. and Description of Additional Genomospecies with Gallibacterium Gen. Nov. Int. J. Syst. Evol. Microbiol. 2003, 53, 275–287. [Google Scholar] [CrossRef]
- Yaman, S.; Sahan Yapicier, O. Gallibacterium Anatis: Moleculer Detection of Tetracycline Resistance and Virulence Gene. J. Worlds Poult. Res. 2020, 10, 385–390. [Google Scholar] [CrossRef]
- Bojesen, A.M.; Vazquez, M.E.; Bager, R.J.; Ifrah, D.; Gonzalez, C.; Aarestrup, F.M. Antimicrobial Susceptibility and Tetracycline Resistance Determinant Genotyping of Gallibacterium Anatis. Vet. Microbiol. 2011, 148, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Allahghadry, T.; Ng, D.Y.K.; Dibaei, A.; Bojesen, A.M. Clonal Spread of Multi-Resistant Gallibacterium Anatis Isolates among Iranian Broilers and Layers. Vet. Res. 2021, 52, 27. [Google Scholar] [CrossRef]
- El-Adawy, H.; Bocklisch, H.; Neubauer, H.; Hafez, H.M.; Hotzel, H. Identification, Differentiation and Antibiotic Susceptibility of Gallibacterium Isolates from Diseased Poultry. Ir. Vet. J. 2018, 71, 5. [Google Scholar] [CrossRef] [PubMed]
- Bojesen, A.M.; Bager, R.J.; Ifrah, D.; Aarestrup, F.M. The Rarely Reported Tet(31) Tetracycline Resistance Determinant Is Common in Gallibacterium Anatis. Vet. Microbiol. 2011, 149, 497–499. [Google Scholar] [CrossRef]
- Kristensen, B.M.; Frees, D.; Bojesen, A.M. GtxA from Gallibacterium Anatis, a Cytolytic RTX-Toxin with a Novel Domain Organisation. Vet. Res. 2010, 41, 25. [Google Scholar] [CrossRef]
- Yang, X.; Xia, Y.H.; Wang, J.Y.; Li, Y.T.; Chang, Y.F.; Chang, H.T.; Liu, H.Y.; Chen, L.; Wang, C.Q. The Role of GtxA during Gallibacterium Anatis Infection of Primary Chicken Oviduct Epithelial Cells. Mol. Cell. Probes 2020, 53, 101641. [Google Scholar] [CrossRef]
- Tang, B.; Bojesen, A.M. Immune Suppression Induced by Gallibacterium Anatis Gtxa during Interaction with Chicken Macrophage-like HD11 Cells. Toxins 2020, 12, 536. [Google Scholar] [CrossRef]
- Persson, G.; Pors, S.E.; Thøfner, I.C.N.; Bojesen, A.M. Vaccination with Outer Membrane Vesicles and the Fimbrial Protein FlfA Offers Improved Protection against Lesions Following Challenge with Gallibacterium Anatis. Vet. Microbiol. 2018, 217, 104–111. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).