Molecular Characterizations of the Coagulase-Negative Staphylococci Species Causing Urinary Tract Infection in Tanzania: A Laboratory-Based Cross-Sectional Study
Abstract
1. Introduction
2. Results
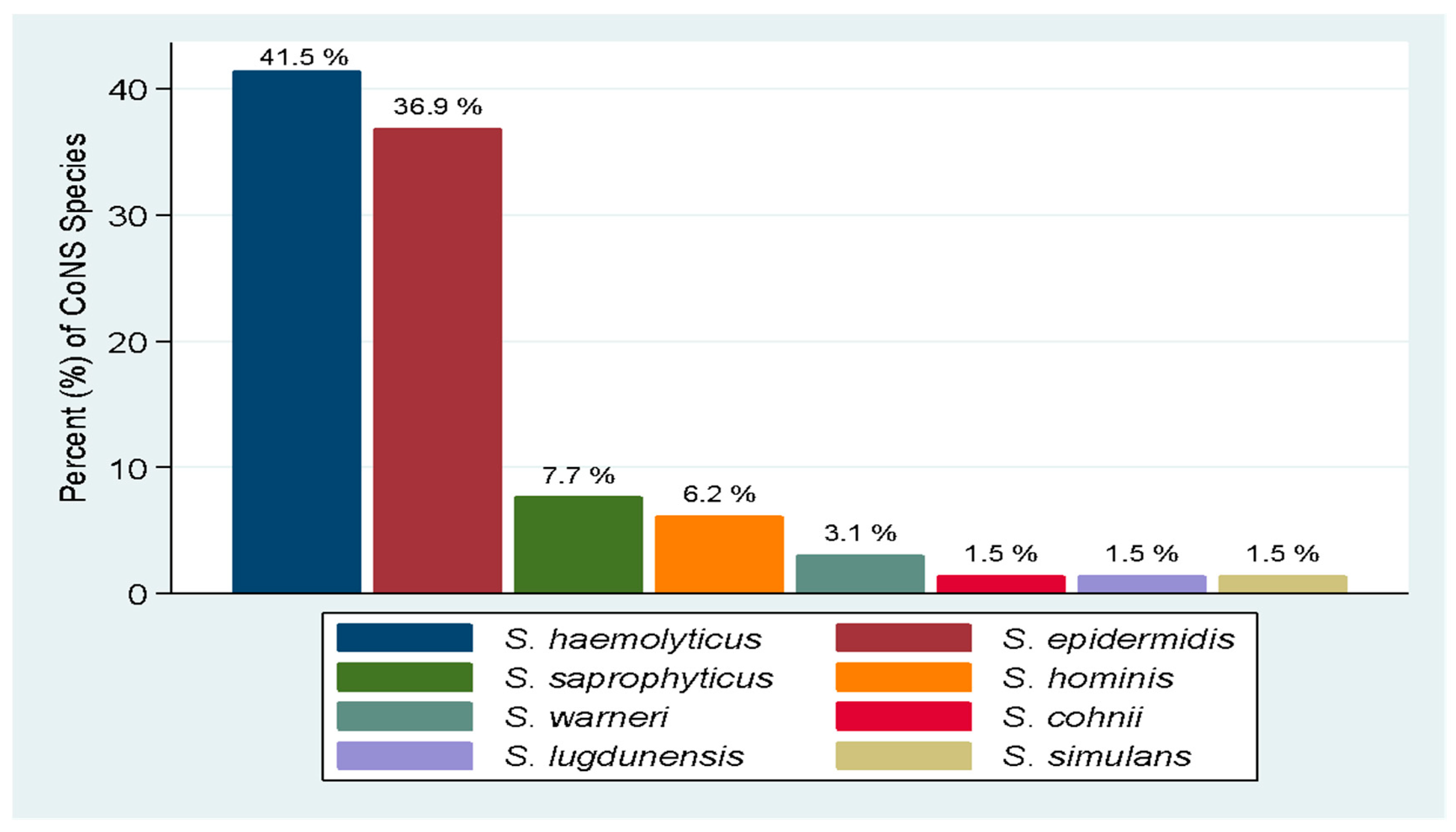
2.1. Bacteria Speciation and Distribution of CoNS Species from Urine Samples
2.2. Multilocus Sequence Types of CoNS Species Causing UTI
2.3. Virulence Genes of CoNS Species Causing UTI
2.4. Antimicrobial Resistance Genes Identified among CoNS Species
3. Discussion
4. Materials and Methods
4.1. Study Design and Area
4.2. Study Population
4.3. Sample Size and Isolate Recovery
4.4. DNA Extraction and Library Preparation
4.5. Illumina Sequencing
4.6. Genome Quality Check and Assembly
4.7. Bacterial Speciation and Multilocus Sequence Typing
4.8. Antimicrobial Resistance Genes Identification and Virulence Genes Annotation
4.9. Statistical Data Analysis
4.10. Ethical Consideration
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Öztürk, R.A. Murt, Epidemiology of urological infections: A global burden. World J. Urol. 2020, 38, 2669–2679. [Google Scholar] [CrossRef] [PubMed]
- Seni, J.; Tito, J.; Makoye, S.; Mbena, H.; Alfred, H.; van der Meer, F.; Pitout, J.; Mshana, S.; DeVinney, R. Multicentre evaluation of significant bacteriuria among pregnant women in the cascade of referral healthcare system in North-western Tanzania: Bacterial pathogens, antimicrobial resistance profiles and predictors. J. Glob. Antimicrob. Resist. 2019, 17, 173–179. [Google Scholar] [CrossRef] [PubMed]
- Chaula, T.; Seni, J.; Ng’Walida, N.; Kajura, A.; Mirambo, M.M.; DeVinney, R.; Mshana, S.E. Urinary Tract Infections among HIV-Positive Pregnant Women in Mwanza City, Tanzania, Are High and Predicted by Low CD4+ Count. Int. J. Microbiol. 2017, 2017, 4042686. [Google Scholar] [CrossRef]
- Mushi, M.F.; Alex, V.G.; Seugendo, M.; Silago, V.; Mshana, S.E. C—reactive protein and urinary tract infection due to Gram-negative bacteria in a pediatric population at a tertiary hospital, Mwanza, Tanzania. Afr. Health Sci. 2019, 19, 3217–3224. [Google Scholar] [CrossRef] [PubMed]
- Msaki, B.P.; Mshana, S.E.; Hokororo, A.; Mazigo, H.D.; Morona, D. Prevalence and predictors of urinary tract infection and severe malaria among febrile children attending Makongoro health centre in Mwanza city, North-Western Tanzania. Arch. Public Health 2012, 70, 4. [Google Scholar] [CrossRef] [PubMed]
- Festo, E.; Kidenya, B.R.; Hokororo, A.; Mshana, S.E. Predictors of urinary tract infection among febrile children attending at Bugando Medical Centre Northwestern, Tanzania. Arch. Clin. Microbiol. 2011, 2, 5. [Google Scholar]
- Koneman, E.W.; Allen, S.D.; Janda, W.M.; Schreckenberger, P.C.; Winn, W.C. Diagnostic microbiology. In The Nonfermentative Gram-Negative Bacilli; Lippincott-Raven Publishers: Philadelphia, PA, USA, 1997; pp. 253–320. [Google Scholar]
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary tract infections: Epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef]
- Hashmi, A.; Abdullah, F.E.; Abdullah, N.E.; Kazmi, S.U. Species Identification and Antibiotic Susceptibilities of Coagulase- Negative Staphylococci Isolated from Urinary Tract Infection Specimens. J. Coll. Physicians Surg. Pak. 2016, 26, 581–584. [Google Scholar]
- Stamm, W.E.; Norrby, S.R. Urinary Tract Infections: Disease Panorama and Challenges. J. Infect. Dis. 2001, 183 (Suppl. S1), S1–S4. [Google Scholar] [CrossRef]
- Qiao, L.-D.; Chen, S.; Yang, Y.; Zhang, K.; Zheng, B.; Guo, H.-F.; Yang, B.; Niu, Y.-J.; Wang, Y.; Shi, B.-K.; et al. Characteristics of urinary tract infection pathogens and their in vitro susceptibility to antimicrobial agents in China: Data from a multicenter study. BMJ Open 2013, 3, e004152. [Google Scholar] [CrossRef]
- John, J.F., Jr.; Gramling, P.K.; O’dell, N.M. Species identification of coagulase-negative staphylococci from urinary tract isolates. J. Clin. Microbiol. 1978, 8, 435–437. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.-H.; Naber, K.; Hacker, J.; Ziebuhr, W. Detection of the icaADBC gene cluster and biofilm formation in Staphylococcus epidermidis isolates from catheter-related urinary tract infections. Int. J. Antimicrob. Agents 2002, 19, 570–575. [Google Scholar] [CrossRef] [PubMed]
- Iorio, N.; Azevedo, M. Methicillin-resistant Staphylococcus epidermidis carrying biofilm formation genes: Detection of clinical isolates by multiplex PCR. Int. Microbiol. 2011, 14, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Nasaj, M.; Saeidi, Z.; Asghari, B.; Roshanaei, G.; Arabestani, M.R. Identification of hemolysin encoding genes and their association with antimicrobial resistance pattern among clinical isolates of coagulase-negative Staphylococci. BMC Res. Notes 2020, 13, 1–6. [Google Scholar] [CrossRef]
- Mbanga, J.; Masuku, S.; Luphahla, S. Antibiotic Resistance Patterns and Virulence Factors of Coagulase Negative Staphylococcus Associated with Urinary Tract Infections in Bulawayo Province, Zimbabwe. Br. J. Med. Med. Res. 2016, 11, 1–9. [Google Scholar] [CrossRef]
- Chen, J.; Fang, Q. Whole-Genome Sequencing Analysis of Methicillin-Resistant Staphylococcus simulans Causing Surgical Site Infection. Genome Announc. 2016, 4, e00555-16. [Google Scholar] [CrossRef]
- Widerström, M. Molecular Epidemiology of Coagulase-Negative Staphylococci in Hospitals and in the Community; Umeå University: Umeå, Sweden, 2010. [Google Scholar]
- Chassain, B.; Lemée, L.; Didi, J.; Thiberge, J.-M.; Brisse, S.; Pons, J.-L.; Pestel-Caron, M. Multilocus Sequence Typing Analysis of Staphylococcus lugdunensis Implies a Clonal Population Structure. J. Clin. Microbiol. 2012, 50, 3003–3009. [Google Scholar] [CrossRef]
- Ehlers, M.M.; Strasheim, W.; Lowe, M.; Ueckermann, V.; Kock, M.M. Molecular Epidemiology of Staphylococcus epidermidis Implicated in Catheter-Related Bloodstream Infections at an Academic Hospital in Pretoria, South Africa. Front. Microbiol. 2018, 9, 417. [Google Scholar] [CrossRef]
- Eligius, F.L.; Sabrina, J.M.; Ewaldo, V.K.; Martin, H. Prevalence, antimicrobial resistance and associated risk factors for bacteriuria in diabetic women in Dar es Salaam, Tanzania. Afr. J. Microbiol. Res. 2011, 5, 683–689. [Google Scholar]
- Francis, F.; Joel, M.F.; Maulidi, F.; Samuel, Y.M. Aetiology, antimicrobial susceptibility and predictors of urinary tract infection among febrile under-fives at Muhimbili National Hospital, Dar es Salaam-Tanzania. Afr. J. Microbiol. Res. 2013, 7, 1029–1034. [Google Scholar]
- Silago, V.; Moremi, N.; Mtebe, M.; Komba, E.; Masoud, S.; Mgaya, F.X.; Mirambo, M.M.; Nyawale, H.A.; Mshana, S.E.; Matee, M.I. Multidrug-Resistant Uropathogens Causing Community Acquired Urinary Tract Infections among Patients Attending Health Facilities in Mwanza and Dar es Salaam, Tanzania. Antibiotics 2022, 11, 1718. [Google Scholar] [CrossRef] [PubMed]
- Hagler, S.; Dobkin, D. Urinary Tract Infection in the Male Caused by Staphylococcus epidermidis Associated with Diverticulum of the Bladder. Clin. Pediatr. 1990, 29, 527–528. [Google Scholar] [CrossRef] [PubMed]
- Kline, K.A.; Lewis, A.L. Gram-Positive Uropathogens, Polymicrobial Urinary Tract Infection, and the Emerging Microbiota of the Urinary Tract. Microbiol. Spectr. 2016, 4, 459–502. [Google Scholar] [CrossRef] [PubMed]
- Megged, O. Coagulase-negative Staphylococci: A rare cause of urinary tract infections in children with consequences on clinical practice. Eur. J. Pediatr. 2022, 181, 1099–1104. [Google Scholar] [CrossRef] [PubMed]
- Kumari, N.; Rai, A.; Jaiswal, C.P.; Xess, A.; Shahi, S.K. Coagulase negative Staphylococci as causative agents of urinary tract infections-prevalence and resistance status in IGIMS, Patna. Indian J. Pathol. Microbiol. 2001, 44, 415–419. [Google Scholar]
- Crude, N.; Tveten, Y.; Kristiansen, B.-E. Urinary tract infections in Norway: Bacterial etiology and susceptibility. A retrospective study of clinical isolates. Clin. Microbiol. Infect. 2001, 7, 543–547. [Google Scholar] [CrossRef]
- Caierão, J.; Superti, S.; Dias, C.A.; D’Azevedo, P.A. Automated systems in the identification and determination of methicillin resistance among coagulase negative staphylococci. Memórias do Instituto Oswaldo Cruz 2006, 101, 277–280. [Google Scholar] [CrossRef]
- Takeuchi, F.; Watanabe, S.; Baba, T.; Yuzawa, H.; Ito, T.; Morimoto, Y.; Kuroda, M.; Cui, L.; Takahashi, M.; Ankai, A.; et al. Whole-Genome Sequencing of Staphylococcus haemolyticus Uncovers the Extreme Plasticity of Its Genome and the Evolution of Human-Colonizing Staphylococcal Species. J. Bacteriol. 2005, 187, 7292–7308. [Google Scholar] [CrossRef]
- Tristan, A.; Lina, G.; Etienne, J.; Vandenesch, F. Biology and Pathogenicity of Staphylococci Other than Staphylococcus aureus and Staphylococcus epidermidis. Gram-Posit. Pathog. 2006, ch47, 572–586. [Google Scholar] [CrossRef]
- Bouchami, O.; de Lencastre, H.; Miragaia, M. Impact of Insertion Sequences and Recombination on the Population Structure of Staphylococcus haemolyticus. PLoS ONE 2016, 11, e0156653. [Google Scholar] [CrossRef]
- Al-Naqshbandi, A.A.; Chawsheen, M.A.; Abdulqader, H.H. Prevalence and antimicrobial susceptibility of bacterial pathogens isolated from urine specimens received in rizgary hospital—Erbil. J. Infect. Public Health 2019, 12, 330–336. [Google Scholar] [CrossRef] [PubMed]
- Haider, J.; Hasan, A.; Bin-Tahir, K. Frequency of urinary tract bacterial infection and their susceptibility patterns among hemodialysis patients in Zliten hospital. J. Microbiol. Exp. 2016, 3, 93–97. [Google Scholar] [CrossRef]
- Orrett, F.A.; Shurland, S.M. Significance of coagulase-negative staphylococci in urinary tract infections in a developing country. Connect. Med. 1998, 62, 199–203. [Google Scholar]
- Widerström, M.; Wiström, J.; Sjöstedt, A.; Monsen, T. Coagulase-negative staphylococci: Update on the molecular epidemiology and clinical presentation, with a focus on Staphylococcus epidermidis and Staphylococcus saprophyticus. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Raz, R.; Colodner, R.; Kunin, C.M. Who are you—Staphylococcus saprophyticus? Clin. Infect. Dis. 2005, 40, 896–898. [Google Scholar] [CrossRef] [PubMed]
- Latham, R.H.; Running, K.; Stamm, W.E. Urinary tract infections in young adult women caused by Staphylococcus saprophyticus. JAMA 1983, 250, 3063–3066. [Google Scholar] [CrossRef] [PubMed]
- Maiden, M.C.J.; Bygraves, J.A.; Feil, E.; Morelli, G.; Russell, J.E.; Urwin, R.; Zhang, Q.; Zhou, J.; Zurth, K.; Caugant, D.A.; et al. Multilocus sequence typing: A portable approach to the identification of clones within populations of pathogenic microorganisms. Proc. Natl. Acad. Sci. USA 1998, 95, 3140–3145. [Google Scholar] [CrossRef] [PubMed]
- Panda, S.; Jena, S.; Sharma, S.; Dhawan, B.; Nath, G.; Singh, D.V. Identification of Novel Sequence Types among Staphylococcus haemolyticus Isolated from Variety of Infections in India. PLoS ONE 2016, 11, e0166193. [Google Scholar] [CrossRef]
- Miragaia, M.; De Lencastre, H.; Perdreau-Remington, F.; Chambers, H.F.; Higashi, J.; Sullam, P.M.; Lin, J.; Wong, K.I.; King, K.A.; Otto, M.; et al. Genetic Diversity of Arginine Catabolic Mobile Element in Staphylococcus epidermidis. PLoS ONE 2009, 4, e7722. [Google Scholar] [CrossRef]
- Soumya, K.R.; Philip, S.; Sugathan, S.; Mathew, J.; Radhakrishnan, E.K. Virulence factors associated with Coagulase Negative Staphylococci isolated from human infections. 3 Biotech 2017, 7, 140. [Google Scholar] [CrossRef]
- Fey, P.D.; Olson, M.E. Current concepts in biofilm formation of Staphylococcus epidermidis. Futur. Microbiol. 2010, 5, 917–933. [Google Scholar] [CrossRef] [PubMed]
- Solati, S.M.; Tajbakhsh, E.; Khamesipour, F.; Gugnani, H.C. Prevalence of virulence genes of biofilm producing strains of Staphylococcus epidermidis isolated from clinical samples in Iran. AMB Express 2015, 5, 134. [Google Scholar] [CrossRef] [PubMed]
- Dworkin, M.S.; Williamson, J.; Jones, J.L.; Kaplan, J.E.; Adult and Adolescent Spectrum of HIV Disease Project. Prophylaxis with Trimethoprim-Sulfamethoxazole for Human Immunodeficiency Virus–Infected Patients: Impact on Risk for Infectious Diseases. Clin. Infect. Dis. 2001, 33, 393–398. [Google Scholar] [CrossRef] [PubMed]
- Duran, N.; Ozer, B.; Duran, G.G.; Onlen, Y.; Demir, C. Antibiotic resistance genes & susceptibility patterns in staphylococci. Indian J. Med. Res. 2012, 135, 389. [Google Scholar] [PubMed]
- Zhang, H.Z.; Hackbarth, C.J.; Chansky, K.M.; Chambers, H.F. A Proteolytic Transmembrane Signaling Pathway and Resistance to β-Lactams in Staphylococci. Science 2001, 291, 1962–1965. [Google Scholar] [CrossRef]
- Chambers, H.F. Methicillin resistance in staphylococci: Molecular and biochemical basis and clinical implications. Clin. Microbiol. Rev. 1997, 10, 781–791. [Google Scholar] [CrossRef]
- Shrestha, L.B.; Bhattarai, N.R.; Rai, K.; Khanal, B. Antibiotic Resistance and mecA Gene Characterization of Coagulase-negative Staphylococci Isolated from Clinical Samples in Nepal. Infect. Drug Resist. 2020, 13, 3163. [Google Scholar] [CrossRef]
- Asiimwe, B.B.; Kiiru, J.; Mshana, S.; Neema, S.; Keenan, K.; Kesby, M.; Mwanga, J.R.; Sloan, D.J.; Mmbaga, B.T.; Smith, V.A.; et al. Protocol for an interdisciplinary cross-sectional study investigating the social, biological and community-level drivers of antimicrobial resistance (AMR): Holistic Approach to Unravel Antibacterial Resistance in East Africa (HATUA). BMJ Open 2021, 11, e041418. [Google Scholar] [CrossRef]
- Micci, F.; Thorsen, J.; Panagopoulos, I.; Nyquist, K.B.; Zeller, B.; Tierens, A.; Heim, S. High-throughput sequencing identifies an NFIA/CBFA2T3 fusion gene in acute erythroid leukemia with t(1;16)(p31;q24). Leukemia 2013, 27, 980–982. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Kang, H.J.; Kawasawa, Y.I.; Cheng, F.; Zhu, Y.; Xu, X.; Li, M.; Sousa, A.M.M.; Pletikos, M.; Meyer, K.A.; Sedmak, G.; et al. Spatio-temporal transcriptome of the human brain. Nature 2011, 478, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Antipov, D.; Hartwick, N.; Shen, M.; Raiko, M.; Lapidus, A.; Pevzner, P.A. plasmidSPAdes: Assembling plasmids from whole genome sequencing data. Bioinformatics 2016, 32, 3380–3387. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Pevzner, P.A. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid Prokaryotic Genome Annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- De Koning, W.; Miladi, M.; Hiltemann, S.; Heikema, A.; Hays, J.P.; Flemming, S.; Stubbs, A.P. NanoGalaxy: Nanopore long-read sequencing data analysis in Galaxy. GigaScience 2020, 9, giaa105. [Google Scholar] [CrossRef]
| Species | ST | Number |
|---|---|---|
| S. haemolyticus | 30 | 8 |
| 56 | 2 | |
| 1 | 2 | |
| 49 | 2 | |
| 145 * | 2 | |
| 38 | 1 | |
| 66 | 1 | |
| 138 * | 1 | |
| 139 * | 1 | |
| 140 * | 1 | |
| 141 * | 1 | |
| 142 * | 1 | |
| 143 * | 1 | |
| 144 * | 1 | |
| S. epidermidis | 490 | 7 |
| 329 | 1 | |
| 150 | 1 | |
| 1184 * | 1 | |
| 1185 * | 1 | |
| 1186 * | 1 | |
| 1187 * | 1 | |
| 1188 * | 1 | |
| 1189 * | 1 | |
| 1190 * | 1 | |
| Unassigned ST | 8 | |
| S. simulans | 56 | 1 |
| S. lugdunensis | 3 | 1 |
| Gene | Product | CoNS Species | Frequency (%) |
|---|---|---|---|
| fbe | Fibrinogen binding protein | S. haemolyticus, S. epidermidis, S. simulans | 13 (12.9) |
| icaB | Poly-beta-1, 6, N-acetyl-D-glucosamine deacetylase | S. haemolyticus, S. epidermidis, S. simulans | 13 (12.9) |
| icaD | Poly-beta-1,6-N-acetyl-D-glucosamine synthesis protein IcaD | S. haemolyticus, S. epidermidis, S. simulans | 14 (13.9) |
| icaA | Poly-beta-1, 6, N-acetyl-D-glucosamine synthase | S. haemolyticus, S. epidermidis, S. simulans | 14 (13.9) |
| icaC | Poly-beta-1, 6, N-acetyl-D-glucosamine export protein | S. haemolyticus, S. epidermidis, S. hominis, S. cohnii | 47 (46.5) |
| Gene(s) singly or in combination | |||
| fbe | S. haemolyticus (1) | 2 (3.4) | |
| S. epidermidis (1) | |||
| icaC | S. epidermidis (8), S. haemolyticus (19) | 34 (57.6) | |
| S. saprophyticus (5), S. hominis (1), S. cohnii (1) | |||
| icaC, fbe | S. haemolyticus (1) | 1 (1.7) | |
| icaA, icaB, fbe | S. epidermidis (1) | 1 (1.7) | |
| icaA, icaC, icaD | S. haemolyticus (1) | 1 (1.7) | |
| icaA, icaB, icaC, icaD | S. epidermidis (5) | 5 (8.5) | |
| S. simulans (1) | |||
| icaA, icaB, icaC, icaD, fbe | S. epidermidis (9), S. haemolyticus (5) | 15 (25.4) | |
| Total | 59 (100.0) | ||
| CoNS Species | Genes [Predicted AMR Phenotype] | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| dfrG [SXT] | blaZ [AMP] | mecA [MET] | aac (6’) aph (2”) [CN] | tetK [TET] | msrA [E/AZT] | erm(C) [E/AZT] | mphC [E] | fosB [FO] | cat(pc221) [C] | tetM [TET] | tetL [TET] | cat(pc233) [C] | cat [C] | |
| S. haemolyticus (27) | 24 | 21 | 21 | 18 | 8 | 11 | 8 | 6 | 4 | 7 | 7 | 3 | 2 | |
| S. epidermidis (23) | 22 | 7 | 1 | 2 | 8 | 5 | 3 | 2 | 12 | 4 | 1 | |||
| S. saprophyticus (2) | 2 | 2 | 1 | 1 | 1 | 2 | 1 | 1 | ||||||
| S. hominis (4) | 2 | 2 | 2 | 1 | 2 | 2 | ||||||||
| S. warneri (2) | 2 | 1 | 2 | 2 | 1 | 1 | 1 | |||||||
| S. cohnii (1) | 1 | 1 | ||||||||||||
| S. simulans (1) | 1 | |||||||||||||
| Total | 53 | 32 | 26 | 24 | 21 | 21 | 14 | 9 | 12 | 9 | 8 | 7 | 4 | 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Phillip, S.; Mushi, M.F.; Decano, A.G.; Seni, J.; Mmbaga, B.T.; Kumburu, H.; Konje, E.T.; Mwanga, J.R.; Kidenya, B.R.; Msemwa, B.; et al. Molecular Characterizations of the Coagulase-Negative Staphylococci Species Causing Urinary Tract Infection in Tanzania: A Laboratory-Based Cross-Sectional Study. Pathogens 2023, 12, 180. https://doi.org/10.3390/pathogens12020180
Phillip S, Mushi MF, Decano AG, Seni J, Mmbaga BT, Kumburu H, Konje ET, Mwanga JR, Kidenya BR, Msemwa B, et al. Molecular Characterizations of the Coagulase-Negative Staphylococci Species Causing Urinary Tract Infection in Tanzania: A Laboratory-Based Cross-Sectional Study. Pathogens. 2023; 12(2):180. https://doi.org/10.3390/pathogens12020180
Chicago/Turabian StylePhillip, Shukrani, Martha F. Mushi, Arun Gonzales Decano, Jeremiah Seni, Blandina T. Mmbaga, Happiness Kumburu, Eveline T. Konje, Joseph R. Mwanga, Benson R. Kidenya, Betrand Msemwa, and et al. 2023. "Molecular Characterizations of the Coagulase-Negative Staphylococci Species Causing Urinary Tract Infection in Tanzania: A Laboratory-Based Cross-Sectional Study" Pathogens 12, no. 2: 180. https://doi.org/10.3390/pathogens12020180
APA StylePhillip, S., Mushi, M. F., Decano, A. G., Seni, J., Mmbaga, B. T., Kumburu, H., Konje, E. T., Mwanga, J. R., Kidenya, B. R., Msemwa, B., Gillespie, S., Maldonado-Barragan, A., Sandeman, A., Sabiti, W., Holden, M. T. G., & Mshana, S. E. (2023). Molecular Characterizations of the Coagulase-Negative Staphylococci Species Causing Urinary Tract Infection in Tanzania: A Laboratory-Based Cross-Sectional Study. Pathogens, 12(2), 180. https://doi.org/10.3390/pathogens12020180







