Biofilm through the Looking Glass: A Microbial Food Safety Perspective
Abstract
1. Introduction
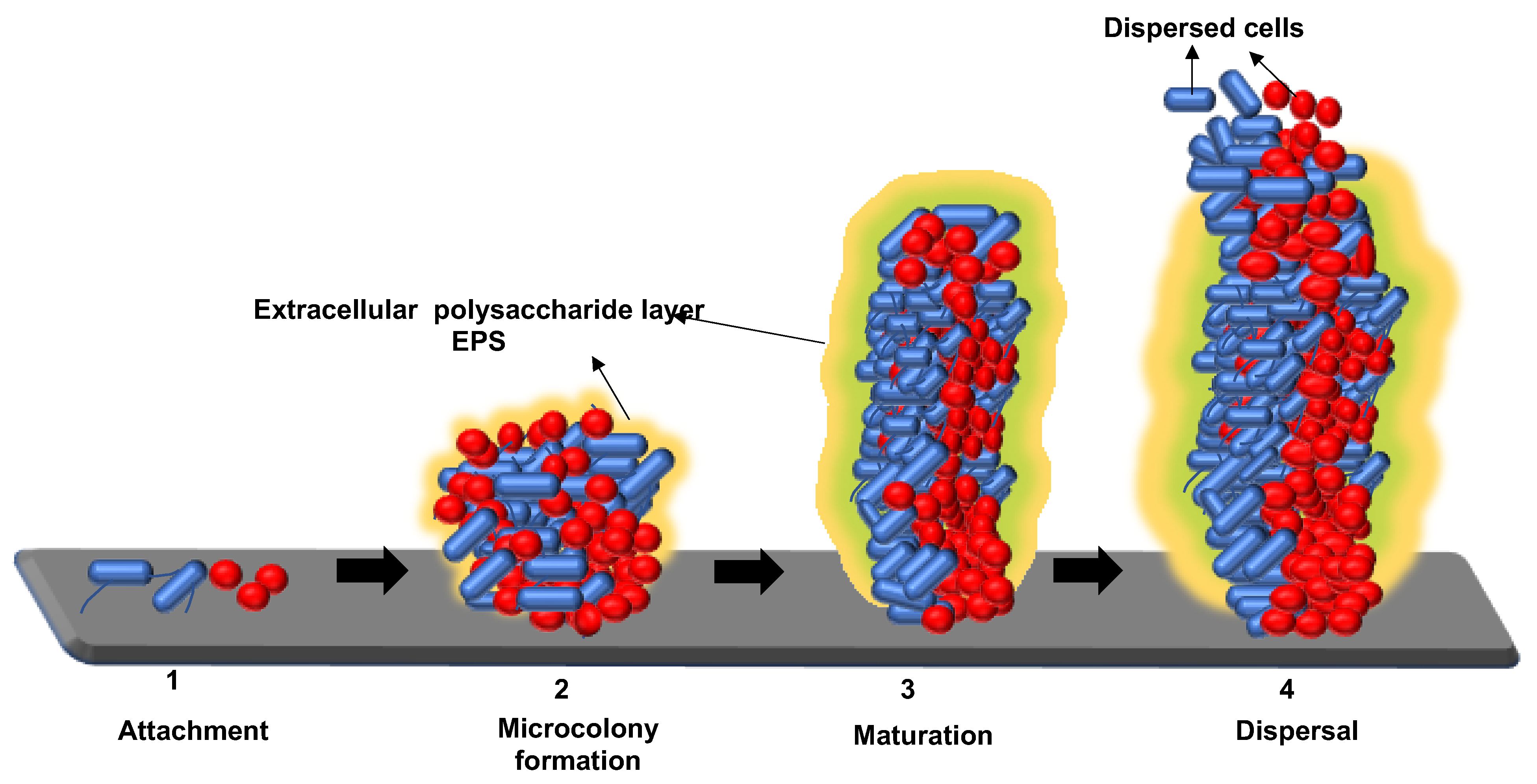
2. Mechanisms for Biofilm Formation: Attachment, Maturation, and Dispersion of Microorganisms to the Food-Processing Environment Surface
2.1. Attachment of Biofilm to the Substratum
2.2. Maturation of Biofilm
2.3. Dispersion
3. Architecture of Biofilm
4. Interactions of Microorganism in a Biofilm Community
4.1. Signaling in Interspecies, Intraspecies, and Cross-Kingdom Communication
4.2. Metabolic Interactions
4.3. Enhanced Biofilm Formation through Microbial Interactions
5. Molecular Basis of Sanitizer Tolerance
6. Biofilm Community and Genetic Element Exchange
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Williams, P. Quorum sensing, communication and cross-kingdom signalling in the bacterial world. Microbiology 2007, 153, 3923–3938. [Google Scholar] [CrossRef] [PubMed]
- Roder, H.L.; Raghupathi, P.K.; Herschend, J.; Brejnrod, A.; Knochel, S.; Sorensen, S.J.; Burmolle, M. Interspecies interactions result in enhanced biofilm formation by co-cultures of bacteria isolated from a food processing environment. Food Microbiol. 2015, 51, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Frey-Klett, P.; Burlinson, P.; Deveau, A.; Barret, M.; Tarkka, M.; Sarniguet, A. Bacterial-fungal interactions: Hyphens between agricultural, clinical, environmental, and food microbiologists. Microbiol. Mol. Biol. Rev. 2011, 75, 583–609. [Google Scholar] [CrossRef] [PubMed]
- Abram, F. Systems-based approaches to unravel multi-species microbial community functioning. Comput. Struct. Biotechnol. J. 2015, 13, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.V.; Gunawan, C.; Mann, R. We Are One: Multispecies Metabolism of a Biofilm Consortium and Their Treatment Strategies. Front. Microbiol. 2021, 12, 635432. [Google Scholar] [CrossRef] [PubMed]
- McNeilly, O.; Mann, R.; Hamidian, M.; Gunawan, C. Emerging Concern for Silver Nanoparticle Resistance in Acinetobacter baumannii and Other Bacteria. Front. Microbiol. 2021, 12, 652863. [Google Scholar] [CrossRef] [PubMed]
- Muller, E.E.L.; Faust, K.; Widder, S.; Herold, M.; Martínez Arbas, S.; Wilmes, P. Using metabolic networks to resolve ecological properties of microbiomes. Curr. Opin. Syst. Biol. 2018, 8, 73–80. [Google Scholar] [CrossRef]
- Yang, J.; Barrila, J.; Mark Ott, C.; King, O.; Bruce, R.; McLean, R.J.C.; Nickerson, C.A. Longitudinal characterization of multispecies microbial populations recovered from spaceflight potable water. NPJ Biofilms Microbiomes 2021, 7, 70. [Google Scholar] [CrossRef] [PubMed]
- Donlan, M.R. Biofilms: Microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Oliveira, N.M.; Martinez-Garcia, E.; Xavier, J.; Durham, W.M.; Kolter, R.; Kim, W.; Foster, K.R. Biofilm Formation As a Response to Ecological Competition. PLoS Biol. 2015, 13, e1002191. [Google Scholar] [CrossRef] [PubMed]
- Burmolle, M.; Ren, D.; Bjarnsholt, T.; Sorensen, S.J. Interactions in multispecies biofilms: Do they actually matter? Trends Microbiol. 2014, 22, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Federle, M.J.; Bassler, B.L. Interspecies communication in bacteria. J. Clin. Investig. 2003, 112, 1291–1299. [Google Scholar] [CrossRef] [PubMed]
- Hughes, D.T.; Sperandio, V. Inter-kingdom signalling: Communication between bacteria and their hosts. Nat. Rev. Microbiol. 2008, 6, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Zammuto, V.; Rizzo, M.G.; Spanò, A.; Genovese, G.; Morabito, M.; Spagnuolo, D.; Capparucci, F.; Gervasi, C.; Smeriglio, A.; Trombetta, D.; et al. In vitro evaluation of antibiofilm activity of crude extracts from macroalgae against pathogens relevant in aquaculture. Aquaculture 2022, 549, 737729. [Google Scholar] [CrossRef]
- Srey, S.; Jahid, I.K.; Ha, S.-D. Biofilm formation in food industries: A food safety concern. Food Control 2013, 31, 572–585. [Google Scholar] [CrossRef]
- Van Houdt, R.; Michiels, C.W. Biofilm formation and the food industry, a focus on the bacterial outer surface. J. Appl. Microbiol. 2010, 109, 1117–1131. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.; Zhu, X. Biofilm formation and food safety in food industries. Trends Food Sci. Technol. 2009, 20, 407–413. [Google Scholar] [CrossRef]
- Kumar, C.G.; Anand, S.K. Significane of microbial biofilms in food industry: A review. Int. J. Food Microbiol. 1998, 42, 9–27. [Google Scholar] [CrossRef]
- Scallan, E.; Hoekstra, R.M.; Angulo, F.J.; Tauxe, R.V.; Widdowson, M.A.; Roy, S.L.; Jones, J.L.; Griffin, P.M. Foodborne illness acquired in the United States—Major pathogens. Emerg. Infect. Dis. 2011, 17, 7–15. [Google Scholar] [CrossRef]
- CDC. Surveillance for Foodborne Diseases Outbreaks United States, 2015; Annual Report; CDC: Atlanta, GA, USA, 2015.
- NIoH (NIH). Research on Microbial Biofilms; National Institutes of Health: Bethesda, MA, USA, 2002.
- Bridier, A.; Sanchez-Vizuete, P.; Guilbaud, M.; Piard, J.C.; Naitali, M.; Briandet, R. Biofilm-associated persistence of food-borne pathogens. Food Microbiol. 2015, 45, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Garrett, T.R.; Bhakoo, M.; Zhang, Z. Bacterial adhesion and biofilms on surfaces. Prog. Nat. Sci. 2008, 18, 1049–1056. [Google Scholar] [CrossRef]
- Hermansson, M. The DLVO theory in microbial adhesion. Colloids Surf. B Biointerfaces 1999, 14, 105–119. [Google Scholar] [CrossRef]
- Jacob, N.I.; Israelachvili, N. Intermolecular and Surface Forces; Academic Press: San Diego, CA, USA, 1992. [Google Scholar]
- Wilson, L.G.; Everett, L.G.; Cullen, S.J. Handbook of Vadose Zone Characterization & Monitoring; CRC Press: Boca Raton, FI, USA, 2018. [Google Scholar]
- Chang, Y.-I.; Chang, P.-K. The role of hydration force on the stability of the suspension of Saccharomyces cerevisiae—Application of the extended DLVO theory. Colloids Surf. A Physicochem. Eng. Asp. 2002, 211, 67–77. [Google Scholar] [CrossRef]
- Ryan, R.P.; Dow, J.M. Diffusible signals and interspecies communication in bacteria. Microbiology 2008, 154, 1845–1858. [Google Scholar] [CrossRef] [PubMed]
- Gracia-Gonzalo, D.; Pagán, R. Influence of environmental factors on bacterial biofilms formation in the food industry: A review. J. Postdr. Res. 2015, 3, 3–12. [Google Scholar]
- de Jesus Pimentel-Filho, N.; de Freitas Martins, M.C.; Nogueira, G.B.; Mantovani, H.C.; Vanetti, M.C.D. Bovicin HC5 and nisin reduce Staphylococcus aureus adhesion to polystyrene and change the hydrophobicity profile and Gibbs free energy of adhesion. Int. J. Food Microbiol. 2014, 190, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Liu, Y.; Wu, H.; Hóiby, N.; Molin, S.; Song, Z.J. Current understanding of multi-species biofilms. Int. J. Oral Sci. 2011, 3, 74–81. [Google Scholar] [CrossRef]
- Madsen, J.S.; Røder, H.L.; Russel, J.; Sørensen, H.; Burmølle, M.; Sørensen, S.J. Coexistence facilitates interspecific biofilm formation in complex microbial communities. Environ. Microbiol. 2016, 18, 2565–2574. [Google Scholar] [CrossRef] [PubMed]
- Giaouris, E.; Heir, E.; Hebraud, M.; Chorianopoulos, N.; Langsrud, S.; Moretro, T.; Habimana, O.; Desvaux, M.; Renier, S.; Nychas, G.J. Attachment and biofilm formation by foodborne bacteria in meat processing environments: Causes, implications, role of bacterial interactions and control by alternative novel methods. Meat Sci. 2014, 97, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Kuchma, S.L.; Connolly, J.P.; O’Toole, G.A. A three-component regulatory system regulates biofilm maturation and type III secretion in Pseudomonas aeruginosa. J. Bacteriol. 2005, 187, 1441–1454. [Google Scholar] [CrossRef] [PubMed]
- Stojicic, S.; Shen, Y.; Haapasalo, M. Effect of the source of biofilm bacteria, level of biofilm maturation, and type of disinfecting agent on the susceptibility of biofilm bacteria to antibacterial agents. J. Endod. 2013, 39, 473–477. [Google Scholar] [CrossRef] [PubMed]
- Davey, M.E.; Caiazza, N.C.; O’Toole, G.A. Rhamnolipid Surfactant Production Affects Biofilm Architecture in Pseudomonas aeruginosa PAO1. J. Bacteriol. 2003, 185, 1027–1036. [Google Scholar] [CrossRef] [PubMed]
- Rickard, A.H.; Gilbert, P.; High, N.J.; Kolenbrander, P.E.; Handley, P.S. Bacterial coaggregation: An integral process in the development of multi-species biofilms. Trends Microbiol. 2003, 11, 94–100. [Google Scholar] [CrossRef]
- Chua, S.L.; Liu, Y.; Yam, J.K.H.; Chen, Y.; Vejborg, R.M.; Tan, B.G.; Kjelleberg, S.; Tolker-Nielsen, T.; Givskov, M.; Yang, L. Dispersed cells represent a distinct stage in the transition from bacterial biofilm to planktonic lifestyles. Nat. Commun. 2014, 5, 4462. [Google Scholar] [CrossRef] [PubMed]
- Wang, R. Biofilms and Meat Safety: A Mini-Review. J. Food Prot. 2019, 82, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Chitlapilly Dass, S.; Anandappa, A. Food Factory Genomics: Where Big Data Drives Quality and Food Safety. Food Prot. Trends 2017, 37, 368–374. [Google Scholar]
- Steenackers, H.; Hermans, K.; Vanderleyden, J.; De Keersmaecker, S.C.J. Salmonella biofilms: An overview on occurrence, structure, regulation and eradication. Food Res. Int. 2012, 45, 502–531. [Google Scholar] [CrossRef]
- Vivian, R.C. The Evaluation of Biofilm Formation and Sensitivity to Peracetic Acid of Salmonella spp. Isolated from Poultry Abattoir. Ph.D. Dissertation, Universidade Estadual Paulista, São Paulo, Brazil, 2014. [Google Scholar]
- Watnick, P.; Kolter, R. Minireview: Biofilm, city of microbes. J. Bacteriol. 2000, 182, 2675–2679. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef] [PubMed]
- Sofos, J.N.; Geornaras, I. Overview of current meat hygiene and safety risks and summary of recent studies on biofilms, and control of Escherichia coli O157:H7 in nonintact, and Listeria monocytogenes in ready-to-eat, meat products. Meat Sci. 2010, 86, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Silagyi, K.; Kim, S.-H.; Martin Lo, Y.; Wei, C.-I. Production of biofilm and quorum sensing by Escherichia coli O157:H7 and its transfer from contact surfaces to meat, poultry, ready-to-eat deli, and produce products. Food Microbiol. 2009, 26, 514–519. [Google Scholar] [CrossRef] [PubMed]
- Cloak Orla, M.; Solow Barbara, T.; Briggs Connie, E.; Chen, C.-Y.; Fratamico Pina, M. Quorum Sensing and Production of Autoinducer-2 in Campylobacter spp., Escherichia coli O157:H7, and Salmonella enterica Serovar Typhimurium in Foods. Appl. Environ. Microbiol. 2002, 68, 4666–4671. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Lee, Y.; Lee, S.; Kim, S.; Lee, J.; Ha, J.; Oh, H.; Shin, I.-S.; Yoon, Y. Microbial contamination including Vibrio cholerae in fishery auction markets in West Sea, South Korea. Fish. Aquat. Sci. 2019, 22, 26. [Google Scholar] [CrossRef]
- Wieczorek, K.; Bomba, A.; Osek, J. Whole-Genome Sequencing-Based Characterization of Listeria monocytogenes from Fish and Fish Production Environments in Poland. Int. J. Mol. Sci. 2020, 21, 9419. [Google Scholar] [CrossRef] [PubMed]
- Logan Savannah, L.; Thomas, J.; Yan, J.; Baker Ryan, P.; Shields Drew, S.; Xavier Joao, B.; Hammer Brian, K.; Parthasarathy, R. The Vibrio cholerae type VI secretion system can modulate host intestinal mechanics to displace gut bacterial symbionts. Proc. Natl. Acad. Sci. USA 2018, 115, E3779–E3787. [Google Scholar] [CrossRef]
- Chitlapilly Dass, S.; Abu-Ghannam, N.; Antony-Babu, S.; Cummins, E.J. Ecology and molecular typing of L. monocytogenes in a processing plant for cold-smoked salmon in the Republic of Ireland. Food Res. Int. 2010, 43, 1529–1536. [Google Scholar] [CrossRef]
- Chitlapilly Dass, S.; Cummins, E.J.; Abu-Ghannam, N. Prevalence and Typing of Listeria Monocytogenes Strains in Retail Vacuum-Packed Cold-Smoked Salmon in the Republic of Ireland. J. Food Saf. 2011, 31, 21–27. [Google Scholar] [CrossRef]
- Fox, E.M.; Solomon, K.; Moore, J.E.; Wall, P.G.; Fanning, S. Phylogenetic profiles of in-house microflora in drains at a food production facility: Comparison and biocontrol implications of Listeria-Positive and -Negative bacterial populations. Appl. Environ. Microbiol. 2014, 80, 3369–3374. [Google Scholar] [CrossRef] [PubMed]
- CDC. Multistate Outbreak of Shiga Toxin-Producing Escherichia coli O157:H7 Infections Linked to Organic Spinach and Spring Mix Blend; CDC: Atlanta, GA, USA, 2015.
- CDC. Multistate Outbreak of Shiga Toxin-Producing Escherichia coli O157 Infections Linked to Alfalfa Sprouts Produced by Jack & The Green Sprouts; CDC: Atlanta, GA, USA, 2016.
- CDC. Multistate Outbreak of Shiga Toxin-Producing Escherichia coli O157:H7 Infections Linked to Leafy Greens (Final Update); CDC: Atlanta, GA, USA, 2018.
- Bogino, P.C.; Oliva, M.D.I.M.; Sorroche, F.G.; Giordano, W. The role of bacterial biofilms and surface components in plant-bacterial associations. Int. J. Mol. Sci. 2013, 14, 15838–15859. [Google Scholar] [CrossRef]
- Truong, H.-N.; Garmyn, D.; Gal, L.; Fournier, C.; Sevellec, Y.; Jeandroz, S.; Piveteau, P. Plants as a realized niche for Listeria monocytogenes. MicrobiologyOpen 2021, 10, e1255. [Google Scholar] [CrossRef] [PubMed]
- Holden, N.J.; Wright, K.M.; Marshall, J.; Holmes, A. Functional Analysis of Shiga Toxin-Producing Escherichia coli Biofilm Components in Plant Leaves. In Shiga Toxin-Producing, E. coli: Methods and Protocols; Schüller, S., Bielaszewska, M., Eds.; Springer: New York, NY, USA, 2021; pp. 163–175. [Google Scholar]
- Sun, Y.; Ma, Y.; Guan, H.; Liang, H.; Zhao, X.; Wang, D. Adhesion mechanism and biofilm formation of Escherichia coli O157:H7 in infected cucumber (Cucumis sativus L.). Food Microbiol. 2021, 103885. [Google Scholar] [CrossRef]
- Ávila-Quezada, G.; Sánchez, E.; Gardea-Béjar, A.A.; Acedo-Félix, E. Salmonella spp. and Escherichia coli: Survival and growth in plant tissue. N. Z. J. Crop Hortic. Sci. 2010, 38, 47–55. [Google Scholar] [CrossRef]
- Esteban-Cuesta, I.; Labrador, M.; Hunt, K.; Reese, S.; Fischer, J.; Schwaiger, K.; Gareis, M. Phenotypic and Genetic Comparison of a Plant-Internalized and an Animal-Isolated Salmonella Choleraesuis Strain. Microorganisms 2021, 9, 1554. [Google Scholar] [CrossRef] [PubMed]
- Grivokostopoulos, N.C.; Makariti, I.P.; Hilaj, N.; Apostolidou, Z.; Skandamis, P.N. Internalization of Salmonella in leafy greens and the impact on acid tolerance. Appl. Environ. Microbiol. 2022, aem.02249-21. [Google Scholar] [CrossRef] [PubMed]
- Arthur, T.M.; Bosilevac, J.M.; Brichta-Harhay, D.M.; Guerini, M.N.; Kalchayanand, N.; Shackelford, S.D.; Wheeler, T.L.; Koohmaraie, M. Transportation and lairage environment effects on prevalence, numbers, and diversity of Escherichia coli O157:H7 on hides and carcasses of beef cattle at processing. J. Food Prot. 2007, 70, 280–286. [Google Scholar] [CrossRef] [PubMed]
- Arthur, T.M.; Bosilevac, J.M.; Nou, X.; Shackelford, S.D.; Wheeler, T.L.; Koohmaraie, M. Comparison of the molecular genotypes of Escherichia coli O157:H7 from the hides of beef cattle in different regions of North America. J. Food Prot. 2007, 70, 1622–1626. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rivera-Betancourt, M.; Shackelford, S.D.; Arthur, T.M.; Westmoreland, K.E.; Bellinger, G.; Rossman, M.; Reagan, J.O.; Koohmaraie, M. Prevalence of Escherichia coli O157:H7, Listeria monocytogenes, and Salmonella in two geographically distant commercial beef processing plants in the United States. J. Food Prot. 2004, 67, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Stopforth, J.D.; Samelis, J.; Sofos, J.N.; Kendall, P.; Smith, G.C. Influence of organic acid concentration on survival of Listeria monocytogenes and Escherichia coli 0157:H7 in beef carcass wash water and on model equipment surfaces. Food Microbiol. 2003, 20, 651–660. [Google Scholar] [CrossRef]
- Stopforth, J.D.; Samelis, J.; Sofos, J.N.; Kendall, P.A.; Smith, G.C. Influence of extended acid stressing in fresh beef decontamination runoff fluids on sanitizer resistance of acid-adapted Escherichia coli O157:H7 in biofilms. J. Food Prot. 2003, 66, 2258–2266. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Kalchayanand, N.; King, D.A.; Luedtke, B.E.; Bosilevac, J.M.; Arthur, T.M. Biofilm formation and sanitizer resistance of Escherichia coli O157:H7 strains isolated from “high event period” meat contamination. J. Food Prot. 2014, 77, 1982–1987. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Luedtke, B.E.; Bosilevac, J.M.; Schmidt, J.W.; Kalchayanand, N.; Arthur, T.M. Escherichia coli O157:H7 Strains Isolated from High-Event Period Beef Contamination Have Strong Biofilm-Forming Ability and Low Sanitizer Susceptibility, Which Are Associated with High pO157 Plasmid Copy Number. J. Food Prot. 2016, 79, 1875–1883. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Schmidt, J.W.; Harhay, D.M.; Bosilevac, J.M.; King, D.A.; Arthur, T.M. Biofilm Formation, Antimicrobial Resistance, and Sanitizer Tolerance of Salmonella enterica Strains Isolated from Beef Trim. Foodborne Pathog. Dis. 2017, 14, 687–695. [Google Scholar] [CrossRef] [PubMed]
- Stoodley, P.; Sauer, K.; Davies, D.G.; Costerton, J.W. Biofilms as complex differentiated communities. Annu. Rev. Microbiol. 2002, 56, 187–209. [Google Scholar] [CrossRef]
- da Silva Fernandes, M.; Coelho Alvares, A.C.; Martins Manoel, J.G.; Ramires Esper, L.M.; Kabuki, D.Y.; Kuaye, A.Y. Formation of multi-species biofilms by Enterococcus faecium, Enterococcus faecalis, and Bacillus cereus isolated from ricotta processing and effectiveness of chemical sanitation procedures. Int. Dairy J. 2017, 72, 23–28. [Google Scholar] [CrossRef]
- Wu, H.; Moser, C.; Wang, H.Z.; Hoiby, N.; Song, Z.J. Strategies for combating bacterial biofilm infections. Int. J. Oral Sci. 2015, 7, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, H.; Xing, T.; Wu, N.; Xu, X.; Zhou, G. Removal of Salmonella biofilm formed under meat processing environment by surfactant in combination with bio-enzyme. LWT Food Sci. Technol. 2016, 66, 298–304. [Google Scholar] [CrossRef]
- Turnbull, L.; Toyofuku, M.; Hynen, A.L.; Kurosawa, M.; Pessi, G.; Petty, N.K.; Osvath, S.R.; Carcamo-Oyarce, G.; Gloag, E.S.; Shimoni, R.; et al. Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat. Commun. 2016, 7, 11220. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.Y.-E.; Chew, S.C.; Tan, S.Y.-Y.; Givskov, M.; Yang, L. Emerging frontiers in detection and control of bacterial biofilms. Curr. Opin. Biotechnol. 2014, 26, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Madsen, J.S.; Sorensen, S.J.; Burmolle, M. High prevalence of biofilm synergy among bacterial soil isolates in cocultures indicates bacterial interspecific cooperation. ISME J. 2015, 9, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Afonso, A.C.; Gomes, I.B.; Saavedra, M.J.; Giaouris, E.; Simões, L.C.; Simões, M. Bacterial coaggregation in aquatic systems. Water Res. 2021, 196, 117037. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; Chen, Y.-W.; Scheible, M.; Chang, C.; Wittchen, M.; Lee, J.H.; Luong, T.T.; Tiner, B.L.; Tauch, A.; Das, A.; et al. Genetic and molecular determinants of polymicrobial interactions in Fusobacterium nucleatum. Proc. Natl. Acad. Sci. USA 2021, 118, e2006482118. [Google Scholar] [CrossRef] [PubMed]
- Ledder, R.G.; Timperley, A.S.; Friswell, M.K.; Macfarlane, S.; McBain, A.J. Coaggregation between and among human intestinal and oral bacteria. FEMS Microbiol. Ecol. 2008, 66, 630–636. [Google Scholar] [CrossRef]
- Taga, E.M.; Bassler, B.L. Chemical communication among bacteria. Proc. Natl. Acad. Sci. USA 2003, 100, 14549–14554. [Google Scholar] [CrossRef]
- Kuhl, M.; Rickelt, L.F.; Thar, R. Combined imaging of bacteria and oxygen in biofilms. Appl. Environ. Microbiol. 2007, 73, 6289–6295. [Google Scholar] [CrossRef] [PubMed]
- Karampatzakis, A.; Sankaran, J.; Kandaswamy, K.; Rice, S.A.; Cohen, Y.; Wohland, T. Measurement of oxygen concentrations in bacterial biofilms using transient state monitoring by single plane illumination microscopy. Biomed. Phys. Eng. Express 2017, 3, 035020. [Google Scholar] [CrossRef]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Kurniawan, A.; Yamamoto, T. Accumulation of NH4+ and NO3− inside Biofilms of Natural Microbial Consortia: Implication on Nutrients Seasonal Dynamic in Aquatic Ecosystems. Int. J. Microbiol. 2019, 2019, 6473690. [Google Scholar] [CrossRef] [PubMed]
- Fuqua, C.; Parsek, M.R.; Greenberg, E.P. Regulation of gene expression in cell to cell communication. Annu. Rev. Genet. 2001, 35, 439–468. [Google Scholar] [CrossRef] [PubMed]
- Sztajer, H.; Szafranski, S.P.; Tomasch, J.; Reck, M.; Nimtz, M.; Rohde, M.; Wagner-Dobler, I. Cross-feeding and interkingdom communication in dual-species biofilms of Streptococcus mutans and Candida albicans. ISME J. 2014, 8, 2256–2271. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.H.; Lee, K.W.K.; Burmolle, M.; Kjelleberg, S.; Rice, S.A. All together now: Experimental multispecies biofilm model systems. Environ. Microbiol. 2017, 19, 42–53. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, E.P. Bacterial communication and group behavior. J. Clin. Investig. 2003, 112, 1288–1290. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, E.P. Bacterial communication: Tiny teamwork. Nature 2003, 424, 134. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588. [Google Scholar] [CrossRef] [PubMed]
- Joint, I.; Tait, K.; Callow, M.E.; Callow, J.A.; Milton, D.; Williams, P.; Camara, M. Cell-to-cell communication across the prokaryote-eukaryote boundary. Science 2002, 298, 1207. [Google Scholar] [CrossRef] [PubMed]
- Brenner, K.; Karig, D.K.; Weiss, R.; Arnold, F.H. Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc. Natl. Acad. Sci. USA 2007, 104, 17300–17304. [Google Scholar] [CrossRef] [PubMed]
- Barathi, S.; Aruljothi, K.N.; Karthik, C.; Padikasan, I.A.; Ashokkumar, V. Biofilm mediated decolorization and degradation of reactive red 170 dye by the bacterial consortium isolated from the dyeing industry wastewater sediments. Chemosphere 2022, 286, 131914. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.W.K.; Periasamy, S.; Mukherjee, M.; Xie, C.; Kjelleberg, S.; Rice, S.A. Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J. 2014, 8, 894–907. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef]
- Daniels, R.; Vanderleyden, J.; Michiels, J. Quorum sensing and swarming migration in bacteria. FEMS Microbiol. Rev. 2004, 28, 261–289. [Google Scholar] [CrossRef] [PubMed]
- Morris, J.J.; Lenski, R.E.; Zinser, E.R. The Black Queen Hypothesis: Evolution of dependencies through adaptive gene loss. mBio 2012, 3, e00036-12. [Google Scholar] [CrossRef] [PubMed]
- McMurry, L.M.; Oethinger, M.; Levy, S.B. Triclosan targets lipid synthesis. Nature 1998, 394, 531–532. [Google Scholar] [CrossRef] [PubMed]
- Farr, S.B.; Kogoma, T. Oxidative stress responses in Escherichia coli and Salmonella typhimurium. Microbiol. Rev. 1991, 55, 561–585. [Google Scholar] [CrossRef] [PubMed]
- Seymour, R.L.; Mishra, P.V.; Khan, M.A.; Spector, M.P. Essential roles of core starvation-stress response loci in carbon-starvation-inducible cross-resistance and hydrogen peroxide-inducible adaptive resistance to oxidative challenge in Salmonella typhimurium. Mol. Microbiol. 1996, 20, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Condell, O.; Iversen, C.; Cooney, S.; Power, K.A.; Walsh, C.; Burgess, C.; Fanning, S. Efficacy of biocides used in the modern food industry to control Salmonella enterica, and links between biocide tolerance and resistance to clinically relevant antimicrobial compounds. Appl. Environ. Microbiol. 2012, 78, 3087–3097. [Google Scholar] [CrossRef] [PubMed]
- Karatzas, K.A.G.; Webber, M.A.; Jorgensen, F.; Woodward, M.J.; Piddock, L.J.V.; Humphrey, T.J. Prolonged treatment of Salmonella enterica serovar Typhimurium with commercial disinfectants selects for multiple antibiotic resistance, increased efflux and reduced invasiveness. J. Antimicrob. Chemother. 2007, 60, 947–955. [Google Scholar] [CrossRef] [PubMed]
- White, D.G.; Goldman, J.D.; Demple, B.; Levy, S.B. Role of the acrAB locus in organic solvent tolerance mediated by expression of marA, soxS, or robA in Escherichia coli. J. Bacteriol. 1997, 179, 6122–6126. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.Y.; Sheng, H.; Seo, K.S.; Park, Y.H.; Hovde, C.J. Characterization of an Escherichia coli O157:H7 plasmid O157 deletion mutant and its survival and persistence in cattle. Appl. Environ. Microbiol. 2007, 73, 2037–2047. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.Y.; La, H.J.; Sheng, H.; Forney, L.J.; Hovde, C.J. Influence of Plasmid pO157 on Escherichia coli O157:H7 Sakai Biofilm Formation. Appl. Environ. Microbiol. 2010, 76, 963–966. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.Y.; Hong, J.B.; Sheng, H.; Shringi, S.; Kaul, R.; Besser, T.E.; Hovde, C.J. Phenotypic diversity of Escherichia coli O157:H7 strains associated with the plasmid O157. J. Microbiol. 2010, 48, 347–357. [Google Scholar] [CrossRef]
- Soumet, C.; Fourreau, E.; Legrandois, P.; Maris, P. Resistance to phenicol compounds following adaptation to quaternary ammonium compounds in Escherichia coli. Vet. Microbiol. 2012, 158, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Braoudaki, M.; Hilton, A.C. Adaptive resistance to biocides in Salmonella enterica and Escherichia coli O157 and cross-resistance to antimicrobial agents. J. Clin. Microbiol. 2004, 42, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Futoma-Kołoch, B.; Książczyk, M.; Korzekwa, K.; Migdał, I.; Pawlak, A.; Jankowska, M.; Kędziora, A.; Dorotkiewicz-Jach, A.; Bugla-Płoskońska, G. Selection and electrophoretic characterization of Salmonella enterica subsp. enterica biocide variants resistant to antibiotics. Pol. J. Vet. Sci. 2015, 18, 725–732. [Google Scholar] [CrossRef] [PubMed]
- Karatzas, K.A.G.; Randall, L.P.; Webber, M.; Piddock, L.J.V.; Humphrey, T.J.; Woodward, M.J.; Coldham, N.G. Phenotypic and Proteomic Characterization of Multiply Antibiotic-Resistant Variants of Salmonella enterica Serovar Typhimurium Selected Following Exposure to Disinfectants. Appl. Environ. Microbiol. 2008, 74, 1508–1516. [Google Scholar] [CrossRef] [PubMed]
- Molin, S.; Tolker-Nielsen, T. Gene transfer occurs with enhanced efficiency in biofilms and induces enhanced stabilisation of the biofilm structure. Curr. Opin. Biotechnol. 2003, 14, 255–261. [Google Scholar] [CrossRef]
- Savage, V.J.; Chopra, I.; O’Neill, A.J. Staphylococcus aureus biofilms promote horizontal transfer of antibiotic resistance. Antimicrob. Agents Chemother. 2013, 57, 1968–1970. [Google Scholar] [CrossRef] [PubMed]
- Licht, T.R.; Christensen, B.B.; Krogfelt, K.A.; Molin, S. Plasmid transfer in the animal intestine and other dynamic bacterial populations: The role of community structure and environment. Microbiology 1999, 145, 2615–2622. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, L.J.; Bouwer, E.J. Rp4 plasmid transfer among species of pseudomonas in a biofilm reactor. Water Sci. Technol. 1999, 39, 163–171. [Google Scholar] [CrossRef]
- Tsen, S.D.; Fang, S.S.; Chen, M.J.; Chien, J.Y.; Lee, C.C.; Tsen, D.H.L. Natural plasmid transformation in Escherichia coli. J. Biomed. Sci. 2002, 9, 246–252. [Google Scholar] [CrossRef] [PubMed]
- Jeanson, S.; Floury, J.; Gagnaire, V.; Lortal, S.; Thierry, A. Bacterial Colonies in Solid Media and Foods: A Review on Their Growth and Interactions with the Micro-Environment. Front. Microbiol. 2015, 6, 1284. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Ma, Q.; Wood, T.K. The R1 Conjugative Plasmid Increases Escherichia coli Biofilm Formation through an Envelope Stress Response. Appl. Environ. Microbiol. 2008, 74, 2690–2699. [Google Scholar] [CrossRef] [PubMed]
- Ong, C.-L.Y.; Beatson, S.A.; McEwan, A.G.; Schembri, M.A. Conjugative Plasmid Transfer and Adhesion Dynamics in an Escherichia coli Biofilm. Appl. Environ. Microbiol. 2009, 75, 6783–6791. [Google Scholar] [CrossRef] [PubMed]
- Burmølle, M.; Bahl, M.I.; Jensen, L.B.; Sørensen, S.J.; Hansen, L.H. Type 3 fimbriae, encoded by the conjugative plasmid pOLA52, enhance biofilm formation and transfer frequencies in Enterobacteriaceae strains. Microbiology 2008, 154, 187–195. [Google Scholar] [CrossRef]
- Teodósio, J.S.; Simões, M.; Mergulhão, F.J. The influence of nonconjugative Escherichia coli plasmids on biofilm formation and resistance. J. Appl. Microbiol. 2012, 113, 373–382. [Google Scholar] [CrossRef] [PubMed]
- May, T.; Ito, A.; Okabe, S. Induction of multidrug resistance mechanism in Escherichia coli biofilms by interplay between tetracycline and ampicillin resistance genes. Antimicrob. Agents Chemother. 2009, 53, 4628–4639. [Google Scholar] [CrossRef] [PubMed]
- Cook, L.; Chatterjee, A.; Barnes, A.; Yarwood, J.; Hu, W.-S.; Dunny, G. Biofilm growth alters regulation of conjugation by a bacterial pheromone. Mol. Microbiol. 2011, 81, 1499–1510. [Google Scholar] [CrossRef] [PubMed]
- Król, J.E.; Nguyen, H.D.; Rogers, L.M.; Beyenal, H.; Krone, S.M.; Top, E.M. Increased transfer of a multidrug resistance plasmid in Escherichia coli biofilms at the air-liquid interface. Appl. Environ. Microbiol. 2011, 77, 5079–5088. [Google Scholar] [CrossRef]
- Mo, S.S.; Sunde, M.; Ilag, H.K.; Langsrud, S.; Heir, E. Transfer Potential of Plasmids Conferring Extended-Spectrum-Cephalosporin Resistance in Escherichia coli from Poultry. Appl. Environ. Microbiol. 2017, 83, e00654-17. [Google Scholar] [CrossRef] [PubMed]
- Maheshwari, M.; Ahmad, I.; Althubiani, A.S. Multidrug resistance and transferability of blaCTX-M among extended-spectrum β-lactamase-producing enteric bacteria in biofilm. J. Glob. Antimicrob. Resist. 2016, 6, 142–149. [Google Scholar] [CrossRef]
- Tanner, W.D.; Atkinson, R.M.; Goel, R.K.; Toleman, M.A.; Benson, L.S.; Porucznik, C.A.; VanDerslice, J.A. Horizontal transfer of the blaNDM-1 gene to Pseudomonas aeruginosa and Acinetobacter baumannii in biofilms. FEMS Microbiol. Lett. 2017, 364, fnx048. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.W.; Murray, S.A.; Dickey, A.M.; Wheeler, T.L.; Harhay, D.M.; Arthur, T.M. Twenty-Four-Month Longitudinal Study Suggests Little to No Horizontal Gene Transfer In Situ between Third-Generation Cephalosporin-Resistant Salmonella and Third-Generation Cephalosporin-Resistant Escherichia coli in a Beef Cattle Feedyard. J. Food Prot. 2022, 85, 323–335. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chitlapilly Dass, S.; Wang, R. Biofilm through the Looking Glass: A Microbial Food Safety Perspective. Pathogens 2022, 11, 346. https://doi.org/10.3390/pathogens11030346
Chitlapilly Dass S, Wang R. Biofilm through the Looking Glass: A Microbial Food Safety Perspective. Pathogens. 2022; 11(3):346. https://doi.org/10.3390/pathogens11030346
Chicago/Turabian StyleChitlapilly Dass, Sapna, and Rong Wang. 2022. "Biofilm through the Looking Glass: A Microbial Food Safety Perspective" Pathogens 11, no. 3: 346. https://doi.org/10.3390/pathogens11030346
APA StyleChitlapilly Dass, S., & Wang, R. (2022). Biofilm through the Looking Glass: A Microbial Food Safety Perspective. Pathogens, 11(3), 346. https://doi.org/10.3390/pathogens11030346




