Structural Analysis of the Outer Membrane Lipoprotein BBA14 (OrfD) and the Corresponding Paralogous Gene Family 143 (PFam143) from Borrelia burgdorferi
Abstract
1. Introduction
2. Results and Discussion
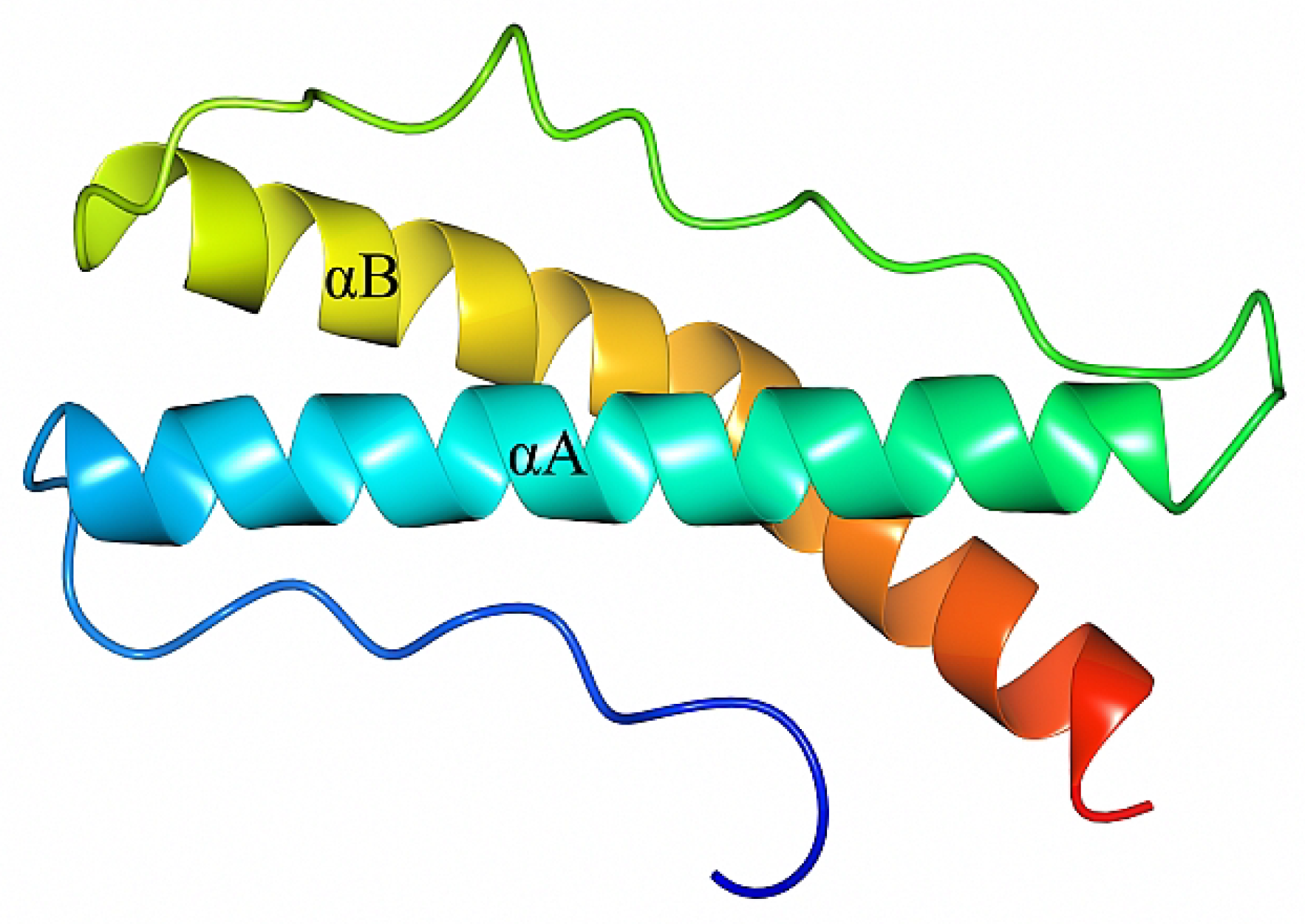
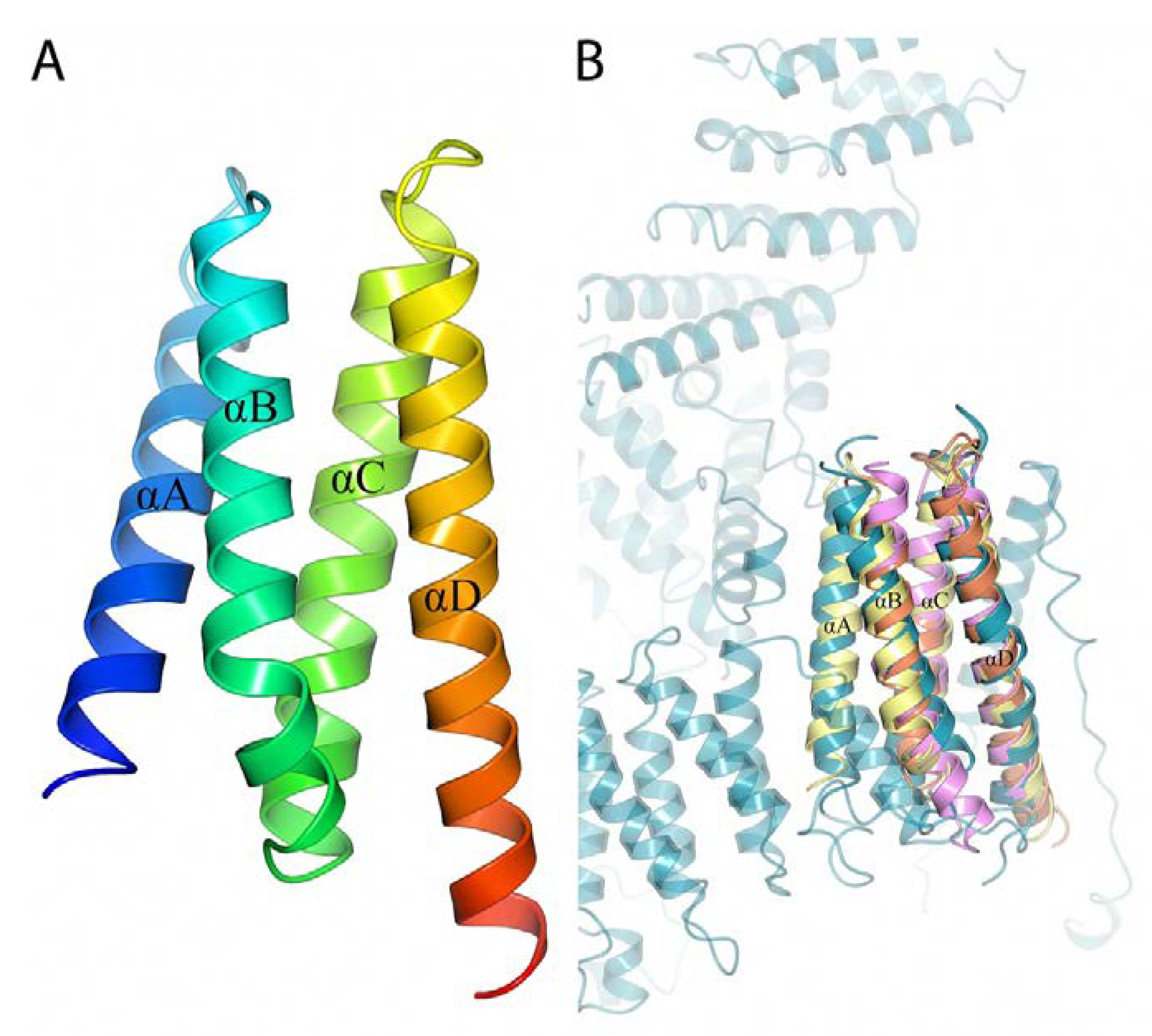
2.1. Crystal Structure of B. burgdorferi BBA14
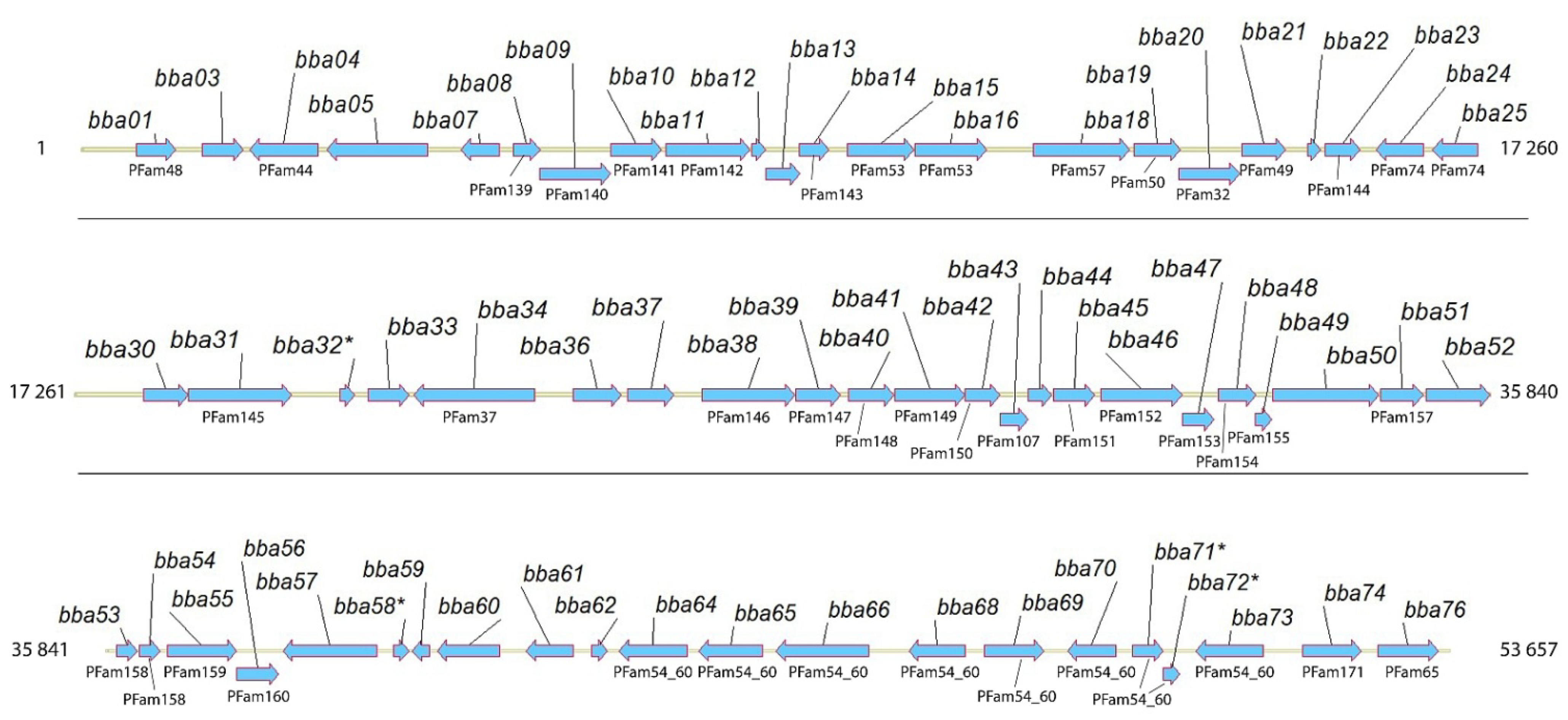
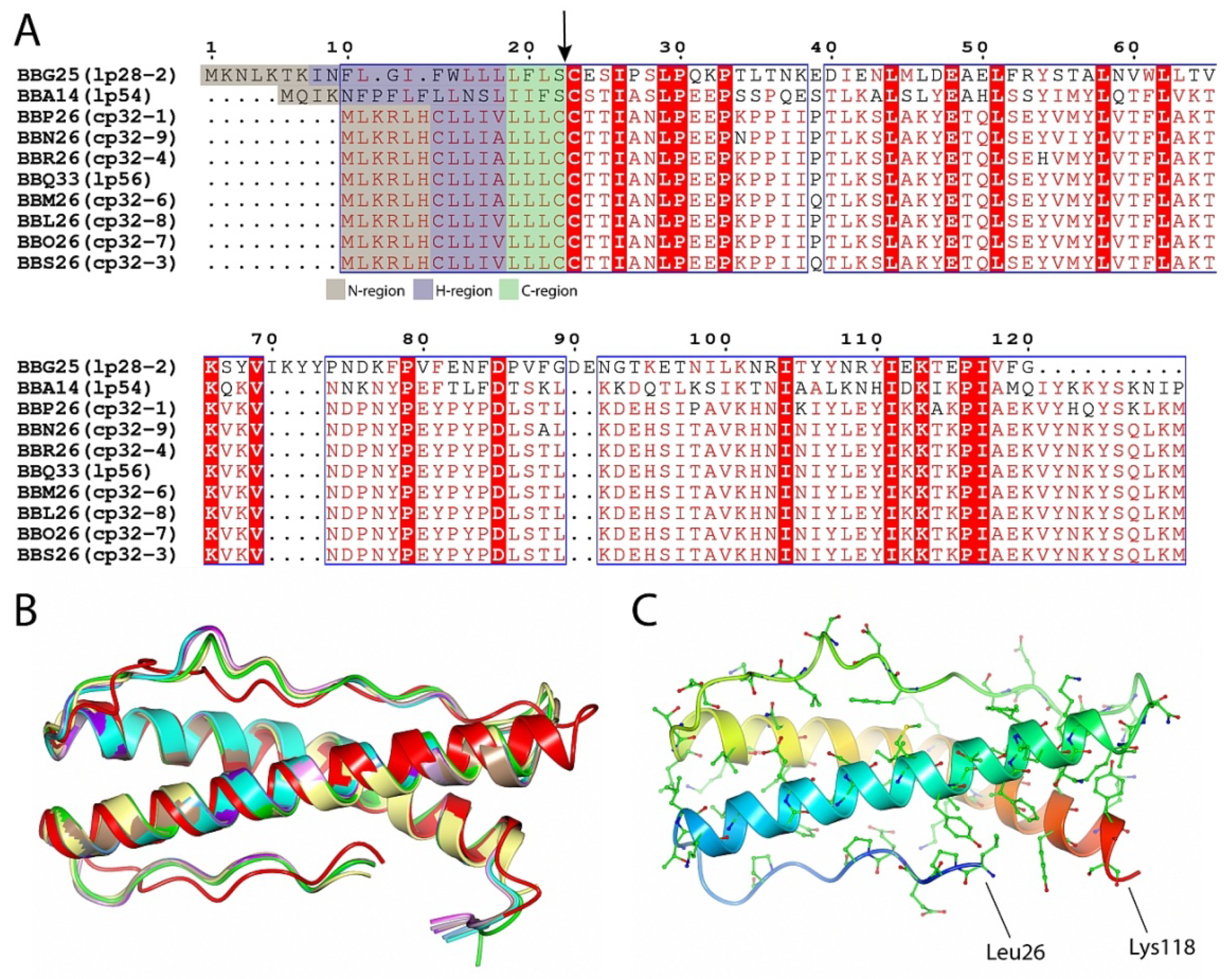
2.2. BBA14 as a Member of the Paralogous Gene Family 143
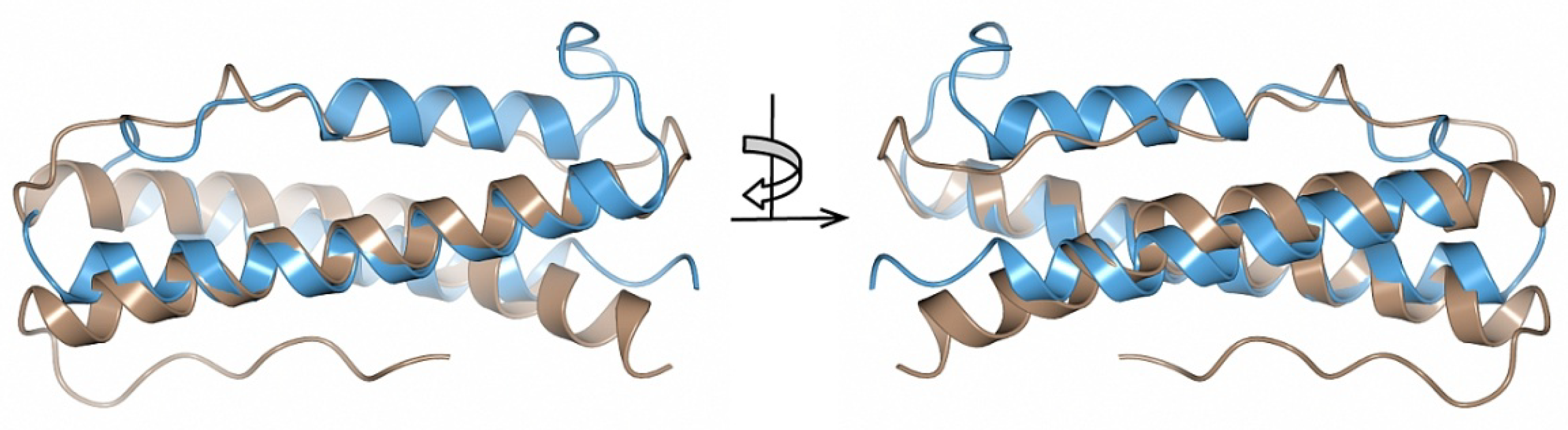
2.3. Structural Similarity
3. Materials and Methods
3.1. Cloning and Expression of BBA14
3.2. Purification of Recombinant BBA14
3.3. Crystallization of BBA14
3.4. Data Collection and Structure Determination
3.5. Protein Structure Prediction
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rudenko, N.; Golovchenko, M.; Grubhoffer, L.; Oliver, J.H., Jr. Updates on Borrelia burgdorferi sensu lato complex with respect to public health. Ticks Tick Borne Dis. 2011, 2, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.M.; Wilske, B.; Fingerle, V.; Lobet, Y.; Gern, L. Transmission of Borrelia garinii OspA serotype 4 to BALB/c mice by Ixodes ricinus ticks collected in the field. J. Clin. Microbiol. 2001, 39, 1169–1171. [Google Scholar] [CrossRef] [PubMed]
- Margos, G.; Wilske, B.; Sing, A.; Hizo-Teufel, C.; Cao, W.C.; Chu, C.; Scholz, H.; Straubinger, R.K.; Fingerle, V. Borrelia bavariensis sp. nov. is widely distributed in Europe and Asia. Int. J. Syst. Evol. Microbiol. 2013, 63 Pt 11, 4284–4288. [Google Scholar] [CrossRef]
- Baranton, G.; Postic, D.; Saint Girons, I.; Boerlin, P.; Piffaretti, J.C.; Assous, M.; Grimont, P.A. Delineation of Borrelia burgdorferi sensu stricto, Borrelia garinii sp. nov., and group VS461 associated with Lyme borreliosis. Int. J. Syst. Bacteriol. 1992, 42, 378–383. [Google Scholar] [CrossRef] [PubMed]
- Stanek, G.; Reiter, M. The expanding Lyme Borrelia complex--clinical significance of genomic species? Clin. Microbiol. Infect. 2011, 17, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Pritt, B.S.; Respicio-Kingry, L.B.; Sloan, L.M.; Schriefer, M.E.; Replogle, A.J.; Bjork, J.; Liu, G.; Kingry, L.C.; Mead, P.S.; Neitzel, D.F.; et al. Borrelia mayonii sp. nov., a member of the Borrelia burgdorferi sensu lato complex, detected in patients and ticks in the upper midwestern United States. Int. J. Syst. Evol. Microbiol. 2016, 66, 4878–4880. [Google Scholar] [CrossRef]
- Stanek, G.; Strle, F. Lyme borreliosis-from tick bite to diagnosis and treatment. FEMS Microbiol. Rev. 2018, 42, 233–258. [Google Scholar] [CrossRef]
- Radolf, J.D.; Caimano, M.J.; Stevenson, B.; Hu, L.T. Of ticks, mice and men: Understanding the dual-host lifestyle of Lyme disease spirochaetes. Nat. Rev. Microbiol. 2012, 10, 87–99. [Google Scholar] [CrossRef]
- Casjens, S.; Palmer, N.; van Vugt, R.; Huang, W.M.; Stevenson, B.; Rosa, P.; Lathigra, R.; Sutton, G.; Peterson, J.; Dodson, R.J.; et al. A bacterial genome in flux: The twelve linear and nine circular extrachromosomal DNAs in an infectious isolate of the Lyme disease spirochete Borrelia burgdorferi. Mol. Microbiol. 2000, 35, 490–516. [Google Scholar] [CrossRef]
- Fraser, C.M.; Casjens, S.; Huang, W.M.; Sutton, G.G.; Clayton, R.; Lathigra, R.; White, O.; Ketchum, K.A.; Dodson, R.; Hickey, E.K.; et al. Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature 1997, 390, 580–586. [Google Scholar] [CrossRef]
- Casjens, S.R.; Mongodin, E.F.; Qiu, W.G.; Luft, B.J.; Schutzer, S.E.; Gilcrease, E.B.; Huang, W.M.; Vujadinovic, M.; Aron, J.K.; Vargas, L.C.; et al. Genome stability of Lyme disease spirochetes: Comparative genomics of Borrelia burgdorferi plasmids. PLoS ONE 2012, 7, e33280. [Google Scholar] [CrossRef]
- Schwan, T.G.; Burgdorfer, W.; Garon, C.F. Changes in infectivity and plasmid profile of the Lyme disease spirochete, Borrelia burgdorferi, as a result of in vitro cultivation. Infect. Immun. 1988, 56, 1831–1836. [Google Scholar] [CrossRef]
- Xu, Y.; Kodner, C.; Coleman, L.; Johnson, R.C. Correlation of plasmids with infectivity of Borrelia burgdorferi sensu stricto type strain B31. Infect. Immun. 1996, 64, 3870–3876. [Google Scholar] [CrossRef]
- Purser, J.E.; Norris, S.J. Correlation between plasmid content and infectivity in Borrelia burgdorferi. Proc. Natl. Acad. Sci. USA 2000, 97, 13865–13870. [Google Scholar] [CrossRef]
- Labandeira-Rey, M.; Skare, J.T. Decreased infectivity in Borrelia burgdorferi strain B31 is associated with loss of linear plasmid 25 or 28-1. Infect. Immun. 2001, 69, 446–455. [Google Scholar] [CrossRef]
- Setubal, J.C.; Reis, M.; Matsunaga, J.; Haake, D.A. Lipoprotein computational prediction in spirochaetal genomes. Microbiology 2006, 152 Pt 1, 113–121. [Google Scholar] [CrossRef]
- Zuckert, W.R. Secretion of bacterial lipoproteins: Through the cytoplasmic membrane, the periplasm and beyond. Biochim. Biophys. Acta 2014, 1843, 1509–1516. [Google Scholar] [CrossRef]
- Kraiczy, P.; Hellwage, J.; Skerka, C.; Becker, H.; Kirschfink, M.; Simon, M.M.; Brade, V.; Zipfel, P.F.; Wallich, R. Complement resistance of Borrelia burgdorferi correlates with the expression of BbCRASP-1, a novel linear plasmid-encoded surface protein that interacts with human factor H and FHL-1 and is unrelated to Erp proteins. J. Biol. Chem. 2004, 279, 2421–2429. [Google Scholar] [CrossRef]
- Hartmann, K.; Corvey, C.; Skerka, C.; Kirschfink, M.; Karas, M.; Brade, V.; Miller, J.C.; Stevenson, B.; Wallich, R.; Zipfel, P.F.; et al. Functional characterization of BbCRASP-2, a distinct outer membrane protein of Borrelia burgdorferi that binds host complement regulators factor H and FHL-1. Mol. Microbiol. 2006, 61, 1220–1236. [Google Scholar] [CrossRef]
- Kraiczy, P. Hide and Seek: How Lyme Disease Spirochetes Overcome Complement Attack. Front. Immunol. 2016, 7, 385. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, Y.; Adusumilli, S.; Liu, L.; Narasimhan, S.; Dai, J.; Zhao, Y.O.; Fikrig, E. Molecular interactions that enable movement of the Lyme disease agent from the tick gut into the hemolymph. PLoS Pathog. 2011, 7, e1002079. [Google Scholar] [CrossRef]
- Gilmore, R.D., Jr.; Howison, R.R.; Dietrich, G.; Patton, T.G.; Clifton, D.R.; Carroll, J.A. The bba64 gene of Borrelia burgdorferi, the Lyme disease agent, is critical for mammalian infection via tick bite transmission. Proc. Natl. Acad. Sci. USA 2010, 107, 7515–7520. [Google Scholar] [CrossRef]
- Kumar, M.; Yang, X.; Coleman, A.S.; Pal, U. BBA52 facilitates Borrelia burgdorferi transmission from feeding ticks to murine hosts. J. Infect. Dis. 2010, 201, 1084–1095. [Google Scholar] [CrossRef]
- Zhang, X.; Yang, X.; Kumar, M.; Pal, U. BB0323 function is essential for Borrelia burgdorferi virulence and persistence through tick-rodent transmission cycle. J. Infect. Dis. 2009, 200, 1318–1330. [Google Scholar] [CrossRef]
- Yang, X.; Qin, J.; Promnares, K.; Kariu, T.; Anderson, J.F.; Pal, U. Novel microbial virulence factor triggers murine lyme arthritis. J. Infect. Dis. 2013, 207, 907–918. [Google Scholar] [CrossRef]
- Shi, Y.; Xu, Q.; McShan, K.; Liang, F.T. Both decorin-binding proteins A and B are critical for the overall virulence of Borrelia burgdorferi. Infect. Immun. 2008, 76, 1239–1246. [Google Scholar] [CrossRef]
- Kenedy, M.R.; Lenhart, T.R.; Akins, D.R. The role of Borrelia burgdorferi outer surface proteins. FEMS Immunol. Med. Microbiol. 2012, 66, 1–19. [Google Scholar] [CrossRef]
- Xu, Y.; Bruno, J.F.; Luft, B.J. Profiling the humoral immune response to Borrelia burgdorferi infection with protein microarrays. Microb. Pathog. 2008, 45, 403–407. [Google Scholar] [CrossRef]
- Brangulis, K.; Akopjana, I.; Petrovskis, I.; Kazaks, A.; Tars, K. Structural analysis of the outer surface proteins from Borrelia burgdorferi paralogous gene family 54 that are thought to be the key players in the pathogenesis of Lyme disease. J. Struct. Biol. 2020, 210, 107490. [Google Scholar] [CrossRef]
- Kaaijk, P.; Luytjes, W. Vaccination against Lyme disease: Are we ready for it? Hum. Vaccin. Immunother. 2016, 12, 757–762. [Google Scholar] [CrossRef]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Porcella, S.F.; Popova, T.G.; Akins, D.R.; Li, M.; Radolf, J.D.; Norgard, M.V. Borrelia burgdorferi supercoiled plasmids encode multicopy tandem open reading frames and a lipoprotein gene family. J. Bacteriol. 1996, 178, 3293–3307. [Google Scholar] [CrossRef][Green Version]
- Damman, C.J.; Eggers, C.H.; Samuels, D.S.; Oliver, D.B. Characterization of Borrelia burgdorferi BlyA and BlyB proteins: A prophage-encoded holin-like system. J. Bacteriol. 2000, 182, 6791–6797. [Google Scholar] [CrossRef]
- Porcella, S.F.; Fitzpatrick, C.A.; Bono, J.L. Expression and immunological analysis of the plasmid-borne mlp genes of Borrelia burgdorferi strain B31. Infect. Immun. 2000, 68, 4992–5001. [Google Scholar] [CrossRef]
- Dowdell, A.S.; Murphy, M.D.; Azodi, C.; Swanson, S.K.; Florens, L.; Chen, S.; Zuckert, W.R. Comprehensive Spatial Analysis of the Borrelia burgdorferi Lipoproteome Reveals a Compartmentalization Bias toward the Bacterial Surface. J. Bacteriol. 2017, 199, e00658. [Google Scholar] [CrossRef]
- Krissinel, E.; Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol. Crystallogr. 2004, 60 Pt 12 Pt 1, 2256–2268. [Google Scholar] [CrossRef]
- Mutschler, H.; Gebhardt, M.; Shoeman, R.L.; Meinhart, A. A novel mechanism of programmed cell death in bacteria by toxin-antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 2011, 9, e1001033. [Google Scholar] [CrossRef]
- Meinhart, A.; Alonso, J.C.; Strater, N.; Saenger, W. Crystal structure of the plasmid maintenance system epsilon/zeta: Functional mechanism of toxin zeta and inactivation by epsilon 2 zeta 2 complex formation. Proc. Natl. Acad. Sci. USA 2003, 100, 1661–1666. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Park, J.H.; Inouye, M. Toxin-antitoxin systems in bacteria and archaea. Annu. Rev. Genet. 2011, 45, 61–79. [Google Scholar] [CrossRef] [PubMed]
- Bukowski, M.; Rojowska, A.; Wladyka, B. Prokaryotic toxin-antitoxin systems--the role in bacterial physiology and application in molecular biology. Acta Biochim. Pol. 2011, 58, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Kozlov, G.; Wong, K.; Gehring, K. Crystal structure of the Legionella effector Lem22. Proteins 2018, 86, 263–267. [Google Scholar] [CrossRef] [PubMed]
- Fang, S.; Li, L.; Cui, B.; Men, S.; Shen, Y.; Yang, X. Structural insight into plant programmed cell death mediated by BAG proteins in Arabidopsis thaliana. Acta Crystallogr. D Biol. Crystallogr. 2013, 69 Pt 6, 934–945. [Google Scholar] [CrossRef]
- Ceccaldi, R.; Sarangi, P.; D’Andrea, A.D. The Fanconi anaemia pathway: New players and new functions. Nat. Rev. Mol. Cell Biol. 2016, 17, 337–349. [Google Scholar] [CrossRef]
- Wang, R.; Wang, S.; Dhar, A.; Peralta, C.; Pavletich, N.P. DNA clamp function of the monoubiquitinated Fanconi anaemia ID complex. Nature 2020, 580, 278–282. [Google Scholar] [CrossRef]
- Pandey, D.P.; Gerdes, K. Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 2005, 33, 966–976. [Google Scholar] [CrossRef]
- Bestor, A.; Stewart, P.E.; Jewett, M.W.; Sarkar, A.; Tilly, K.; Rosa, P.A. Use of the Cre-lox recombination system to investigate the lp54 gene requirement in the infectious cycle of Borrelia burgdorferi. Infect. Immun. 2010, 78, 2397–2407. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Drozdetskiy, A.; Cole, C.; Procter, J.; Barton, G.J. JPred4: A protein secondary structure prediction server. Nucleic Acids Res. 2015, 43, W389–W394. [Google Scholar] [CrossRef]
- Mueller, U.; Darowski, N.; Fuchs, M.R.; Forster, R.; Hellmig, M.; Paithankar, K.S.; Puhringer, S.; Steffien, M.; Zocher, G.; Weiss, M.S. Facilities for macromolecular crystallography at the Helmholtz-Zentrum Berlin. J. Synchrotron. Radiat. 2012, 19 Pt 3, 442–449. [Google Scholar] [CrossRef]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 2011, 67 Pt 4, 235–242. [Google Scholar] [CrossRef]
- Evans, P.R. An introduction to data reduction: Space-group determination, scaling and intensity statistics. Acta Crystallogr. D Biol. Crystallogr. 2011, 67 Pt 4, 282–292. [Google Scholar] [CrossRef]
- Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 2010, 66 Pt 2, 125–132. [Google Scholar] [CrossRef]
- Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 2008, 64 Pt 1, 112–122. [Google Scholar] [CrossRef]
- Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 2006, 62 Pt 9, 1002–1011. [Google Scholar] [CrossRef]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 2004, 60 Pt 12 Pt 1, 2126–2132. [Google Scholar] [CrossRef]
- Murshudov, G.N.; Vagin, A.A.; Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 1997, 53 Pt 3, 240–255. [Google Scholar] [CrossRef]
| Dataset | BBA14 |
|---|---|
| X-ray diffraction data | |
| PDB entry | 7QDV |
| Beamline | 14.1 BESSY II, Helmholtz-Zentrum, Berlin |
| Space group | P 41212 |
| a, b, c (Å) | 36.4, 36.4, 129.9 |
| α, β, γ (°) | 90.0, 90.0, 90.0 |
| Wavelength (Å) | 0.97961 |
| Resolution (Å) | 35.08–1.90 |
| Highest resolution bin (Å) | 1.95–1.90 |
| No. of reflections | 182,843 |
| No. of unique reflections | 7534 |
| Completeness (%) | 99.8 (100.0) |
| Rmerge | 0.06 (0.38) |
| CC1/2 | 1.00 (0.99) |
| I/σ (I) | 32.8 (9.3) |
| Multiplicity | 24.3 (25.7) |
| Refinement | |
| Rwork | 0.189 (0.234) |
| Rfree | 0.245 (0.292) |
| Average B-factor (Å2) | |
| Overall | 43.0 |
| From Wilson plot | 27.9 |
| No. of atoms | |
| Protein | 774 |
| Water | 36 |
| RMS deviations from ideal | |
| Bond lengths (Å) | 0.009 |
| Bond angles (o) | 1.565 |
| Ramachandran outliers (%) | |
| Residues in most favored regions (%) | 97.87 |
| Residues in allowed regions (%) | 2.13 |
| Outliers (%) | 0.00 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akopjana, I.; Brangulis, K. Structural Analysis of the Outer Membrane Lipoprotein BBA14 (OrfD) and the Corresponding Paralogous Gene Family 143 (PFam143) from Borrelia burgdorferi. Pathogens 2022, 11, 154. https://doi.org/10.3390/pathogens11020154
Akopjana I, Brangulis K. Structural Analysis of the Outer Membrane Lipoprotein BBA14 (OrfD) and the Corresponding Paralogous Gene Family 143 (PFam143) from Borrelia burgdorferi. Pathogens. 2022; 11(2):154. https://doi.org/10.3390/pathogens11020154
Chicago/Turabian StyleAkopjana, Inara, and Kalvis Brangulis. 2022. "Structural Analysis of the Outer Membrane Lipoprotein BBA14 (OrfD) and the Corresponding Paralogous Gene Family 143 (PFam143) from Borrelia burgdorferi" Pathogens 11, no. 2: 154. https://doi.org/10.3390/pathogens11020154
APA StyleAkopjana, I., & Brangulis, K. (2022). Structural Analysis of the Outer Membrane Lipoprotein BBA14 (OrfD) and the Corresponding Paralogous Gene Family 143 (PFam143) from Borrelia burgdorferi. Pathogens, 11(2), 154. https://doi.org/10.3390/pathogens11020154





