Infection with a Recently Discovered Gammaherpesvirus Variant in European Badgers, Meles meles, is Associated with Higher Relative Viral Loads in Blood
Abstract
1. Introduction
- (1)
- acquire longer sequences of the MusGHV-1 DNA polymerase gene to investigate for additional mutations/genetic differences between the two variants;
- (2)
- establish the relative prevalence of the two MusGHV-1 variants within the population, investigate the rate of coinfection, and whether potential differences in variant prevalence are associated with demographic (i.e., host sex and age) and/or socio-geographic (i.e., badger sett and social group) patterns;
- (3)
- analyse potential differences in viral loads in blood samples between individuals infected with each or both MusGHV-1 variants.
2. Materials and Methods
2.1. Animal Sampling
2.2. Genotyping of MusGHV-1 DNA
2.3. MusGHV-1 DNA Polymerase Gene Mapping and Phylogenic Analysis
2.4. Geographic Distribution of MusGHV-1 Strains and Social Group-Specific Prevalence
2.5. Blood Viral Load Quantification
2.6. Statistical Analyses
3. Results
3.1. Substitutions in the DNApol Gene of the Two MusGHV-1 Variants
3.2. Phylogenetic Relationships between the Two MusGHV-1 Variants and Other Gammaherpesvirus Species
3.3. Novel MusGHV-1 Variant Infections Were Clustered within Just Three Social Groups
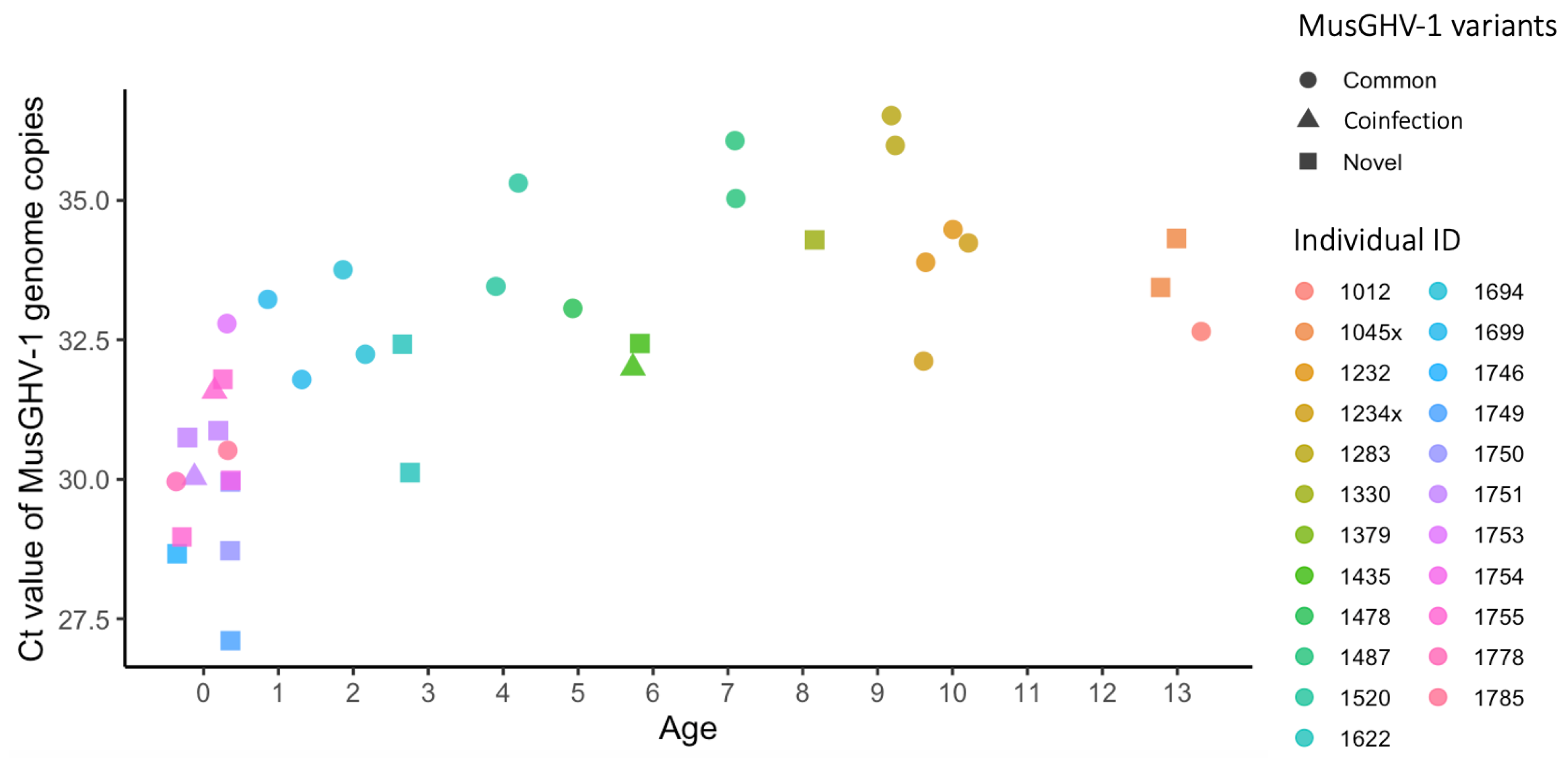
3.4. The MusGHV-1 Novel Variant Was Associated with Higher Viral Load in the Blood Stream
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Xiao, X.; Newman, C.; Buesching, C.D.; Macdonald, D.W.; Zhou, Z.M. Animal sales from Wuhan wet markets immediately prior to the COVID-19 pandemic. Sci. Rep. 2021, 11, 11898. [Google Scholar] [CrossRef] [PubMed]
- Worobey, M.; Levy, J.I.; Serrano, L.M.; Crits-Christoph, A.; Pekar, J.E.; Goldstein, S.A.; Rasmussen, A.L.; Kraemer, M.U.G.; Newman, C.; Koopmans, M.P.G.; et al. The Huanan Seafood Wholesale Market in Wuhan was the early epicenter of the COVID-19 pandemic. Science 2022, 959, abp8715. [Google Scholar] [CrossRef] [PubMed]
- Biek, R.; Pybus, O.G.; Lloyd-Smith, J.O.; Didelot, X. Measurably evolving pathogens in the genomic era. Trends Ecol. Evol. 2015, 30, 306–313. [Google Scholar] [CrossRef] [PubMed]
- Duffy, S.; Shackelton, L.A.; Holmes, E.C. Rates of evolutionary change in viruses: Patterns and determinants. Nat. Rev. Genet. 2008, 9, 267–276. [Google Scholar] [CrossRef] [PubMed]
- Scheinberg, L. The Red Queen. Arch. Neurol. 1983, 40, 189. [Google Scholar] [CrossRef]
- Carius, H.J.; Little, T.J.; Ebert, D. Genetic variation in a host-parasite association: Potential for coevolution and frequency-dependent selection. Evolution 2001, 55, 1136–1145. [Google Scholar] [CrossRef]
- Lively, C.M.; Dybdahl, M.F. Parasite adaptation to locally common host genotypes. Nature 2000, 405, 679–681. [Google Scholar] [CrossRef]
- Salathé, M.; Kouyos, R.D.; Bonhoeffer, S. The state of affairs in the kingdom of the Red Queen. Trends Ecol. Evol. 2008, 23, 439–445. [Google Scholar] [CrossRef]
- Escalera-Zamudio, M.; Rojas-Anaya, E.; Kolokotronis, S.-O.; Taboada, B.; Loza-Rubio, E.; Méndez-Ojeda, M.L.; Arias, C.F.; Osterrieder, N.; Greenwood, A.D. Bats, Primates, and the Evolutionary Origins and Diversification of Mammalian Gammaherpesviruses. mBio 2016, 7, e01425-16. [Google Scholar] [CrossRef]
- Davison, A.J. Evolution of the herpesviruses. Vet. Microbiol. 2002, 86, 69–88. [Google Scholar] [CrossRef]
- Brito, A.F.; Baele, G.; Nahata, K.D.; Grubaugh, N.D.; Pinney, J.W. Intrahost speciations and host switches played an important role in the evolution of herpesviruses. Virus Evol. 2021, 7, veab025. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, P.J.; Pope, L.C.; Greig, C.; Dawson, D.A.; Rogers, L.M.; Erven, K.; Wilson, G.J.; Delahay, R.J.; Cheeseman, C.L.; Burke, T. Mating system of the Eurasian badger, Meles meles, in a high density population. Mol. Ecol. 2005, 14, 273–284. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.D.P.; Macdonald, D.W.; Dickman, A.J. An analysis and review of models of the sociobiology of the Mustelidae. Mamm. Rev. 2000, 30, 171–196. [Google Scholar] [CrossRef]
- Woodroffe, R.; Macdonald, D.W. Badger sociality—Models of spatial grouping. Mamm. Predators 1993, 65, 145–169. [Google Scholar]
- Macdonald, D.W.; Newman, C. Population dynamics of badgers (Meles meles) in Oxfordshire, UK: Numbers, density and cohort life histories, and a possible role of climate change in population growth. J. Zool. 2002, 256, 121–138. [Google Scholar] [CrossRef]
- Banks, M.; King, D.P.; Daniells, C.; Stagg, D.A.; Gavier-Widen, D. Partial characterization of a novel gammaherpesvirus isolated from a European badger (Meles meles). J. Gen. Virol. 2002, 83, 1325–1330. [Google Scholar] [CrossRef]
- Barton, E.; Mandal, P.; Speck, S.H. Pathogenesis and Host Control of Gammaherpesviruses: Lessons from the Mouse. Annu. Rev. Immunol. 2011, 29, 351–397. [Google Scholar] [CrossRef]
- Sin, Y.W.; Annavi, G.; Dugdale, H.L.; Newman, C.; Burke, T.; MacDonald, D.W. Pathogen burden, co-infection and major histocompatibility complex variability in the European badger (Meles meles). Mol. Ecol. 2014, 23, 5072–5088. [Google Scholar] [CrossRef]
- King, D.P.; Mutukwa, N.; Lesellier, S.; Cheeseman, C.; Chambers, M.A.; Banks, M. Detection of Mustelid Herpesvirus-1 Infected European Badgers (Meles meles) in the British Isles. J. Wildl. Dis. 2004, 40, 99–102. [Google Scholar] [CrossRef]
- Tsai, M.-S.; François, S.; Newman, C.; Macdonald, D.W.; Buesching, C.D. Patterns of genital tract mustelid gammaherpesvirus 1 (Musghv-1) reactivation are linked to stressors in european badgers (Meles meles). Biomolecules 2021, 11, 716. [Google Scholar] [CrossRef]
- Tsai, M.; Fogarty, U.; Byrne, A.W.; O’Keeffe, J.; Newman, C.; Macdonald, D.W.; Buesching, C.D. Effects of Mustelid gammaherpesvirus 1 (MusGHV-1) Reactivation in European Badger (Meles meles) Genital Tracts on Reproductive Fitness. Pathogens 2020, 9, 769. [Google Scholar] [CrossRef] [PubMed]
- Kent, A.; Ehlers, B.; Mendum, T.; Newman, C.; Macdonald, D.W.; Chambers, M.; Buesching, C.D. Genital tract screening finds widespread infection with mustelid gammaherpesvirus 1 in the European badger (Meles meles). J. Wildl. Dis. 2018, 54, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Buesching, C.D.; Clarke, J.R.; Ellwood, S.A.; King, C.; Newman, C.; Macdonald, D.W. The Mammals of Wytham Woods. In Wytham Woods: Oxford’s Ecological Laboratory; Oxford University Press: Oxford, UK, 2011; ISBN 9780191810039. [Google Scholar]
- Dourmishev, L.A.; Dourmishev, A.L.; Palmeri, D.; Schwartz, R.A.; Lukac, D.M. Molecular Genetics of Kaposi’s Sarcoma-Associated Herpesvirus (Human Herpesvirus 8) Epidemiology and Pathogenesis. Microbiol. Mol. Biol. Rev. 2003, 67, 175–212. [Google Scholar] [CrossRef] [PubMed]
- Trimpert, J.; Osterrieder, N. Herpesvirus DNA Polymerase Mutants—How Important Is Faithful Genome Replication? Curr. Clin. Microbiol. Rep. 2019, 6, 240–248. [Google Scholar] [CrossRef]
- Tian, W.; Hwang, Y.T.; Lu, Q.; Hwang, C.B.C. Finger Domain Mutation Affects Enzyme Activity, DNA Replication Efficiency, and Fidelity of an Exonuclease-Deficient DNA Polymerase of Herpes Simplex Virus Type 1. J. Virol. 2009, 83, 7194–7201. [Google Scholar] [CrossRef] [PubMed]
- Lunn, D.P.; Davis-Poynter, N.; Flaminio, M.J.B.F.; Horohov, D.W.; Osterrieder, K.; Pusterla, N.; Townsend, H.G.G. Equine herpesvirus-1 consensus statement. J. Vet. Intern. Med. 2009, 23, 450–461. [Google Scholar] [CrossRef]
- Nugent, J.; Smith, K.C.; Mumford, J.A.; Newton, J.R.; Bowden, R.J.; Allen, G.P. Analysis of Equid Herpesvirus 1 Strain Variation Reveals a Point Mutation of the DNA Polymerase Strongly Associated with Neuropathogenic versus Nonneuropathogenic Disease Outbreaks. J. Virol. 2006, 80, 4047–4060. [Google Scholar] [CrossRef]
- Stasiak, K.; Rola, J.; Ploszay, G.; Socha, W.; Zmudzinski, J.F. Detection of the neuropathogenic variant of equine herpesvirus 1 associated with abortions in mares in Poland. BMC Vet. Res. 2015, 11, 102. [Google Scholar] [CrossRef]
- Goodman, L.B.; Loregian, A.; Perkins, G.A.; Nugent, J.; Buckles, E.L.; Mercorelli, B.; Kydd, J.H.; Palù, G.; Smith, K.C.; Osterrieder, N.; et al. A point mutation in a herpesvirus polymerase determines neuropathogenicity. PLoS Pathog. 2007, 3, 1583–1592. [Google Scholar] [CrossRef]
- Pusterla, N.; Wilson, W.D.; Mapes, S.; Finno, C.; Isbell, D.; Arthur, R.M.; Ferraro, G.L. Characterization of viral loads, strain and state of equine herpesvirus-1 using real-time PCR in horses following natural exposure at a racetrack in California. Vet. J. 2009, 179, 230–239. [Google Scholar] [CrossRef]
- Pusterla, N.; Mapes, S.; Wilson, W.D. Prevalence of equine herpesvirus type 1 in trigeminal ganglia and submandibular lymph nodes of equids examined postmortem. Vet. Rec. 2010, 167, 376–379. [Google Scholar] [CrossRef] [PubMed]
- Savill, P.; Perrins, C.; Kirby, K.; Fisher, N. Wytham Woods: Oxford’s Ecological Laboratory; Oxford University Press: Oxford, UK, 2010; Volume 48. [Google Scholar]
- MacDonald, D.W.; Newman, C.; Nouvellet, P.M.; Buesching, C.D. An analysis of Eurasian badger (Meles meles) population dynamics: Implications for regulatory mechanisms. J. Mammal. 2009, 90, 1392–1403. [Google Scholar] [CrossRef]
- Macdonald, D.W.; Newman, C.; Buesching, C.D. Badgers in the rural landscape—conservation paragon or farmland pariah? Lessons from the Wytham Badger Project. Wildl. Conserv. Farml. 2015, 2, 65–95. [Google Scholar] [CrossRef]
- Sugianto, N.A.; Buesching, C.D.; Macdonald, D.W.; Newman, C. The Importance of Refining Anaesthetic Regimes to Mitigate Adverse Effects in Very Young and Very Old Wild Animals: The European Badger (Meles meles). J. Zool. Res. 2019, 3, 10–17. [Google Scholar]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Okonechnikov, K.; Golosova, O.; Fursov, M.; Varlamov, A.; Vaskin, Y.; Efremov, I.; German Grehov, O.G.; Kandrov, D.; Rasputin, K.; Syabro, M.; et al. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef]
- Nei, M.; Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 1986, 3, 418–426. [Google Scholar] [CrossRef]
- Miyata, T.; Katoh, K.; Misawa, K.; Kuma, K. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar]
- Sugianto, N.A.; Newman, C.; Macdonald, D.W.; Buesching, C.D. Heterochrony of puberty in the European badger (Meles meles) can be explained by growth rate and group-size: Evidence for two endocrinological phenotypes. PLoS ONE 2019, 14, e0203910. [Google Scholar] [CrossRef]
- Annavi, G.; Newman, C.; Dugdale, H.L.; Buesching, C.D.; Sin, Y.W.; Burke, T.; Macdonald, D.W. Neighbouring-group composition and within-group relatedness drive extra-group paternity rate in the European badger (Meles meles). J. Evol. Biol. 2014, 27, 2191–2203. [Google Scholar] [CrossRef]
- Hancox, M. Field age determination in the european badger. Rev. D’écologie 1988, 43, 399–404. [Google Scholar]
- R-Development-Core-Team R: A Language and Environment for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 28 August 2022).
- RStudio Team. RStudio: Integrated Development for R. RStudio. Available online: https://rstudio.com (accessed on 28 August 2022).
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef] [PubMed]
- Knopf, C.W. Evolution of viral DNA-dependent DNA polymerases. Virus Genes 1998, 16, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, B.; Dural, G.; Yasmum, N.; Lembo, T.; de Thoisy, B.; Ryser-Degiorgis, M.-P.; Ulrich, R.G.; McGeoch, D.J. Novel Mammalian Herpesviruses and Lineages within the Gammaherpesvirinae: Cospeciation and Interspecies Transfer. J. Virol. 2008, 82, 3509–3516. [Google Scholar] [CrossRef]
- Bauer, C.C.; Aberle, S.W.; Popow-Kraupp, T.; Kapitan, M.; Hofmann, H.; Puchhammer-Stöckl, E. Serum Epstein-Barr virus DNA load in primary Epstein-Barr virus infection. J. Med. Virol. 2005, 75, 54–58. [Google Scholar] [CrossRef]
- Tsai, M.; Newman, C.; Macdonald, D.W.; Buesching, C.D. Stress-Related Herpesvirus Reactivation in Badgers Can Result in Clostridium Proliferation. Ecohealth 2021, 18, 440–450. [Google Scholar] [CrossRef] [PubMed]
- Hendrikse, L.D.; Kambli, A.; Kayko, C.; Canuti, M.; Rodrigues, B.; Stevens, B.; Vashon, J.; Lang, A.S.; Needle, D.B.; Troyer, R.M. Identification of a novel gammaherpesvirus in Canada lynx (Lynx canadensis). Viruses 2019, 11, 363. [Google Scholar] [CrossRef] [PubMed]
- Thorsteinsdóttir, L.; Torfason, E.G.; Torsteinsdóttir, S.; Svansson, V. Genetic diversity of equine gammaherpesviruses (γ-EHV) and isolation of a syncytium forming EHV-2 strain from a horse in Iceland. Res. Vet. Sci. 2013, 94, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Marmi, J.; López-Giráldez, F.; Macdonald, D.W.; Calafell, F.; Zholnerovskaya, E.; Domingo-Roura, X. Mitochondrial DNA reveals a strong phylogeographic structure in the badger across Eurasia. Mol. Ecol. 2006, 15, 1007–1020. [Google Scholar] [CrossRef] [PubMed]
- Allen, A.; Guerrero, J.; Byrne, A.; Lavery, J.; Presho, E.; Courcier, E.; O’Keeffe, J.; Fogarty, U.; Delahay, R.; Wilson, G.; et al. Genetic evidence further elucidates the history and extent of badger introductions from Great Britain into Ireland. R. Soc. Open Sci. 2020, 7, 200288. [Google Scholar] [CrossRef]
- Domingo-Roura, X.; Macdonald, D.W.; Roy, M.S.; Marmi, J.; Terradas, J.; Woodroffe, R.; Burke, T.; Wayne, R.K. Confirmation of low genetic diversity and multiple breeding females in a social group of Eurasian badgers from microsatellite and field data. Mol. Ecol. 2003, 12, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Alizon, S.; Hurford, A.; Mideo, N.; Van Baalen, M. Virulence evolution and the trade-off hypothesis: History, current state of affairs and the future. J. Evol. Biol. 2009, 22, 245–259. [Google Scholar] [CrossRef] [PubMed]
- Abdul-Rahman, F.; Tranchina, D.; Gresham, D. Fluctuating Environments Maintain Genetic Diversity through Neutral Fitness Effects and Balancing Selection. Mol. Biol. Evol. 2021, 38, 4362–4375. [Google Scholar] [CrossRef] [PubMed]
- Gibson, A.K.; Xu, J.Y.; Lively, C.M. Within-population covariation between sexual reproduction and susceptibility to local parasites. Evolution 2016, 70, 2049–2060. [Google Scholar] [CrossRef] [PubMed]
- Tollenaere, C.; Bryja, J.; Galan, M.; Cadet, P.; Deter, J.; Chaval, Y.; Berthier, K.; Ribas Salvador, A.; Voutilainen, L.; Laakkonen, J.; et al. Multiple parasites mediate balancing selection at two MHC class II genes in the fossorial water vole: Insights from multivariate analyses and population genetics. J. Evol. Biol. 2008, 21, 1307–1320. [Google Scholar] [CrossRef] [PubMed]
- Acevedo, M.A.; Dillemuth, F.P.; Flick, A.J.; Faldyn, M.J.; Elderd, B.D. Virulence-driven trade-offs in disease transmission: A meta-analysis. Evolution 2019, 73, 636–647. [Google Scholar] [CrossRef]
- De Roode, J.C.; Yates, A.J.; Altizer, S. Virulence-transmission trade-offs and population divergence in virulence in a naturally occurring butterfly parasite. Proc. Natl. Acad. Sci. USA 2008, 105, 7489–7494. [Google Scholar] [CrossRef]
- Ben-Shachar, R.; Koelle, K. Transmission-clearance trade-offs indicate that dengue virulence evolution depends on epidemiological context. Nat. Commun. 2018, 9, 2355. [Google Scholar] [CrossRef]
- Rappocciolo, G.; Birch, J.; Ellis, S.A. Down-regulation of MHC class I expression by equine herpesvirus-1. J. Gen. Virol. 2003, 84, 293–300. [Google Scholar] [CrossRef]
- Ma, G.; Feineis, S.; Osterrieder, N.; Van de Walle, G.R. Identification and Characterization of Equine Herpesvirus Type 1 pUL56 and Its Role in Virus-Induced Downregulation of Major Histocompatibility Complex Class I. J. Virol. 2012, 86, 3554–3563. [Google Scholar] [CrossRef][Green Version]
- Itzhaki, R.F. Corroboration of a major role for herpes simplex virus type 1 in Alzheimer’s disease. Front. Aging Neurosci. 2018, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Charonis, S.; James, L.M.; Georgopoulos, A.P. In silico assessment of binding affinities of three dementia-protective Human Leukocyte Antigen (HLA) alleles to nine human herpes virus antigens. Curr. Res. Transl. Med. 2020, 68, 211–216. [Google Scholar] [CrossRef] [PubMed]
- Hulaniuk, M.L.; Mojsiejczuk, L.; Jauk, F.; Remondegui, C.; Mammana, L.; Bouzas, M.B.; Zapiola, I.; Ferro, M.V.; Ajalla, C.; Blejer, J.; et al. Genetic diversity and phylogeographic analysis of human herpesvirus type 8 (HHV-8) in two distant regions of Argentina: Association with the genetic ancestry of the population. Infect. Genet. Evol. 2020, 85, 104523. [Google Scholar] [CrossRef] [PubMed]



| Start Position | End Position | Primer Sequence (5′-3′) | Product Length | |
|---|---|---|---|---|
| pol5-F | 3 | 22 | GGCAGGGAATTTTTATAACC | 699 |
| pol5-R | 681 | 701 | CCACCCAAAAGTAGAAAATCC | |
| pol6-F | 345 | 364 | CCCCTCTGGAACTGTGCTAA | 637 |
| pol6-R | 962 | 981 | AGGGTCACATGTCCCCAAAT | |
| pol7-F | 863 | 846 | ATGTCTGGGGGAAAATGG | 486 |
| pol7-R | 1329 | 1348 | GACCTCCTATGCACTGCTTG | |
| pol10-F | 1185 | 1204 | TGAAGTTCACACACCCCAGA | 361 |
| pol10-R | 1524 | 1545 | TCCATCGGTCAGCACTCTC | |
| pol3-F | 1328 | 1347 | CCAAGCAGTGCATAGGAGGT | 771 |
| pol3-R | 2072 | 2098 | TGGACTTCTCCAACATGCGTCGCCCTT | |
| pol11-F | 1684 | 1703 | CCGATCTTGGTGGTTGATTT | 627 |
| pol11-R | 2291 | 2310 | CTTAATTGGCTCCTCGAACA | |
| pol12-F | 2015 | 2034 | CAGGTGTGTCCTCGGGTATT | 561 |
| pol12-R | 2556 | 2575 | TCACTTTGAAAAGTGGAAGTGG | |
| pol9-F | 2393 | 2412 | TGATGAAGGGAGTGGATCTC | 597 |
| pol9-R | 2936 | 2989 | TCACAGCTTTGTCTGCACTG |
| Tattoo Number | Sex | Age | Spring | Summer | Autumn |
|---|---|---|---|---|---|
| 1012 | Male | 13 | v | ||
| 1045x | Female | 13 | v | v | |
| 1232 | Female | 10 | v | v | |
| 1234x | Female | 10 | v | v | |
| 1283 | Male | 9 | v | v | |
| 1330 | Male | 8 | v | ||
| 1379 | Male | >8 * | v | v | |
| 1435 | Female | 6 | v | v | |
| 1478 | Female | 5 | v | ||
| 1487 | Male | 7 | v | v | |
| 1520 | Female | 4 | v | v | |
| 1622 | Male | 3 | v | v | |
| 1694 | Female | 2 | v | v | |
| 1699 | Male | 1 | v | v | |
| 1735 | Male | 0 | v | ||
| 1746 | Male | 0 | v | ||
| 1749 | Female | 0 | v | ||
| 1750 | Female | 0 | v | v | |
| 1751 | Female | 0 | v | v | v |
| 1753 | Female | 0 | v | ||
| 1754 | Male | 0 | v | ||
| 1755 | Male | 0 | v | v | v |
| 1778 | Male | 0 | v | ||
| 1785 | Female | 0 | v |
| Position | Reference (AF275657 a) | Common Variant | Novel Variant | Conserved Protein Domain Family b | In Conserved Region c | In Conserved Region within Host Order d |
|---|---|---|---|---|---|---|
| 253 | P | L | L | 3′-5′ exonuclease Exo I | Yes | Yes |
| 274 | N | Y | Y | 3′-5′ exonuclease Exo I | Yes | Yes |
| 591 | K | K | R | Polymerase | No | No |
| 649 | N | N | K | Polymerase | No | Yes |
| 819 | E | E | G | Polymerase | No | No |
| 872 | T | T | N | Polymerase | No | No |
| 889 | S | S | L | Polymerase | No | Yes |
| Variable | Estimate | Std. Error | df | t Value | p Value | |
|---|---|---|---|---|---|---|
| (Intercept) | 34.855 | 0.464 | 28.794 | 75.072 | ||
| Genotype | ||||||
| Common | (Reference) | |||||
| Novel | −1.815 | 0.547 | 19.447 | −3.316 | 0.004 | |
| Coinfection | −0.346 | 0.834 | 32.885 | −0.415 | 0.681 | |
| Age Group | ||||||
| Adult | (Reference) | |||||
| Juvenile | −3.030 | 0.543 | 18.615 | −5.583 | <0.001 | |
| Season | ||||||
| Spring | (Reference) | |||||
| Summer | −1.379 | 0.453 | 23.179 | −3.045 | 0.006 | |
| Autumn | 0.012 | 0.416 | 17.381 | 0.028 | 0.978 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsai, M.-s.; François, S.; Newman, C.; Macdonald, D.W.; Buesching, C.D. Infection with a Recently Discovered Gammaherpesvirus Variant in European Badgers, Meles meles, is Associated with Higher Relative Viral Loads in Blood. Pathogens 2022, 11, 1154. https://doi.org/10.3390/pathogens11101154
Tsai M-s, François S, Newman C, Macdonald DW, Buesching CD. Infection with a Recently Discovered Gammaherpesvirus Variant in European Badgers, Meles meles, is Associated with Higher Relative Viral Loads in Blood. Pathogens. 2022; 11(10):1154. https://doi.org/10.3390/pathogens11101154
Chicago/Turabian StyleTsai, Ming-shan, Sarah François, Chris Newman, David W. Macdonald, and Christina D. Buesching. 2022. "Infection with a Recently Discovered Gammaherpesvirus Variant in European Badgers, Meles meles, is Associated with Higher Relative Viral Loads in Blood" Pathogens 11, no. 10: 1154. https://doi.org/10.3390/pathogens11101154
APA StyleTsai, M.-s., François, S., Newman, C., Macdonald, D. W., & Buesching, C. D. (2022). Infection with a Recently Discovered Gammaherpesvirus Variant in European Badgers, Meles meles, is Associated with Higher Relative Viral Loads in Blood. Pathogens, 11(10), 1154. https://doi.org/10.3390/pathogens11101154







