Microbiological Quality of Nuts, Dried and Candied Fruits, Including the Prevalence of Cronobacter spp.
Abstract
:1. Introduction
2. Results and Discussion
2.1. Prevalence of Total Plate Count (TPC) of Bacteria, Yeasts, and Molds in Tested Samples
2.2. Presence of Cronobacter spp. in the Tested Food Products and Phenotypic Identification of Presumptive Isolates
2.3. Identification of the Isolates from the Genus Cronobacter
3. Materials and Methods
3.1. Sample Collection
- samples of different types of nut: Italian and Brazil nuts, hazelnuts, almonds, pecan, cashew, pine nuts, and macadamia (20 samples);
- samples of dried fruit: prunes, raisins, cherries, sour cherries, figs, banana, dates, apricot, black currant, cranberry, Goji and Chia berries (24 samples);
- samples of candied fruit: mango, pineapple, Jackfruit, plums, and passion fruit (8 samples);
- samples of seed: sunflower and pumpkin seeds (4 samples);
- samples of mixes of seed, dried fruit and nut (mixes in various proportions of: raisins, dried cranberries, walnuts, hazelnuts, cashews, almonds, and sunflower seeds) (8 samples).
3.2. Microbiological Analysis of Samples
- total plate count of bacteria (TPC) on the PCA medium (Merck, Warsaw, Poland); incubation conditions 72 h at 30 °C [59];
- yeasts and molds counts on the YGC medium (Merck, Warsaw, Poland); incubation conditions 5 days at 25 °C [60];
- occurrence of bacteria from the genus Cronobacter [53].
3.3. Phenotypic Identification of Presumptive Cronobacter spp. Isolates
3.4. Genetic Identification of Cronobacter spp. Isolates
3.4.1. Extraction of DNA
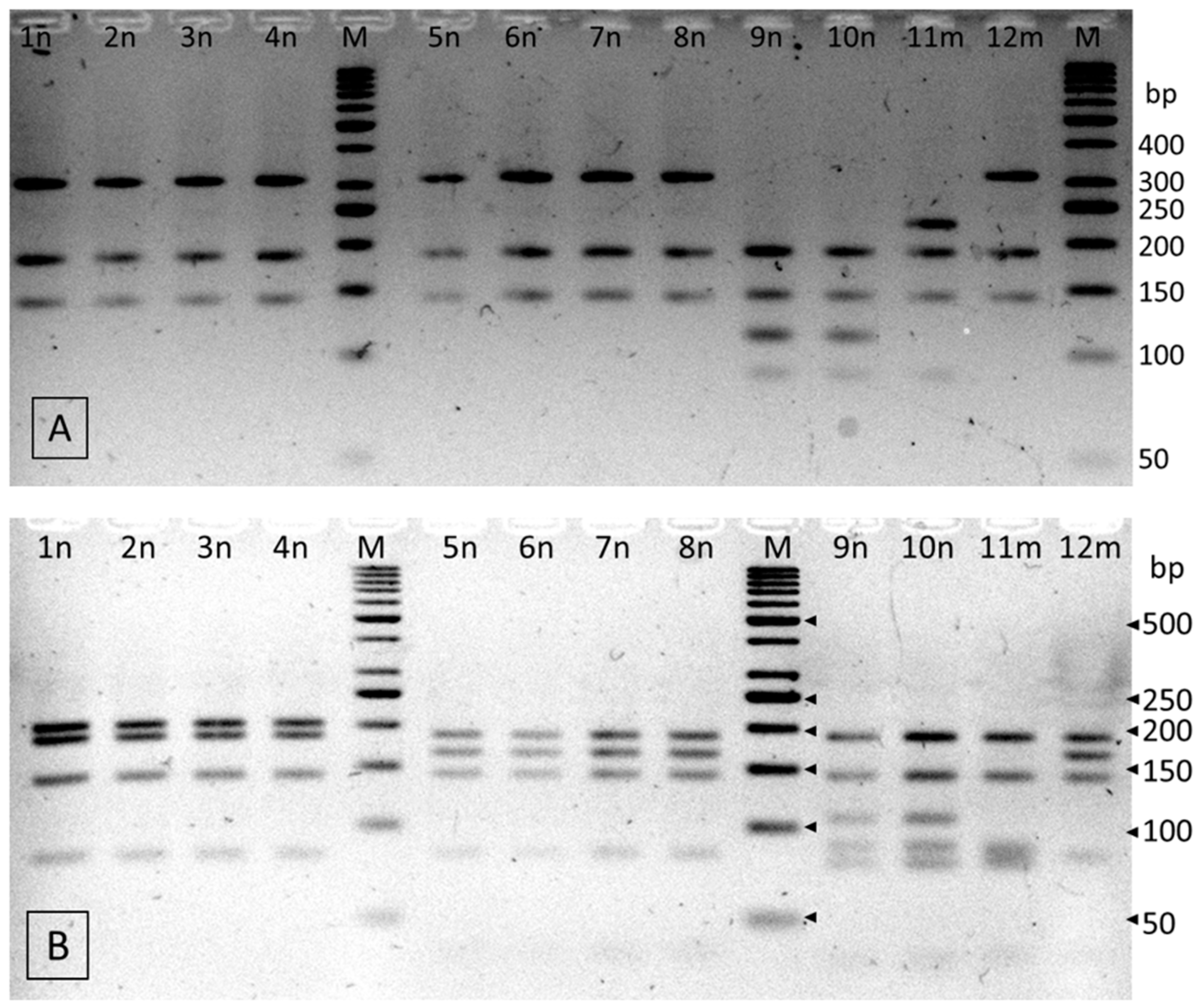
3.4.2. PCR-RFLP
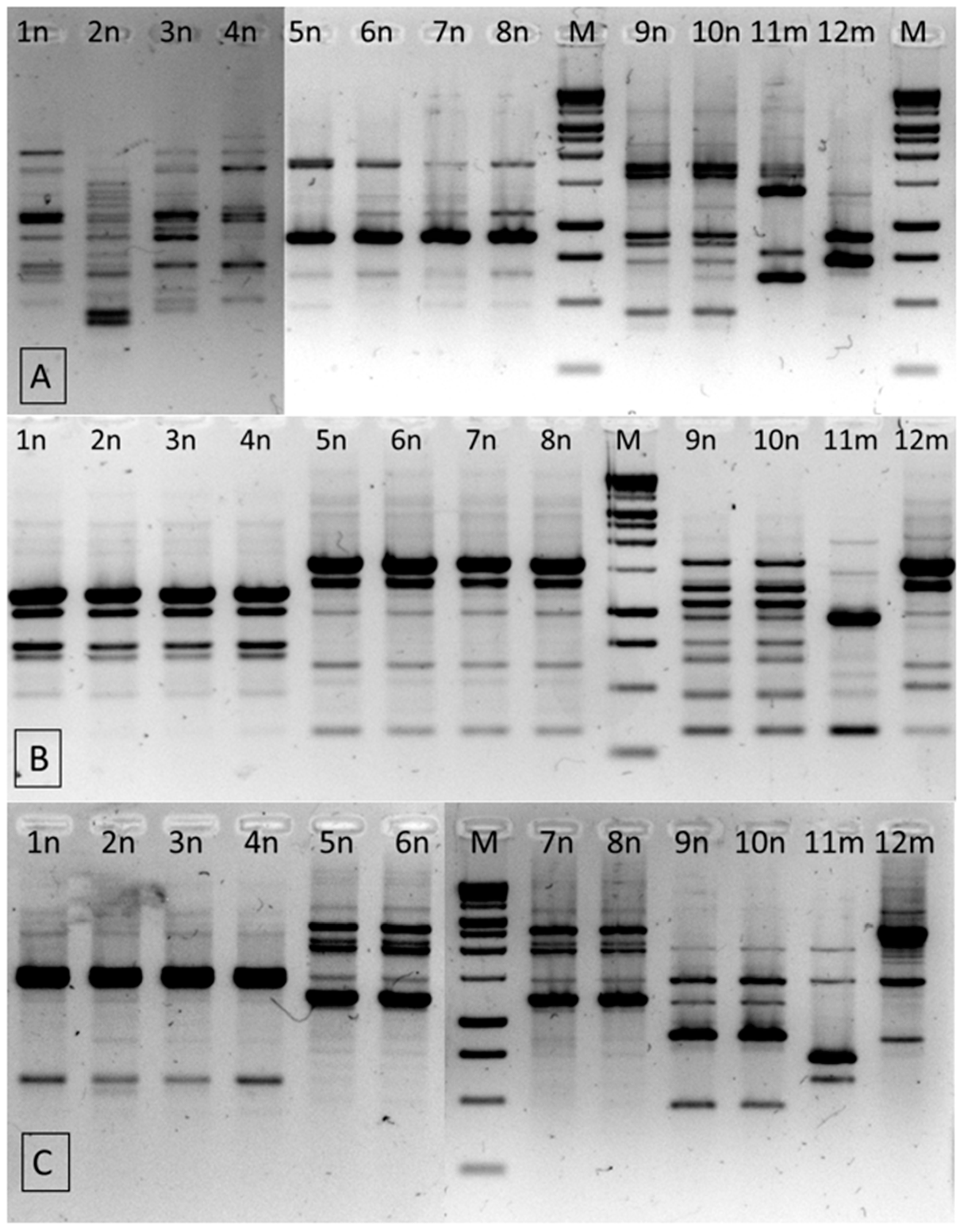
3.4.3. RAPD-PCR
3.5. Statistical Analysis of Results
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- FAO/WHO. Ranking of Low Moisture Foods in Support of Microbiological Risk Management: Preliminary report of FAO/WHO expert Consultation on Ranking of Low Moisture Foods. Part I—Main Report; FAO/WHO: Rome, Italy; Geneva, Switzerland, 2014. [Google Scholar]
- Ntuli, V.; Chatanga, P.; Kwiri, R.; Gadaga, H.T.; Gere, J.; Matsepo, T.; Potloane Rethabile, P. Microbiological quality of selected dried fruits and vegetables in Maseru, Lesotho. Afr. J. Microbiol. Res. 2017, 11, 185–193. [Google Scholar] [CrossRef] [Green Version]
- Danyluk, M.; Jones, T.M.; Abd, S.J.; Schlitt-Dittrich, F.; Jacobs, M.; Harris, L.J. Prevalence and amounts of Salmonella found on raw California almonds. J. Food Prot. 2007, 70, 820–827. [Google Scholar] [CrossRef] [PubMed]
- Frelka, J.; Davidson, G.; Harris, L.J. Changes in aerobic plate and Escherichia coli—coliform counts and in populations of inoculated foodborne pathogens on inshell walnuts during storage. J. Food Prot. 2016, 79, 1143–1153. [Google Scholar] [CrossRef] [PubMed]
- FAOSTAT. Food and Agriculture Organization of the United Nations 2021. Available online: http://www.fao.org/faostat/en/#data (accessed on 29 June 2021).
- Santo, D.; Graça, A.; Nunes, C.; Quintas, C. Escherichia coli and Cronobacter sakazakii in ‘Tommy Atkins’ minimally processed mangos: Survival, growth and effect of UV-C and electrolyzed water. Food Microbiol. 2018, 70, 49–54. [Google Scholar] [CrossRef]
- Beatty, M.E.; Laporte, T.N.; Phan, Q.; Van Duyne, S.V.; Braden, C. A multistate outbreak of Salmonella enterica serotype Saintpaul infections linked to mango consumption: A recurrent theme. Clin. Infect. Dis. 2004, 38, 1337–1338. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.F.; Tauxe, R.V.; Hedberg, C.W. The growing burden of foodborne outbreaks due to contaminated fresh produce: Risks and opportunities. Epidemiol. Infect. 2009, 137, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Davidson, G.R.; Frelka, J.C.; Yang, M.; Jones, T.M.; Harris, L.J. Prevalence of Escherichia coli O157:H7 and Salmonella on inshell California Walnuts. J. Food Prot. 2015, 78, 1547–1553. [Google Scholar] [CrossRef]
- Miller, B.D.; Rigdon, C.E.; Ball, J.; Rounds, J.M.; Klos, R.F.; Brennan, B.M.; Arends, K.D.; Kennelly, P.; Hedberg, C.; Smith, K.E. Use of traceback methods to confirm the source of a multistate Escherichia coli O157:H7 outbreak due to in-shell hazelnuts. J. Food Prot. 2012, 75, 320–327. [Google Scholar] [CrossRef] [PubMed]
- Podolak, R.; Enache, E.; Stone, W.; Black, D.G.; Elliott, P.H. Sources and risk factors for contamination, survival, persistence, and heat resistance of Salmonella in low-moisture foods. J. Food Prot. 2010, 73, 1919–1936. [Google Scholar] [CrossRef]
- Alsonosi, A.; Hariri, S.; Kajsík, M.; Oriešková, M.; Hanulík, V.; Röderová, M.; Petrželová, J.; Kollárová, H.; Drahovská, H.; Forsythe, S.; et al. The speciation and genotyping of Cronobacter isolates from hospitalised patients. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 1979–1988. [Google Scholar] [CrossRef] [Green Version]
- Mullane, N.; Iversen, C.; Healy, B.; Walsh, C.; Whyte, P.; Wall, P.; Quinn, T.; Fanning, S. Enterobacter sakazakii an emerging bacterial pathogen with implications for infant health. Minerva Pediatr. 2007, 59, 137–148. [Google Scholar] [PubMed]
- Healy, B.; Cooney, S.; O’Brien, S.; Iversen, C.; Whyte, P.; Nally, J.; Callanan, J.J.; Fanning, S. Cronobacter (Enterobacter sakazakii): An opportunistic foodborne pathogen. Foodborne Pathog. Dis. 2010, 7, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Forsythe, S.J. Updates on the Cronobacter Genus. Annu. Rev. Food Sci. Technol. 2018, 9, 23–44. [Google Scholar] [CrossRef] [PubMed]
- Iversen, C.; Mullane, N.; McCardell, B.; Tall, B.; Lehner, A.; Fanning, S.; Stephan, R.; Joosten, H. Cronobacter gen. nov., a new genus to accommodate the biogroups of Enterobacter sakazakii, and proposal of Cronobacter sakazakii gen. nov., comb. nov., Cronobacter malonaticus sp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov., Cronobacter genomospecies 1, and of three subspecies, Cronobacter dublinensis subsp. dublinensis subsp. nov., Cronobacter dublinensis subsp. lausannensis subsp. nov. and Cronobacter dublinensis subsp. lactardi subsp. nov. Int. J. Syst. Evol. Microbiol. 2008, 58, 1442–1447. [Google Scholar]
- Stoop, B.; Lehner, A.; Iversen, C.; Fanning, S.; Stephan, R. Development and evaluation of rpoB-based PCR systems to differentiate the six proposed species within the genus Cronobacter. Int. J. Food Microbiol. 2009, 136, 165–168. [Google Scholar] [CrossRef] [Green Version]
- Joseph, S.; Forsythe, S.J. Predominance of Cronobacter sakazakii sequence type 4 in neonatal infections. Emerg. Infect. Dis. 2011, 17, 1713–1715. [Google Scholar] [CrossRef] [PubMed]
- Masood, N.; Moore, K.; Farbos, A.; Hariri, S.; Paszkiewicz, K.; Dickins, B.; McNally, A.; Forsythe, S. Draft genome sequences of three newly identified species in the genus Cronobacter, C. helveticus LMG23732, C. pulveris LMG24059, and C. zurichensis LMG23730. Genome Announc. 2013, 1, e01038-13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Drudy, D.; Mullane, N.R.; Quinn, T.; Wall, P.G.; Fanning, S. Enterobacter sakazakii: An emerging pathogen in powdered infant formula. Clin. Infect. Dis. 2006, 42, 996–1002. [Google Scholar] [CrossRef] [Green Version]
- Kim, K.; Jang, S.; Kim, S.; Park, J.; Heu, S.; Ryu, S. Prevalence and genetic diversity of Enterobacter sakazakii in ingredients of infant foods. Int. J. Food Microbiol. 2008, 122, 196–203. [Google Scholar] [CrossRef]
- Muller, A.; Stephan, R.; Fricker-Feer, C.; Lehner, A. Genetic diversity of Cronobacter sakazakii isolates collected from a Swiss infant formula production facility. J. Food Prot. 2013, 76, 883–887. [Google Scholar] [CrossRef]
- Akineden, Ö.; Murata, K.J.; Gross, M.; Usleber, E. Microbiological quality of raw dried pasta from the German market, with special emphasis on Cronobacter species. J. Food Sci. 2015, 80, 2860–2867. [Google Scholar] [CrossRef]
- Li, Y.; Chen, Q.; Zhao, J.; Jiang, H.; Lu, F.; Bie, X.; Lu, Z. Isolation, identification and antimicrobial resistance of Cronobacter spp. isolated from various foods in China. Food Control 2014, 37, 109–114. [Google Scholar] [CrossRef]
- Lou, X.; Si, G.; Yu, H.; Qi, J.; Liu, T.; Fang, Z. Possible reservoir and routes of transmission of Cronobacter (Enterobacter sakazakii) via wheat flour. Food Control 2014, 43, 258–262. [Google Scholar] [CrossRef]
- Berthold-Pluta, A.; Garbowska, M.; Stefańska, I.; Pluta, A. Microbiological quality of selected ready-to-eat leaf vegetables, sprouts and non-pasteurized fresh fruit-vegetable juices including the presence of Cronobacter spp. Food Microbiol. 2017, 65, 221–230. [Google Scholar] [CrossRef]
- Kim, J.-B.; Park, Y.-B.; Kang, S.-H.; Lee, M.-J.; Kim, K.-C.; Jeong, H.-R.; Kim, D.-H.; Yoon, M.-H.; Lee, J.-B.; Oh, D.-H. Prevalence, genetic diversity, and antibiotic susceptibility of Cronobacter spp. (Enterobacter sakazakii) isolated from sunshik, its ingredients and soils. Food Sci. Biotechnol. 2011, 20, 941–948. [Google Scholar] [CrossRef]
- Weiss, C.; Becker, B.; Holzapfel, W. Application and acceptability of three commercial systems for detection of Enterobacter sakazakii in ready-to-eat vegetable salads. Arch. Lebensmittelhyg. 2005, 56, 34–38. [Google Scholar]
- Vojkovska, H.; Karpiskova, R.; Orieskova, M.; Drahovska, H. Characterization of Cronobacter spp. isolated from food of plant origin and environmental samples collected from farms and from supermarkets in the Czech Republic. Int. J. Food Microbiol. 2016, 217, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Brandão, M.L.L.; Umeda, N.S.; Jackson, E.; Forsythe, S.J.; de Filippis, I. Isolation, molecular and phenotypic characterization, and antibiotic susceptibility of Cronobacter spp. from Brazilian retail foods. Food Microbiol. 2017, 63, 129–138. [Google Scholar] [CrossRef]
- Garbowska, M.; Berthold-Pluta, A.; Stasiak-Różańska, L. Microbiological quality of selected spices and herbs including the presence of Cronobacter spp. Food Microbiol. 2015, 49, 1–5. [Google Scholar] [CrossRef]
- Heperkan, D.; Dalkilic-Kaya, G.; Juneja, V.K. Cronobacter sakazakii in baby foods and baby food ingredients of dairy origin and microbiological profile of positive samples. LWT Food Sci. Technol. 2017, 75, 402–407. [Google Scholar] [CrossRef]
- Singh, N.; Goel, G.; Raghav, M. Prevalence and characterization of Cronobacter spp. from various foods, medicinal plants, and environmental samples. Curr. Microbiol. 2015, 71, 31–38. [Google Scholar] [CrossRef]
- Kandhai, M.C.; Heuvelink, A.E.; Reij, M.W.; Beumer, R.R.; Dijk, R.; van Tilburg, J.J.H.C.; van Schothorst, M.; Gorris, L.G.M. A study into the occurrence of Cronobacter spp. in the Netherlands between 2001 and 2005. Food Control 2010, 21, 1127–1136. [Google Scholar] [CrossRef]
- Mohammed, M.A.; Sallam, K.I.; Tamura, T. Prevalence, identification and molecular characterization of Cronobacter sakazakii isolated from retail meat products. Food Control 2015, 53, 206–211. [Google Scholar] [CrossRef]
- Friedemann, M. Enterobacter sakazakii in food and beverages (other than infant formula and milk powder). Int. J. Food Microbiol. 2007, 116, 1–10. [Google Scholar] [CrossRef]
- Vasconcellos, L.; Carvalho, C.T.; Tavares, R.O.; de Mello Medeiros, V.; de Oliveira Rosas, C.; Silva, J.N.; dos Reis Lopes, S.M.; Forsythe, S.J.; Brandão, M.L.L. Isolation, molecular and phenotypic characterization of Cronobacter spp. in ready-to-eat salads and foods from Japanese cuisine commercialized in Brazil. Food Res. Int. 2018, 107, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Craven, H.M.; McAuley, C.M.; Duffy, L.L.; Fegan, N. Distribution, prevalence and persistence of Cronobacter (Enterobacter sakazakii) in the nonprocessing and processing environments of five milk powder factories. J. Appl. Microbiol. 2010, 109, 1044–1052. [Google Scholar] [CrossRef]
- Feeney, A.; Johnston, C.D.; Govender, R.; O’Mahony, J.; Coffey, A.; Sleator, R.D. Analysis of the role of the Cronobacter sakazakii ProP homologues in osmotolerance. Gut Pathog. 2014, 6, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Feeney, A.; Kropp, K.A.; O’Connor, R.; Sleator, R.D. Cronobacter sakazakii: Stress survival and virulence potential in an opportunistic foodborne pathogen. Gut Microbes 2014, 5, 711–718. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hurrell, E.; Kucerova, E.; Loughlin, M.; Caubilla-Barron, J.; Forsythe, S.J. Biofilm formation on enteral feeding tubes by Cronobacter sakazakii, Salmonella serovars and other Enterobacteriaceae. Int. J. Food Microbiol. 2009, 136, 227–231. [Google Scholar] [CrossRef]
- Beuchat, L.R.; Kim, H.; Gurtler, J.B.; Lin, L.-C.; Ryu, J.-H.; Richards, G.M. Cronobacter sakazakii in foods and factors affecting its survival, growth, and inactivation. Int. J. Food Microbiol. 2009, 136, 204–213. [Google Scholar] [CrossRef]
- Lehner, A.; Riedel, K.; Eberl, L.; Breeuwer, P.; Diep, B.; Stephan, R. Biofilm formation, extracellular polysaccharide production, and cell-to-cell signaling in various Enterobacter sakazakii strains: Aspects promoting environmental persistence. J. Food Prot. 2005, 68, 2287–2294. [Google Scholar] [CrossRef] [PubMed]
- Singh, N.; Patil, A.; Prabhune, A.; Goel, G. Inhibition of quorum-sensing-mediated biofilm formation in Cronobacter sakazakii strains. Microbiology 2016, 162, 1708–1714. [Google Scholar] [CrossRef] [PubMed]
- Ling, N.; Forsythe, S.; Wu, Q.; Ding, Y.; Zhang, J.; Zeng, H. Insights into Cronobacter sakazakii Biofilm Formation and Control Strategies in the Food Industry. Engineering 2020, 4, 393–405. [Google Scholar] [CrossRef]
- Iversen, C.; Forsythe, S.J. Isolation of Enterobacter sakazakii and other Enterobacteriaceae from powdered infant formula milk and related products. J. Food Microbiol. 2004, 21, 771–777. [Google Scholar] [CrossRef]
- Patrick, M.E.; Mahon, B.E.; Greene, S.A.; Rounds, J.; Cronquist, A.; Wymore, K.; Bowen, A. Incidence of Cronobacter spp. infections, United States, 2003–2009. Emerg. Infect. Dis. 2014, 20, 1536–1539. [Google Scholar] [CrossRef]
- Holy, O.; Forsythe, S.J. Cronobacter species as emerging causes of healthcare-associated infection. J. Hosp. Infect. 2014, 86, 169–177. [Google Scholar] [CrossRef] [Green Version]
- Eglezos, S.; Huang, B.; Stuttard, E. A Survey of the bacteriological quality of preroasted peanut, almond, cashew, hazelnut, and Brazil nut kernels received into three Australian nut-processing facilities over a period of 3 years. J. Food Prot. 2008, 71, 402–404. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.J.; Lieberman, V.; Mashiana, R.P.; Atwill, E.; Yang, M.; Chandler, J.C.; Bisha, B.; Jones, T. Prevalence and amounts of Salmonella found on raw California inshell pistachios. J. Food Prot. 2016, 79, 1304–1315. [Google Scholar] [CrossRef] [Green Version]
- Eglezos, S. The bacteriological quality of retail-level peanut, almond, cashew, hazelnut, Brazil, and mixed nut kernels produced in two Australian nut-processing facilities over a period of 3 years. Foodborne Pathog. Dis. 2010, 7, 863–866. [Google Scholar] [CrossRef]
- Witthuhn, R.; Engelbrecht, S.; Joubert, E.; Britz, T. Microbial content of commercial South African high-moisture dried fruits. J. Appl. Microbiol. 2005, 98, 722–726. [Google Scholar] [CrossRef]
- International Organization for Standardization. ISO 22964:2017. Microbiology of the Food Chain—Horizontal Method for the Detection of Cronobacter spp.; ISS Facility Services: Warsaw, Poland, 2017. [Google Scholar]
- Joseph, S.; Forsythe, S.J. Multilocus Sequence Typing (MLST) for Cronobacter spp. Methods Mol. Biol. 2017, 1616, 241–248. [Google Scholar]
- Vlach, J.; Javůrková, B.; Karamonová, L.; Blažková, M.; Fukal, L. Novel PCR-RFLP system based on rpoB gene for differentiation of Cronobacter species. Food Microbiol. 2017, 62, 1–8. [Google Scholar] [CrossRef]
- Jošić, D.; Stojanovic, M.; Lepšanović, Z.; Katić, V. Molecular characterization of Cronobacter sakazakii isolated from different herbal teas and mixtures in Serbia. Genetika 2017, 49, 921–934. [Google Scholar] [CrossRef]
- Freire, F.d.C.O.; Offord, L. Bacterial and yeast counts in Brazilian commodities and spices. Braz. J. Microbiol. 2002, 33, 145–148. [Google Scholar] [CrossRef] [Green Version]
- Hochel, I.; Růžčková, H.; Krásný, L.; Demnerová, K. Occurrence of Cronobacter spp. in retail foods. J. Appl. Microbiol. 2012, 112, 1257–1265. [Google Scholar] [CrossRef] [PubMed]
- International Organization for Standardization. ISO 4833-1:2013. Microbiology of the Food Chain—Horizontal Method for the Enumeration of Microorganisms—Part 1: Colony Count at 30 Degrees C by the Pour Plate Technique; ISS Facility Services: Warsaw, Poland, 2013. [Google Scholar]
- International Organization for Standardization. ISO 7954:1987. Microbiology—General Guidance for Enumeration of Yeasts and Moulds—Colony Count Technique at 25 °C; ISS Facility Services: Warsaw, Poland, 1987. [Google Scholar]
- Yamagishi, J.; Sato, Y.; Shinozaki, N.; Ye, B.; Tsuboi, A.; Nagasaki, M.; Yamashita, R. Comparison of Boiling and Robotics Automation Method in DNA Extraction for Metagenomic Sequencing of Human Oral Microbes. PLoS ONE 2016, 11, e0154389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Food Products (Number of Analyzed Samples) | Number of Samples | |||||||
|---|---|---|---|---|---|---|---|---|
| Total Plate Count of Bacteria [log Colony Forming Units (CFU) g−1] | ||||||||
| Not Detected in 0.1 g | > 1–2 | > 2–3 | > 3–4 | > 4–5 | > 5–6 | Range | Mean ± Standard Deviation | |
| Nuts (20) | 4 | 5 | 4 | 5 | 2 | 0 | 1.2–4.4 | 2.80 ab ± 1.03 |
| Dried fruits (24) | 10 | 2 | 6 | 4 | 0 | 2 | 1.9–5.3 | 3.07 b ± 1.06 |
| Candied fruits (8) | 0 | 3 | 5 | 0 | 0 | 0 | 1.5–2.8 | 2.20 a ± 0.46 |
| Seeds (4) | 0 | 0 | 2 | 1 | 1 | 0 | 2.4–4.4 | 3.17 b ± 0.80 |
| Mixes of dried fruits, seeds and nuts (8) | 1 | 2 | 3 | 2 | 0 | 0 | 1.4–3.2 | 2.47 ab ± 0.62 |
| Total (64) | 15 | 12 | 20 | 12 | 3 | 2 | 1.2–5.3 | 2.26 ± 1.37 |
| Food Products (Number of Analyzed Samples) | Number of Samples | ||||
|---|---|---|---|---|---|
| Total Count of Yeasts [log CFU g−1] | |||||
| Not Detected in 0.1 g | >1–2 | >2–3 | Range | Mean ± Standard Deviation | |
| Nuts (20) | 16 | 3 | 1 | 1.4–2.6 | 1.88 a ± 0.47 |
| Dried fruits (24) | 23 | 1 | 0 | 1.8 | 1.80 a ± 0.00 |
| Candied fruits (8) | 8 | 0 | 0 | – | – |
| Seeds (4) | 4 | 0 | 0 | – | – |
| Mixes of dried fruits, seeds and nuts (8) | 6 | 2 | 0 | 1.5–1.9 | 1.70 a ± 0.22 |
| Total (64) | 57 | 6 | 1 | 1.4–2.6 | 1.82 ± 0.37 |
| Food Products (Number of Analyzed Samples) | Number of Samples | |||||
|---|---|---|---|---|---|---|
| Total Count of Molds [log CFU g−1] | ||||||
| Not Detected in 0.1 g | >1–2 | >2–3 | >3–4 | Range | Mean ± Standard Deviation | |
| Nuts (20) | 2 | 8 | 10 | 0 | 1.3–2.6 | 1.99 a ± 0.41 |
| Dried fruits (24) | 8 | 4 | 10 | 2 | 1.8–4.0 | 2.49 b ± 0.59 |
| Candied fruits (8) | 7 | 1 | 0 | 0 | 1.5 | 1.50 a ± 0.01 |
| Seeds (4) | 0 | 2 | 1 | 1 | 1.0–3.8 | 2.13 ab ± 1.09 |
| Mixes of dried fruits, seeds and nuts (8) | 4 | 0 | 2 | 2 | 2.7–3.8 | 3.22 c ± 0.48 |
| Total (64) | 21 | 15 | 23 | 5 | 1.0–4.0 | 2.29 ± 0.69 |
| RFLP-Pattern | Strain | Fragment Length (bp) * | Species Identification | References |
|---|---|---|---|---|
| A | 1n, 2n, 3n, 4n | 312, 186, 142 (HinP1I) 206, 186, 142, 83 (HinP1I + Csp6I) | C. turicensis | C. turicensis LMG 23827 |
| B | 5n, 6n, 7n, 8n, 12m | 312, 186, 142 (HinP1I) 186, 167, 142, 83 (HinP1I + Csp6I) | C. malonaticus | C. malonaticus LMG 23826 |
| C | 9n, 10n | 186, 142, 114, 110, 88 (HinP1I) 186, 142, 106, 88, 79 (HinP1I+ Csp6I) | C. sakazakii | C. sakazakii NTCT 8155 |
| D | 11m | 224, 186, 142, 88 (HinP1I) 186, 142, 88, 83, 79 (HinP1I+ Csp6I) | C. sakazakii | C. sakazakii ATCC 29544 |
| Isolate No. | Origin | Species Identification by PCR-RFLP | PCR-RFLP Pattern | RAPD-PCR * Patterns | ||
|---|---|---|---|---|---|---|
| RP | UBC245 | UBC282 | ||||
| 1n | almonds | C. turicensis | A | 1 | 1 | 1 |
| 2n | hazelnuts | C. turicensis | A | 2 | 1 | 1 |
| 3n | almonds | C. turicensis | A | 1 | 1 | 1 |
| 4n | almonds | C. turicensis | A | 1 | 1 | 1 |
| 5n | hazelnuts | C. malonaticus | B | 3 | 2 | 2 |
| 6n | cashew nuts | C. malonaticus | B | 3 | 2 | 2 |
| 7n | pini nuts | C. malonaticus | B | 3 | 2 | 2 |
| 8n | macadamia nuts | C. malonaticus | B | 3 | 2 | 2 |
| 9n | Brazilian nuts | C. sakazakii | C | 4 | 3 | 3 |
| 10n | Brazilian nuts | C. sakazakii | C | 4 | 3 | 3 |
| 11m | mixes of dried fruits, seeds and nuts | C. sakazakii | D | 5 | 4 | 4 |
| 12m | mixes of dried fruits, seeds and nuts | C. malonaticus | B | 6 | 5 | 5 |
| Food Product | Presence of Bacteria from the Genus Cronobacter * [Positive Samples Count/Analyzed Samples Count (% of Positive Samples)] | Presence of C. sakazakii ** [Positive Samples Count/Analyzed Samples Count (% of Positive Samples)] | Presence of C. turicensis ** [Positive Samples Count/Analyzed Samples Count (% of Positive Samples)] | Presence of C. malonaticus ** [Positive Samples Count/Analyzed Samples Count (% of Positive Samples)] |
|---|---|---|---|---|
| Nuts | 10 1/20 (50.0) | 2 2 /20 (10.0) | 4 3/20 (20.0) | 4 4 /20 (20.0) |
| Dried fruits | 0/24 (0.0) | 0/24 (0.0) | 0/24 (0.0) | 0/24 (0.0) |
| Candied fruits | 0/8 (0.0) | 0/8 (0.0) | 0/8 (0.0) | 0/8 (0.0) |
| Seeds | 0/4 (0.0) | 0/4 (0.0) | 0/4 (0.0) | 0/4 (0.0) |
| Mixes of dried fruits, seeds and nuts | 2 1/8 (25.0) | 1 2/8 (12.5) | 0/8 (0.0) | 1 4/8 (12.5) |
| Total | 12/64 (18.8) | 3/64 (4.7) | 4/64 (6.3) | 5/64 (7.8) |
| Species | GenBank Accession No. | Predicted Fragments in In Silico Analysis (bp) |
|---|---|---|
| C. sakazakii ATCC 29544 | FJ717657 | 186, 142, 106, 88, 79, 39, 15, 2 |
| C. sakazakii NTCT 8155 | CP012253 | 186, 142, 88, 83, 79, 35, 23, 15, 4, 2 |
| C. malonaticus LMG 23826 | CP013940 | 186, 167, 142, 83, 39, 23, 15, 2 |
| C. turicensis LMG 23827 | NC_013282 | 206, 186, 142, 83, 23, 15, 2 |
| C. dubliniensis LMG 23823 | CP012266 | 211, 167, 142, 87, 35, 15, 2 |
| C. muytjensii ATCC 51329 | CP012268 | 188, 167, 96, 87, 46, 35, 23, 15, 2 |
| C. condimenti LMG 26250 | CP012264 | 211, 167, 142, 122, 15, 2 |
| C. universalis NCTC 9529 | CP012257 | 167, 157, 142, 83, 39, 31, 23, 15, 2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berthold-Pluta, A.; Garbowska, M.; Stefańska, I.; Stasiak-Różańska, L.; Aleksandrzak-Piekarczyk, T.; Pluta, A. Microbiological Quality of Nuts, Dried and Candied Fruits, Including the Prevalence of Cronobacter spp. Pathogens 2021, 10, 900. https://doi.org/10.3390/pathogens10070900
Berthold-Pluta A, Garbowska M, Stefańska I, Stasiak-Różańska L, Aleksandrzak-Piekarczyk T, Pluta A. Microbiological Quality of Nuts, Dried and Candied Fruits, Including the Prevalence of Cronobacter spp. Pathogens. 2021; 10(7):900. https://doi.org/10.3390/pathogens10070900
Chicago/Turabian StyleBerthold-Pluta, Anna, Monika Garbowska, Ilona Stefańska, Lidia Stasiak-Różańska, Tamara Aleksandrzak-Piekarczyk, and Antoni Pluta. 2021. "Microbiological Quality of Nuts, Dried and Candied Fruits, Including the Prevalence of Cronobacter spp." Pathogens 10, no. 7: 900. https://doi.org/10.3390/pathogens10070900
APA StyleBerthold-Pluta, A., Garbowska, M., Stefańska, I., Stasiak-Różańska, L., Aleksandrzak-Piekarczyk, T., & Pluta, A. (2021). Microbiological Quality of Nuts, Dried and Candied Fruits, Including the Prevalence of Cronobacter spp. Pathogens, 10(7), 900. https://doi.org/10.3390/pathogens10070900






