2. Results
A search was carried out in various biotopes of clinically healthy cats for promising strains of indigenous microorganisms: candidates for probiotic preparations for the treatment and prevention of surgical infection. The results of the conducted bacteriological analysis are shown in
Table 1.
The data presented in the table indicate that in total, we isolated 214 cultures of microorganisms, most of which (102, 47.7%) were isolated from the contents of the intestinal tract of the studied animals. Most often, E. coli 63 (29.4%), S. saprophyticus 17 (7.9%), S. faecalis 16 (7.5%), and L. acidophilus 11 (5.1%) were isolated from different biotopes of cats.
In total, we isolated 37 (17.3%) strains of indigenous microorganisms, which were isolated only from a biopsy of the skin and samples of feces from the experimental animals. Thus, we most often isolated
L. rhamnosus 6 (11.8%) and
L. acidophilus 4 (7.8%) from the total number of isolated cultures from a skin biopsy.
L. plantarum and
L. acidophilus in seven (6.8%) cultures and
B. bifidum and
L. rhamnosus in four (3.9%) strains of the total number of isolated microorganisms were more often isolated from fecal samples. For a more thorough identification of the isolated
Escherichia coli cultures, their serological typification was carried out, the results of which are shown in
Table 2.
The data presented in the table show that the largest number of E. coli were isolated from fecal samples, namely, 37 (58.7%) microorganisms from the total number of isolated E. coli cultures. It should be noted that the isolated serogroups of E. coli isolates varied significantly among different biotopes of healthy animals. Thus, we most often isolated O111 (three, 30.0%) and O101 and O119 (two each, 20.0%) serogroups of E. coli from a skin biopsy; O116 (four, 28.6%), O113 (three, 21.4%), and O25, O114, and O119 (two each, 14.3%) serogroups from the oral cavity; O83 (eight, 21.7%), and O1, O4, and O22 (six each, 16.2%), and O2 and O18 (four each, 10.8%) serogroups from fecal samples; and O2 and O18 (one each, 50.0%) serogroups from peripheral blood samples of the total number of Escherichia coli cultures. It should be noted that none of the 63 E. coli isolates were classified as lactose-negative or hemolysin-producing E. coli. According to the results of the biological test, it was established that all microorganisms that were isolated from the contents of the oral cavity, feces, skin biopsy, and peripheral blood samples from 18 clinically healthy animals did not cause death of white mice.
Below, we present a detailed analysis of the biological marker properties of candidate cultures for the selection of the most promising strains in probiotic preparations for the treatment and prevention of surgical infections in cats.
First of all, the sensitivity of isolated strains of indigenous bacteria to antimicrobial drugs was determined. The sensitivity of isolated strains of microorganisms of the genus
Bifidobacterium to antibacterial agents is shown in
Figure 1 and
Figure 2. Isolated cultures of
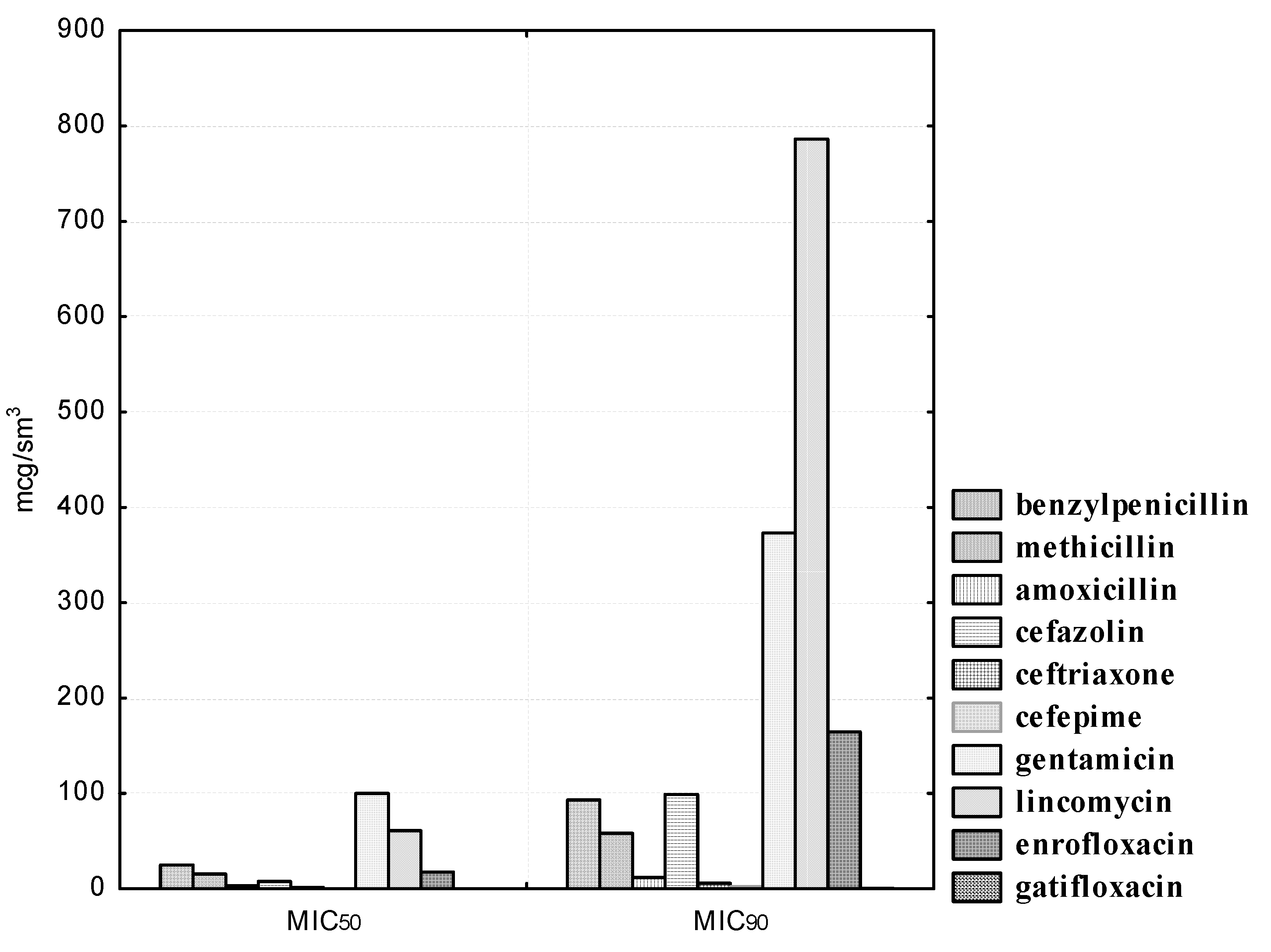
B. bifidum showed the greatest sensitivity to drugs of the β-lactam group (penicillins and cephalosporins). Thus, the growth of four (100.0%)
B. bifidum isolates was suppressed by benzylpenicillin concentrations in the range from 100 to 400; methicillin concentrations were suppressed in the range from 62.5 to 500; the concentration of amoxicillin was suppressed in the range from 12.5 to 25.0; the concentration of cefazolin was suppressed in the range from 12.5 to 50.0; the concentration of ceftriaxone was suppressed in the range from 6.25 to 25.0; cefepime concentrations were suppressed in the range from 1.56 to 3.12 μg/mL.
We noted a high degree of resistance of the isolated B. bifidum cultures in relation to the antibiotic of the aminoglycoside group, gentamicin. Thus, doses of gentamicin in the range from 1.56 to 25.0 μg/mL did not affect the experimental isolates of bifidobacteria; 100.0% of the strains were resistant to the antibiotic. However, a concentration of 50.0 μg/mL inhibited the growth of one (25.0%); a concentration of 100.0 μg/mL, two (50.0%); and a concentration of 200.0 μg/mL, three (75.0%) B. bifidum cultures. The MIC50 of gentamicin to B. bifidum isolates was 100.0, and MIC90 was 373.22 μg/mL.
We noted heterogeneous sensitivity in the isolated cultures of B. bifidum in relation to the antibiotic of the lincosamide group, lincomycin. Thus, doses in the range from 1.56 to 6.25 μg/mL did not affect the experimental strains; the concentration of lincomycin of 12.5 μg/mL inhibited the growth of one bifidobacteria (25.0%); a concentration of 50.0 μg/mL inhibited the growth of two bifidobacteria (50.0%); only the highest dilution of the antibiotic 100.0 μg/mL inhibited the growth of all four isolated cultures of bifidobacteria. The MIC50 of lincomycin against B. bifidum isolates was 60.99, and MIC90 was 786.22 μg/mL. It should be noted that the studied strains of B. bifidum were resistant to an antimicrobial agent from the group of fluorinated quinolones, enrofloxacin. Thus, the concentration of 6.25 μg/mL inhibited the growth of one bifidobacteria (25.0%); a concentration of enrofloxacin 12.5 and 25.0 μg/mL inhibited the growth of two bifidobacteria (50.0%); and three (75.0%) experimental cultures of bifidobacteria were sensitive to the lowest dilution of 50.0 μg/mL. However, none of the B. bifidum strains were resistant to gatifloxacin, a representative of a new generation of fluoroquinolones with anti-anaerobic properties (the MIC50 was 0.12, and the MIC90 was 0.47 μg/mL).
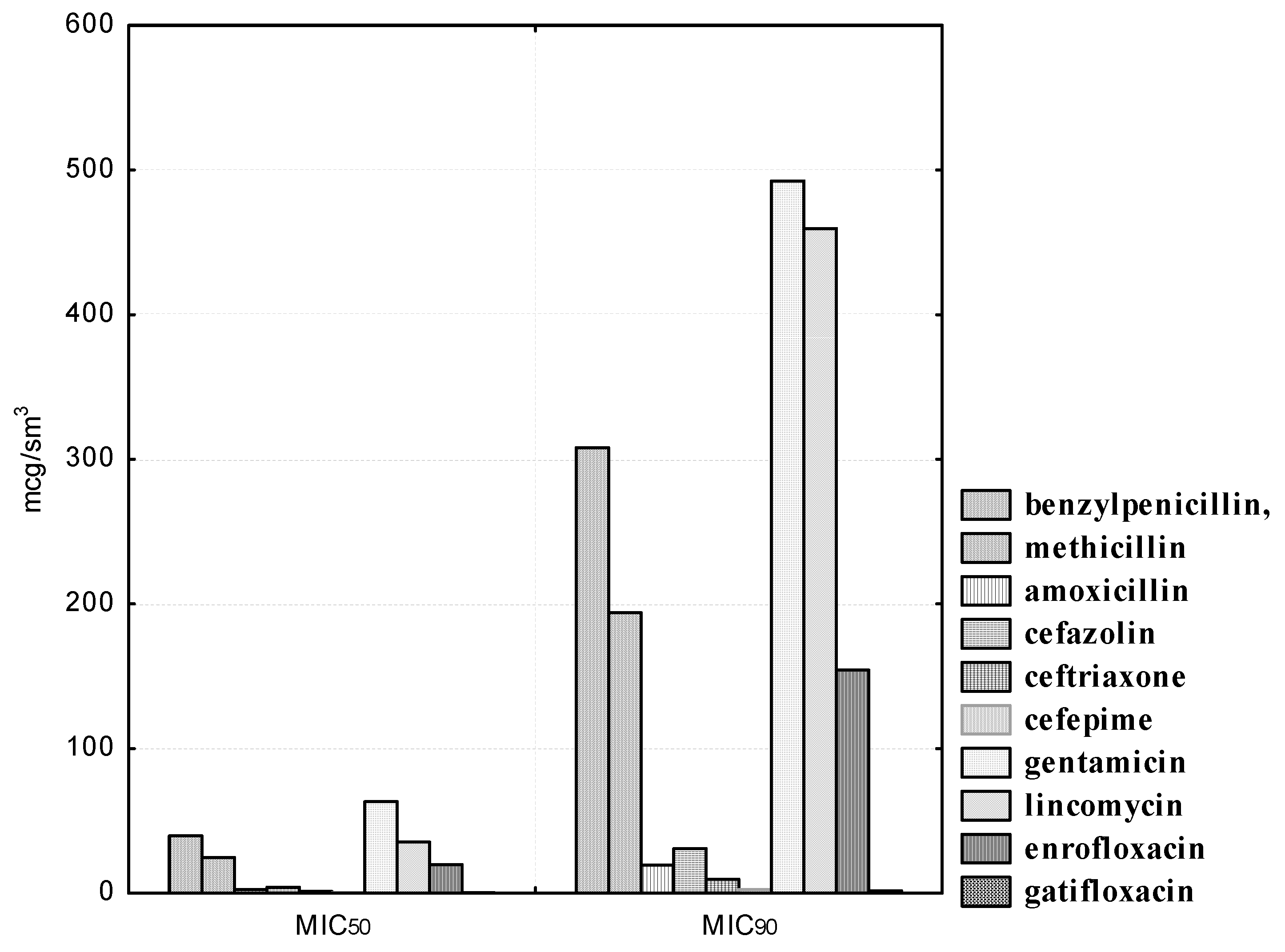
B. adolescentis strains isolated from clinically healthy cats also exhibited the highest sensitivity to antibiotics of the β-lactam group. Thus, the concentration of benzylpenicillin was from 200.0 to 400.0, amoxicillin from 12.5 to 25.0, methicillin from 125.0 to 500.0; cefazolin 12.5 to 50.0, ceftriaxone from 6.25 to 25.0, and cefepime from 1.56 to 3.12 μg/mL inhibited the growth of three (100.0%) experimental cultures of bifidobacteria. In addition, it was found that gatifloxacin was also a highly active agent for experimental isolates of B. adolescentis. The MIC50 of the preparation was 0.19, and the MIC90 was 1.57 μg/mL. It should be noted that B. adolescentis isolates exhibited heterogeneous sensitivity to gentamicin. Thus, all experimental strains were resistant at antibiotic concentrations in the range from 1.56 to 12.5 μg/mL; two (66.6%) isolates were stable at drug dilutions of 25.0 and 50.0 μg/mL; only at a minimal dilution of gentamicin of 200.0 μg/mL were all three isolated cultures of B. adolescentis susceptible to it. The MIC50 of gentamicin was 63.19, and the MIC90 was 492.38 μg/mL. B. adolescentis isolates are highly resistant to lincomycin. Thus, antibiotic concentrations from 1.56 to 6.25 μg/mL did not affect the experimental strains; at drug dilutions of 12.5 and 25.0 μg/mL, the sensitivity was recorded in one (33.3%) of the studied cultures; and the maximum concentrations of lincomycin of 50.0 and 100.0 μg/mL were active against two (66.6%) isolated cultures of bifidobacteria. The MIC50 of the antibiotic was 35.35, and the MIC90 was 459.68 μg/mL. We have also established a high resistance of experimental indigenous microorganisms to enrofloxacin. Thus, at maximum drug dilutions from 0.78 to 6.25 μg/mL, we recorded resistance in three (100.0%) studied strains; at a dilution of 12.5 μg/mL, two (66.6%) cultures of B. adolescentis possessed enrofloxacin resistance; at its maximum concentrations of 25.0 and 50.0 μg/mL, we found sensitivity in two (66.6%) cultures of ‘ifidobacterial. The MIC50 of enrofloxacin was 19.77, and the MIC90 was 154.27 μg/mL.
The antibacterial susceptibility of the isolated
L. acidophilus cultures is shown in
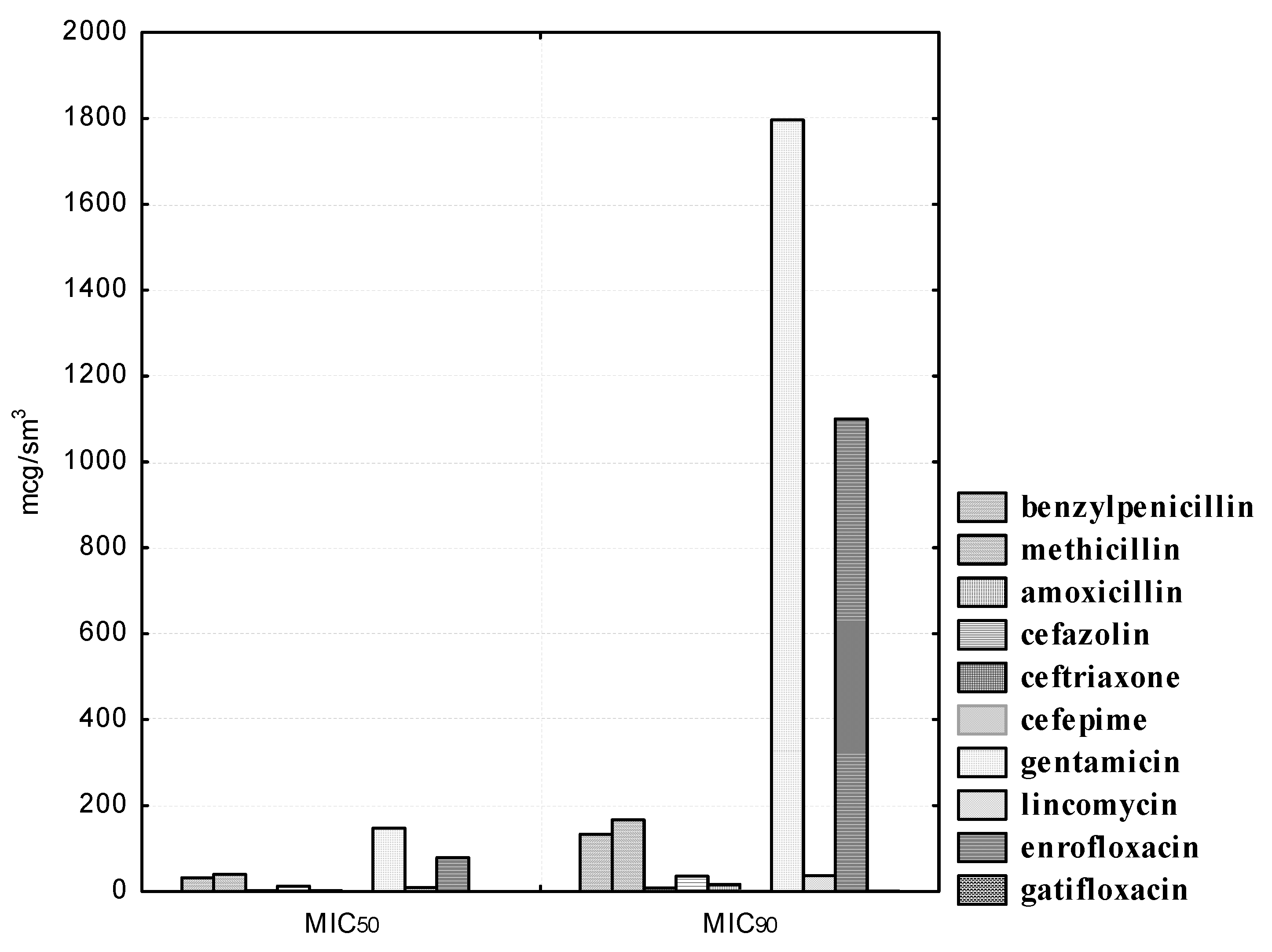
Figure 3.
Experienced strains of lactobacilli exhibit heterogeneous sensitivity to antibiotics of the β-lactam group. Thus, L. acidophilus isolates were susceptible to penicillin (MIC50 was 31.89, and MIC90 was 133.26 μg/mL), methicillin (MIC50 was 39.86, and MIC90 was 166.64 μg/mL), amoxicillin (MIC50 was 1.99, and MIC90 was 8.31 μg/mL), as well as to cefepime (the MIC50 of the drug was 0.26, and the MIC90 was 1.29 μg/mL). However, isolated strains of lactobacilli were resistant to antibiotics of the cephalosporin series I and III, generations of cefazolin and ceftriaxone, respectively. Thus, even at the maximum concentrations of cefazolin (25.0–50.0) and ceftriaxone (12.5–25.0 μg/mL), one (9.1%) strain of lactobacilli, namely, L. acidophilus No. 24, was resistant. The MIC50 of cefazolin was 12.60, and the MIC90 was 35.50 μg/mL. The MIC50 of ceftriaxone was 2.54, and the MIC90 was 18.86 μg/mL.
We also noted a high degree of resistance in the experimental cultures of lactobacilli to gentamicin, lincomycin, and enrofloxacin. Thus, 100.0% of the studied L. acidophilus strains were resistant at gentamicin concentrations ranging from 1.56 to 12.5 μg/mL; nine (81.8%) isolates were resistant at 25.0 μg/mL dilution of the drug; eight (72.7%) microorganisms showed antibiotic resistance at a dilution of 50.0 μg/mL. Even with a minimal dilution of gentamicin of 200.0 μg/mL, only six (54.5%) L. acidophilus isolates were sensitive to the drug. Lincomycin concentrations from 1.56 to 3.12 μg/mL did not affect the experimental strains; at drug dilutions of 6.25 μg/mL, the sensitivity was recorded in four (36.4%) isolated cultures of lactobacilli; at a dilution of 12.5 μg/mL, six (54.5%) isolates were sensitive; at dilutions of 25.0 and 50.0 μg/ml, we registered one (9.1%) strain of lactobacilli, namely, L. acidophilus No. 24, which was resistant to the drug; only at its maximum concentration of 100.0 μg/mL was the antibiotic effective against 100.0% of the studied bacteria. It was found that, at the maximum dilution of enrofloxacin in the range of 0.78–3.12 μg/mL, all 11 (100.0%) isolates showed resistance to the drug; even with the minimum dilutions of the antibiotic at 25.0 and 50.0 μg/mL, seven (63.6%) isolated cultures of L. acidophilus remained resistant. The MIC50 of gentamicin was 147.50, and the MIC90 was 1796.29 μg/mL. The MIC50 of lincomycin was 9.32, and the MIC90 was 36.67 μg/mL. The MIC50 of enrofloxacin was 78.50, and the MIC90 was 1100.19 μg/mL. It should be noted that L. acidophilus strains isolated by us showed sensitivity to gatifloxacin, a representative of a new generation of fluorinated quinolones. The MIC50 of gatifloxacin was 0.20, and the MIC90 was 0.46 μg/mL.
The sensitivity of isolated
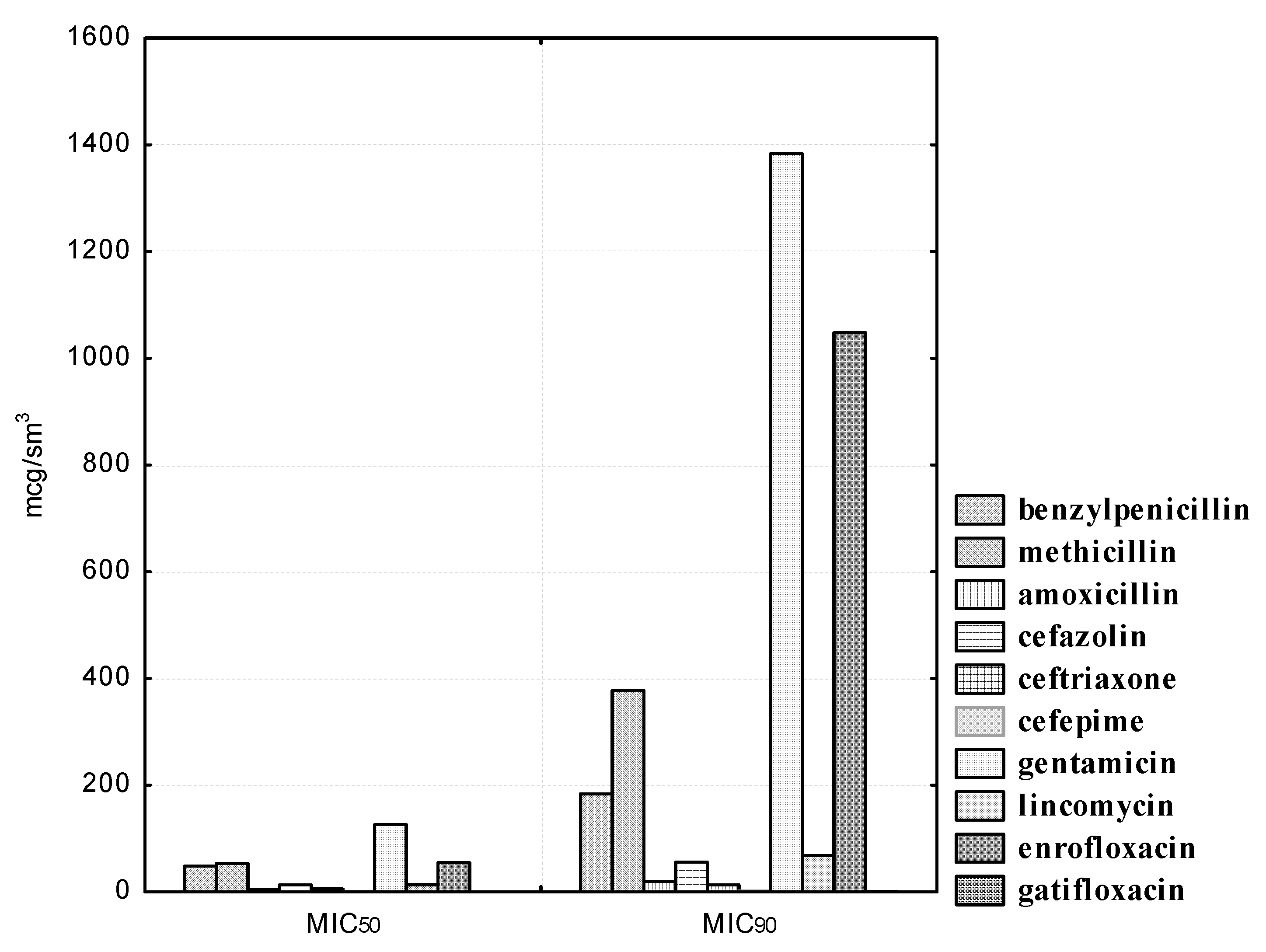
L. plantarum strains to antibacterial agents is reflected in
Figure 4. Isolated
L. plantarum strains showed heterogeneous sensitivity to penicillins. Thus, the experimental cultures were sensitive to methicillin, while antibiotic concentrations in the range from 125.0 to 500.0 μg/mL inhibited the growth of nine (100.0%) isolated bacteria. One (11.1%) strain of lactobacilli, namely,
L. plantarum “Victoria” No. 22, remained resistant to benzylpenicillin at a concentration of 200.0 μg/mL; however, at the highest antibiotic dilution of 400.0 μg/mL, all nine (100.0%) experimental isolates were sensitive to it. It should be noted that resistance to amoxicillin, even at its maximum concentration of 25.0 μg/mL, was established in one (11.1%) isolate, namely, in
L. plantarum “Victoria” No. 22.
We have also established a heterogeneous sensitivity of experienced cultured lactobacilli to cephalosporins. Thus, the concentration of cefazolin in the range from 0.78 to 3.12 μg/mL did not affect all the isolated strains; at a dilution of the drug of 6.25 μg/mL, resistance was found in seven (77.8%) cultures of lactobacteria; when using the antibiotic at 12.5 μg/ml, five (55.5%) isolates were susceptible to it; at a dilution of 25.0 μg/mL, sensitivity was found in six (66.7%) microorganisms; even at a concentration of 50.0 μg/mL, resistance to cefazolin was found in one (11.1%) isolated culture of L. plantarum, namely, L. plantarum “Victoria” No. 22. It should be noted that the L. plantarum “Victoria” No. 22 strain showed resistance to ceftriaxone and cefepime at concentrations of 12.5 and 1.56 μg/mL, respectively. However, the smallest dilutions of ceftriaxone 25.0 and cefepime 3.12 μg/mL inhibited the growth of all nine (100.0%) lactobacilli isolates. The MIC50 of cefazolin was 13.9, and the MIC90 was 55.84 μg/mL. The MIC50 of ceftriaxone was 5.46, and the MIC90 was 13.45 μg/mL. The MIC50 of cefepime was 0.68 and the MIC90 was 1.67 μg/mL.
Experimental cultures of L. plantarum show resistance to lincomycin. Thus, even at the maximum antibiotic concentrations of 50.0 and 100.0 μg/mL, we found resistance in one (11.1%) isolate, namely, L. plantarum “Victoria” No. 22. The MIC50 of lincomycin was 13.79 and the MIC90 was 67.54 μg/mL. We also established a high degree of resistance in the experimental isolates of lactobacilli to gentamicin and enrofloxacin. Thus, gentamicin concentrations in the range from 1.56 to 12.5 μg/mL did not retard the growth of nine (100.0%) isolated cultures of microorganisms; at a dilution of 25.0 μg/mL, we observed resistance in eight (88.9%) strains; at concentrations of 50.0 and 100.0 μg/mL, sensitivity was observed in four (44.4%) isolates; even at a minimal dilution of the antibiotic of 200.0 μg/mL, we found resistance in four (44.4%) isolated cultures of L. plantarum. The maximum enrofloxacin concentrations of 25.0 and 50.0 μg/mL inhibited the growth of five (55.5%) isolated strains of microorganisms. The MIC50 of gentamicin was 126.15, and the MIC90 was 1383.31 μg/mL. The MIC50 of enrofloxacin was 54.87, and the MIC90 was 1048.11 μg/mL. It is necessary to pay attention to the fact that we established a high sensitivity of L. plantarum isolates to gatifloxacin. Thus, the MIC50 of gatifloxacin was 0.13, and the MIC90 was 0.35 μg/mL.
All
L. rhamnosus isolates were susceptible to benzylpenicillin (the MIC
50 of the antibiotic was 55.3, and the MIC
90 was 204.77 μg/mL), ceftriaxone (the MIC
50 of the drug was 3.12, and the MIC
90 was 8.98 μg/mL), and gatifloxacin (the MIC
50 of fluoroquinolone was 0.24, and the MIC
90 was 0.73 μg/mL). Isolated strains of lactobacilli showed heterogeneous sensitivity to cefepime. Thus, cefepime concentrations in the range from 0.04 to 0.18 μg/mL did not retard the growth of all 10 isolated bacterial cultures; at a dilution of 0.39 μg/mL, we observed sensitivity in three (30.0%) isolates; at a concentration of 0.78 μg/mL, the sensitivity was observed in seven (70.0%) isolated microorganisms; at a concentration of 1.56 μg/mL, antibiotic resistance was established in only one (10.0%) strain, namely,
L. rhamnosus No. 26; however, all 10 (100.0%) tested lactobacilli were sensitive to the IV generation cephalosporin at a minimum dilution of 3.12 μg/mL. The MIC
50 of cefepime was 0.56, and the MIC
90 was 1.47 μg/mL. It was found that experimental
L. rhamnosus isolates have resistance to cefazolin. Thus, even at a maximum concentration of the antibiotic of 50.0 μg/mL, we observed resistance to it in two (20.0%) isolated strains, namely,
L. rhamnosus No. 5 and
L. rhamnosus No. 26. The MIC
50 of cefazolin was 14.75, and the MIC
90 was 72.47 μg/mL. In addition, we registered resistance in lactobacilli to amoxicillin. Thus, at the minimum dilutions of the antibiotic of 12.5 and 25.0 μg/mL, resistance was also observed in two (20.0%) experimental isolates, namely,
L. rhamnosus No. 20 and
L. rhamnosus No. 26. The MIC
50 of amoxicillin was 6.80, and the MIC
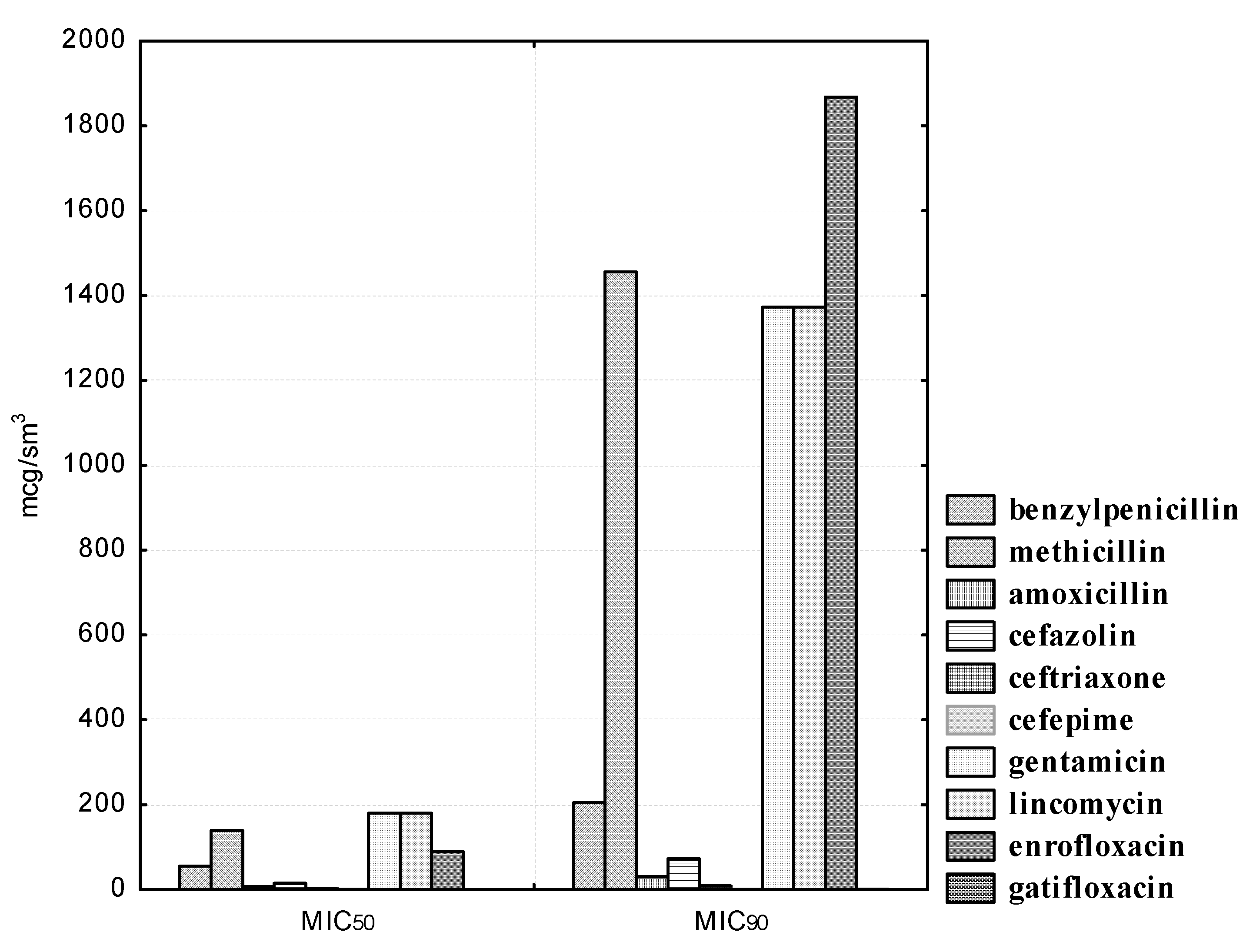
90 was 30.26 μg/mL. A high degree of resistance was also registered in the experimental strains of microorganisms to methicillin, gentamicin, lincomycin, and enrofloxacin. Thus, even at the maximum methicillin concentrations of 250.0 and 500.0 μg/mL, three (30.0%) strains had resistance to it, namely,
L. rhamnosus No. 5,
L. rhamnosus No. 20, and
L. rhamnosus No. 26. The MIC
50 of methicillin was 139.47 and the MIC
90 was 1455.76 μg/mL. At a minimum dilution of gentamicin of 200.0 μg/mL, only five (50.0%) studied cultures of lactobacilli were sensitive to the drug. The MIC
50 of gentamicin was 179.89, and the MIC
90 was 1372.70 μg/mL. At maximum lincomycin concentrations of 50.0 and 100.0 μg/mL, resistance was recorded in six (60.0%) isolated
L. rhamnosus cultures. The MIC
50 of lincomycin was 179.89, and the MIC
90 was 1372.70 μg/mL. The maximum enrofloxacin concentration of 50.0 μg/mL also had no effect on six (60.0%) isolates of the experimental microorganisms. The MIC
50 of enrofloxacin was 89.71, and the MIC
90 was 1867.97 μg/mL (see
Figure 5).
Thus, the cultures of bifidobacteria and lactobacilli isolated by us revealed a high sensitivity to antibiotics of the β-lactam group (with the exception of L. acidophilus No. 24, L. plantarum “Victoria” No. 22, L. rhamnosus No. 5, L. rhamnosus No. 20, and L. rhamnosus No. 26, which showed significant variability in sensitivity to antibacterial drugs of this group, which indicates the great potential of these microorganisms) and resistance to aminoglycosides, lincosamides, and fluoroquinolones (with the exception of gatifloxacin, which showed high efficacy against all lactic acid microorganisms).
Furthermore, the adhesive properties of the probiotic microbiota were determined. The results of determining the adhesive properties of strains of bacteria of the genus
Bifidobacterium isolated from clinically healthy cats are presented in
Table 3.
It can be seen from the data presented that the strains of bifidobacteria isolated by us had an average adhesive activity, since the AAI to erythrocytes of cats in the sample was 2.56 (from 1.56 to 4.24). It should be noted that the majority of the representatives of bifidoflora (four, 57.1%) isolates showed an average ability of adhesion, and only one (14.3%) isolated strain, namely,
B. adolescentis No. 23, had highly adhesive properties, while the AAR index was 4.24 ± 0.41 tank./er.; K was 88.0%; and MAI was 4.81. The results of determining the adhesive properties of the isolated strains of bacteria of the
L. plantarum species are shown in
Table 4.
The presented data indicate that all L. plantarum isolates exhibit high and medium adhesion capacity, namely, five (55.6%) and four (44.4%) isolated strains, respectively. The AAI indicator, which in the sample is 4.40 (from 2.24 to 8.28), also indicates the highly adhesive properties of the isolated cultures of lactobacilli. It should be noted that L. plantarum No. 8 had the highest adhesive activity; the AAR index was 5.32 ± 0.26 bact/er., and for L. plantarum “Victoria” No. 22, the AAR index was 8.28 ± 0.17 bact/er., and the K index for these bacteria in both cases was 100.0%. It should be noted that we recorded high MAI values in these isolates, namely, for L. plantarum No. 8 it was 5.32, and for L. plantarum “Victoria” No. 22 it was 8.28. At the same time, in the tested cultures of L. plantarum No. 8 and L. plantarum “Victoria” No. 22, we also observed the highest IR indices: 33 (24.8%) and 43 (20.3%) bacteria in the same field of view of the microscope, respectively.
The results of determining the adhesive properties of
L. rhamnosus bacterial strains isolated from clinically healthy cats are reflected in
Table 5.
The data in
Table 5 indicate that only two (20.0%)
L. rhamnosus cultures were classified as low-adhesive strains, while the rest of the microorganisms had medium and high adhesion capacity: four (40.0%) isolates each. The AAI index to erythrocytes of cats in the sample was 3.47 (from 1.36 to 6.16), which is regarded as the ability of adhesion in the isolated cultures of medium-grade microorganisms. The data given in the table also indicate that the
L. rhamnosus No. 6 strains had the greatest adhesive activity; at the same time, the SPA indicator was 5.04 ± 0.40 tank/er., the K indicator was 92.0%, and the MAI indicator was 5.47. For
L. rhamnosus No. 26, the AAR index was 6.16 ± 0.51 baht/er., the K index was 88.0%, and the MAI index was 7.00. It should be noted that the IR index for these cultures was 19 (15.1%) and 36 (23.4%) bacteria in one microscope field of view, respectively.
The results of determining the adhesive properties of bacterial strains of the
L. acidophilus species are shown in
Table 6.
These data indicate that all L. acidophilus strains isolated from clinically healthy cats were capable of adhesion and were characterized by moderate to high adhesion. Thus, eight (72.7%) isolated cultures of lactoflora had highly adhesive properties, and the last three (27.3%) strains had an average ability to adhere. The AAI index in the sample of isolated cultures of microorganisms is 4.49 (from 3.72 to 6.24), which is regarded as a high adhesion capacity. We observed the highest rates of adhesive activity in L. acidophilus strain No. 12; the AAR index was 5.00 ± 0.33 bact/er., the K index was 96.0%, and the MAI index was 5.20. For L. acidophilus No. 24 isolate, the AAR indicator was 6.24 ± 0.46 bact/er., the K indicator was 92.0%, and the MAI indicator was 6.78. It should be noted that the IR index for these cultures was 29 (23.2%) and 33 (21.1%) bacteria in one microscope field of view, respectively.
The results of the study made it possible to reveal one of the main mechanisms of the protective action of the normal flora isolated by us, namely, their adhesive activity. It has been proven that the adhesive properties of the isolated lactobacteria and bifidobacteria are quite variable, even within the same species. It was shown that the highest adhesive activity was for the Bifidobacterium strain B. adolescentis No. 23 genus, as well as the Lactobacillus strains L. plantarum No. 8, L. plantarum “Victoria” No. 22, L. rhamnosus No. 6, L. rhamnosus No. 26, L. acidophilus No. 12, and L. acidophilus No. 24 genus.
The characteristics of the antagonistic activity of the isolated strains of indigenous bacteria to the main causative agents of surgical infection in cats are presented in
Table 7. The data obtained indicate that antagonistic properties were inherent to varying degrees in all experimental strains of probiotic microorganisms, but the most active were representatives of the genera
B. adolescentis (
n = 3),
L. plantarum (
n = 9), and
L. acidophilus (
n = 11). Thus, 60 (23.5%), 197 (25.8%), and 291 (31.1%) cultures were sensitive to the isolates of these genera, and 170 (66.7%), 415 (54.2%), and 484 (51.8%) test cultures of microorganisms were highly sensitive, respectively, from the total number of test reactions performed. It should be noted that the lowest inhibitory activity was observed in representatives of the
B. bifidum genus (
n = 4) and
L. rhamnosus genus (
n = 10). Thus, 129 (37.9%) and 126 (14.8%) cultures were insensitive to the strains of these genera, and 105 (30.9%) and 142 (16.7%) test cultures, respectively, of the total number of test reactions performed were sensitive.
For greater clarity of the obtained material, as well as for a comparative analysis of the antagonistic activity of individual strains of indigenous bacteria within each species, we present the results obtained in separate tables. The results of determining the antagonistic activity of the isolated strains of bacteria of the
Bifidobacterium genus to the main causative agents of surgical infection in cats are shown in
Table 8.
It was found that the strains of the B. adolescentis species, namely, B. adolescentis No. 1, B. adolescentis No. 23, and B. adolescentis No. 28, were the most capable of eliminating both Gram-positive and Gram-negative microflora.
The results of determining the antagonistic activity of
L. plantarum on the causative agents of purulent–inflammatory processes in cats are shown in
Table 9.
The cultures of
L. plantarum No. 3,
L. plantarum No. 8, and
L. plantarum No. 27 also demonstrated relatively high growth retardation zones. Thus,
L. plantarum isolate
No. 3 almost completely suppressed the growth of
S. intermedius (growth delay zone—30.60 ± 1.07 mm),
S. pyogenes (growth delay zone—32.00 ± 0.70 mm), and
K. pneumoniae (growth delay zone—29.60 ± 0.50 mm);
L. plantarum strain
No. 8 actively inhibited
S. pyogenes cultures (growth delay zone—30.00 ± 0.31 mm) and
Escherichia coli cultures (growth delay zone in the range of 18.40 ± 0.50 mm up to 26.20 ± 0.58 mm);
L. plantarum isolate
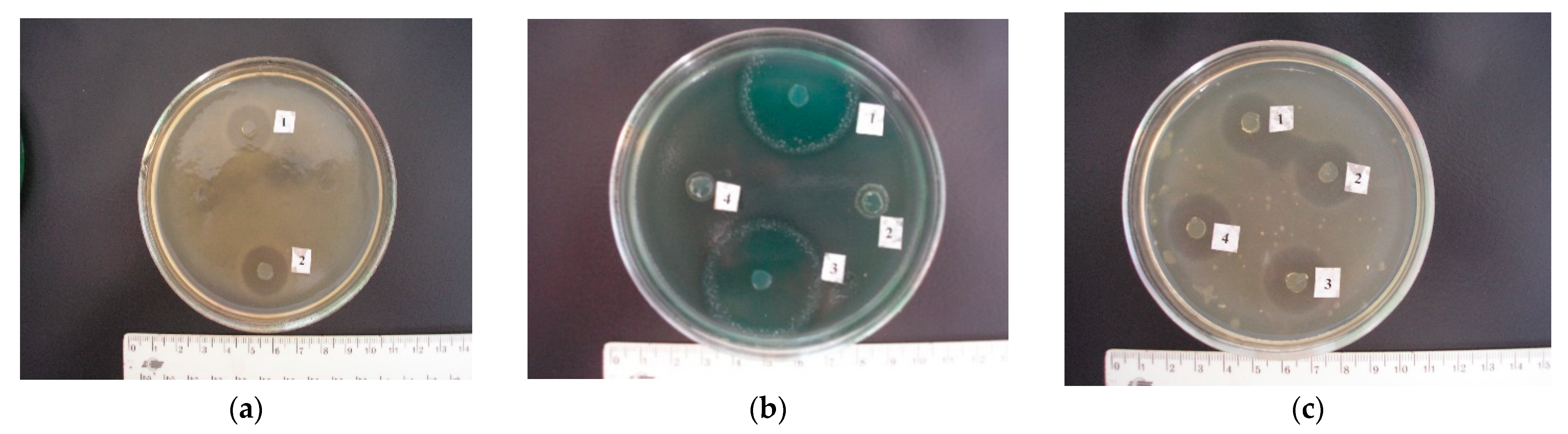
No. 27 also actively eliminated both Gram-positive (growth delay zone in the range from 8.60 ± 0.50 to 20.80 ± 0.86 mm) and Gram-negative (growth delay zone in the range from 6.60 ± 0.50 to 30.40 ± 2.83 mm) microflora (
Figure 6a,b). At the same time, the IIAA for these cultures was 20.8, 19.9, and 15.5 mm, respectively.
The least antagonistic activity was recorded in the probiotic
L. plantarum No. 25 strain on
S. uberis (growth inhibition zone—2.40 ± 0.50 mm),
E. coli O18 (growth inhibition zone—2.80 ± 0.37 mm),
P. aeruginosa (growth inhibition zone—3.60 ± 0.50 mm),
E. aerogenes (growth inhibition zone—1.00 ± 0.44 mm),
P. vulgaris (growth inhibition zone—2.40 ± 0.50 mm),
P. mirabilis (growth inhibition zone—2.00 ± 0.31 mm), and
C. albicans (growth retardation zone—2.80 ± 0.37 mm) (see
Figure 6b.). Only
S. intermedius, S. pyogenes, and
E. coli O111 exhibited moderate sensitivity to it (the growth retardation zone in these cultures ranged from 11.60 ± 0.74 to 13.00 ± 0.54 mm). The IIAA index for
L. plantarum No. 25 was 6.2 mm.
The results of determining the antagonistic activity of isolated strains of
L. rhamnosus bacteria on the main pathogens of purulent–inflammatory processes in cats are reflected in
Table 10.
The research results given in the table indicate that the
L. rhamnosus No. 26 strain was the most capable of eliminating both Gram-positive and Gram-negative microflora. Thus, this isolate inhibited the growth of coccal microflora in the range from 20.40 ± 0.50 to 30.00 ± 0.70 mm;
Escherichia coli in the range from 23.40 ± 0.50 to 29.20 ± 0.86 mm (
Figure 6c);
Pseudomonas aeruginosa—31.40 ± 1.20 mm;
Klebsiella—37.40 ± 1.32 mm;
Enterobacteriaceae—27.00 ± 1.00 mm; citrobacters—15.40 ± 1.07 mm; proteus from 14.20 ± 1.49 to 16.80 ± 0.58 mm, as well as fungi of the
Candida genus—22.20 ± 0.86 mm. In this case, the IIAA index was 24.6 mm. We also observed a relatively high antagonistic activity in the
L. rhamnosus No. 6,
L. rhamnosus No. 29 and
L. rhamnosus No. 30 strains. Thus, the listed isolates inhibited the growth of Gram-positive microflora in the range from 16.40 ± 0.50 to 28.40 ± 0.92; Gram-negative bacteria, from 6.00 ± 0.70 to 32.60 ± 1.24 (see
Figure 6c); and fungi of the
Candida genus, from 16.00 ± 0.70 to 17.00 ± 0.70 mm. At the same time, the IIAA index for these strains was 22.4, 21.5, and 19.7 mm, respectively. It should be noted that L. rhamnosus No. 33 isolate shows the least antagonistic properties to
S. intermedius, S. epidermidis, S. faecalis, P. aeruginosa, K. pneumoniae, E. aerogenes, C. freundii, P. vulgaris, P. mirabilis, and
C. albicans; the growth retardation zones were 4.60 ± 0.51, 4.40 ± 0.50, 3.60 ± 0.50, 1.00 ± 0.54, 4.60 ± 0.50, 3.60 ± 0.50, 0, 0, 3.40 ± 0.50, and 1.80 ± 0.58 mm, respectively; and the AIAA index was 5.3 mm.
The results of determining the antagonistic activity of the isolated strains of bacteria of the
L. acidophilus species against the main causative agents of surgical infections in cats are presented in
Table 11.
These data indicate that among the bacteria of the L. acidophilus species isolated by us, the L. acidophilus No. 24 strain had the greatest antagonistic activity. Thus, it actively inhibited the growth of Gram-positive (in the range from 20.20 ± 0.66 to 37.00 ± 1.64 mm) and Gram-negative (in the range from 17.40 ± 1.40 to 37.60 ± 0.92 mm) microflora, as well as representatives of C. albicans (18.20 ± 1.28 mm). At the same time, the IIAA indicator for it was 26.8 mm. We noted a relatively high intensity of antagonistic activity in the L. acidophilus No. 12 isolate, which eliminated both Gram-positive (in the range from 19.60 ± 0.50 to 33.20 ± 2.03 mm) and Gram-negative (in the range from 12.20 ± 1.28 to 29.60 ± 1.72 mm) microflora, as well as fungi of the genus Candida (14.80 ± 0.86 mm), while the IIAA for it was 21.4 mm. It should be noted that L. acidophilus No. 14, L. acidophilus No. 16, and L. acidophilus No. 21 had the least antagonistic properties. Thus, the claimed isolates moderately suppressed the growth of Gram-positive (in the range from 6.60 ± 0.50 to 12.60 ± 1.16 mm) and Gram-negative (in the range from 5.40 ± 0.92 to 13.80 ± 1.28 mm) bacteria, as well as representatives of C. albicans (in the range from 5.20 ± 0.58 to 5.80 ± 0.86 mm). In this case, the IIAA for them was equal to 9.8, 9.3, and 9.1 mm, respectively.
Thus, in order to select the most promising strains of probiotic microflora, we analyzed the indicators of the intensity of antagonistic activity, which were determined on the basis of comparing the values of the growth inhibition zones of pathogens of purulent–inflammatory processes in cats to candidate cultures. It was shown that among the tested probiotic strains, L. plantarum “Victoria” No. 22, L. rhamnosus No. 26, and L. acidophilus No. 24 had the most pronounced antagonistic properties in which the IIAA indices were 26.0, 24.6, and 26.8 mm, respectively.
We also conducted studies to determine the correlation between the adhesive and antagonistic properties of strains of lactic acid microorganisms isolated from clinically healthy cats. It was found that the adhesive activity of isolated bifidobacteria and lactobacilli clearly correlates with their antagonistic properties (г = 0.9; p < 0.001 and г = 0.88; p < 0.001, respectively).
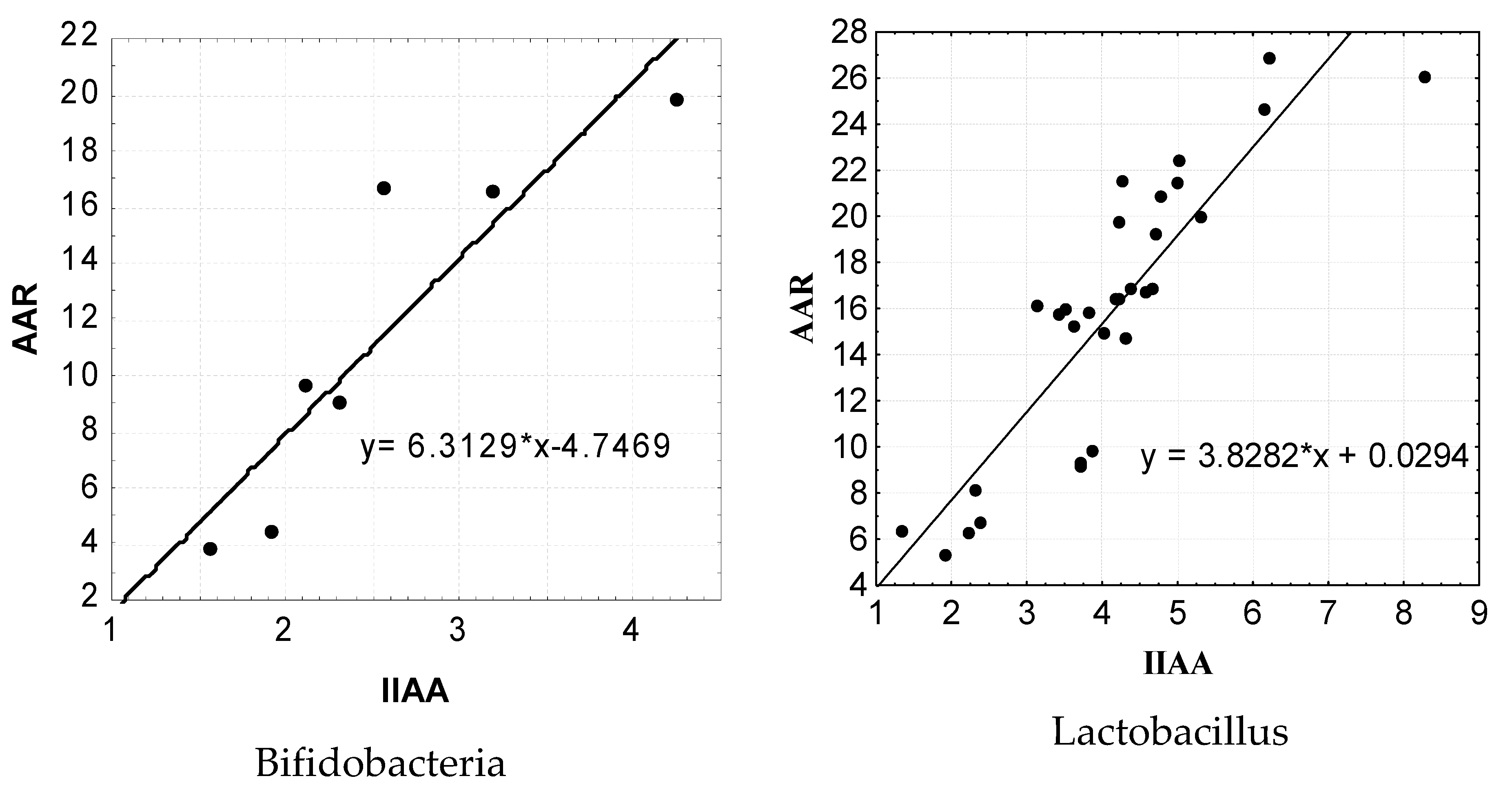
For a more detailed statistical analysis of the dependence of the level of the antagonistic activity of isolated lactic acid microorganisms on the level of their adhesive properties, we applied a regression analysis using a linear function, the results of which are shown in
Figure 7.
The results of the performed regression analysis show that there is a clear correlation between the AAR level and the IIAA level. Thus, the regression equation for the indicators of AAR and IIAA in isolated bifidobacteria has the following form: y = 6.3129 * x −4.7469. It should be noted that the regression equation of the marker indices of AAR and IIAA in the isolated strains of lactobacilli also showed the presence of a rather significant pattern, which had the following form: y = 3.8282 * x +0.0294. It should be noted that the calculation of the reliability of certain mathematical models of the AAR levels in the representatives of bifidoflora and lactoflora isolated by us from the IIAA indicators showed the maximum level (p < 0.001).
In the future, to determine the severity of the antagonistic activity of industrial strains of lactic acid bacteria, as well as to select the concentration of lactobacilli in probiotic preparations, we developed a more accurate method for determining the level of quantitative antagonistic activity of lactic acid microorganisms, in which the method of serial dilution of the probiotic strain in MRS-2 is used, and sowing of test cultures is carried out on a two-layer solid nutrient medium. The results of determining the quantitative antagonistic activity of industrial strains of lactobacilli against the main causative agents of surgical infection in cats are shown in
Table 12.
It can be seen from the above data that the studied L. plantarum “Victoria” No. 22, L. rhamnosus No. 26, and L. acidophilus No. 24 strains had the highest quantitative antagonistic activity in relation to cultures of S. pyogenes and P. aeruginosa; namely, the MIC of these were 2.61 ± 0.04 log, 3.02 ± 0.08 log, 2.49 ± 0.04 log and 2.43 ± 0.03 log, 2.87 ± 0.07 log, 2.46 ± 0.03 log CFU/cm3, respectively. It should be noted that the studied strains of lactobacilli had high antimicrobial activity against all representatives of Gram-positive microflora. Thus, the MIC values in the experimental isolates of L. plantarum “Victoria” No. 22, L. rhamnosus No. 26, and L. acidophilus No. 24 to test cultures of staphylococci and streptococci ranged from 2.49 ± 0.04 lg to 4.96 ± 0.06 lg CFU/cm3. However, it was found that industrial strains of lactobacilli have an unequal ability to influence the growth and viability of representatives of Gram-negative microflora. At the same time, MIK values varied over a wider range, namely, from 2.43 ± 0.03 log to 5.86 ± 0.09 log CFU/cm3. It was shown that the investigated L. plantarum “Victoria” No. 22, L. rhamnosus No. 26, and L. acidophilus No. 24 strains have relatively low antifungal activity; the MIKA indices for C. albicans test cultures were 6.47 ± 0.10 lg, 6.74 ± 0.08 lg, and 6.44 ± 0.06 lg CFU/cm3, respectively. Thus, our proposed method for determining the level of antagonistic activity of lactic acid microorganisms makes it possible to carry out not only a qualitative but also a quantitative assessment of their antagonistic activity.