Occurrence of Listeria monocytogenes in Artisanal Cheeses from Poland and Its Identification by MALDI-TOF MS
Abstract
1. Introduction
2. Results
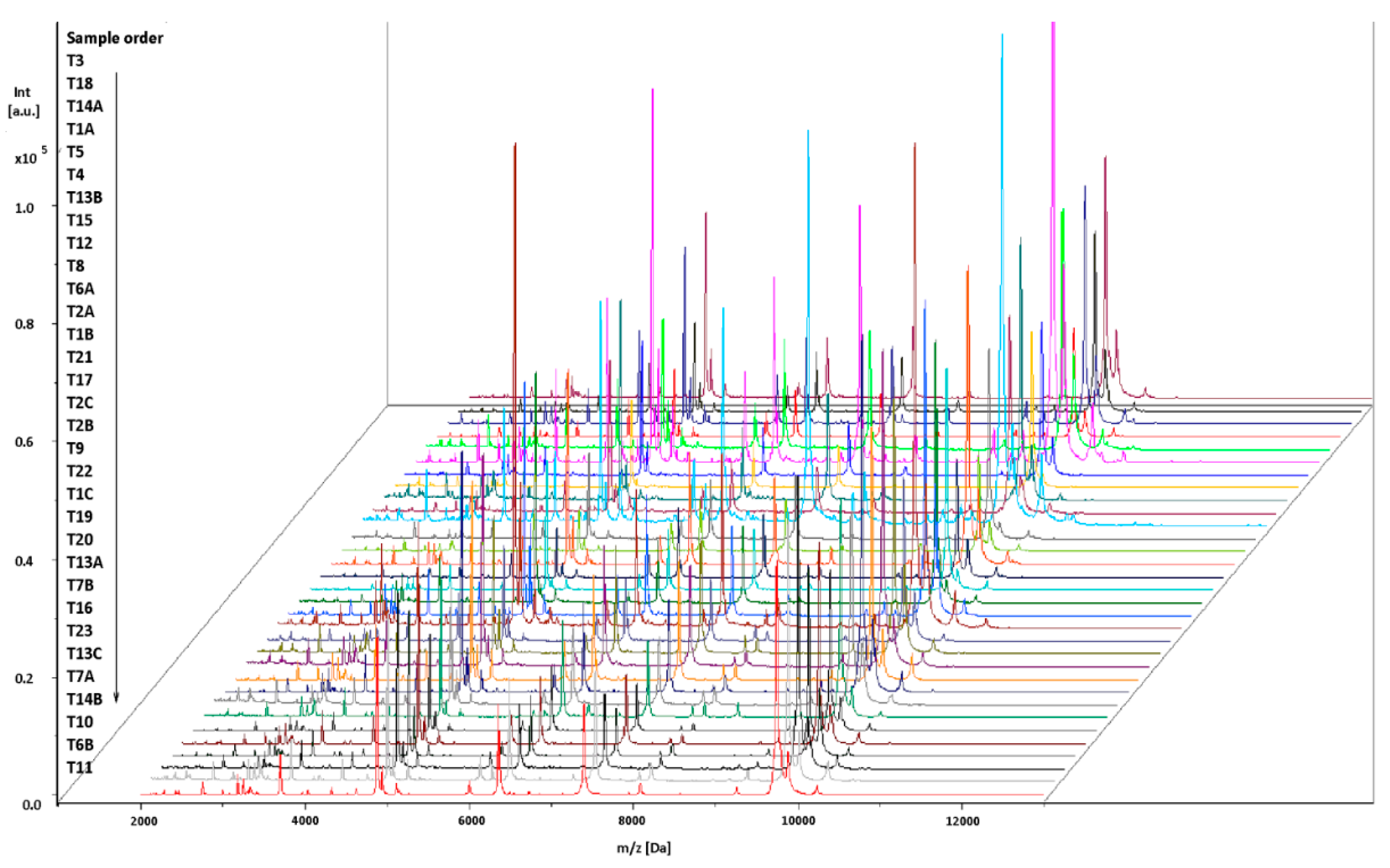
2.1. Bacteriological Analysis and Identification by MALDI-TOF MS
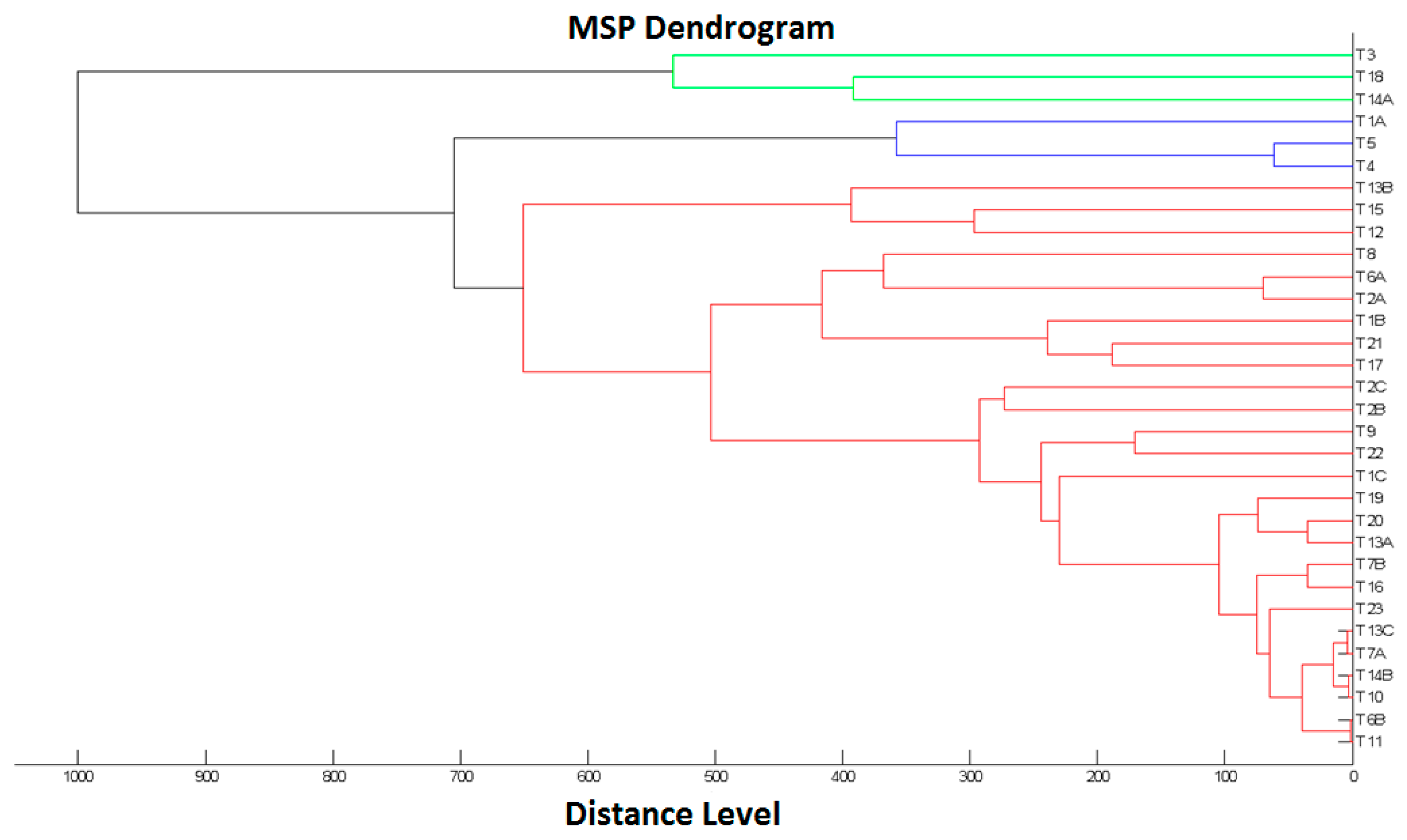
2.2. Dendrogram for the L. monocytogenes Isolates
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Bacteriological Analysis
4.3. Identification of L. monocytogenes Isolates by MALDI-TOF MS
4.4. Dendrogram Construction for L. monocytogenes Isolates
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- EFSA and ECDC (European Food Safety Authority and European Centre for Disease Prevention and Control). The European Union one health 2018 zoonoses report. EFSA J. 2019, 17, e05926. [Google Scholar] [CrossRef]
- Bille, J.; Blanc, D.S.; Schmid, H.; Boubaker, K.; Baumgartner, A.; Siegrist, H.H.; Tritten, M.L.; Lienhard, R.; Berner, D.; Anderau, R.; et al. Outbreak of human listeriosis associated with tomme cheese in northwest Switzerland, 2005. Eurosurveillance 2006, 11, 11–12. [Google Scholar] [CrossRef]
- Choi, M.J.; Jackson, K.A.; Medus, C.; Beal, J.; Rigdon, C.E.; Cloyd, T.C.; Forstner, M.J.; Ball, J.; Bosch, S.; Bottichio, L.; et al. Notes from the field: Multistate outbreak of listeriosis linked to soft-ripened cheese—United States, 2013. Morb. Mortal. Wkly. Rep. 2014, 63, 294–295. [Google Scholar]
- Jackson, K.A.; Biggerstaff, M.; Tobin-D’Angelo, M.; Sweat, D.; Klos, R.; Nosari, J.; Garrison, O.; Boothe, E.; Saathoff-Huber, L.; Hainstock, L.; et al. Multistate outbreak of Listeria monocytogenes associated with mexican-style cheese made from pasteurized milk among pregnant, hispanic women. J. Food Prot. 2011, 74, 949–953. [Google Scholar] [CrossRef] [PubMed]
- Jackson, K.A.; Gould, L.H.; Hunter, J.C.; Kucerova, Z.; Jackson, B. Listeriosis outbreaks associated with soft cheeses, United States, 1998–20141. Emerg. Infect. Dis. 2018, 24, 1116–1118. [Google Scholar] [CrossRef]
- Koch, J.; Dworak, R.; Prager, R.; Becker, B.; Brockmann, S.; Wicke, A.; Wichmann-Schauer, H.; Hof, H.; Werber, D.; Stark, K. Large listeriosis outbreak linked to cheese made from pasteurized milk, Germany, 2006–2007. Foodborne Pathog. Dis. 2010, 7, 1581–1584. [Google Scholar] [CrossRef] [PubMed]
- McIntyre, L.; Wilcott, L.; Naus, M. Listeriosis outbreaks in British Columbia, Canada, caused by soft ripened cheese contaminated from environmental sources. BioMed. Res. Int. 2015, 2015, 131623. [Google Scholar] [CrossRef] [PubMed]
- Margalho, L.P.; Feliciano, M.D.; Silva, C.E.; Abreu, J.S.; Piran, M.V.F.; Sant’Ana, A.S. Brazilian artisanal cheeses are rich and diverse sources of nonstarter lactic acid bacteria regarding technological, biopreservative, and safety properties—Insights through multivariate analysis. J. Dairy Sci. 2020, 103, 7908–7926. [Google Scholar] [CrossRef]
- Aldalur, A.; Bustamante, M.Á.; Barron, L.J.R. Characterization of curd grain size and shape by 2-dimensional image analysis during the cheesemaking process in artisanal sheep dairies. J. Dairy Sci. 2019, 102, 1083–1095. [Google Scholar] [CrossRef] [PubMed]
- Johler, S.; Macori, G.; Bellio, A.; Acutis, P.; Gallina, S.; Decastelli, L. Short communication: Characterization of staphylococcus aureus isolated along the raw milk cheese production process in artisan dairies in Italy. J. Dairy Sci. 2018, 101, 2915–2920. [Google Scholar] [CrossRef]
- Meng, Z.; Zhang, L.; Xin, L.; Lin, K.; Yi, H.; Han, X. Technological characterization of Lactobacillus in semihard artisanal goat cheeses from different Mediterranean areas for potential use as nonstarter lactic acid bacteria. J. Dairy Sci. 2018, 101, 2887–2896. [Google Scholar] [CrossRef]
- Carrascosa, C.; Millán, R.; Saavedra, P.; Jaber, J.R.; Raposo, A.; Sanjuán, E. Identification of the risk factors associated with cheese production to implement the hazard analysis and critical control points (HACCP) system on cheese farms. J. Dairy Sci. 2016, 99, 2606–2616. [Google Scholar] [CrossRef] [PubMed]
- Mata, G.M.S.C.; Martins, E.; Machado, S.G.; Pinto, M.S.; De Carvalho, A.F.; Vanetti, M.C.D. Performance of two alternative methods for Listeria detection throughout Serro Minas cheese ripening. Braz. J. Microbiol. 2016, 47, 749–756. [Google Scholar] [CrossRef] [PubMed]
- European Commission. Commission Regulation (EC) No 2073/2005 of 15 November 2005 on Microbiological Criteria for Foodstuffs. Off. J. Eur. Union 2005, 338, 1–26. [Google Scholar]
- European Commission. Regulation (EC) No 178/2002 of the European Parliament and of the Council of 28 January 2002 Laying down the General Principles and Requirements of Food Law, Establishing the European Food Safety Authority and Laying down Procedures in Matters of Food Safety. Off. J. Eur. Union 2002, 31, 1–24. [Google Scholar]
- Barbuddhe, S.B.; Maier, T.; Schwarz, G.; Kostrzewa, M.; Hof, H.; Domann, E.; Chakraborty, T.; Hain, T. Rapid identification and typing of listeria species by matrix-assisted laser desorption ionization-time of flight mass spectrometry. Appl. Environ. Microbiol. 2008, 74, 5402–5407. [Google Scholar] [CrossRef] [PubMed]
- Thouvenot, P.; Vales, G.; Bracq-Dieye, H.; Tessaud-Rita, N.; Maury, M.M.; Moura, A.; Lecuit, M.; Leclercq, A. MALDI-TOF mass spectrometry-based identification of Listeria species in surveillance: A prospective study. J. Microbiol. Methods 2018, 144, 29–32. [Google Scholar] [CrossRef]
- De Koster, C.G.; Brul, S. MALDI-TOF MS identification and tracking of food spoilers and food-borne pathogens. Curr. Opin. Food Sci. 2016, 10, 76–84. [Google Scholar] [CrossRef]
- Pusztahelyi, T.; Szabó, J.; Dombrádi, Z.; Kovács, S.; Pócsi, I. Foodborne Listeria monocytogenes: A real challenge in quality control. Scientifica 2016, 2016, 5768526. [Google Scholar] [CrossRef][Green Version]
- European Food Safety Authority and European Centre for Disease Prevention and Control (EFSA and ECDC). The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2017. EFSA J. 2018, 16, e05500. [Google Scholar] [CrossRef]
- Kevenk, T.O.; Gulel, G.T. Prevalence, antimicrobial resistance and serotype distribution of Listeria monocytogenes isolated from raw milk and dairy products. J. Food Saf. 2016, 36, 11–18. [Google Scholar] [CrossRef]
- Arslan, S.; Özdemir, F. Prevalence and antimicrobial resistance of Listeria spp. in homemade white cheese. Food Control 2008, 19, 360–363. [Google Scholar] [CrossRef]
- Barría, C.; Singer, R.S.; Bueno, I.; Estrada, E.; Rivera, D.; Ulloa, S.; Fernández, J.; Mardones, F.O.; Moreno-Switt, A.I. Tracing Listeria monocytogenes contamination in artisanal cheese to the processing environments in cheese producers in southern chile. Food Microbiol. 2020, 90, 103499. [Google Scholar] [CrossRef]
- Hunt, K.; Drummond, N.; Murphy, M.; Butler, F.; Buckley, J.; Jordan, K. A case of bovine raw milk contamination with Listeria monocytogenes. Ir. Vet. J. 2012, 65, 13. [Google Scholar] [CrossRef]
- Kroplewska, M.; Paluszak, Z.; Breza-Boruta, B.; Marosz, A. The study on the survival skills of Listeria monocytogenes inhabit-ing a food processing plant. Res. Pap. Wrocław Univ. Econ. 2017, 494, 114–122. [Google Scholar]
- Mansouri-Najand, L.; Kianpour, M.; Sami, M.; Jajarmi, M. Prevalence of Listeria monocytogenes in raw milk in Kerman, Iran. Vet. Res. Forum Int. 2015, 6, 223–226. [Google Scholar]
- Tahoun, A.B.; Elez, R.M.A.; Abdelfatah, E.N.; Elsohaby, I.; El-Gedawy, A.A.; Elmoslemany, A.M. Listeria monocytogenes in raw milk, milking equipment and dairy workers: Molecular characterization and antimicrobial resistance patterns. J. Glob. Antimicrob. Resist. 2017, 10, 264–270. [Google Scholar] [CrossRef]
- Culak, R.A.; Fang, M.; Simon, S.B.; Dekio, I.; Rajakaruna, L.K.; Shah, H.N. Changes in the matrix markedly enhance the resolution and accurate identification of human pathogens by MALDI-TOF MS. J. Anal. Bioanal. Tech. 2014, S2, 2. [Google Scholar] [CrossRef]
- PN-EN ISO 11290-1:1999. Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes—Part 1: Detection Method; PKN: Warszawa, Poland, 1999.
- PN-EN ISO 11290-1: 2017-07. Microbiology of the Food Chain—Horizontal Method for the Detection and Enumeration of Listeria monocytogenes and of Listeria spp.—Part 1: Detection Method; PKN: Warszawa, Poland, 2017.
- Sauer, S.; Freiwald, A.; Maier, T.; Kube, M.; Reinhardt, R.; Kostrzewa, M.; Geider, K. Classification and identification of bacteria by mass spectrometry and computational analysis. PLoS ONE 2008, 3, e2843. [Google Scholar] [CrossRef]
| MALDI–TOF MS. | |||||
|---|---|---|---|---|---|
| Production facilities | Year of sampling | Sample code | Score values | Species ID according to MALDI Biotyper 3.1 | |
| G1 | 2014 | 1A | 2.041–2.272 | L. monocytogenes | DSM 20600T DSM |
| G1 | 2014 | 1B | 2.028–-2.055 | L. monocytogenes | DSM 20600T DSM |
| G1 | 2015 | 1C | 2.087–2.193 | L. monocytogenes | Mb 19348_1 CHB |
| D4 | 2014 | 16 | 1.803–2.036 | L. monocytogenes | Mb 19348_1 CHB |
| J | 2015 | 17 | 2.05–2.086 | L. monocytogenes | Mb 19348_1 CHB |
| I | 2015 | 2A | 1.915–2.141 | L. monocytogenes | DSM 20600T DSM |
| I | 2015 | 2B | 1.935–2.033 | L. monocytogenes | DSM 20600T DSM |
| I | 2015 | 2C | 2.033–2.152 | L. monocytogenes | Mb 19348_1 CHB |
| I | 2016 | 3 | 1.997–2.28 | L. monocytogenes | Mb 19348_1 CHB |
| D1 | 2016 | 4 | 2.239–2.29 | L. monocytogenes | DSM 20600T DSM |
| D2 | 2016 | 5 | 2.052–2.191 | L. monocytogenes | Mb 19348_1 CHB |
| A | 2016 | 18 | 2.249–2.268 | L. monocytogenes | Mb 19348_1 CHB |
| C | 2016 | 19 | 1.976–1.988 | L. monocytogenes | Mb 19348_1 CHB |
| G2 | 2016 | 6A | 2.101–2.137 | L. monocytogenes | DSM 20600T DSM |
| G2 | 2016 | 6B | 1.946–2.043 | L. monocytogenes | DSM 20600T DSM |
| F | 2017 | 7A | 2.042–2.122 | L. monocytogenes | DSM 20600T DSM |
| F | 2017 | 7B | 2.031–2.095 | L. monocytogenes | Mb 19348_1 CHB |
| J | 2017 | 21 | 2.052–2.153 | L. monocytogenes | Mb 19348_1 CHB |
| G2 | 2017 | 22 | 2.045–2.101 | L. monocytogenes | Mb 19348_1 CHB |
| C | 2017 | 23 | 1.905–2.02 | L. monocytogenes | CCUG 315227 CCUG |
| F | 2018 | 8 | 2.027–2.234 | L. monocytogenes | Mb 19348_1 CHB |
| G3 | 2018 | 9 | 2.127–2.164 | L. monocytogenes | Mb 19348_1 CHB |
| D3 | 2018 | 10 | 1.996–2.182 | L. monocytogenes | Mb 19348_1 CHB |
| K | 2018 | 12 | 1.901–1.929 | L. monocytogenes | Mb 19348_1 CHB |
| H | 2018 | 13A | 2.023–2.048 | L. monocytogenes | DSM 20600T DSM |
| H | 2018 | 13B | 1.903–2.112 | L. monocytogenes | Mb 19348_1 CHB |
| H | 2018 | 13C | 1.991–2.065 | L. monocytogenes | Mb 19348_1 CHB |
| I | 2018 | 14A | 2.306–2.368 | L. monocytogenes | Mb 19348_1 CHB |
| I | 2018 | 14B | 2.013–2.106 | L. monocytogenes | Mb 19348_1 CHB |
| I | 2018 | 15 | 1.812–2.048 | L. monocytogenes | Mb 19348_1 CHB |
| E | 2018 | 20 | 1.922–1.955 | L. monocytogenes | DSM 20600T DSM |
| B | 2018 | 11 | 1.96–1.977 | L. monocytogenes | DSM 20600T DSM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pyz-Łukasik, R.; Gondek, M.; Winiarczyk, D.; Michalak, K.; Paszkiewicz, W.; Piróg-Komorowska, A.; Policht, A.; Ziomek, M. Occurrence of Listeria monocytogenes in Artisanal Cheeses from Poland and Its Identification by MALDI-TOF MS. Pathogens 2021, 10, 632. https://doi.org/10.3390/pathogens10060632
Pyz-Łukasik R, Gondek M, Winiarczyk D, Michalak K, Paszkiewicz W, Piróg-Komorowska A, Policht A, Ziomek M. Occurrence of Listeria monocytogenes in Artisanal Cheeses from Poland and Its Identification by MALDI-TOF MS. Pathogens. 2021; 10(6):632. https://doi.org/10.3390/pathogens10060632
Chicago/Turabian StylePyz-Łukasik, Renata, Michał Gondek, Dagmara Winiarczyk, Katarzyna Michalak, Waldemar Paszkiewicz, Anna Piróg-Komorowska, Agata Policht, and Monika Ziomek. 2021. "Occurrence of Listeria monocytogenes in Artisanal Cheeses from Poland and Its Identification by MALDI-TOF MS" Pathogens 10, no. 6: 632. https://doi.org/10.3390/pathogens10060632
APA StylePyz-Łukasik, R., Gondek, M., Winiarczyk, D., Michalak, K., Paszkiewicz, W., Piróg-Komorowska, A., Policht, A., & Ziomek, M. (2021). Occurrence of Listeria monocytogenes in Artisanal Cheeses from Poland and Its Identification by MALDI-TOF MS. Pathogens, 10(6), 632. https://doi.org/10.3390/pathogens10060632







