Investigating the Circulation of Ebola Viruses in Bats during the Ebola Virus Disease Outbreaks in the Equateur and North Kivu Provinces of the Democratic Republic of Congo from 2018
Abstract
1. Introduction
2. Results
2.1. Samples and Species per Collection Site
2.2. Molecular Screening for Filoviruses
2.3. Serological Screening for Presence of Antibodies against Ebolaviruses
3. Discussion
4. Materials and Methods
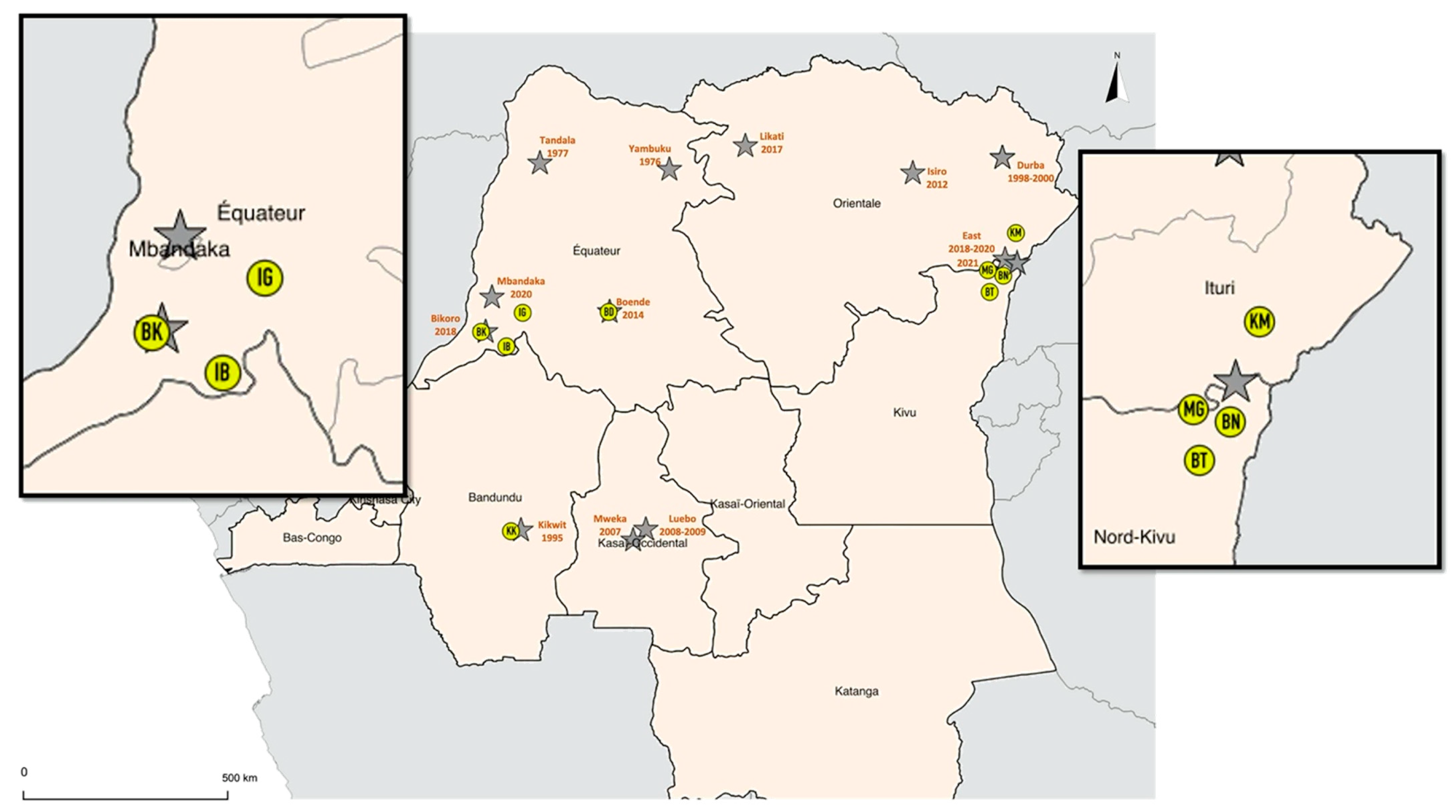
4.1. Collection Sites
4.2. Screening for Ebola Virus Antibodies
4.3. Cut-Off Determination
4.4. Molecular Confirmation of Bat Species
4.5. Nucleic Acid Extraction, RT-PCR Screening and Nanopore Sequencing for Detection of Filoviruses
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Languon, S.; Quaye, O. Filovirus Disease Outbreaks: A Chronological Overview. Virol. Res. Treat. 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Mahase, E. New Ebola Outbreak in DRC Has Killed 20 People so Far. BMJ 2020, 370, m2867. [Google Scholar] [CrossRef]
- Adepoju, P. Ebola Returns to Guinea and DR Congo. Lancet 2021, 397, 781. [Google Scholar] [CrossRef]
- LaBrunda, M.; Amin, N. The Emerging Threat of Ebola. Glob. Health Secur. 2019, 103–139. [Google Scholar] [CrossRef]
- Mbala Kingebeni, P.; Villabona-Arenas, C.-J.; Vidal, N.; Likofata, J.; Nsio-Mbeta, J.; Makiala-Mandanda, S.; Mukadi, D.; Mukadi, P.; Kumakamba, C.; Djokolo, B.; et al. Rapid Confirmation of the Zaire Ebola Virus in the Outbreak of the Equateur Province in the Democratic Republic of Congo: Implications for Public Health Interventions. Clin. Infect. Dis. 2018, 68, 330–333. [Google Scholar] [CrossRef]
- Wise, J. Ebola: DRC’s 10th Outbreak Is Declared Over. BMJ 2020, 369, m2601. [Google Scholar] [CrossRef] [PubMed]
- End of the 11th Ebola Outbreak-Democratic Republic of the Congo. Available online: https://www.afro.who.int/health-topics/ebola-virus-disease/end-ebola-11-drc (accessed on 6 December 2020).
- Epidémie de La Maladie à Virus Ebola Province Du Nord Kivu, République Démocratique Du Congo Sitrep N°50/2021 (Date de Publication: 29 Mars 2021)-Democratic Republic of the Congo. Available online: https://reliefweb.int/report/democratic-republic-congo/epid-mie-de-la-maladie-virus-ebola-province-du-nord-kivu-r-30 (accessed on 30 March 2021).
- Keita, A.K.; Düx, A.; Diallo, H.; Calvignac-Spencer, S.; Sow, M.S.; Keita, M.B.; Sidibe, Y.; Ayouba, A.; Lemarcis, F.; Marí Saéz, A.; et al. Resurgence of Ebola Virus in Guinea after 5 Years Calls for Careful Attention to Survivors without Creating Further Stigmatization. Available online: https://virological.org/t/guinea-2021-ebov-genomes/651 (accessed on 30 March 2021).
- Mbala-Kingebeni, P.; Pratt, C.; Mutafali-Ruffin, M.; Pauthner, M.G.; Bile, F.; Nkuba-Ndaye, A.; Black, A.; Kinganda-Lusamaki, E.; Faye, M.; Aziza, A.; et al. Ebola Virus Transmission Initiated by Relapse of Systemic Ebola Virus Disease. N. Engl. J. Med. 2021, 384, 1240–1247. [Google Scholar] [CrossRef]
- Leroy, E.M.; Rouquet, P.; Formenty, P.; Souquière, S.; Kilbourne, A.; Froment, J.-M.; Bermejo, M.; Smit, S.; Karesh, W.; Swanepoel, R.; et al. Multiple Ebola Virus Transmission Events and Rapid Decline of Central African Wildlife. Science 2004, 303, 387–390. [Google Scholar] [CrossRef] [PubMed]
- Leroy, E.M.; Epelboin, A.; Mondonge, V.; Pourrut, X.; Gonzalez, J.-P.; Muyembe-Tamfum, J.-J.; Formenty, P. Human Ebola Outbreak Resulting from Direct Exposure to Fruit Bats in Luebo, Democratic Republic of Congo, 2007. Vector Borne Zoonotic Dis. 2009, 9, 723–728. [Google Scholar] [CrossRef] [PubMed]
- Leroy, E.M.; Kumulungui, B.; Pourrut, X.; Rouquet, P.; Hassanin, A.; Yaba, P.; Délicat, A.; Paweska, J.T.; Gonzalez, J.-P.; Swanepoel, R. Fruit Bats as Reservoirs of Ebola Virus. Nature 2005, 438, 575–576. [Google Scholar] [CrossRef]
- Marí Saéz, A.; Weiss, S.; Nowak, K.; Lapeyre, V.; Zimmermann, F.; Düx, A.; Kühl, H.S.; Kaba, M.; Regnaut, S.; Merkel, K.; et al. Investigating the Zoonotic Origin of the West African Ebola Epidemic. EMBO Mol. Med. 2015, 7, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Amman, B.R.; Bird, B.H.; Bakarr, I.A.; Bangura, J.; Schuh, A.J.; Johnny, J.; Sealy, T.K.; Conteh, I.; Koroma, A.H.; Foday, I.; et al. Isolation of Angola-like Marburg Virus from Egyptian Rousette Bats from West Africa. Nat. Commun. 2020, 11, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Pourrut, X.; Souris, M.; Towner, J.S.; Rollin, P.E.; Nichol, S.T.; Gonzalez, J.-P.; Leroy, E. Large Serological Survey Showing Cocirculation of Ebola and Marburg Viruses in Gabonese Bat Populations, and a High Seroprevalence of Both Viruses in Rousettus Aegyptiacus. BMC Infect. Dis. 2009, 9, 159. [Google Scholar] [CrossRef] [PubMed]
- Towner, J.S.; Pourrut, X.; Albariño, C.G.; Nkogue, C.N.; Bird, B.H.; Grard, G.; Ksiazek, T.G.; Gonzalez, J.-P.; Nichol, S.T.; Leroy, E.M. Marburg Virus Infection Detected in a Common African Bat. PLoS ONE 2007, 2, e764. [Google Scholar] [CrossRef] [PubMed]
- Towner, J.S.; Amman, B.R.; Sealy, T.K.; Carroll, S.A.R.; Comer, J.A.; Kemp, A.; Swanepoel, R.; Paddock, C.D.; Balinandi, S.; Khristova, M.L.; et al. Isolation of Genetically Diverse Marburg Viruses from Egyptian Fruit Bats. PLoS Pathog. 2009, 5, e1000536. [Google Scholar] [CrossRef]
- Amman, B.R.; Carroll, S.A.; Reed, Z.D.; Sealy, T.K.; Balinandi, S.; Swanepoel, R.; Kemp, A.; Erickson, B.R.; Comer, J.A.; Campbell, S.; et al. Seasonal Pulses of Marburg Virus Circulation in Juvenile Rousettus Aegyptiacus Bats Coincide with Periods of Increased Risk of Human Infection. PLoS Pathog. 2012, 8, e1002877. [Google Scholar] [CrossRef] [PubMed]
- Amman, B.R.; Nyakarahuka, L.; McElroy, A.K.; Dodd, K.A.; Sealy, T.K.; Schuh, A.J.; Shoemaker, T.R.; Balinandi, S.; Atimnedi, P.; Kaboyo, W.; et al. Marburgvirus Resurgence in Kitaka Mine Bat Population after Extermination Attempts, Uganda. Emerg. Infect. Dis. 2014, 20, 1761–1764. [Google Scholar] [CrossRef]
- Swanepoel, R.; Smit, S.B.; Rollin, P.E.; Formenty, P.; Leman, P.A.; Kemp, A.; Burt, F.J.; Grobbelaar, A.A.; Croft, J.; Bausch, D.G.; et al. Studies of Reservoir Hosts for Marburg Virus. Emerg. Infect. Dis. 2007, 13, 1847–1851. [Google Scholar] [CrossRef]
- Kuzmin, I.V.; Niezgoda, M.; Franka, R.; Agwanda, B.; Markotter, W.; Breiman, R.F.; Shieh, W.J.; Zaki, S.R.; Rupprecht, C.E. Marburg Virus in Fruit Bat, Kenya. Emerg. Infect. Dis. 2010, 16, 352–354. [Google Scholar] [CrossRef] [PubMed]
- Storm, N.; Jansen Van Vuren, P.; Markotter, W.; Paweska, J.T. Antibody Responses to Marburg Virus in Egyptian Rousette Bats and Their Role in Protection against Infection. Viruses 2018, 10. [Google Scholar] [CrossRef]
- Kajihara, M.; Hang’ombe, B.M.; Changula, K.; Harima, H.; Isono, M.; Okuya, K.; Yoshida, R.; Mori-Kajihara, A.; Eto, Y.; Orba, Y.; et al. Marburgvirus in Egyptian Fruit Bats, Zambia. Emerg. Infect. Dis. 2019, 25, 1577–1580. [Google Scholar] [CrossRef] [PubMed]
- Pawęska, J.T.; Jansen van Vuren, P.; Kemp, A.; Storm, N.; Grobbelaar, A.A.; Wiley, M.R.; Palacios, G.; Markotter, W. Marburg Virus Infection in Egyptian Rousette Bats, South Africa, 2013–20141. Emerg. Infect. Dis. 2018, 24, 1134–1137. [Google Scholar] [CrossRef] [PubMed]
- Negredo, A.; Palacios, G.; Vázquez-Morón, S.; González, F.; Dopazo, H.; Molero, F.; Juste, J.; Quetglas, J.; Savji, N.; de la Cruz Martínez, M.; et al. Discovery of an Ebolavirus-Like Filovirus in Europe. PLoS Pathog. 2011, 7, e1002304. [Google Scholar] [CrossRef]
- Kemenesi, G.; Kurucz, K.; Dallos, B.; Zana, B.; Földes, F.; Boldogh, S.; Görföl, T.; Carroll, M.W.; Jakab, F. Re-Emergence of Lloviu Virus in Miniopterus Schreibersii Bats, Hungary, 2016. Emerg. Microbes. Infect. 2018, 7, 66. [Google Scholar] [CrossRef]
- Yuan, J.; Zhang, Y.; Li, J.; Zhang, Y.; Wang, L.-F.; Shi, Z. Serological Evidence of Ebolavirus Infection in Bats, China. Virol. J. 2012, 9, 236. [Google Scholar] [CrossRef]
- Yang, X.-L.; Zhang, Y.-Z.; Jiang, R.-D.; Guo, H.; Zhang, W.; Li, B.; Wang, N.; Wang, L.; Waruhiu, C.; Zhou, J.-H.; et al. Genetically Diverse Filoviruses in Rousettus and Eonycteris Spp. Bats, China, 2009 and 2015. Emerg. Infect. Dis. 2017, 23, 482–486. [Google Scholar] [CrossRef]
- Taniguchi, S.; Watanabe, S.; Masangkay, J.S.; Omatsu, T.; Ikegami, T.; Alviola, P.; Ueda, N.; Iha, K.; Fujii, H.; Ishii, Y.; et al. Reston Ebolavirus Antibodies in Bats, the Philippines. Emerg. Infect. Dis. 2011, 17, 1559. [Google Scholar] [CrossRef]
- Jayme, S.I.; Field, H.E.; de Jong, C.; Olival, K.J.; Marsh, G.; Tagtag, A.M.; Hughes, T.; Bucad, A.C.; Barr, J.; Azul, R.R.; et al. Molecular Evidence of Ebola Reston Virus Infection in Philippine Bats. Virol. J. 2015, 12, 107. [Google Scholar] [CrossRef]
- Goldstein, T.; Anthony, S.J.; Gbakima, A.; Bird, B.H.; Bangura, J.; Tremeau-Bravard, A.; Belaganahalli, M.N.; Wells, H.L.; Dhanota, J.K.; Liang, E.; et al. The Discovery of Bombali Virus Adds Further Support for Bats as Hosts of Ebolaviruses. Nat. Microbiol. 2018, 3, 1084–1089. [Google Scholar] [CrossRef] [PubMed]
- Forbes, K.M. Bombali Ebola Virus in Mops Condylurus Bat, Kenya. Emerg. Infect. Dis. J. CDC 2019, 25. [Google Scholar] [CrossRef]
- Karan, L.S.; Makenov, M.T.; Korneev, M.G.; Sacko, N.; Boumbaly, S.; Yakovlev, S.A.; Kourouma, K.; Bayandin, R.B.; Gladysheva, A.V.; Shipovalov, A.V.; et al. Bombali Virus in Mops Condylurus Bats, Guinea. Emerg. Infect. Dis. 2019, 25. [Google Scholar] [CrossRef]
- Amman, B.R.; Jones, M.E.B.; Sealy, T.K.; Uebelhoer, L.S.; Schuh, A.J.; Bird, B.H.; Coleman-McCray, J.D.; Martin, B.E.; Nichol, S.T.; Towner, J.S. Oral Shedding of Marburg Virus in Experimentally Infected Egyptian Fruit Bats (Rousettus Aegyptiacus). J. Wildl. Dis. 2015, 51, 113–124. [Google Scholar] [CrossRef]
- Jones, M.E.B.; Schuh, A.J.; Amman, B.R.; Sealy, T.K.; Zaki, S.R.; Nichol, S.T.; Towner, J.S. Experimental Inoculation of Egyptian Rousette Bats (Rousettus Aegyptiacus) with Viruses of the Ebolavirus and Marburgvirus Genera. Viruses 2015, 7, 3420–3442. [Google Scholar] [CrossRef] [PubMed]
- Pourrut, X.; Délicat, A.; Rollin, P.E.; Ksiazek, T.G.; Gonzalez, J.-P.; Leroy, E.M. Spatial and Temporal Patterns of Zaire Ebolavirus Antibody Prevalence in the Possible Reservoir Bat Species. J. Infect. Dis. 2007, 196, S176–S183. [Google Scholar] [CrossRef] [PubMed]
- Hayman, D.T.S.; Yu, M.; Crameri, G.; Wang, L.-F.; Suu-Ire, R.; Wood, J.L.N.; Cunningham, A.A. Ebola Virus Antibodies in Fruit Bats, Ghana, West Africa. Emerg. Infect. Dis. 2012, 18, 1207–1209. [Google Scholar] [CrossRef]
- Ogawa, H.; Miyamoto, H.; Nakayama, E.; Yoshida, R.; Nakamura, I.; Sawa, H.; Ishii, A.; Thomas, Y.; Nakagawa, E.; Matsuno, K.; et al. Seroepidemiological Prevalence of Multiple Species of Filoviruses in Fruit Bats (Eidolon Helvum) Migrating in Africa. J. Infect. Dis. 2015, 212, S101–S108. [Google Scholar] [CrossRef]
- De Nys, H.M.; Kingebeni, P.M.; Keita, A.K.; Butel, C.; Thaurignac, G.; Villabona-Arenas, C.-J.; Lemarcis, T.; Geraerts, M.; Vidal, N.; Esteban, A.; et al. Survey of Ebola Viruses in Frugivorous and Insectivorous Bats in Guinea, Cameroon, and the Democratic Republic of the Congo, 2015–2017. Emerg. Infect. Dis. 2018, 24. [Google Scholar] [CrossRef] [PubMed]
- Breman, J.G.; Teams, E.V.S.; Johnson, K.M.; Teams, E.V.S.; van der Groen, G.; Teams, E.V.S.; Robbins, C.B.; Teams, E.V.S.; Szczeniowski, M.V.; Teams, E.V.S.; et al. A Search for Ebola Virus in Animals in the Democratic Republic of the Congo and Cameroon: Ecologic, Virologic, and Serologic Surveys, 1979–1980. J. Infect. Dis. 1999, 179, S139–S147. [Google Scholar] [CrossRef]
- Leirs, H.; Mills, J.N.; Krebs, J.W.; Childs, J.E.; Akaibe, D.; Woollen, N.; Ludwig, G.; Peters, C.J.; Ksiazek, T.G.; Members, O.S.G. Search for the Ebola Virus Reservoir in Kikwit, Democratic Republic of the Congo: Reflections on a Vertebrate Collection. J. Infect. Dis. 1999, 179, S155–S163. [Google Scholar] [CrossRef]
- Gryseels, S.; Mbala-Kingebeni, P.; Akonda, I.; Angoyo, R.; Ayouba, A.; Baelo, P.; Mukadi, D.B.; Bugentho, E.; Bushmaker, T.; Butel, C.; et al. Role of Wildlife in Emergence of Ebola Virus in Kaigbono (Likati), Democratic Republic of the Congo, 2017. Emerg. Infect. Dis. 2020, 26, 2205–2209. [Google Scholar] [CrossRef] [PubMed]
- Kamins, A.O.; Rowcliffe, J.M.; Ntiamoa-Baidu, Y.; Cunningham, A.A.; Wood, J.L.N.; Restif, O. Characteristics and Risk Perceptions of Ghanaians Potentially Exposed to Bat-Borne Zoonoses through Bushmeat. EcoHealth 2015, 12, 104–120. [Google Scholar] [CrossRef] [PubMed]
- Han, H.-J.; Wen, H.; Zhou, C.-M.; Chen, F.-F.; Luo, L.-M.; Liu, J.; Yu, X.-J. Bats as Reservoirs of Severe Emerging Infectious Diseases. Virus Res. 2015, 205, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Baudel, H.; de Nys, H.; Mpoudi Ngole, E.; Peeters, M.; Desclaux, A. Understanding Ebola Virus and Other Zoonotic Transmission Risks through Human-Bat Contacts: Exploratory Study on Knowledge, Attitudes and Practices in Southern Cameroon. Zoonoses Public Health 2019, 66, 288–295. [Google Scholar] [CrossRef]
- Formenty, P.; Boesch, C.; Wyers, M.; Steiner, C.; Donati, F.; Dind, F.; Walker, F.; Le Guenno, B. Ebola Virus Outbreak among Wild Chimpanzees Living in a Rain Forest of Côte d’Ivoire. J. Infect. Dis. 1999, 179, S120–S126. [Google Scholar] [CrossRef]
- Tapanes, E.; Detwiler, K.M.; Cords, M. Bat Predation by Cercopithecus Monkeys: Implications for Zoonotic Disease Transmission. EcoHealth 2016, 13, 405–409. [Google Scholar] [CrossRef]
- Bermejo, M.; Rodríguez-Teijeiro, J.D.; Illera, G.; Barroso, A.; Vilà, C.; Walsh, P.D. Ebola Outbreak Killed 5000 Gorillas. Science 2006, 314, 1564. [Google Scholar] [CrossRef]
- Hayman, D.T.S.; Emmerich, P.; Yu, M.; Wang, L.-F.; Suu-Ire, R.; Fooks, A.R.; Cunningham, A.A.; Wood, J.L.N. Long-Term Survival of an Urban Fruit Bat Seropositive for Ebola and Lagos Bat Viruses. PLoS ONE 2010, 5, e11978. [Google Scholar] [CrossRef]
- Schuh, A.J.; Amman, B.R.; Jones, M.E.B.; Sealy, T.K.; Uebelhoer, L.S.; Spengler, J.R.; Martin, B.E.; Coleman-McCray, J.A.D.; Nichol, S.T.; Towner, J.S. Modelling Filovirus Maintenance in Nature by Experimental Transmission of Marburg Virus between Egyptian Rousette Bats. Nat. Commun. 2017, 8, 14446. [Google Scholar] [CrossRef]
- Forbes, K.M.; Webala, P.W.; Jääskeläinen, A.J.; Abdurahman, S.; Ogola, J.; Masika, M.M.; Kivistö, I.; Alburkat, H.; Plyusnin, I.; Levanov, L.; et al. Bombali Virus in Mops Condylurus Bat, Kenya. Emerg. Infect. Dis. 2019, 25, 955–957. [Google Scholar] [CrossRef]
- Swanepoel, R.; Leman, P.A.; Burt, F.J.; Zachariades, N.A.; Braack, L.E.; Ksiazek, T.G.; Rollin, P.E.; Zaki, S.R.; Peters, C.J. Experimental Inoculation of Plants and Animals with Ebola Virus. Emerg. Infect. Dis. 1996, 2, 321–325. [Google Scholar] [CrossRef] [PubMed]
- Leendertz, S.A.J.; Gogarten, J.F.; Düx, A.; Calvignac-Spencer, S.; Leendertz, F.H. Assessing the Evidence Supporting Fruit Bats as the Primary Reservoirs for Ebola Viruses. EcoHealth 2016, 13, 18–25. [Google Scholar] [CrossRef]
- Lacroix, A.; Vidal, N.; Keita, A.K.; Thaurignac, G.; Esteban, A.; de Nys, H.; Diallo, R.; Toure, A.; Goumou, S.; Soumah, A.K.; et al. Wide Diversity of Coronaviruses in Frugivorous and Insectivorous Bat Species: A Pilot Study in Guinea, West Africa. Viruses 2020, 12. [Google Scholar] [CrossRef] [PubMed]
- Ayouba, A.; Touré, A.; Butel, C.; Keita, A.K.; Binetruy, F.; Sow, M.S.; Foulongne, V.; Delaporte, E.; Peeters, M. Development of a Sensitive and Specific Serological Assay Based on Luminex Technology for Detection of Antibodies to Zaire Ebola Virus. J. Clin. Microbiol. 2017, 55, 165–176. [Google Scholar] [CrossRef]
- Peel, A.J.; McKinley, T.J.; Baker, K.S.; Barr, J.A.; Crameri, G.; Hayman, D.T.S.; Feng, Y.-R.; Broder, C.C.; Wang, L.-F.; Cunningham, A.A.; et al. Use of Cross-Reactive Serological Assays for Detecting Novel Pathogens in Wildlife: Assessing an Appropriate Cutoff for Henipavirus Assays in African Bats. J. Virol. Methods 2013, 193, 295–303. [Google Scholar] [CrossRef]
- Ayouba, A.; Ahuka-Mundeke, S.; Butel, C.; Mbala Kingebeni, P.; Loul, S.; Tagg, N.; Arenas, C.-J.V.; Lacroix, A.; Ndimbo-Kumugo, S.-P.; Keita, A.K.; et al. Extensive Serological Survey of Multiple African Non-Human Primate Species Reveals Low Prevalence of IgG Antibodies to Four Ebola Virus Species. J. Infect. Dis. 2019. [Google Scholar] [CrossRef]
- Irwin, D.M.; Kocher, T.D.; Wilson, A.C. Evolution of the Cytochromeb Gene of Mammals. J. Mol. Evol. 1991, 32, 128–144. [Google Scholar] [CrossRef] [PubMed]
- Kocher, T.D.; Thomas, W.K.; Meyer, A.; Edwards, S.V.; Pääbo, S.; Villablanca, F.X.; Wilson, A.C. Dynamics of Mitochondrial DNA Evolution in Animals: Amplification and Sequencing with Conserved Primers. Proc. Natl. Acad. Sci. USA 1989, 86, 6196–6200. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A Tool for Phylogenetic Analysis and Post-Analysis of Large Phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Nesi, N.; Nakouné, E.; Cruaud, C.; Hassanin, A. DNA Barcoding of African Fruit Bats (Mammalia, Pteropodidae). The Mitochondrial Genome Does Not Provide a Reliable Discrimination between Epomophorus Gambianus and Micropteropus Pusillus. Comptes Rendus Biol. 2011, 334, 544–554. [Google Scholar] [CrossRef] [PubMed]
| Equateur | North-Kivu/Ituri | Kikwit | Boende | Total | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| BK | IB | IG | subtotal | BN | BT | KM | MG | subtotal | KK | subtotal | BD | subtotal | ||
| FRUGIVOROUS BATS | ||||||||||||||
| Family PTEROPODIDAE | ||||||||||||||
| Casinycteris argynnis | 1 | 11 | - | 12 | - | - | - | - | - | - | - | 4 | 4 | 16 |
| Eidolon helvum | 2 | - | 3 | 5 | 2 | - | 2 | 10 | 14 | - | - | 130 | 130 | 149 |
| Epomophorus sp. | 2 | - | - | 2 | 185 | 34 | 4 | 43 | 266 | - | - | - | - | 268 |
| Epomops franqueti | 20 | 57 | 27 | 104 | - | - | 21 | 35 | 56 | 2 | 2 | 17 | 17 | 179 |
| Hypsignathus monstrosus | - | - | - | - | - | - | 1 | 5 | 6 | - | - | 5 | 5 | 11 |
| Lissonycteris angolensis | - | - | - | - | 23 | 1 | 2 | - | 26 | - | - | - | - | 26 |
| Megaloglossus woermanni | - | 4 | - | 4 | - | - | 1 | - | 1 | 5 | 5 | 1 | 1 | 11 |
| Micropteropus pusillus | 118 | - | - | 118 | 3 | - | 3 | - | 6 | 33 | 33 | 3 | 3 | 160 |
| Myonycteris torquata | 11 | 16 | 2 | 29 | - | - | 1 | 1 | 2 | 30 | 30 | 5 | 5 | 66 |
| Rousettus aegyptiacus | - | - | - | - | - | 1 | 4 | - | 5 | - | - | - | - | 5 |
| Scotonycteris bergmansi | - | 1 | - | 1 | - | - | - | - | - | - | - | - | - | 1 |
| Subtotal frugivorous bats | 154 | 89 | 32 | 275 | 213 | 36 | 39 | 94 | 382 | 70 | 70 | 165 | 165 | 892 |
| INSECTIVOROUS BATS | ||||||||||||||
| Family EMBALLONURIDAE | ||||||||||||||
| Taphozous mauritianus | - | - | - | - | - | - | 7 | - | 7 | - | - | - | - | 7 |
| Family HIPPOSIDERIDAE | ||||||||||||||
| Hipposideros caffer | - | - | - | - | 61 | - | - | - | 61 | - | - | - | - | 61 |
| Hipposideros sp. | - | - | - | - | - | - | - | - | - | 1 | 1 | - | - | 1 |
| Family MOLOSSIDAE | ||||||||||||||
| Chaerephon sp. | - | - | - | - | - | - | - | - | - | 6 | 6 | 1 | 1 | 7 |
| indeterminate | 2 | - | - | 2 | - | - | - | - | - | - | - | - | - | 2 |
| Mops sp. | 11 | - | - | 11 | - | - | - | - | - | - | - | - | - | 11 |
| Family NYCTERIDAE | ||||||||||||||
| Nycteris arge | - | - | - | - | - | - | - | - | - | - | - | 1 | 1 | 1 |
| Nycteris sp. | - | - | - | - | - | - | - | - | - | - | - | 1 | 1 | 1 |
| Family VESPERTILIONIDAE | ||||||||||||||
| Neoromicia nanus | - | - | - | - | - | - | - | - | - | 2 | 2 | - | - | 2 |
| Neoromicia sp. | - | - | - | - | - | - | - | - | - | 4 | 4 | - | - | 4 |
| Scotophilus dinganii | - | - | - | - | - | - | 3 | - | 3 | 14 | 14 | - | - | 17 |
| Scotophilus nux | - | - | - | - | - | - | - | - | - | 1 | 1 | - | - | 1 |
| Subtotal insectivorous bats | 13 | - | - | 13 | 61 | - | 10 | - | 71 | 28 | 28 | 3 | 3 | 115 |
| TOTAL | 167 | 89 | 32 | 288 | 274 | 36 | 49 | 94 | 453 | 98 | 98 | 168 | 168 | 1007 |
| N Tested | Oral Swab | Oral+Rectal Swab | Rectal Swab | |
|---|---|---|---|---|
| FRUGIVOROUS BATS | ||||
| Family PTEROPODIDAE | ||||
| Casinycteris argynnis | 12 | - | - | 12 |
| Eidolon helvum | 17 | - | 15 | 2 |
| Epomophorus sp. | 263 | - | 261 | 2 |
| Epomops franqueti | 126 | 2 | 63 | 61 |
| Hypsignathus monstrosus | 5 | - | 5 | - |
| Lissonycteris angolensis | 24 | - | 22 | 2 |
| Megaloglossus woermanni | 4 | - | - | 4 |
| Micropteropus pusillus | 121 | - | 48 | 73 |
| Myonycteris torquata | 30 | - | 5 | 25 |
| Rousettus aegyptiacus | 1 | - | 1 | - |
| Scotonycteris bergmansi | 1 | - | - | 1 |
| Subtotal frugivorous bats | 604 | 2 | 420 | 182 |
| INSECTIVOROUS BATS | ||||
| Family EMBALLONURIDAE | ||||
| Taphozous mauritianus | 0 | - | - | - |
| Family HIPPOSIDERIDAE | ||||
| Hipposideros caffer | 59 | 56 | 3 | - |
| Hipposideros sp. | 0 | - | - | |
| Family MOLOSSIDAE | ||||
| Chaerephon sp. | 0 | - | - | - |
| indeterminate | 2 | - | 2 | - |
| Mops sp. | 11 | - | - | 11 |
| Family NYCTERIDAE | ||||
| Nycteris arge | 0 | - | - | - |
| Nycteris sp. | 0 | - | - | - |
| Family VESPERTILIONIDAE | ||||
| Neoromicia nanus | 0 | - | - | - |
| Neoromicia sp. | 0 | - | - | - |
| Scotophilus dinganii | 0 | - | - | - |
| Scotophilus nux | 0 | - | - | - |
| Subtotal insectivorous bats | 72 | 56 | 5 | 11 |
| TOTAL | 676 | 58 | 425 | 193 |
| Kikwit | Boende | Equateur | North Kivu/Ituri | Total | ||
|---|---|---|---|---|---|---|
| n = 98 | n = 168 | n = 287 | n = 372 | n = 925 | ||
| N (%) pos | N (%) pos | N (%) pos | N (%) pos | N (%) pos | ||
| EBOV | NP | 0-0 (0-0) | 0-0 (0-0) | 1-1 (0.3-0.3) | 0-0 (0-0) | 1-1 (0.1-0.1) |
| GP-K | 1-3 (1-3) | 1-21 (0.6-30.9) | 0-4 (0-1.4) | 3-19 (0.8-5.1) | 5-47 (0.5-5.1) | |
| GP-M | 0-4 (0-4) | 1-8 (0.6-4.8) | 0-2 (0.0.7) | 2-7 (0.5-1.9) | 3-21 (0.3-2.3) | |
| VP40 | 0-0 (0-0) | 0-4 (0.0-2.4) | 0-2 (0.0.7) | 0-0 (0-0) | 0-6 (0-0.6) | |
| NP+GP | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | |
| SUDV | NP | 0-1 (0-1) | 0-0 (0-0) | 0-1 (0-0.3) | 1-2 (0.3-0.5) | 1-4 (0.1-0.4) |
| GP | 0-3 (0-3) | 4-42 (2.4-25) | 0-3 (0-1.1) | 3-5 (0.4-1.3) | 7-53 (0.7-5.7) | |
| VP40 | 0-2 (0-2) | 0-1 (0-0.6) | 0-1 (0-0.3) | 0-4 (0-1.1) | 2-8 (0.2-0.8) | |
| NP+GP | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-1 (0-0.3) | 0-1 (0-0.1) | |
| BDBV | GP | 0-2 (0-2) | 1-18 (0.6-10.7) | 0-3 (0-1.1) | 1-14 (0.3-3.7) | 2-37 (0.2-4.0) |
| VP40 | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-2 (0-0.2) | |
| RESTV | GP | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) | 0-0 (0-0) |
| Equateur | North Kivu/Ituri | Kikwit | Boende | Total | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| N Tested | N (%) pos | N Tested | N (%) pos | N Tested | N (%) pos | N Tested | N (%) pos | N Tested | N (%) pos | |
| FRUGIVOROUS BATS | ||||||||||
| Family PTEROPODIDAE | ||||||||||
| Casinycteris argynnis | 11 | 0 (0) | na | na | 4 | 0 (0) | 15 | 0 (0) | ||
| Eidolon helvum | 5 | 0 (0) | 14 | 2–6 (14.3–42.9) | na | 130 | 5–44 (3.8–33.8) | 149 | 7–50 (4.7–33.6) | |
| Epomophorus sp. | 2 | 0 (0) | 187 | 0–22 (0–11.8) | na | na | 189 | 0–22 (0–11.6) | ||
| Epomops franqueti | 104 | 0–3 (0–2.9) | 56 | 0–5 (0–8.9) | 2 | 0 (0) | 17 | 0 (0) | 179 | 0–8 (0–4.5) |
| Hypsignathus monstrosus | na | 6 | 0 (0) | na | 5 | 0 (0) | 11 | 0 (0) | ||
| Lissonycteris angolensis | na | 26 | 0 (0) | na | na | 26 | 0 (0) | |||
| Megaloglossus woermanni | 4 | 0 (0) | 1 | 0 (0) | 5 | 0 (0) | 1 | 0 (0) | 11 | 0 (0) |
| Micropteropus pusillus | 118 | 1–4 (0.8–3.4) | 4 | 0–2 (0–50) | 33 | 1–4 (3–12.1) | 3 | 0 (0) | 158 | 2–10 (1.2–6.3) |
| Myonycteris torquata | 29 | 0–1 (0–3.4) | 2 | 0 (0) | 30 | 1–4 (3.3–13.3) | 5 | 0–1 (0–20) | 66 | 1–6 (1.5–9.1) |
| Rousettus aegyptiacus | na | 5 | 1–4 (20–80) | na | na | 5 | 1–4 (20–80) | |||
| Scotonycteris bergmansi | 1 | 0 (0) | na | na | na | 1 | 0 (0) | |||
| Subtotal frugivorous bats | 274 | 1–8 (0.4–2.9) | 301 | 3–39 (1–13) | 70 | 2–8 (2.9–11.4) | 165 | 5–45 (3–27.3) | 810 | 11–100 (1.4–12.3) |
| INSECTIVOROUS BATS | ||||||||||
| Family EMBALLONURIDAE | ||||||||||
| Taphozous mauritianus | na | 7 | 0 (0) | na | na | 7 | 0 (0) | |||
| Family HIPPOSIDERIDAE | ||||||||||
| Hipposideros caffer | na | 61 | 0 (0) | na | na | 61 | 0 (0) | |||
| Hipposideros sp. | 0 | na | 1 | 0 (0) | na | 1 | 0 (0) | |||
| Family MOLOSSIDAE | ||||||||||
| Chaerephon sp. | na | na | 6 | 0 (0) | 1 | 0 (0) | 7 | 0 (0) | ||
| Indeterminate | 2 | 0 (0) | na | na | na | 2 | 0 (0) | |||
| Mops sp. | 11 | 0 (0) | na | na | na | 0 (0) | ||||
| Family NYCTERIDAE | ||||||||||
| Nycteris arge | na | na | na | 1 | 0 (0) | 1 | 0 (0) | |||
| Nycteris sp. | na | na | na | 1 | 0 (0) | 1 | 0 (0) | |||
| Family VESPERTILIONIDAE | ||||||||||
| Neoromicia nanus | na | na | 2 | 0 (0) | na | 2 | 0 (0) | |||
| Neoromicia sp. | na | na | 4 | 0 (0) | na | 4 | 0 (0) | |||
| Scotophilus dinganii | na | 3 | 0 (0) | 14 | 0 (0) | na | 17 | 0 (0) | ||
| Scotophilus nux | na | na | 1 | 0 (0) | na | 1 | 0 (0) | |||
| Subtotal insectivorous bats | 13 | 0 (0) | 71 | 0 | 28 | 0 (0) | 3 | 0 (0) | 115 | 0 (0) |
| Total | 287 | 1–8 (0.3–2.8) | 372 | 3–39 (0.8–10.5) | 98 | 2–8 (2–8.2) | 168 | 5–45 (3–26.8) | 925 | 11–100 (1.2–10.8) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lacroix, A.; Mbala Kingebeni, P.; Ndimbo Kumugo, S.P.; Lempu, G.; Butel, C.; Serrano, L.; Vidal, N.; Thaurignac, G.; Esteban, A.; Mukadi Bamuleka, D.; et al. Investigating the Circulation of Ebola Viruses in Bats during the Ebola Virus Disease Outbreaks in the Equateur and North Kivu Provinces of the Democratic Republic of Congo from 2018. Pathogens 2021, 10, 557. https://doi.org/10.3390/pathogens10050557
Lacroix A, Mbala Kingebeni P, Ndimbo Kumugo SP, Lempu G, Butel C, Serrano L, Vidal N, Thaurignac G, Esteban A, Mukadi Bamuleka D, et al. Investigating the Circulation of Ebola Viruses in Bats during the Ebola Virus Disease Outbreaks in the Equateur and North Kivu Provinces of the Democratic Republic of Congo from 2018. Pathogens. 2021; 10(5):557. https://doi.org/10.3390/pathogens10050557
Chicago/Turabian StyleLacroix, Audrey, Placide Mbala Kingebeni, Simon Pierre Ndimbo Kumugo, Guy Lempu, Christelle Butel, Laetitia Serrano, Nicole Vidal, Guillaume Thaurignac, Amandine Esteban, Daniel Mukadi Bamuleka, and et al. 2021. "Investigating the Circulation of Ebola Viruses in Bats during the Ebola Virus Disease Outbreaks in the Equateur and North Kivu Provinces of the Democratic Republic of Congo from 2018" Pathogens 10, no. 5: 557. https://doi.org/10.3390/pathogens10050557
APA StyleLacroix, A., Mbala Kingebeni, P., Ndimbo Kumugo, S. P., Lempu, G., Butel, C., Serrano, L., Vidal, N., Thaurignac, G., Esteban, A., Mukadi Bamuleka, D., Likofata, J., Delaporte, E., Muyembe Tamfum, J.-J., Ayouba, A., Peeters, M., & Ahuka Mundeke, S. (2021). Investigating the Circulation of Ebola Viruses in Bats during the Ebola Virus Disease Outbreaks in the Equateur and North Kivu Provinces of the Democratic Republic of Congo from 2018. Pathogens, 10(5), 557. https://doi.org/10.3390/pathogens10050557







