Correlation between Acinetobacter baumannii Resistance and Hospital Use of Meropenem, Cefepime, and Ciprofloxacin: Time Series Analysis and Dynamic Regression Models
Abstract
1. Introduction
2. Results
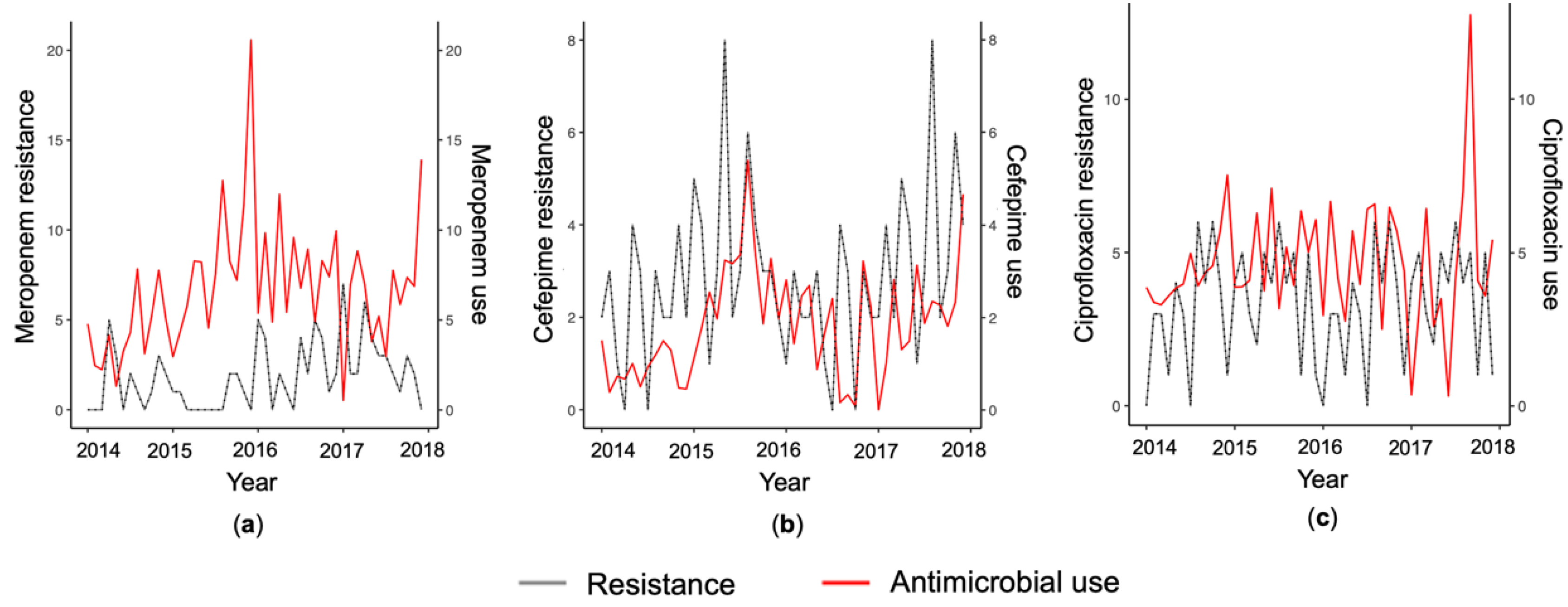
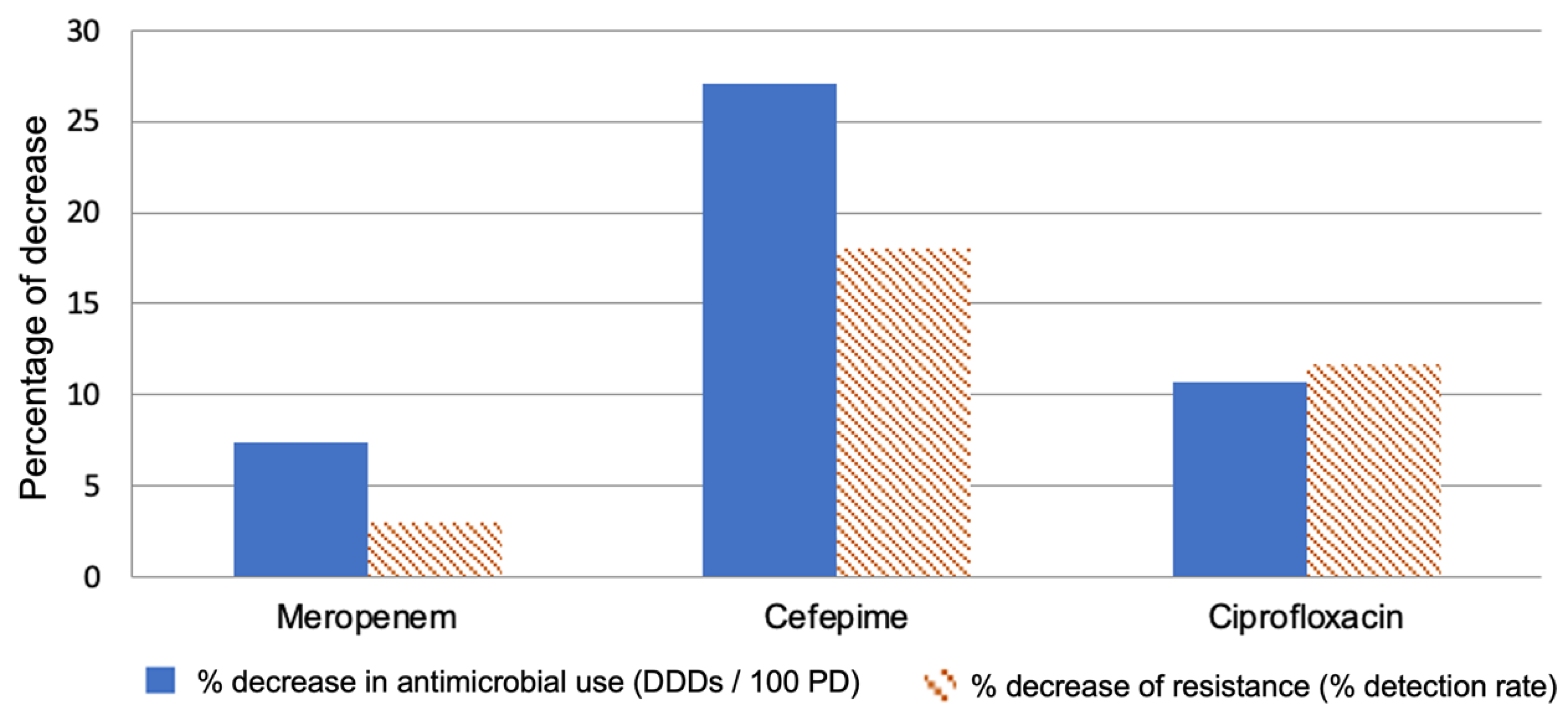
2.1. Meropenem
2.2. Cefepime
2.3. Ciprofloxacin
3. Discussion
4. Materials and Methods
4.1. Clinical Setting
4.2. Antibiotic Consumption
4.3. Microbiological Data
4.4. Data Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Antimicrobial Agent/Class | Order 1 | p-Value 2 | AIC 3 | R2 4 |
|---|---|---|---|---|
| Imipenem | 0 | 0.058 | 155.46 | 0.286 |
| Ceftazidime | 0 | 0.305 | 174.69 | 0.042 |
| Piperacillin/Tazobactam | 0 | 0.386 | 162.17 | 0.22 |
| Tigecyclin | 0 | 0.201 | 180.3 | 0.149 |
References
- Fournier, P.E.; Richet, H. The epidemiology and control of A. baumannii in health care facilities. Clin. Infect. Dis. 2006, 42, 692–699. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Seifert, H.; Paterson, D.L. A. baumannii: Emergence of a successful pathogen. Clin. Microbiol. Rev. 2008, 21, 538–582. [Google Scholar] [CrossRef]
- Bergogne-Berezin, E.; Towner, K.J. Acinetobacter spp. as nosocomial pathogens: Microbiological, clinical, and epidemiological features. Clin. Microbiol. Rev. 1996, 9, 148–165. [Google Scholar] [CrossRef] [PubMed]
- Evans, B.; Hamouda, A.; Amyes, S. The rise of carbapenem-resistant A. baumannii. Curr. Pharm. Des. 2013, 19, 223–238. [Google Scholar] [CrossRef] [PubMed]
- Dijkshoorn, L.; Nemec, A.; Seifert, H. An increasing threat in hospitals: Multidrug-resistant A. baumannii. Nat. Rev. Microbiol. 2007, 5, 939–951. [Google Scholar] [CrossRef] [PubMed]
- Vila, J.; Pachon, J. Therapeutic options for A. baumannii infections. Expert Opin. Pharmacother. 2008, 9, 587–599. [Google Scholar] [CrossRef]
- Baron, S.; Hadjadj, L.; Rolain, J.M.; Olaitan, A.O. Molecular mechanisms of polymyxin resistance: Knowns and unknowns. Int. J. Antimicrob. Agents. 2016, 48, 583–591. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L. The epidemiological profile of infections with multidrug-resistant Pseudomonas aeruginosa and Acinetobacter species. Clin. Inf. Dis. 2006, 43, S43–S48. [Google Scholar] [CrossRef] [PubMed]
- Hsueh, P.R.; Teng, L.J.; Chen, C.Y.; Chen, W.H.; Ho, S.W.; Luh, K.T. Pandrug-resistant A. baumannii causing nosocomial infections in a university hospital, Taiwan. Emerg. Infect. Dis. 2002, 8, 827–832. [Google Scholar] [CrossRef] [PubMed]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Paterson, D.L. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Kanafani, Z.; Kanj, S. Acinetobacter Infection: Treatment and Prevention. UpToDate. 2015. Available online: https://www.uptodate.com/contents/acinetobacter-infection-treatment-and-prevention (accessed on 28 February 2021).
- Ye, J.J.; Huang, C.T.; Shie, S.S.; Huang, P.Y.; Su, L.H.; Chiu, C.H.; Chiang, P.C. Multidrug resistant A. baumannii: Risk factors for appearance of imipenem resistant strains on patients formerly with susceptible strains. PLoS ONE 2010, 5, e9947. [Google Scholar] [CrossRef]
- Falagas, M.E.; Kopterides, P. Risk factors for the isolation of multi-drug resistant A. baumannii and Pseudomonas aeruginosa: A systematic review of the literature. J. Hosp. Infect. 2006, 64, 7–15. [Google Scholar] [CrossRef]
- Kim, Y.J.; Kim, S.I.; Hong, K.W.; Kim, Y.R.; Park, Y.J.; Kang, M.W. Risk factors for mortality in patients with carbapenem-resistant A. baumannii bacteremia: Impact of appropriate antimicrobial therapy. J. Korean Med. Sci. 2012, 27, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Meric, M.; Kasap, M.; Gacar, G.; Budak, F.; Dundar, D.; Kolayli, F.; Vahaboglu, H. Emergence and spread of carbapenem-resistant A. baumannii in a tertiary care hospital in Turkey. FEMS Microbiol. Lett. 2008, 282, 214–218. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Ouellette, M. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327. [Google Scholar] [CrossRef]
- Coyne, S.; Rosenfeld, N.; Lambert, T.; Courvalin, P.; Perichon, B. Overexpression of Resistance-Nodulation-Cell Division Pump AdeFGH Confers Multidrug Resistance in A. baumannii. Antimicrob. Agents Chemother. 2010, 54, 4389–4393. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A.; Strahilevitz, J.; Hooper, D.C. Plasmid-mediated quinolone resistance. Microbiol. Spectrum 2014, 2, 10.1128. [Google Scholar] [CrossRef] [PubMed]
- Hamed, S.M.; Elkhatib, W.F.; El-Mahallawy, H.A.; Helmy, M.M.; Ashour, M.S.; Aboshanab, K.M. Multiple mechanisms contributing to ciprofloxacin resistance among Gram negative bacteria causing infections to cancer patients. Sci. Rep. 2018, 8, 1–10. [Google Scholar]
- Butler, D.A.; Biagi, M.; Tan, X.; Qasmieh, S.; Bulman, Z.P.; Wenzler, E. Multidrug Resistant A. baumannii: Resistance by Any Other Name Would Still be Hard to Treat. Curr. Infect. Dis. Rep. 2019, 21, 46. [Google Scholar] [CrossRef]
- Pavlovic, R.R.; Jankovic, S.M. Inverse correlation of Acinetobacter spp. resistance rate and ciprofloxacin utilization. J. Antib. 2014, 67, 273–275. [Google Scholar] [CrossRef][Green Version]
- Yang, P.; Chen, Y.; Jiang, S.; Shen, P.; Lu, X.; Xiao, Y. Association between antibiotic consumption and the rate of carbapenem-resistant Gram-negative bacteria from China based on 153 tertiary hospitals data in 2014. Antimicrob. Resist. Infect. Control. 2018, 7, 137. [Google Scholar] [CrossRef] [PubMed]
- Yoon, Y.K.; Yang, K.S.; Lee, S.E.; Kim, H.J.; Sohn, J.W.; Kim, M.J. Effects of Group 1 versus Group 2 carbapenems on the susceptibility of Acinetobacter baumannii to carbapenems: A before and after intervention study of carbapenem-use stewardship. PLoS ONE 2014, 9, e99101. [Google Scholar] [CrossRef]
- López-Lozano, J.M.; Lawes, T.; Nebot, C.; Beyaert, A.; Bertrand, X.; Hocquet, D.; Aldeyab, M.; Scott, M.; Conlon-Bingham, G.; Farren, D.; et al. THRESHOLDS study group. A nonlinear time-series analysis approach to identify thresholds in associations between population antibiotic use and rates of resistance. Nat. Microbiol. 2019, 4, 1160–1172. [Google Scholar] [CrossRef] [PubMed]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing: 30th Informational Supplement; CLSI Document M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2020; Available online: https://clsi.org/media/3481/m100ed30_sample.pdf (accessed on 6 April 2021).
- WHO Collaborating Centre for Drug Statistics Methodology, ATC Classification Index with DDDs, 2021; Norwegian Institute of Public Health: Oslo, Norway, 2020.
- Falagas, M.E.; Mourtzoukou, E.G.; Polemis, M.; The Greek System for Surveillance of Antimicrobial Resistance; Vatopoulos, A.C. Trends in antimicrobial resistance of Acinetobacter baumannii clinical isolates from hospitalised patients in Greece and treatment implications. Clin. Microbiol. Infect. 2007, 13, 816–819. [Google Scholar] [CrossRef] [PubMed]
- Polemis, M.; Tryfinopoulou, K.; Giakkoupi, P.; WHONET-Greece Study Group; Vatopoulos, A. Eight-year trends in the relative isolation frequency and antimicrobial susceptibility among bloodstream isolates from Greek hospitals: Data from the Greek Electronic System for the Surveillance of Antimicrobial Resistance—WHONET-Greece, 2010 to 2017. Eurosurveillance 2020, 25, 1900516. [Google Scholar] [CrossRef]
- Karaiskos, I.; Lagou, S.; Pontikis, K.; Rapti, V.; Poulakou, G. The “Old” and the “New” Antibiotics for MDR Gram-Negative Pathogens: For Whom, When, and How. Front Public Health 2019, 7, 151. [Google Scholar] [CrossRef] [PubMed]
- Kengkla, K.; Kongpakwattana, K.; Saokaew, S.; Apisarnthanarak, A.; Chaiyakunapruk, N. Comparative efficacy and safety of treatment options for MDR and XDR Acinetobacter baumannii infections: A systematic review and network meta-analysis. Antimicrob. Chemother. 2018, 73, 22–32. [Google Scholar] [CrossRef]
- Tansarli, G.S.; Papaparaskevas, J.; Balaska, M.; Samarkos, M.; Pantazatou, A.; Markogiannakis, A.; Daikos, G.L. Colistin resistance in carbapenemase-producing Klebsiella pneumoniae bloodstream isolates: Evolution over 15 years and temporal association with colistin use by time series analysis. Int. J. Antimicrob. Agents. 2018, 52, 397–403. [Google Scholar] [CrossRef]
- Monnet, D.L.; López-Lozano, J.M.; Campillos, P.; Burgos, A.; Yagüe, A.; Gonzalo, N. Making sense of antimicrobial use and resistance surveillance data: Application of ARIMA and transfer function models. Clin. Microbiol. Infect. 2001, 7, 29–36. [Google Scholar] [CrossRef]
- López-Lozano, J.M.; Monnet, D.L.; Yagüe, A.; Burgos, A.; Gonzalo, N.; Campillos, P.; Saez, M. Modelling and forecasting antimicrobial resistance and its dynamic relationship to antimicrobial use: A time series analysis. Int. J. Antimicrob. Agents. 2000, 14, 21–31. [Google Scholar] [CrossRef]
- World Health Organization. Global Antimicrobial Resistance Surveillance System (GLASS): Molecular Methods for Antimicrobial Resistance (AMR) Diagnostics to Enhance the Global Antimicrobial Resistance Surveillance System. 2019. WHO/WSI/AMR/2019.1. Available online: https://apps.who.int/iris/handle/10665/310993 (accessed on 6 April 2021).
- Bard, J.D.; Lee, F. Why Can’t We Just Use PCR? The Role of Genotypic versus Phenotypic Testing for Antimicrobial Resistance Testing. Clin. Microbiol. Newsl. 2018, 40, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Allard, R. Use of time-series analysis in infectious disease surveillance. Bull. World Health Organ. 1998, 76, 327–333. [Google Scholar] [PubMed]
- Mozes, J.; Ebrahimi, F.; Goracz, O.; Miszti, C.; Kardos, G. Effect of carbapenem consumption patterns on the molecular epidemiology and carbapenem resistance of Acinetobacter baumannii. J. Med. Microbiol. 2014, 63, 1654–1662. [Google Scholar] [CrossRef]
- Guo, W.; Sun, F.; Liu, F.; Cao, L.; Yang, J.; Chen, Y. Antimicrobial resistance surveillance and prediction of Gram-negative bacteria based on antimicrobial consumption in a hospital setting: A 15-year retrospective study. Medicine 2019, 98, e17157. [Google Scholar] [CrossRef] [PubMed]
- Box, G.; Jenkins, G.M. Time Series Analysis: Forecasting and Control, 4th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2008; pp. 21–192. [Google Scholar]

| Antimicrobial Agent | Year | |||
|---|---|---|---|---|
| 2014 | 2015 | 2016 | 2017 | |
| Imipenem | 1.29 | 1.10 | 0.41 | 0.02 |
| Meropenem | 4.28 | 8.49 | 7.77 | 6.42 |
| Ceftazidime | 0.51 | 0.31 | 0.62 | 0.48 |
| Cefepime | 0.88 | 2.75 | 1.69 | 2.08 |
| Ciprofloxacin | 4.41 | 4.89 | 4.86 | 4.44 |
| Piperacillin/Tazobactam | 3.59 | 3.88 | 4.45 | 3.84 |
| Tigecyclin | 0.91 | 1.72 | 2.08 | 2.69 |
| Per Specimen | Percent of Isolates | Per Department | Percent of Isolates |
|---|---|---|---|
| Blood | 14.63% | Medical wards | 52.44% |
| Urine | 15.24% | Surgical wards | 25% |
| Broncho-Alveolar Lavage | 21.95% | Intensive Care Units | 16.46% |
| Sputum | 14.63% | Oncology/Hematological wards | 4.88% |
| Trauma | 11.59% | Mixed medical/surgical ward | 1.22% |
| Other | 21.95% |
| Estimate | Model Parameter | Standard Error | p-Value |
|---|---|---|---|
| A. A. baumannii resistance | |||
| ar1 | −0.517 | 0.118 | 0.000 |
| ar2 | −0.578 | 0.114 | <0.001 |
| AIC | 186.93 | ||
| R2 | 0.531 | ||
| B. Meropenem use (in DDD/100 PD) | |||
| ar1 | −0.831 | 0.122 | <0.001 |
| ar2 | −0.637 | 0.144 | <0.001 |
| ar3 | −0.569 | 0.117 | <0.001 |
| AIC | 242.91 | ||
| R2 | 0.638 | ||
| C. Impact of meropenem use on A. baumannii resistance | |||
| ar1 | −0.564 | 0.126 | <0.001 |
| ar2 | −0.610 | 0.124 | <0.001 |
| mer2 | 0.130 | 0.057 | 0.024 |
| AIC | 166.25 | ||
| R2 | 0.626 | ||
| Estimate | Model Parameter | Standard Error | p-Value |
|---|---|---|---|
| A. A. baumannii resistance | |||
| ar1 | −0.677 | 0.129 | <0.001 |
| ar2 | −0.710 | 0.122 | <0.001 |
| AIC | 100.56 | ||
| R2 | 0.580 | ||
| B. Cefepime use (in DDD/100 PD) | |||
| ar1 | −0.463 | 0.137 | <0.001 |
| ar2 | −0.466 | 0.133 | <0.001 |
| AIC | 139.03 | ||
| R2 | 0.619 | ||
| C. Impact of cefepime use on A. baumannii resistance | |||
| ar1 | −0.576 | 0.210 | 0.006 |
| ar2 | −0.559 | 0.222 | 0.011 |
| cef1 | 0.865 | 0.395 | 0.028 |
| AIC | 85.53 | ||
| R2 | 0.660 | ||
| Estimate | Model Parameter | Standard Error | p-Value |
|---|---|---|---|
| A. A. baumannii resistance | |||
| ma1 | −0.900 | 0.181 | <0.001 |
| AIC | 99.38 | ||
| R2 | 0.486 | ||
| B. Ciprofloxacin use (in DDD/100 PD) | |||
| ar1 | −0.527 | 0.147 | <0.001 |
| ar3 | −0.299 | 0.152 | 0.004 |
| AIC | 77.76 | ||
| R2 | 0.550 | ||
| C. Impact of ciprofloxacin use on A. baumannii resistance | |||
| cip1 | 0.733 | 0.081 | <0.001 |
| AIC | 77.52 | ||
| R2 | 0.617 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kousovista, R.; Athanasiou, C.; Liaskonis, K.; Ivopoulou, O.; Ismailos, G.; Karalis, V. Correlation between Acinetobacter baumannii Resistance and Hospital Use of Meropenem, Cefepime, and Ciprofloxacin: Time Series Analysis and Dynamic Regression Models. Pathogens 2021, 10, 480. https://doi.org/10.3390/pathogens10040480
Kousovista R, Athanasiou C, Liaskonis K, Ivopoulou O, Ismailos G, Karalis V. Correlation between Acinetobacter baumannii Resistance and Hospital Use of Meropenem, Cefepime, and Ciprofloxacin: Time Series Analysis and Dynamic Regression Models. Pathogens. 2021; 10(4):480. https://doi.org/10.3390/pathogens10040480
Chicago/Turabian StyleKousovista, Rania, Christos Athanasiou, Konstantinos Liaskonis, Olga Ivopoulou, George Ismailos, and Vangelis Karalis. 2021. "Correlation between Acinetobacter baumannii Resistance and Hospital Use of Meropenem, Cefepime, and Ciprofloxacin: Time Series Analysis and Dynamic Regression Models" Pathogens 10, no. 4: 480. https://doi.org/10.3390/pathogens10040480
APA StyleKousovista, R., Athanasiou, C., Liaskonis, K., Ivopoulou, O., Ismailos, G., & Karalis, V. (2021). Correlation between Acinetobacter baumannii Resistance and Hospital Use of Meropenem, Cefepime, and Ciprofloxacin: Time Series Analysis and Dynamic Regression Models. Pathogens, 10(4), 480. https://doi.org/10.3390/pathogens10040480







