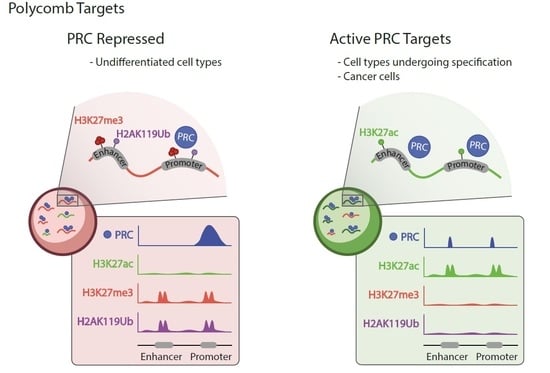
Functions of Polycomb Proteins on Active Targets
Abstract
1. Introduction
2. The Polycomb System
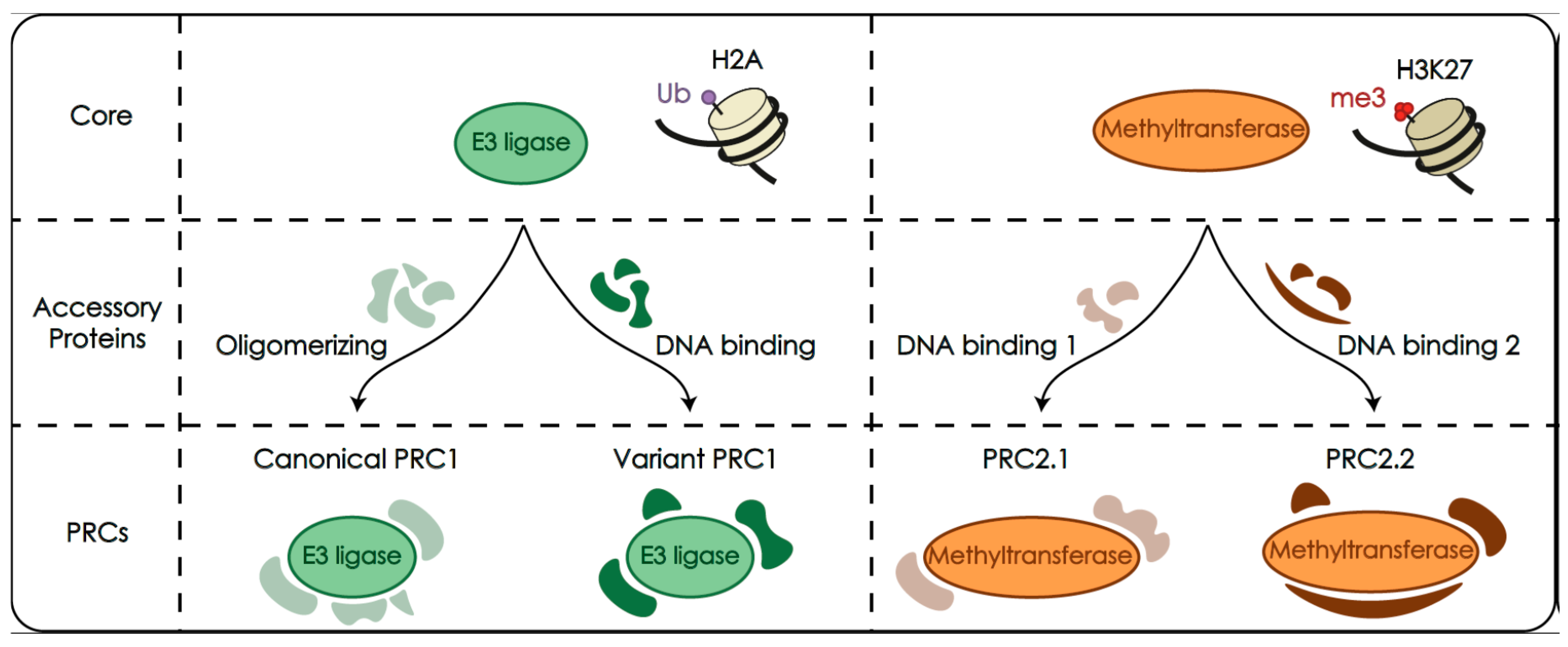
2.1. Polycomb Complexes PRC1 and PRC2 as Histone Modifiers
2.2. PRC1 and PRC2 Are Made of a Catalytic Module and Associated Accessory Subunits
2.3. Catalytic Modifications of Histone Tails
2.4. Recruitment of Polycomb Complexes to Chromatin
2.5. High Order Chromatin Structures Mediated by Polycomb
3. Polycomb and Transcriptionally Active Loci
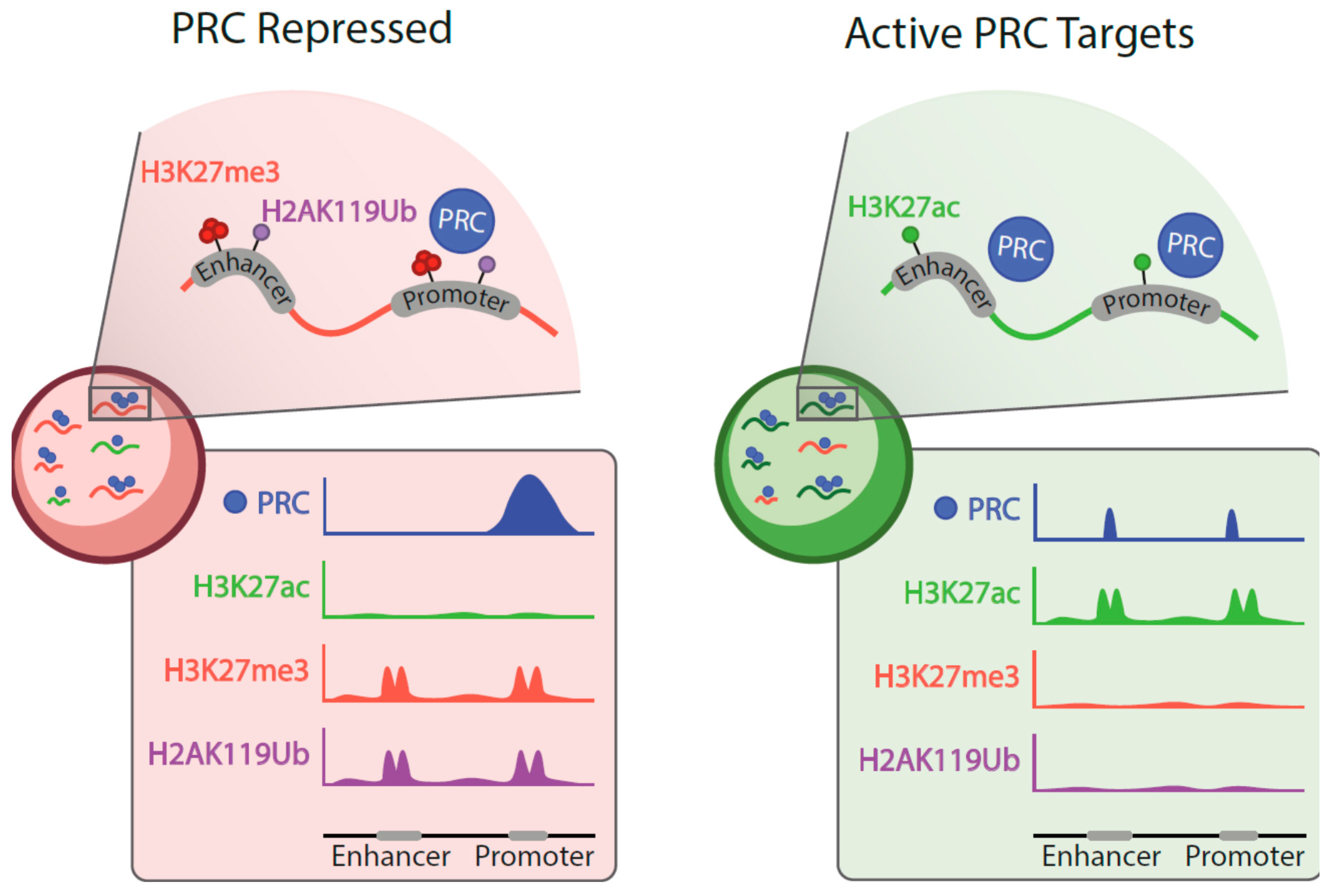
- Presence of PRC1 (invariable, RING1B in particular) and PRC2 subunits;
- Low/undetectable levels of H3k27me3 or H2AK119Ub modifications;
- Chromatin enrichment rates generally lower than at Polycomb-repressed domains;
- In differentiated cell types, the ratio of active to silent targets larger than in cells of fly embryos or ESCs;
- Enhancers and super-enhancers, in addition to promoters, among regulatory sites occupied by Polycomb products.
3.1. Polycomb Occupancy of Active Targets
3.2. Polycomb Proteins on Active Targets
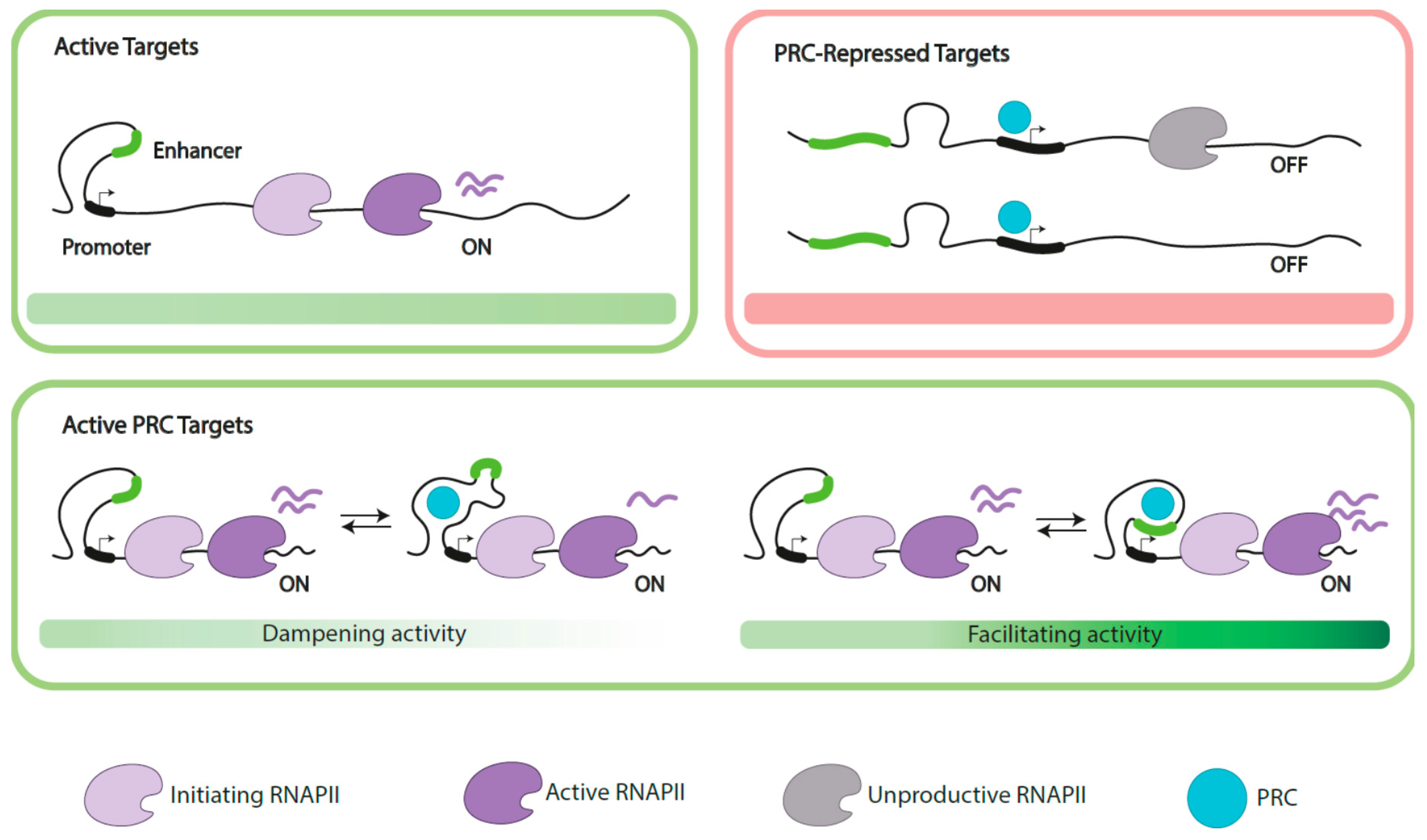
3.3. Polycomb Functions on Active Genes
3.3.1. Dampening Gene Activity
3.3.2. Supporting Gene Expression
3.3.3. Indirect (Positive) Role in Gene Activation
4. Conclusions and Perspectives
Funding
Conflicts of Interest
References
- Klemm, S.L.; Shipony, Z.; Greenleaf, W.J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 2019, 20, 207–220. [Google Scholar] [CrossRef]
- Ernst, J.; Kheradpour, P.; Mikkelsen, T.S.; Shoresh, N.; Ward, L.D.; Epstein, C.B.; Zhang, X.; Wang, L.; Issner, R.; Coyne, M.; et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 2011, 473, 43–49. [Google Scholar] [CrossRef]
- Szabo, Q.; Bantignies, F.; Cavalli, G. Principles of genome folding into topologically associating domains. Sci. Adv. 2019, 5, eaaw1668. [Google Scholar] [CrossRef]
- Rowley, M.J.; Corces, V.G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 2018, 19, 789–800. [Google Scholar] [CrossRef]
- Lewis, E.B. A gene complex controlling segmentation in Drosophila Genes. Dev. Cancer Life Work Edward B Lewis 2007, 276, 229–242. [Google Scholar]
- Schuettengruber, B.; Bourbon, H.-M.; Di Croce, L.; Cavalli, G. Genome Regulation by Polycomb and Trithorax: 70 Years and Counting. Cell 2017, 171, 34–57. [Google Scholar] [CrossRef]
- Whitcomb, S.J.; Basu, A.; Allis, C.D.; Bernstein, E. Polycomb Group proteins: An evolutionary perspective. Trends Genet. 2007, 23, 494–502. [Google Scholar] [CrossRef]
- Shao, Z.; Raible, F.; Mollaaghababa, R.; Guyon, J.R.; Wu, C.-T.; Bender, W.; Kingston, R.E. Stabilization of Chromatin Structure by PRC1, a Polycomb Complex. Cell 1999, 98, 37–46. [Google Scholar] [CrossRef]
- Kuzmichev, A.; Nishioka, K.; Erdjument-Bromage, H.; Tempst, P.; Reinberg, D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 2002, 16, 2893–2905. [Google Scholar] [CrossRef]
- Czermin, B.; Melfi, R.; McCabe, D.; Seitz, V.; Imhof, A.; Pirrotta, V. Drosophila Enhancer of Zeste/ESC Complexes Have a Histone H3 Methyltransferase activity that marks chromosomal polycomb sites. Cell 2002, 111, 185–196. [Google Scholar] [CrossRef]
- Plath, K.; Fang, J.; Mlynarczyk-Evans, S.K.; Cao, R.; Worringer, K.A.; Wang, H.; De La Cruz, C.C.; Otte, A.P.; Panning, B.; Zhang, Y. Role of Histone H3 Lysine 27 Methylation in X Inactivation. Science 2003, 300, 131–135. [Google Scholar] [CrossRef]
- Müller, J.; Hart, C.M.; Francis, N.J.; Vargas, M.L.; Sengupta, A.; Wild, B.; Miller, E.L.; O’Connor, M.; Kingston, R.E.; Simon, J.A. Histone Methyltransferase Activity of a Drosophila Polycomb Group Repressor Complex. Cell 2002, 111, 197–208. [Google Scholar] [CrossRef]
- Boyer, L.A.; Plath, K.; Zeitlinger, J.; Brambrink, T.; Medeiros, L.A.; Lee, T.I.; Levine, S.S.; Wernig, M.; Tajonar, A.; Ray, M.K.; et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 2006, 441, 349–353. [Google Scholar] [CrossRef]
- Endoh, M.; Endo, T.A.; Koseki, H.; Endoh, T.; Fujimura, Y.-I.; Ohara, O.; Toyoda, T.; Otte, A.P.; Okano, M.; Brockdorff, N.; et al. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 2008, 135, 1513–1524. [Google Scholar] [CrossRef]
- Jacobs, J.J.L.; Kieboom, K.; Marino, S.; DePinho, R.A.; Van Lohuizen, M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 1999, 397, 164–168. [Google Scholar] [CrossRef]
- Koppens, M.; Van Lohuizen, M. Context-dependent actions of Polycomb repressors in cancer. Oncogene 2015, 35, 1341–1352. [Google Scholar] [CrossRef]
- Mozgova, I.; Hennig, L. The Polycomb Group Protein Regulatory Network. Annu. Rev. Plant Biol. 2015, 66, 269–296. [Google Scholar] [CrossRef]
- Vidal, M.; Starowicz, K.; Starowicz, K. Polycomb complexes PRC1 and their function in hematopoiesis. Exp. Hematol. 2017, 48, 12–31. [Google Scholar] [CrossRef]
- Huang, C.; Nogales, E.; Ciferri, C. Molecular Architecture of the Polycomb Repressive Complex 2. Polycomb Group Proteins 2017, 45, 165–189. [Google Scholar]
- Aranda, S.; Martín, G.M.; Di Croce, L.D. Regulation of gene transcription by Polycomb proteins. Sci. Adv. 2015, 1, e1500737. [Google Scholar] [CrossRef]
- Kingston, R.E.; Tamkun, J.W. Transcriptional Regulation by Trithorax-Group Proteins. Cold Spring Harb. Perspect. Biol. 2014, 6, a019349. [Google Scholar] [CrossRef] [PubMed]
- Kassis, J.A.; Kennison, J.A.; Tamkun, J.W. Polycomb and Trithorax Group Genes in Drosophila. Genetics 2017, 206, 1699–1725. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wang, L.; Erdjument-Bromage, H.; Vidal, M.; Tempst, P.; Jones, R.S.; Zhang, Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature 2004, 431, 873–878. [Google Scholar] [CrossRef] [PubMed]
- Bentley, M.L.; Corn, J.E.; Dong, K.C.; Phung, Q.; Cheung, T.K.; Cochran, A.G. Recognition of UbcH5c and the nucleosome by the Bmi1/Ring1b ubiquitin ligase complex. EMBO J. 2011, 30, 3285–3297. [Google Scholar] [CrossRef]
- McGinty, R.K.; Henrici, R.C.; Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 2014, 514, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Kagey, M.H.; Melhuish, T.A.; Wotton, D. The Polycomb Protein Pc2 Is a SUMO E3. Cell 2003, 113, 127–137. [Google Scholar] [CrossRef]
- Tsukada, Y.-I.; Fang, J.; Erdjument-Bromage, H.; Warren, M.E.; Borchers, C.H.; Tempst, P.; Zhang, Y. Histone demethylation by a family of JmjC domain-containing proteins. Nature 2005, 439, 811–816. [Google Scholar] [CrossRef]
- Turberfield, A.H.; Kondo, T.; Nakayama, M.; Koseki, Y.; King, H.W.; Koseki, H.; Klose, R.J. KDM2 proteins constrain transcription from CpG island gene promoters independently of their histone demethylase activity. Nucleic Acids Res. 2019, 47, 9005–9023. [Google Scholar] [CrossRef]
- Leeb, M.; Pasini, D.; Novatchkova, M.; Jaritz, M.; Helin, K.; Wutz, A. Polycomb complexes act redundantly to repress genomic repeats and genes. Genes Dev. 2010, 24, 265–276. [Google Scholar] [CrossRef]
- Kahn, T.G.; Dorafshan, E.; Schultheis, D.; Zare, A.; Stenberg, P.; Reim, I.; Pirrotta, V.; Schwartz, Y.B. Interdependence of PRC1 and PRC2 for recruitment to Polycomb Response Elements. Nucleic Acids Res. 2016, 44, 10132–10149. [Google Scholar] [CrossRef]
- Zepeda-Martinez, J.A.; Pribitzer, C.; Wang, J.; Bsteh, D.; Golumbeanu, S.; Zhao, Q.; Burkard, T.R.; Reichholf, B.; Rhie, S.K.; Jude, J.; et al. Parallel PRC2/cPRC1 and vPRC1 pathways silence lineage-specific genes and maintain self-renewal in mouse embryonic stem cells. Sci. Adv. 2020, 6, eaax5692. [Google Scholar] [CrossRef]
- Kalb, R.; Latwiel, S.; Baymaz, H.I.; Jansen, P.W.T.C.; Müller, C.W.; Vermeulen, M.; Müller, J.; Müller, C.W. Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat. Struct. Mol. Biol. 2014, 21, 569–571. [Google Scholar] [CrossRef] [PubMed]
- Blackledge, N.P.; Farcas, A.M.; Kondo, T.; King, H.W.; McGouran, J.; Hanssen, L.L.; Ito, S.; Cooper, S.; Kondo, K.; Koseki, Y.; et al. Variant PRC1 Complex-Dependent H2A Ubiquitylation Drives PRC2 Recruitment and Polycomb Domain Formation. Cell 2014, 157, 1445–1459. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Zhang, J.; Bonasio, R.; Strino, F.; Sawai, A.; Parisi, F.; Kluger, Y.; Reinberg, D. PCGF Homologs, CBX Proteins, and RYBP Define Functionally Distinct PRC1 Family Complexes. Mol. Cell 2012, 45, 344–356. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Taylor, A.B.; Leal, B.Z.; Chadwell, L.V.; Ilangovan, U.; Robinson, A.K.; Schirf, V.; Hart, P.J.; Lafer, E.; Demeler, B.; et al. Polycomb Group Targeting through Different Binding Partners of RING1B C-Terminal Domain. Structure 2010, 18, 966–975. [Google Scholar] [CrossRef] [PubMed]
- Margueron, R.; Justin, N.; Ohno, K.; Sharpe, M.L.; Son, J.; Iii, W.J.D.; Voigt, P.; Martin, S.R.; Taylor, W.R.; De Marco, V.; et al. Role of the polycomb protein EED in the propagation of repressive histone marks. Nature 2009, 461, 762–767. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Justin, N.; Wilson, J.R.; Gamblin, S.J. Comment on “Structural basis of histone H3K27 trimethylation by an active polycomb repressive complex 2”. Science 2016, 354, 1543. [Google Scholar] [CrossRef]
- Lee, C.-H.; Yu, J.-R.; Kumar, S.; Jin, Y.; Leroy, G.; Bhanu, N.; Kaneko, S.; Garcia, B.A.; Hamilton, A.D.; Reinberg, D. Allosteric Activation Dictates PRC2 Activity Independent of Its Recruitment to Chromatin. Mol. Cell 2018, 70, 422–434.e6. [Google Scholar] [CrossRef]
- Holoch, D.; Margueron, R. Mechanisms Regulating PRC2 Recruitment and Enzymatic Activity. Trends Biochem. Sci. 2017, 42, 531–542. [Google Scholar] [CrossRef]
- Conway, E.; Jerman, E.; Healy, E.; Ito, S.; Holoch, D.; Oliviero, G.; Deevy, O.; Glancy, E.; Fitzpatrick, D.J.; Mucha, M.; et al. A Family of Vertebrate-Specific Polycombs Encoded by the LCOR/LCORL Genes Balance PRC2 Subtype Activities. Mol. Cell 2018, 70, 408–421.e8. [Google Scholar] [CrossRef]
- Healy, E.; Mucha, M.; Glancy, E.; Fitzpatrick, D.J.; Conway, E.; Neikes, H.K.; Monger, C.; Van Mierlo, G.; Baltissen, M.P.; Koseki, Y.; et al. PRC2.1 and PRC2.2 Synergize to Coordinate H3K27 Trimethylation. Mol. Cell 2019, 76, 437–452.e6. [Google Scholar] [CrossRef] [PubMed]
- Isono, K.; Endo, T.A.; Ku, M.; Yamada, D.; Suzuki, R.; Sharif, J.; Ishikura, T.; Toyoda, T.; Bernstein, B.E.; Koseki, H. SAM Domain Polymerization Links Subnuclear Clustering of PRC1 to Gene Silencing. Dev. Cell 2013, 26, 565–577. [Google Scholar] [CrossRef] [PubMed]
- Wani, A.H.; Boettiger, A.N.; Schorderet, P.; Ergun, A.; Münger, C.; Sadreyev, R.I.; Zhuang, X.; Kingston, R.E.; Francis, N.J. Chromatin topology is coupled to Polycomb group protein subnuclear organization. Nat. Commun. 2016, 7, 10291. [Google Scholar] [CrossRef] [PubMed]
- Tatavosian, R.; Duc, H.N.; Huynh, T.N.; Fang, N.; Schmitt, B.; Shi, X.; Deng, Y.; Phiel, C.; Yao, T.; Zhang, Z.; et al. Live-cell single-molecule dynamics of PcG proteins imposed by the DIPG H3.3K27M mutation. Nat. Commun. 2018, 9, 2080. [Google Scholar] [CrossRef] [PubMed]
- Plys, A.J.; Davis, C.P.; Kim, J.; Rizki, G.; Keenen, M.M.; Marr, S.K.; Kingston, R.E. Phase separation of Polycomb-repressive complex 1 is governed by a charged disordered region of CBX2. Genes Dev. 2019, 33, 799–813. [Google Scholar] [CrossRef]
- Farcas, A.M.; Blackledge, N.P.; Sudbery, I.; Long, H.K.; McGouran, J.; Rose, N.R.; Lee, S.; Sims, D.; Cerase, A.; Sheahan, T.W.; et al. KDM2B links the Polycomb Repressive Complex 1 (PRC1) to recognition of CpG islands. eLife 2012, 1, 00205. [Google Scholar] [CrossRef]
- He, J.; Shen, L.; Wan, M.; Taranova, O.; Wu, H.; Zhang, Y. Kdm2b maintains murine embryonic stem cell status by recruiting PRC1 complex to CpG islands of developmental genes. Nat. Cell Biol. 2013, 15, 373–384. [Google Scholar] [CrossRef]
- Wu, X.; Johansen, J.V.; Helin, K. Fbxl10/Kdm2b Recruits Polycomb Repressive Complex 1 to CpG Islands and Regulates H2A Ubiquitylation. Mol. Cell. 2013, 49, 1134–1146. [Google Scholar] [CrossRef]
- Stielow, B.; Finkernagel, F.; Stiewe, T.; Nist, A.; Suske, G. MGA, L3MBTL2 and E2F6 determine genomic binding of the non-canonical Polycomb repressive complex PRC1.6. PLoS Genet. 2018, 14, e1007193. [Google Scholar] [CrossRef]
- Choi, J.; Bachmann, A.L.; Tauscher, K.; Benda, C.; Fierz, B.; Müller, J. DNA binding by PHF1 prolongs PRC2 residence time on chromatin and thereby promotes H3K27 methylation. Nat. Struct. Mol. Biol. 2017, 24, 1039–1047. [Google Scholar] [CrossRef]
- Li, H.; Liefke, R.; Jiang, J.; Kurland, J.V.; Tian, W.; Deng, P.; Zhang, W.; He, Q.; Patel, D.J.; Bulyk, M.L.; et al. Polycomb-like proteins link the PRC2 complex to CpG islands. Nature 2017, 549, 287–291. [Google Scholar] [CrossRef] [PubMed]
- Perino, M.; Van Mierlo, G.; Karemaker, I.D.; Van Genesen, S.; Vermeulen, M.; Marks, H.; Van Heeringen, S.J.; Veenstra, G.J.C. MTF2 recruits Polycomb Repressive Complex 2 by helical-shape-selective DNA binding. Nat. Genet. 2018, 50, 1002–1010. [Google Scholar] [CrossRef] [PubMed]
- Pasini, D.; Cloos, P.A.C.; Walfridsson, J.; Olsson, L.; Bukowski, J.-P.; Johansen, J.V.; Bak, M.; Tommerup, N.; Rappsilber, J.; Helin, K. JARID2 regulates binding of the Polycomb repressive complex 2 to target genes in ES cells. Nature 2010, 464, 306–310. [Google Scholar] [CrossRef] [PubMed]
- Asenjo, H.G.; Gallardo, A.; López-Onieva, L.; Tejada, I.; Martorell-Marugán, J.; Carmona-Saez, P.; Landeira, D. Polycomb regulation is coupled to cell cycle transition in pluripotent stem cells. Sci. Adv. 2020, 6, eaay4768. [Google Scholar] [CrossRef]
- Min, J.; Zhang, Y.; Xu, R.-M. Structural basis for specific binding of Polycomb chromodomain to histone H3 methylated at Lys 27. Genes Dev. 2003, 17, 1823–1828. [Google Scholar] [CrossRef]
- Nowak, A.J.; Alfieri, C.; Stirnimann, C.U.; Rybin, V.; Baudin, F.; Ly-Hartig, N.; Lindner, D.; Müller, C.W. Chromatin-modifying complex component Nurf55/p55 associates with histones H3 and H4 and polycomb repressive complex 2 subunit Su(z)12 through partially overlapping binding sites. J. Biol. Chem. 2011, 286, 23388–23396. [Google Scholar] [CrossRef]
- Kloet, S.; Makowski, M.M.; Baymaz, H.I.; Van Voorthuijsen, L.; Karemaker, I.D.; Santanach, A.; Jansen, P.W.; Croce, L.D.; Vermeulen, M. The dynamic interactome and genomic targets of Polycomb complexes during stem-cell differentiation. Nat. Struct. Mol. Biol. 2016, 23, 682–690. [Google Scholar] [CrossRef]
- Lee, H.-G.; Kahn, T.G.; Simcox, A.; Schwartz, Y.B.; Pirrotta, V. Genome-wide activities of Polycomb complexes control pervasive transcription. Genome Res. 2015, 25, 1170–1181. [Google Scholar] [CrossRef]
- Fursova, N.A.; Blackledge, N.P.; Nakayama, M.; Ito, S.; Koseki, Y.; Farcas, A.M.; King, H.W.; Koseki, H.; Klose, R.J. Synergy between Variant PRC1 Complexes Defines Polycomb-Mediated Gene Repression. Mol. Cell 2019, 74, 1020–1036.e8. [Google Scholar] [CrossRef]
- Taherbhoy, A.M.; Huang, O.W.; Cochran, A.G. BMI1–RING1B is an autoinhibited RING E3 ubiquitin ligase. Nat. Commun. 2015, 6, 7621. [Google Scholar] [CrossRef]
- Rose, N.R.; King, H.W.; Blackledge, N.P.; Fursova, N.A.; Ember, K.J.; Fischer, R.; Kessler, B.M.; Klose, R.J. RYBP stimulates PRC1 to shape chromatin-based communication between Polycomb repressive complexes. eLife 2016, 5, 524. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Wang, M.; Zhu, B.; Li, G.; Chang, L.; Yu, J.; Song, A.; Liu, C.; Huang, W.; Zhang, T.; et al. RYBP/YAF2-PRC1 complexes and histone H1-dependent chromatin compaction mediate propagation of H2AK119ub1 during cell division. Nat. Cell Biol. 2020, 22, 439–452. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, K.J.; Scelfo, A.; Jammula, S.; Cuomo, A.; Barozzi, I.; Stützer, A.; Fischle, W.; Bonaldi, T.; Pasini, D. Polycomb-Dependent H3K27me1 and H3K27me2 Regulate Active Transcription and Enhancer Fidelity. Mol. Cell 2014, 53, 49–62. [Google Scholar] [CrossRef]
- Oksuz, O.; Narendra, V.; Lee, C.-H.; Descostes, N.; Leroy, G.; Raviram, R.; Blumenberg, L.; Karch, K.; Rocha, P.P.; Garcia, B.A.; et al. Capturing the Onset of PRC2-Mediated Repressive Domain Formation. Mol. Cell 2018, 70, 1149–1162.e5. [Google Scholar] [CrossRef] [PubMed]
- McCabe, M.T.; Graves, A.P.; Chen, S.B.; Della Pietra, A.; Dul, E.; Hughes, A.M.; Gilbert, S.A.; Thrall, S.H.; Tummino, P.J.; Kruger, R.G.; et al. Mutation of A677 in histone methyltransferase EZH2 in human B-cell lymphoma promotes hypertrimethylation of histone H3 on lysine 27 (H3K27). Proc. Natl. Acad. Sci. USA 2012, 109, 2989–2994. [Google Scholar] [CrossRef] [PubMed]
- Poepsel, S.; Kasinath, V.; Nogales, E. Cryo-EM structures of PRC2 simultaneously engaged with two functionally distinct nucleosomes. Nat. Struct. Mol. Biol. 2018, 25, 154–162. [Google Scholar] [CrossRef]
- Blackledge, N.P.; Fursova, N.A.; Kelley, J.R.; Huseyin, M.K.; Feldmann, A.; Klose, R.J. PRC1 Catalytic Activity Is Central to Polycomb System Function. Mol. Cell 2020, 77, 857–874.e9. [Google Scholar] [CrossRef]
- Yuan, W.; Xu, M.; Huang, C.; Liu, N.; Chen, S.; Zhu, B. H3K36 Methylation Antagonizes PRC2-mediated H3K27 Methylation. J. Biol. Chem. 2011, 286, 7983–7989. [Google Scholar] [CrossRef]
- Yuan, G.; Ma, B.; Yuan, W.; Zhang, Z.; Chen, P.; Ding, X.; Feng, L.; Shen, X.; Chen, S.; Li, G.; et al. Histone H2A Ubiquitination Inhibits the Enzymatic Activity of H3 Lysine 36 Methyltransferases. J. Biol. Chem. 2013, 288, 30832–30842. [Google Scholar] [CrossRef]
- Zhen, C.Y.; Tatavosian, R.; Huynh, T.N.; Duc, H.N.; Das, R.; Kokotovic, M.; Grimm, J.B.; Lavis, L.D.; Lee, J.; Mejia, F.J.; et al. Live-cell single-molecule tracking reveals co-recognition of H3K27me3 and DNA targets polycomb Cbx7-PRC1 to chromatin. eLife 2016, 5, 731. [Google Scholar] [CrossRef]
- Connelly, K.E.; Weaver, T.M.; Alpsoy, A.; Gu, B.X.; Musselman, C.A.; Dykhuizen, E.C. Engagement of DNA and H3K27me3 by the CBX8 chromodomain drives chromatin association. Nucleic Acids Res. 2019, 47, 2289–2305. [Google Scholar] [CrossRef] [PubMed]
- Arrigoni, R.; Alam, S.L.; Wamstad, J.A.; Bardwell, V.J.; Sundquist, W.I.; Schreiber-Agus, N. The Polycomb-associated protein Rybp is a ubiquitin binding protein. FEBS Lett. 2006, 580, 6233–6241. [Google Scholar] [CrossRef] [PubMed]
- Cooper, S.; Grijzenhout, A.; Underwood, E.; Ancelin, K.; Zhang, T.; Nesterova, T.B.; Anil-Kirmizitas, B.; Bassett, A.; Kooistra, S.M.; Agger, K.; et al. Jarid2 binds mono-ubiquitylated H2A lysine 119 to mediate crosstalk between Polycomb complexes PRC1 and PRC2. Nat. Commun. 2016, 7, 13661. [Google Scholar] [CrossRef] [PubMed]
- Agger, K.; Cloos, P.A.C.; Christensen, J.; Pasini, D.; Rose, S.; Rappsilber, J.; Issaeva, I.; Canaani, E.; Salcini, A.E.; Helin, K. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 2007, 449, 731–734. [Google Scholar] [CrossRef]
- Zhu, P.; Zhou, W.; Wang, J.; Puc, J.; Ohgi, K.A.; Erdjument-Bromage, H.; Tempst, P.; Glass, C.K.; Rosenfeld, M.G. A Histone H2A Deubiquitinase Complex Coordinating Histone Acetylation and H1 Dissociation in Transcriptional Regulation. Mol. Cell 2007, 27, 609–621. [Google Scholar] [CrossRef]
- Joo, H.-Y.; Zhai, L.; Yang, C.; Nie, S.; Erdjument-Bromage, H.; Tempst, P.; Chang, C.; Wang, H. Regulation of cell cycle progression and gene expression by H2A deubiquitination. Nature 2007, 449, 1068–1072. [Google Scholar] [CrossRef]
- Campagne, A.; Lee, M.-K.; Zielinski, D.; Michaud, A.; Le Corre, S.; Dingli, F.; Chen, H.; Shahidian, L.Z.; Vassilev, I.; Servant, N.; et al. BAP1 complex promotes transcription by opposing PRC1-mediated H2A ubiquitylation. Nat. Commun. 2019, 10, 348. [Google Scholar] [CrossRef]
- Sneppen, K.; Ringrose, L. Theoretical analysis of Polycomb-Trithorax systems predicts that poised chromatin is bistable and not bivalent. Nat. Commun. 2019, 10, 2133. [Google Scholar] [CrossRef]
- Schwartz, Y.B.; Kahn, T.G.; Nix, D.A.; Li, X.-Y.; Bourgon, R.; Biggin, M.; Pirrotta, V. Genome-wide analysis of Polycomb targets in Drosophila melanogaster. Nat. Genet. 2006, 38, 700–705. [Google Scholar] [CrossRef]
- Tamburri, S.; Lavarone, E.; Fernández-Pérez, D.; Conway, E.; Zanotti, M.; Manganaro, D.; Pasini, D. Histone H2AK119 Mono-Ubiquitination Is Essential for Polycomb-Mediated Transcriptional Repression. Mol. Cell 2020, 77, 840–856.e5. [Google Scholar] [CrossRef]
- Tsuboi, M.; Kishi, Y.; Yokozeki, W.; Koseki, H.; Hirabayashi, Y.; Gotoh, Y. Ubiquitination-Independent Repression of PRC1 Targets during Neuronal Fate Restriction in the Developing Mouse Neocortex. Dev. Cell 2018, 47, 758–772.e5. [Google Scholar] [CrossRef] [PubMed]
- Coleman, R.T.; Struhl, G. Causal role for inheritance of H3K27me3 in maintaining the OFF state of aDrosophilaHOX gene. Science 2017, 356, eaai8236. [Google Scholar] [CrossRef] [PubMed]
- Moussa, H.F.; Bsteh, D.; Yelagandula, R.; Pribitzer, C.; Stecher, K.; Bartalska, K.; Michetti, L.; Wang, J.; Zepeda-Martinez, J.A.; Elling, U.; et al. Canonical PRC1 controls sequence-independent propagation of Polycomb-mediated gene silencing. Nat. Commun. 2019, 10, 1931. [Google Scholar] [CrossRef] [PubMed]
- Simon, J.; Chiang, A.; Bender, W.; Shimell, M.J.; Connor, M.O. Elements of the Drosophila Bithorax Complex That Mediate Repression by Polycomb Group Products. Dev. Biol. 1993, 158, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Orsi, G.A.; Kasinathan, S.; Hughes, K.T.; Saminadin-Peter, S.; Henikoff, S.; Ahmad, K. High-resolution mapping defines the cooperative architecture of Polycomb response elements. Genome Res. 2014, 24, 809–820. [Google Scholar] [CrossRef]
- Brown, J.; Mucci, D.; Whiteley, M.; Dirksen, M.-L.; Kassis, J.A. The Drosophila Polycomb Group Gene pleiohomeotic Encodes a DNA Binding Protein with Homology to the Transcription Factor YY1. Mol. Cell 1998, 1, 1057–1064. [Google Scholar] [CrossRef]
- Brown, J.L.; Sun, M.-A.; Kassis, J.A. Global changes of H3K27me3 domains and Polycomb group protein distribution in the absence of recruiters Spps or Pho. Proc. Natl. Acad. Sci. USA 2018, 115, E1839–E1848. [Google Scholar] [CrossRef]
- Kassis, J.A.; Brown, J.L. Polycomb Group Response Elements in Drosophila and Vertebrates. Genet. Genom. Fish Phenomics 2013, 81, 83–118. [Google Scholar] [CrossRef]
- Erokhin, M.; Georgiev, P.; Chetverina, D. Drosophila DNA-Binding Proteins in Polycomb Repression. Epigenomes 2018, 2, 1. [Google Scholar] [CrossRef]
- Mendenhall, E.M.; Koche, R.P.; Truong, T.; Zhou, V.W.; Issac, B.; Chi, A.S.; Ku, M.; Bernstein, B.E. GC-Rich Sequence Elements Recruit PRC2 in Mammalian ES Cells. PLoS Genet. 2010, 6, e1001244. [Google Scholar] [CrossRef]
- Basu, A.; Wilkinson, F.H.; Colavita, K.; Fennelly, C.; Atchison, M.L. YY1 DNA binding and interaction with YAF2 is essential for Polycomb recruitment. Nucleic Acids Res. 2013, 42, 2208–2223. [Google Scholar] [CrossRef]
- Højfeldt, J.W.; Laugesen, A.; Willumsen, B.M.; Damhofer, H.; Hedehus, L.; Tvardovskiy, A.; Mohammad, F.; Jensen, O.N.; Helin, K. Accurate H3K27 methylation can be established de novo by SUZ12-directed PRC2. Nat. Struct. Mol. Biol. 2018, 25, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Brockdorff, N. Noncoding RNA and Polycomb recruitment. RNA 2013, 19, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Davidovich, C.; Cech, T.R. The recruitment of chromatin modifiers by long noncoding RNAs: Lessons from PRC2. RNA 2015, 21, 2007–2022. [Google Scholar] [CrossRef] [PubMed]
- Creyghton, M.P.; Cheng, A.W.; Welstead, G.G.; Kooistra, T.; Carey, B.W.; Steine, E.J.; Hanna, J.H.; Lodato, M.A.; Frampton, G.M.; Sharp, P.A.; et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 2010, 107, 21931–21936. [Google Scholar] [CrossRef]
- Huseyin, M.K.; Klose, R.J. Live-cell single particle tracking of PRC1 reveals a highly dynamic system with low target site occupancy. bioRxiv 2020. [Google Scholar]
- Eagen, K.P.; Aiden, E.L.; Kornberg, R.D. Polycomb-mediated chromatin loops revealed by a subkilobase-resolution chromatin interaction map. Proc. Natl. Acad. Sci. USA 2017, 114, 8764–8769. [Google Scholar] [CrossRef]
- Ogiyama, Y.; Schuettengruber, B.; Papadopoulos, G.L.; Chang, J.-M.; Cavalli, G. Polycomb-Dependent Chromatin Looping Contributes to Gene Silencing during Drosophila Development. Mol. Cell 2018, 71, 73–88.e5. [Google Scholar] [CrossRef]
- Kundu, S.; Ji, F.; Sunwoo, H.; Jain, G.; Lee, J.T.; Sadreyev, R.I.; Dekker, J.; Kingston, R.E. Polycomb Repressive Complex 1 Generates Discrete Compacted Domains that Change during Differentiation. Mol. Cell 2017, 65, 432–446.e5. [Google Scholar] [CrossRef]
- Schoenfelder, S.; Sugar, R.; Dimond, A.; Javierre, B.M.; Armstrong, H.; Mifsud, B.; Dimitrova, E.; Matheson, L.; Tavares-Cadete, F.; Furlan-Magaril, M.; et al. Polycomb repressive complex PRC1 spatially constrains the mouse embryonic stem cell genome. Nat. Genet. 2015, 47, 1179–1186. [Google Scholar] [CrossRef]
- Boyle, S.; Flyamer, I.M.; Williamson, I.; Sengupta, D.; Bickmore, W.A.; Illingworth, R.S. A central role for canonical PRC1 in shaping the 3D nuclear landscape. Genes Dev. 2020, 34, 931–949. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, J.D.; Feldmann, A.; Hernández-Rodríguez, B.; Díaz, N.; Brown, J.M.; Fursova, N.A.; Blackledge, N.P.; Prathapan, P.; Dobrinic, P.; Huseyin, M.K.; et al. Cohesin Disrupts Polycomb-Dependent Chromosome Interactions in Embryonic Stem Cells. Cell Rep. 2020, 30, 820–835.e10. [Google Scholar] [CrossRef] [PubMed]
- Davidson, I.F.; Bauer, B.; Goetz, D.; Tang, W.; Wutz, G.; Peters, J.-M. DNA loop extrusion by human cohesin. Science 2019, 366, 1338–1345. [Google Scholar] [CrossRef] [PubMed]
- Fudenberg, G.; Imakaev, M.; Lu, C.; Goloborodko, A.; Abdennur, N.; Mirny, L.A. Formation of Chromosomal Domains by Loop Extrusion. Cell Rep. 2016, 15, 2038–2049. [Google Scholar] [CrossRef] [PubMed]
- Cheutin, T.; Cavalli, G. Loss of PRC1 induces higher-order opening of Hox loci independently of transcription during Drosophila embryogenesis. Nat. Commun. 2018, 9, 3898. [Google Scholar] [CrossRef] [PubMed]
- Loubiere, V.; Papadopoulos, G.L.; Szabo, Q.; Martinez, A.-M.; Cavalli, G. Widespread activation of developmental gene expression characterized by PRC1-dependent chromatin looping. Sci. Adv. 2020, 6, eaax4001. [Google Scholar] [CrossRef]
- Banani, S.F.; Lee, H.O.; Hyman, A.A.; Rosen, M.K. Biomolecular condensates: Organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 2017, 18, 285–298. [Google Scholar] [CrossRef]
- Boija, A.; Klein, I.A.; Sabari, B.R.; Dall’Agnese, A.; Coffey, E.L.; Zamudio, A.V.; Li, C.H.; Shrinivas, K.; Manteiga, J.C.; Hannett, N.M.; et al. Transcription Factors Activate Genes through the Phase-Separation Capacity of Their Activation Domains. Cell 2018, 175, 1842–1855.e16. [Google Scholar] [CrossRef]
- Sabari, B.R.; Dall’Agnese, A.; Li, C.H.; Guo, Y.E.; Day, D.S.; Schuijers, J.; Vasile, E.; Malik, S.; Hnisz, D.; Lee, T.I.; et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361, eaar3958. [Google Scholar] [CrossRef]
- Larson, A.G.; Elnatan, D.; Keenen, M.M.; Trnka, M.J.; Johnston, J.B.; Burlingame, A.L.; Agard, D.A.; Redding, S.; Narlikar, G.J. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 2017, 547, 236–240. [Google Scholar] [CrossRef]
- Strom, A.R.; Emelyanov, A.V.; Mir, M.; Fyodorov, D.V.; Darzacq, X.; Karpen, G.H. Phase separation drives heterochromatin domain formation. Nature 2017, 547, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Grau, D.J.; Chapman, B.; Garlick, J.D.; Borowsky, M.; Francis, N.J.; Kingston, R.E. Compaction of chromatin by diverse Polycomb group proteins requires localized regions of high charge. Genes Dev. 2011, 25, 2210–2221. [Google Scholar] [CrossRef] [PubMed]
- Lau, M.S.; Schwartz, M.G.; Kingston, R.E.; Kundu, S.; Savol, A.J.; Wang, P.I.; Marr, S.K.; Grau, D.J.; Schorderet, P.; Sadreyev, R.I.; et al. Mutation of a nucleosome compaction region disrupts Polycomb-mediated axial patterning. Science 2017, 355, 1081–1084. [Google Scholar] [CrossRef] [PubMed]
- Tatavosian, R.; Kent, S.; Brown, K.; Yao, T.; Duc, H.N.; Huynh, T.N.; Zhen, C.Y.; Ma, B.; Wang, H.; Ren, X. Nuclear condensates of the Polycomb protein chromobox 2 (CBX2) assemble through phase separation. J. Biol. Chem. 2018, 294, 1451–1463. [Google Scholar] [CrossRef] [PubMed]
- De Graaff, W.; Tomotsune, D.; Oosterveen, T.; Takihara, Y.; Koseki, H.; Deschamps, J. Randomly inserted and targeted Hox/reporter fusions transcriptionally silenced in Polycomb mutants. Proc. Natl. Acad. Sci. USA 2003, 100, 13362–13367. [Google Scholar] [CrossRef]
- Shirai, M.; Osugi, T.; Koga, H.; Kaji, Y.; Takimoto, E.; Komuro, I.; Hara, J.; Miwa, T.; Yamauchi-Takihara, K.; Takihara, Y. The Polycomb-group gene Rae28 sustains Nkx2.5/Csx expression and is essential for cardiac morphogenesis. J. Clin. Investig. 2002, 110, 177–184. [Google Scholar] [CrossRef]
- Breiling, A.; O’Neill, L.P.; D’Eliseo, D.; Turner, B.M.; Orlando, V. Epigenome changes in active and inactive Polycomb-group-controlled regions. EMBO Rep. 2004, 5, 976–982. [Google Scholar] [CrossRef]
- Kwong, C.; Adryan, B.; Bell, I.; Meadows, L.; Russell, S.; Manak, J.R.; White, R. Stability and Dynamics of Polycomb Target Sites in Drosophila Development. PLoS Genet. 2008, 4, e1000178. [Google Scholar] [CrossRef]
- Papp, B.; Muller, J. Histone trimethylation and the maintenance of transcriptional ONand OFF states by trxG and PcG proteins. Genes Dev. 2006, 20, 2041–2054. [Google Scholar] [CrossRef]
- Langlais, K.K.; Brown, J.L.; Kassis, J.A. Polycomb Group Proteins Bind an engrailed PRE in Both the “ON” and “OFF” Transcriptional States of engrailed. PLoS ONE 2012, 7, e48765. [Google Scholar] [CrossRef]
- Fujimura, Y.-I.; Vidal, M.; Koseki, H.; Isono, K.-I.; Endoh, M.; Kajita, H.; Mizutani-Koseki, Y.; Takihara, Y.; Van Lohuizen, M.; Otte, A.; et al. Distinct roles of Polycomb group gene products in transcriptionallyrepressed and active domains of Hoxb8. Development 2006, 133, 2371–2381. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Wu, Z.J.; Groner, A.C.; He, H.H.; Cai, C.; Lis, R.T.; Wu, X.; Stack, E.C.; Loda, M.; Liu, T.; et al. EZH2 Oncogenic Activity in Castration-Resistant Prostate Cancer Cells Is Polycomb-Independent. Science 2012, 338, 1465–1469. [Google Scholar] [CrossRef] [PubMed]
- Chung, C.-Y.; Sun, Z.; Mullokandov, G.; Bosch, A.; Qadeer, Z.A.; Cihan, E.; Rapp, Z.; Parsons, R.; Aguirre-Ghiso, J.A.; Farias, E.F.; et al. Cbx8 Acts Non-canonically with Wdr5 to Promote Mammary Tumorigenesis. Cell Rep. 2016, 16, 472–486. [Google Scholar] [CrossRef] [PubMed]
- Loubiere, V.; Delest, A.; Thomas, A.; Bonev, B.; Schuettengruber, B.; Sati, S.; Martinez, A.-M.; Cavalli, G. Coordinate redeployment of PRC1 proteins suppresses tumor formation during Drosophila development. Nat. Genet. 2016, 48, 1436–1442. [Google Scholar] [CrossRef] [PubMed]
- Morey, L.; Aloia, L.; Cozzuto, L.; Benitah, S.A.; Di Croce, L. RYBP and Cbx7 Define Specific Biological Functions of Polycomb Complexes in Mouse Embryonic Stem Cells. Cell Rep. 2013, 3, 60–69. [Google Scholar] [CrossRef]
- Brookes, E.; De Santiago, I.; Hebenstreit, D.; Morris, K.J.; Carroll, T.; Xie, S.Q.; Stock, J.K.; Heidemann, M.; Eick, D.; Nozaki, N.; et al. Polycomb Associates Genome-wide with a Specific RNA Polymerase II Variant, and Regulates Metabolic Genes in ESCs. Cell Stem Cell 2012, 10, 157–170. [Google Scholar] [CrossRef]
- Frangini, A.; Sjöberg, M.; Roman-Trufero, M.; Dharmalingam, G.; Haberle, V.; Bartke, T.; Lenhard, B.; Malumbres, M.; Vidal, M.; Dillon, N. The Aurora B Kinase and the Polycomb Protein Ring1B Combine to Regulate Active Promoters in Quiescent Lymphocytes. Mol. Cell 2013, 51, 647–661. [Google Scholar] [CrossRef]
- Creppe, C.; Palau, A.; Malinverni, R.; Valero, V.; Buschbeck, M. A Cbx8-Containing Polycomb Complex Facilitates the Transition to Gene Activation during ES Cell Differentiation. PLoS Genet. 2014, 10, e1004851. [Google Scholar] [CrossRef]
- Schaaf, C.A.; Misulovin, Z.; Gause, M.; Koenig, A.; Gohara, D.W.; Watson, A.; Dorsett, D. Cohesin and Polycomb Proteins Functionally Interact to Control Transcription at Silenced and Active Genes. PLoS Genet. 2013, 9, e1003560. [Google Scholar] [CrossRef]
- Pemberton, H.; Anderton, E.; Patel, H.; Brookes, S.; Chandler, H.; Palermo, R.; Stock, J.; Rodriguez-Niedenführ, M.; Racek, T.; De Breed, L.; et al. Genome-wide co-localization of Polycomb orthologs and their effects on gene expression in human fibroblasts. Genome Biol. 2014, 15, R23. [Google Scholar] [CrossRef]
- Gao, Z.; Lee, P.; Stafford, J.M.; Von Schimmelmann, M.; Schaefer, A.; Reinberg, D. An AUTS2–Polycomb complex activates gene expression in the CNS. Nature 2014, 516, 349–354. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Shao, Z.; Li, D.; Xie, H.; Kim, W.; Huang, J.; Taylor, J.E.; Pinello, L.; Glass, K.; Jaffe, J.D.; et al. Developmental control of polycomb subunit composition by GATA factors mediates a switch to non-canonical functions. Mol. Cell 2015, 57, 304–316. [Google Scholar] [CrossRef] [PubMed]
- Morey, L.; Santanach, A.; Blanco, E.; Aloia, L.; Nora, E.P.; Bruneau, B.G.; Di Croce, L. Polycomb Regulates Mesoderm Cell Fate-Specification in Embryonic Stem Cells through Activation and Repression Mechanisms. Cell Stem Cell 2015, 17, 300–315. [Google Scholar] [CrossRef] [PubMed]
- Rai, K.; Akdemir, K.C.; Kwong, L.N.; Fiziev, P.; Wu, C.-J.; Keung, E.Z.; Sharma, S.; Samant, N.S.; Williams, M.; Axelrad, J.B.; et al. Dual Roles of RNF2 in Melanoma Progression. Cancer Discov. 2015, 5, 1314–1327. [Google Scholar] [CrossRef] [PubMed]
- Boom, V.V.D.; Maat, H.; Geugien, M.; López, A.R.; Sotoca, A.M.; Jaques, J.; Brouwers-Vos, A.Z.; Fusetti, F.; Groen, R.W.J.; Yuan, H.; et al. Non-canonical PRC1.1 Targets Active Genes Independent of H3K27me3 and Is Essential for Leukemogenesis. Cell Rep. 2016, 14, 332–346. [Google Scholar] [CrossRef] [PubMed]
- Maezawa, S.; Hasegawa, K.; Yukawa, M.; Sakashita, A.; Alavattam, K.G.; Andreassen, P.R.; Vidal, M.; Koseki, H.; Barski, A.; Namekawa, S.H. Polycomb directs timely activation of germline genes in spermatogenesis. Genes Dev. 2017, 31, 1693–1703. [Google Scholar] [CrossRef]
- Pherson, M.; Misulovin, Z.; Gause, M.; Mihindukulasuriya, K.A.; Swain, A.; Dorsett, D. Polycomb repressive complex 1 modifies transcription of active genes. Sci. Adv. 2017, 3, e1700944. [Google Scholar] [CrossRef]
- Cohen, I.; Zhao, D.; Zheng, D.; Ezhkova, E. 1358 PRC1 fine-tunes gene repression and activation to safeguard skin epithelium development and stem cell specification. J. Investig. Dermatol. 2018, 138, S231. [Google Scholar] [CrossRef]
- Cohen, I.; Zhao, D.-J.; Menon, G.; Nakayama, M.; Koseki, H.; Zheng, D.; Ezhkova, E. PRC1 preserves epidermal tissue integrity independently of PRC2. Genes Dev. 2018, 33, 55–60. [Google Scholar] [CrossRef]
- Chan, H.L.; Beckedorff, F.; Zhang, Y.; Garcia-Huidobro, J.; Jiang, H.; Colaprico, A.; Bilbao, D.; Figueroa, M.E.; Lacava, J.; Shiekhattar, R.; et al. Polycomb complexes associate with enhancers and promote oncogenic transcriptional programs in cancer through multiple mechanisms. Nat. Commun. 2018, 9, 3377. [Google Scholar] [CrossRef]
- Zhang, Y.; Chan, H.L.; Garcia-Martinez, L.; Slingerland, J.M.; Karl, D.L.; Verdun, R.E.; Morey, L. Estrogen induces dynamic ERα and RING1B recruitment to control gene and enhancer activities in luminal breast cancer. Sci. Adv. 2020, 6, 7249–7254. [Google Scholar] [CrossRef] [PubMed]
- Bradner, J.E.; Hnisz, D.; Young, R.A. Transcriptional Addiction in Cancer. Cell 2017, 168, 629–643. [Google Scholar] [CrossRef] [PubMed]
- Vo, L.T.; Kinney, M.A.; Liu, X.; Zhang, Y.; Barragan, J.; Sousa, P.M.; Jha, D.K.; Han, A.; Cesana, M.; Shao, Z.; et al. Regulation of embryonic haematopoietic multipotency by EZH1. Nature 2018, 553, 506–510. [Google Scholar] [CrossRef] [PubMed]
- Rada-Iglesias, Á.; Bajpai, R.; Swigut, T.; Brugmann, S.A.; Flynn, R.A.; Wysocka, J. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 2010, 470, 279–283. [Google Scholar] [CrossRef] [PubMed]
- Scelfo, A.; Fernández-Pérez, D.; Tamburri, S.; Zanotti, M.; Lavarone, E.; Soldi, M.; Bonaldi, T.; Ferrari, K.J.; Pasini, D. Functional Landscape of PCGF Proteins Reveals Both RING1A/B-Dependent-and RING1A/B-Independent-Specific Activities. Mol. Cell 2019, 74, 1037–1052.e7. [Google Scholar] [CrossRef]
- Zhao, W.; Huang, Y.; Zhang, J.; Liu, M.; Ji, H.; Wang, C.; Cao, N.; Li, C.; Xia, Y.; Jiang, Q.; et al. Polycomb group RING finger proteins 3/5 activate transcription via an interaction with the pluripotency factor Tex10 in embryonic stem cells. J. Biol. 2017, 292, 21527–21537. [Google Scholar]
- Cordonnier, G.; Mandoli, A.; Cagnard, N.; Hypolite, G.; Lhermitte, L.; Verhoeyen, E.; Asnafi, V.; Dillon, N.; MacIntyre, E.; Martens, J.H.; et al. CBFβ-SMMHC Affects Genome-wide Polycomb Repressive Complex 1 Activity in Acute Myeloid Leukemia. Cell Rep. 2020, 30, 299–307.e3. [Google Scholar] [CrossRef]
- Illingworth, R.S.; Botting, C.H.; Grimes, G.R.; Bickmore, W.A.; Eskeland, R. PRC1 and PRC2 Are Not Required for Targeting of H2A.Z to Developmental Genes in Embryonic Stem Cells. PLoS ONE 2012, 7, e34848. [Google Scholar] [CrossRef]
- Kang, H.; Jung, Y.L.; McElroy, K.A.; Zee, B.M.; Wallace, H.A.; Woolnough, J.L.; Park, P.J.; Kuroda, M.I. Bivalent complexes of PRC1 with orthologs of BRD4 and MOZ/MORF target developmental genes inDrosophila. Genes Dev. 2017, 31, 1988–2002. [Google Scholar] [CrossRef]
- Dou, Y.; Milne, T.; Tackett, A.J.; Smith, E.R.; Fukuda, A.; Wysocka, J.; Allis, C.D.; Chait, B.T.; Hess, J.L.; Roeder, R.G. Physical Association and Coordinate Function of the H3 K4 Methyltransferase MLL1 and the H4 K16 Acetyltransferase MOF. Cell 2005, 121, 873–885. [Google Scholar] [CrossRef]
- Strübbe, G.; Popp, C.; Schmidt, A.; Pauli, A.; Ringrose, L.; Beisel, C.; Paro, R. Polycomb purification by in vivo biotinylation tagging reveals cohesin and Trithorax group proteins as interaction partners. Proc. Natl. Acad. Sci. USA 2011, 108, 5572–5577. [Google Scholar] [CrossRef] [PubMed]
- Kar, G.; Kim, J.K.; Kolodziejczyk, A.A.; Natarajan, K.N.; Triglia, E.T.; Mifsud, B.; Elderkin, S.; Marioni, J.C.; Pombo, A.; Teichmann, S.A. Flipping between Polycomb repressed and active transcriptional states introduces noise in gene expression. Nat. Commun. 2017, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Lenstra, T.L.; Rodriguez, J.; Chen, H.; Larson, D.R. Transcription Dynamics in Living Cells. Annu. Rev. Biophys. 2016, 45, 25–47. [Google Scholar] [CrossRef]
- Fukaya, T.; Lim, B.; Levine, M. Enhancer Control of Transcriptional Bursting. Cell 2016, 166, 358–368. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Zhao, W.; Huang, Y.; Tong, H.; Xia, Y.; Jiang, Q.; Qin, J. Loss of Polycomb Group Protein Pcgf1 Severely Compromises Proper Differentiation of Embryonic Stem Cells. Sci. Rep. 2017, 7, 46276. [Google Scholar] [CrossRef]
- Ahmad, K.; Spens, A.E. Separate Polycomb Response Elements control chromatin state and activation of the vestigial gene. PLoS Genet. 2019, 15, e1007877. [Google Scholar] [CrossRef]
- Kondo, T.; Isono, K.; Kondo, K.; Endo, T.A.; Itohara, S.; Vidal, M.; Koseki, H. Polycomb Potentiates Meis2 Activation in Midbrain by Mediating Interaction of the Promoter with a Tissue-Specific Enhancer. Dev. Cell 2014, 28, 94–101. [Google Scholar] [CrossRef]
- Cruz-Molina, S.; Respuela, P.; Tebartz, C.; Kolovos, P.; Nikolic, M.; Fueyo, R.; Van Ijcken, W.; Grosveld, F.; Frommolt, P.; Bazzi, H.; et al. PRC2 Facilitates the Regulatory Topology Required for Poised Enhancer Function during Pluripotent Stem Cell Differentiation. Cell Stem Cell 2017, 20, 689–705.e9. [Google Scholar] [CrossRef]
- Mikulski, P.; Komarynets, O.; Fachinelli, F.; Weber, A.P.; Schubert, D. Characterization of the Polycomb-Group Mark H3K27me3 in Unicellular Algae. Front. Plant Sci. 2017, 8, 505–512. [Google Scholar] [CrossRef]

| Complex | Mammals | Flies | Protein Motifs | Functions |
|---|---|---|---|---|
| PRC1 core subunits | RING1/RING1A, RNF2/RING1B | Sce | RING finger, RAWUL domain | RING1-PCGF pairs as heterodimeric E3 ligases that monoubiquitylate H2A |
| PCGF1, PCGF2/MEL18, PCGF3, PCGF4/BMI1, PCGF5, PCGF6 | Psc, Su(z)2, L(3)73Ah | RING finger, RAWUL domain | ||
| canonical PRC1 | CBX2, CBX4, CBX6 CBX7, CBX8 | Pc | Chromobox | H3K27me3 recognition |
| PHC1, PHC2, PHC3 | Ph | SAM domain | Oligomerization, high order structures | |
| variant PRC1 | KDM2B | dKDM2 | CXXC motif, jmjC, Fbox, LRR | DNA binding |
| RYBP, YAF2 | dRYBP | Zn finger | H2AUb recognition | |
| MGA-MAX | (1) | heterodimeric DNA binding module | ||
| E2F-TFDP1 | (1) | |||
| L3MBTL2 | dSfmbt | MBT domains | ||
| PRC2 core subunits | EZH1/EZH2 | E(z) | SET domain | H3K27 methyltransferase |
| SUZ12 | Su(z)12 | Several | Allosteric integration, recruitment | |
| EED | esc | WD repeats | H3K27me recognition | |
| RBBP4, RBBP7 | Caf1 | WD repeats | H3, H4 recognition | |
| PRC2.1 | PCL1, PCL2, PCL3 | Pcl | DNA binding | |
| PALI1, PALI2, PALI3 | (1) | Protein-protein interactions | ||
| PRC2.2 | JARID2 | Jarid2 | JmjC, ARID | H2AUb recognition, recruitment |
| AEBP2 | Jing | Zn finger | DNA binding |
| PRC Subunits | Cell Type | Reference |
|---|---|---|
| RING1B, EZH2, SUZ12 | Murine ESCs | [125,126] |
| RING1B, CBX7, EZH2 | Murine quiescent B-cells | [127] |
| CBX8 | Murine neural progenitors | [128] |
| Pc, PSc, Ph | Drosophila imaginal discs | [129] |
| CBX6, 7, 8; RING1A, RING1B | Human fibroblast cell lines | [130] |
| RING1B | Postnatal mouse brain cells | [131] |
| EZH1, EZH2, EED, SUZ12 | Human differentiating erythroid cells | [132] |
| RING1B, PCGF2, CBX2, RYBP | Murine cardiac-mesoderm precursor cells | [133] |
| RING1B | Human melanoma cell lines | [134] |
| RING1B, PCGF2 | Neural progenitors | [57] |
| RING1A, RING1B, CBX2, PCGF1, KDM2B | Human erythroleukemic K562 cell line, AML patient cells | [135] |
| Pc, Ph | Drosophila embryo, imaginal discs | [124] |
| RING1B, PCGF4 | Human fibroblasts, K562 cells | [124] |
| RING1B, PCGF2 | Murine spermatogonia cells | [136] |
| Pc, Ph | Drosophila BG3 cell line | [137] |
| RING1B, RYBP, PCGF4, KDM2B, L3MBTL2 | Murine epidermal progenitors | [138,139] |
| RING1B | Human breast tumor cell lines MCF10A, T47D, MDA-MB-231; Human liver cancer cell line Hep G2, K562 cells | [140] |
| CBX4, PCGF2, PCGF4 | Human breast tumor cell lines | [140,141] |
| RING1B | Human leukemic cell line ME-1 | [142] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giner-Laguarda, N.; Vidal, M. Functions of Polycomb Proteins on Active Targets. Epigenomes 2020, 4, 17. https://doi.org/10.3390/epigenomes4030017
Giner-Laguarda N, Vidal M. Functions of Polycomb Proteins on Active Targets. Epigenomes. 2020; 4(3):17. https://doi.org/10.3390/epigenomes4030017
Chicago/Turabian StyleGiner-Laguarda, Natalia, and Miguel Vidal. 2020. "Functions of Polycomb Proteins on Active Targets" Epigenomes 4, no. 3: 17. https://doi.org/10.3390/epigenomes4030017
APA StyleGiner-Laguarda, N., & Vidal, M. (2020). Functions of Polycomb Proteins on Active Targets. Epigenomes, 4(3), 17. https://doi.org/10.3390/epigenomes4030017