RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions
Abstract
:Simple Summary
Abstract
1. Introduction
2. RNAi in the Western Corn Rootworm
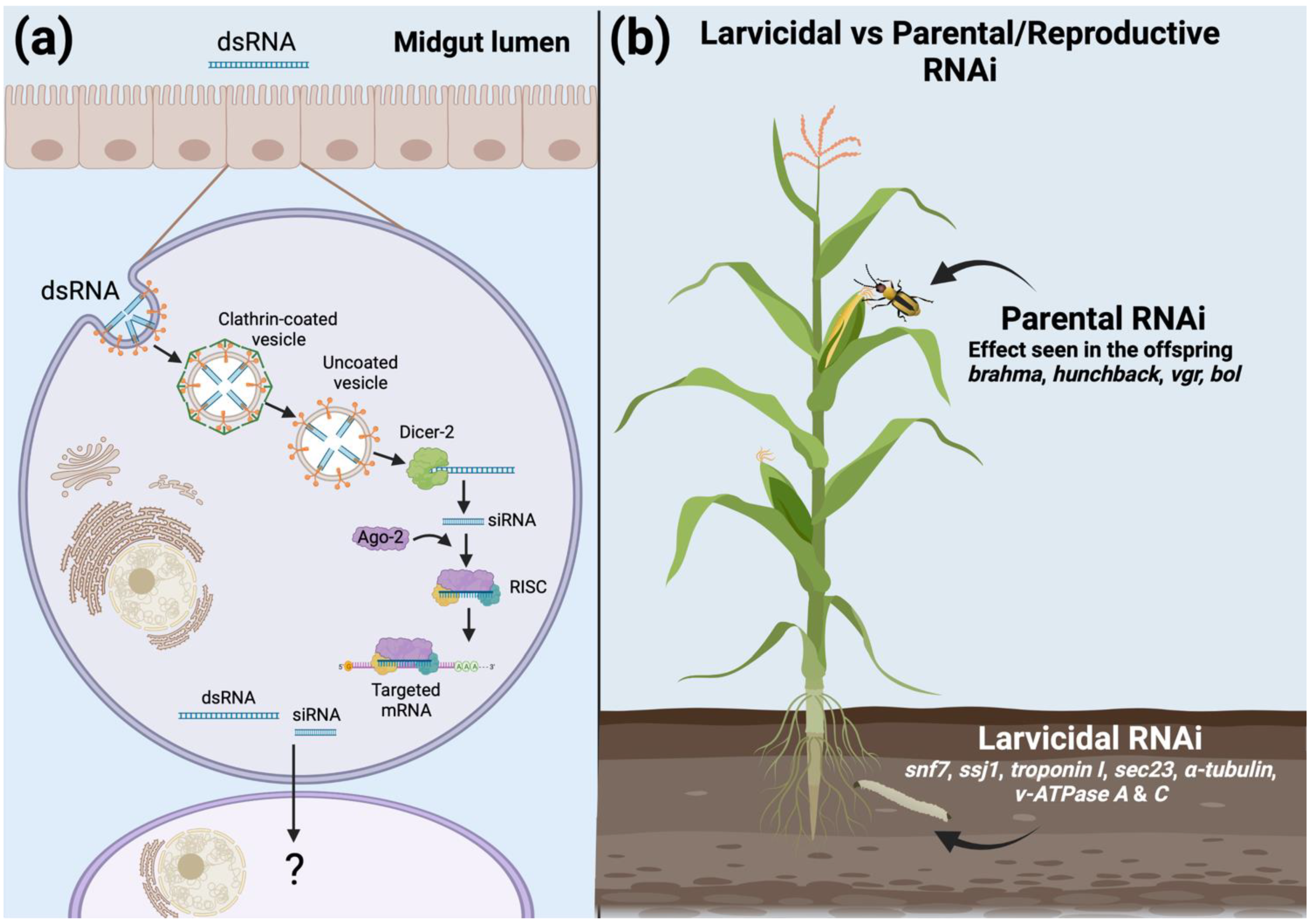
2.1. dsRNA Uptake
2.2. Silencing Mechanism
2.3. Systemic Spread
3. RNAi Traits for Rootworm Control
3.1. RNAi as a New Mode of Action for Rootworm Control
3.2. Target Site Discovery
3.2.1. Larvicidal RNAi
3.2.2. Parental and Reproductive RNAi
3.3. Transgenic dsRNA Events
4. Field Efficacy of RNAi for Insect Control, Insect Resistance Management, and RNAi Resistance
4.1. Field Efficacy of RNAi Traits
4.2. Insect Resistance Management
4.3. Rootworm Resistance to RNAi
5. Environmental Risk Assessment
5.1. Environmental Risk Assessment Principles
5.2. Environmental Risk Assessment for RNAi Traits
5.3. Evaluation of Western Corn Rootworm RNAi Traits on Non-Target Organisms
6. RNAi Mammalian Safety
6.1. RNA and Its History of Safe Consumption
6.2. Biological Barriers to RNA Absorption
6.3. Evaluation of Potential Activity or Adverse Effects of Ingested RNA
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Sappington, T.W.; Siegfried, B.D.; Guillemaud, T. Coordinated Diabrotica genetics research: Accelerating progress on an urgent insect pest problem. Am. Entomol. 2006, 52, 90–97. [Google Scholar] [CrossRef] [Green Version]
- Gray, M.E.; Sappington, T.W.; Miller, N.J.; Moeser, J.; Bohn, M.O. Adaptation and invasiveness of western corn rootworm: Intensifying research on a worsening pest. Annu. Rev. Entomol. 2009, 54, 303–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gillette, C.P. Diabrotica virgifera LeConte as a corn rootworm. J. Econ. Entomol. 1912, 54, 364–366. [Google Scholar] [CrossRef]
- Melhus, I.E.; Painter, R.H.; Smith, F.O. A search for resistance to the injury caused by species of Diabrotica in the corns of Guatemala. Iowa State Coll. J. Sci. 1954, 29, 75–94. [Google Scholar]
- Baca, F. New member of the harmful entomofauna of Yugoslavia Diabrotica virgifera virgifera LeConte (Coleoptera: Chrysomelidae). Zast Bilja 1994, 45, 125–131. [Google Scholar]
- Kiss, J.; Edwards, C.R.; Berger, H.K.; Cate, P.; Cean, M.; Cheek, S.; Derron, J.; Festic, H.; Furlan, L.; Iger-Barcic, J.; et al. Monitoring of western corn rootworm (Diabrotica virgifera virgifera LeConte) in Europe 1992–2003. In Western Corn Rootworm: Ecology and Management; Vidal, S., Kuhlman, U., Eds.; C.R. Edwards, CABI International: Wallingford, UK, 2005; Volume 2, pp. 29–39. [Google Scholar]
- Meinke, L.J.; Souza, D.; Siegfried, B.D. The use of insecticides to manage the western corn rootworm, Diabrotica virgifera virgifera, LeConte: History, field-evolved resistance, and associated mechanisms. Insects 2021, 12, 112. [Google Scholar] [CrossRef] [PubMed]
- Gassmann, A.J. Resistance to Bt maize by western corn rootworm: Effects of pest biology, the pest–crop interaction and the agricultural landscape on resistance. Insects 2021, 12, 136. [Google Scholar] [CrossRef]
- Metcalf, R.L. Implications and prognosis of resistance to insecticides. In Pest Resistance to Pesticides; Georghiou, G.P., Saito, T., Eds.; Plenum Press: New York, NY, USA, 1983; pp. 769–792. [Google Scholar]
- Meinke, L.J.; Siegfried, B.D.; Wright, R.J.; Chandler, L.D. Adult susceptibility of Nebraska populations of western corn rootworm to selected insecticides. J. Econ. Entomol. 1998, 91, 594–600. [Google Scholar] [CrossRef] [Green Version]
- Levine, E.; Spencer, J.L.; Isard, S.A.; Onstad, D.W.; Gray, M.E. Adaptation of the western corn rootworm to crop rotation: Evolution of a new strain in response to a cultural management practice. Am. Entomol. 2002, 48, 94–107. [Google Scholar] [CrossRef] [Green Version]
- Pereira, A.E.; Wang, H.; Zukoff, S.N.; Meinke, L.J.; French, B.W.; Siegfried, B.D. Evidence of field-evolved resistance to bifenthrin in western corn rootworm (Diabrotica virgifera virgifera LeConte) populations in western Nebraska and Kansas. PLoS ONE 2015, 10, e0142299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Souza, D.; Vieira, B.C.; Fritz, B.K.; Hoffmann, W.C.; Peterson, J.A.; Kruger, G.R.; Meinke, L.J. Western corn rootworm pyrethroid resistance confirmed by aerial application simulations of commercial insecticides. Sci. Rep. 2019, 9, 6713. [Google Scholar] [CrossRef] [Green Version]
- Gassmann, A.J.; Petzold-Maxwell, J.L.; Clifton, E.H.; Dunbar, M.W.; Hoffmann, A.M.; Ingber, D.A.; Keweshan, R.S. Field-evolved resistance by western corn rootworm to multiple Bacillus thuringiensis toxins in transgenic maize. Proc. Natl. Acad. Sci. USA 2014, 111, 5141–5146. [Google Scholar] [CrossRef] [Green Version]
- Wechsler, S.; Smith, D. Has resistance taken root in U.S. corn fields? Demand for insect control. Am. J. Agric. Econ. 2018, 100, 1136–1150. [Google Scholar] [CrossRef]
- Crickmore, N.; Berry, C.; Panneerselvam, S.; Mishra, R.; Connor, T.R.; Bonning, B.C. A structure-based nomenclature for Bacillus thuringiensis and other bacteria-derived pesticidal proteins. J. Invertebr. Pathol. 2020, 107438. [Google Scholar] [CrossRef] [PubMed]
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, Corn Event MON 863: Corn Rootworm-Protected Corn. 2003. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/000524-00528-20030224.pdf (accessed on 27 December 2021).
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, Agrisure™ RW Rootworm-Protected Com. 2006. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/067979-00005-20061003.pdf (accessed on 27 December 2021).
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, 5307 Corn. 2012. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/067979-00022-20120731.pdf (accessed on 27 December 2021).
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, Herculex Xtra Insect Protection. 2005. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/029964-00005-20051027.pdf (accessed on 27 December 2021).
- Gassmann, A.J.; Petzold-Maxwell, J.L.; Keweshan, R.S.; Dunbar, M.W. Field-evolved resistance to Bt maize by western corn rootworm. PLoS ONE 2011, 6, e22629. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrader, P.M.; Estes, R.E.; Tinsley, N.A.; Gassmann, A.J.; Gray, M.E. Evaluation of adult emergence and larval root injury for Cry3Bb1-resistant populations of the western corn rootworm. J. Appl. Entomol. 2017, 141, 41–52. [Google Scholar] [CrossRef] [Green Version]
- Reinders, J.D.; Hitt, B.D.; Stroup, W.W.; French, B.W.; Meinke, L.J. Spatial variation in western corn rootworm (Coleoptera: Chrysomelidae) susceptibility to Cry3 toxins in Nebraska. PLoS ONE 2018, 13, e0208266. [Google Scholar] [CrossRef] [PubMed]
- Calles-Torrez, V.; Knodel, J.J.; Boetel, M.A.; French, B.W.; Fuller, B.W.; Ransom, J.K. Field-evolved resistance of northern and western corn rootworm (Coleoptera: Chrysomelidae) populations to corn hybrids expressing single and pyramided Cry3Bb1 and Cry34/35Ab1 Bt proteins in North Dakota. J. Econ. Entomol. 2019, 112, 1875–1886. [Google Scholar] [CrossRef] [PubMed]
- Jakka, S.R.; Shrestha, R.B.; Gassmann, A.J. Broad-spectrum resistance to Bacillus thuringiensis toxins by western corn rootworm (Diabrotica virgifera virgifera). Sci. Rep. 2016, 6, 27860. [Google Scholar] [CrossRef]
- Wangila, D.S.; Gassmann, A.J.; Petzold-Maxwell, J.L.; French, B.W.; Meinke, L.J. Susceptibility of Nebraska western corn rootworm (Coleoptera: Chrysomelidae) populations to Bt corn events. J. Econ. Entomol. 2015, 108, 742–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gassmann, A.J.; Shrestha, R.B.; Kropf, A.L.; St Clair, C.R.; Brenizer, B.D. Field-evolved resistance by western corn rootworm to Cry34/35Ab1 and other Bacillus thuringiensis traits in transgenic maize. Pest Manag. Sci. 2020, 76, 268–276. [Google Scholar] [CrossRef] [PubMed]
- Zukoff, S.N.; Ostlie, K.R.; Potter, B.; Meihls, L.N.; Zukoff, A.L.; French, L.; Ellersieck, M.R.; Wade French, B.; Hibbard, B.E. Multiple assays indicate varying levels of cross resistance in Cry3Bb1-selected field populations of the western corn rootworm to mCry3A, eCry3.1Ab, and Cry34/35Ab1. J. Econ. Entomol. 2016, 109, 1387–1398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gassmann, A.J.; Shrestha, R.B.; Jakka, S.R.; Dunbar, M.W.; Clifton, E.H.; Paolino, A.R.; Ingber, D.A.; French, B.W.; Masloski, K.E.; Dounda, J.W.; et al. Evidence of resistance to Cry34/35Ab1 corn by western corn rootworm (Coleoptera: Chrysomelidae): Root injury in the field and larval survival in plant-based bioassays. J. Econ. Entomol. 2016, 109, 1872–1880. [Google Scholar] [CrossRef]
- Ludwick, D.C.; Meihls, L.N.; Ostlie, K.R.; Potter, B.D.; French, L.; Hibbard, B.E. Minnesota field population of western corn rootworm (Coleoptera: Chrysomelidae) shows incomplete resistance to Cry34Ab1/Cry35Ab1 and Cry3Bb1. J. Appl. Entomol. 2017, 141, 28–40. [Google Scholar] [CrossRef] [Green Version]
- Bolognesi, R.; Ramaseshadri, P.; Anderson, J.; Bachman, P.; Clinton, W.; Flannagan, R.; Ilagan, O.; Lawrence, C.; Levine, S.; Moar, W.; et al. Characterizing the mechanism of action of double-stranded RNA activity against western corn rootworm (Diabrotica virgifera virgifera LeConte). PLoS ONE 2012, 7, e47534. [Google Scholar] [CrossRef]
- Head, G.P.; Carroll, M.W.; Evans, S.P.; Rule, D.M.; Willse, A.R.; Clark, T.L.; Storer, N.P.; Flannagan, R.D.; Samuel, L.W.; Meinke, L.J. Evaluation of SmartStax and SmartStax PRO maize against western corn rootworm and northern corn rootworm: Efficacy and resistance management. Pest Manag. Sci. 2017, 73, 1883–1899. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, SmartStax® PRO Enlist™ Refuge Advanced®. 2017. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/062719-00707-20170608.pdf (accessed on 27 December 2021).
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.V.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA interference: Biology, mechanism, and applications. Microbiol. Mol. Biol. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Timmons, L.; Fire, A. Specific interference by ingested dsRNA. Nature 1998, 395, 854. [Google Scholar] [CrossRef] [PubMed]
- Ghildiyal, M.; Zamore, P.D. Small silencing RNAs: An expanding universe. Nat. Rev. Genet. 2009, 10, 94–108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shabalina, S.A.; Koonin, E.V. Origins and evolution of eukaryotic RNA interference. Trends Ecol. Evol. 2008, 23, 578–587. [Google Scholar] [CrossRef] [Green Version]
- Mukherjee, K.; Campos, H.; Kolaczkowski, B. Evolution of animal and plant dicers: Early parallel duplications and recurrent adaptation of antiviral RNA binding in plants. Mol. Biol. Evol. 2013, 30, 627–641. [Google Scholar] [CrossRef] [Green Version]
- Zhu, K.Y.; Palli, S.R. Mechanisms, Applications, and challenges of insect RNA interference. Annu. Rev. Entomol. 2020, 65, 293–311. [Google Scholar] [CrossRef] [Green Version]
- Ender, C.; Meister, G. Argonaute proteins at a glance. J. Cell Sci. 2010, 123, 1819–1823. [Google Scholar] [CrossRef] [Green Version]
- Tang, R.; Li, L.; Zhu, D.; Hou, D.; Cao, T.; Gu, H.; Zhang, J.; Chen, J.; Zhang, C.Y.; Zen, K. Mouse miRNA-709 directly regulates miRNA-15a/16-1 biogenesis at the posttranscriptional level in the nucleus: Evidence for a microRNA hierarchy system. Cell Res. 2012, 22, 504–515. [Google Scholar] [CrossRef]
- Olsen, P.H.; Ambros, V. The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev. Biol. 1999, 216, 671–680. [Google Scholar] [CrossRef] [Green Version]
- Seggerson, K.; Tang, L.; Moss, E.G. Two genetic circuits repress the Caenorhabditis elegans heterochronic gene lin-28 after translation initiation. Dev. Biol. 2002, 243, 215–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Krützfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef]
- Bagga, S.; Bracht, J.; Hunter, S.; Massirer, K.; Holtz, J.; Eachus, R.; Pasquinelli, A.E. Regulation by let-7 and lin-4 miRNAs results in target mRNA degradation. Cell 2005, 122, 553–563. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.; Belasco, J.G. Micro-RNA regulation of the mammalian lin-28 gene during neuronal differentiation of embryonal carcinoma cells. Mol. Cell. Biol. 2005, 25, 9198–9208. [Google Scholar] [CrossRef] [Green Version]
- Aravin, A.A.; Naumova, N.M.; Tulin, A.V.; Vagin, V.V.; Rozovsky, Y.M.; Gvozdev, V.A. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 2001, 11, 1017–1027. [Google Scholar] [CrossRef] [Green Version]
- Cox, D.N.; Chao, A.; Baker, J.; Chang, L.; Qiao, D.; Lin, H. A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev. 1998, 12, 3715–3727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, N.; Kazazian, H.H. L1 retrotransposition is suppressed by endogenously encoded small interfering RNAs in human cultured cells. Nat. Struct. Mol. Biol. 2006, 13, 763–771. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, B.L.; Langland, J.O. When two strands are better than one: The mediators and modulators of the cellular responses to double-stranded RNA. Virology 1996, 219, 339–349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bellés, X. Beyond Drosophila: RNAi in vivo and functional genomics in insects. Annu. Rev. Entomol. 2010, 55, 111–128. [Google Scholar] [CrossRef] [Green Version]
- Price, D.R.; Gatehouse, J.A. RNAi-mediated crop protection against insects. Trends Biotechnol. 2008, 26, 393–400. [Google Scholar] [CrossRef]
- Baum, J.A.; Bogaert, T.; Clinton, W.; Heck, G.R.; Feldmann, P.; Ilagan, O.; Johnson, S.; Plaetinck, G.; Munyikwa, T.; Pleau, M.; et al. Control of coleopteran insect pests through RNA interference. Nat. Biotechnol. 2007, 25, 1322–1326. [Google Scholar] [CrossRef]
- Alves, A.P.; Lorenzen, M.D.; Beeman, R.W.; Foster, J.E.; Siegfried, B.D. RNA interference as a method for target-site screening in the Western corn rootworm, Diabrotica virgifera virgifera. J. Insect Sci. 2010, 10, 162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Winston, W.M.; Molodowitch, C.; Hunter, C.P. Systemic RNAi in C. elegans requires the putative transmembrane protein SID-1. Science 2002, 295, 2456–2459. [Google Scholar] [CrossRef] [Green Version]
- Feinberg, E.H.; Hunter, C.P. Transport of dsRNA into cells by the transmembrane protein SID-1. Science 2003, 301, 1545–1547. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McEwan, D.L.; Weisman, A.S.; Hunter, C.P. Uptake of extracellular double-stranded RNA by SID-2. Mol. Cell 2012, 47, 746–754. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jose, A.M.; Kim, Y.A.; Leal-Ekman, S.; Hunter, C.P. Conserved tyrosine kinase promotes the import of silencing RNA into Caenorhabditis elegans cells. Proc. Natl. Acad. Sci. USA 2012, 109, 14520–14525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hinas, A.; Wright, A.J.; Hunter, C.P. SID-5 is an endosome-associated protein required for efficient systemic RNAi in C. elegans. Curr. Biol. 2012, 22, 1938–1943. [Google Scholar] [CrossRef] [Green Version]
- Tomoyasu, Y.; Miller, S.C.; Tomita, S.; Schoppmeier, M.; Grossmann, D.; Bucher, G. Exploring systemic RNA interference in insects: A genome-wide survey for RNAi genes in Tribolium. Genome Biol. 2008, 9, R10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miyata, K.; Ramaseshadri, P.; Zhang, Y.; Segers, G.; Bolognesi, R.; Tomoyasu, Y. Establishing an in vivo assay system to identify components involved in environmental RNA interference in the western corn rootworm. PLoS ONE 2014, 9, e101661. [Google Scholar] [CrossRef] [Green Version]
- Pinheiro, D.H.; Vélez, A.M.; Fishilevich, E.; Wang, H.; Carneiro, N.P.; Valencia-Jiménez, A.; Valicente, F.H.; Narva, K.E.; Siegfried, B.D. Clathrin-dependent endocytosis is associated with RNAi response in the western corn rootworm, Diabrotica virgifera virgifera LeConte. PLoS ONE 2018, 13, e0201849. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cappelle, K.; de Oliveira, C.F.; Van Eynde, B.; Christiaens, O.; Smagghe, G. The involvement of clathrin-mediated endocytosis and two Sid-1-like transmembrane proteins in double-stranded RNA uptake in the Colorado potato beetle midgut. Insect Mol. Biol. 2016, 25, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Gruenberg, J. The endocytic pathway: A mosaic of domains. Nat. Rev. Mol. Cell Biol. 2001, 2, 721–730. [Google Scholar] [CrossRef]
- Marsh, M.; Helenius, A. Virus entry: Open sesame. Cell 2006, 124, 729–740. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sorkin, A. Cargo recognition during clathrin-mediated endocytosis: A team effort. Curr. Opin. Cell Biol. 2004, 16, 392–399. [Google Scholar] [CrossRef] [PubMed]
- McMahon, H.T.; Boucrot, E. Molecular mechanism and physiological functions of clathrin-mediated endocytosis. Nat. Rev. Mol. Cell Biol. 2011, 12, 517–533. [Google Scholar] [CrossRef] [PubMed]
- Saleh, M.C.; van Rij, R.P.; Hekele, A.; Gillis, A.; Foley, E.; O’Farrell, P.H.; Andino, R. The endocytic pathway mediates cell entry of dsRNA to induce RNAi silencing. Nat. Cell Biol. 2006, 8, 793–802. [Google Scholar] [CrossRef]
- Xiao, D.; Gao, X.; Xu, J.; Liang, X.; Li, Q.; Yao, J.; Zhu, K.Y. Clathrin-dependent endocytosis plays a predominant role in cellular uptake of double-stranded RNA in the red flour beetle. Insect Biochem. Mol. Biol. 2015, 60, 68–77. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.C.; Fu, K.Y.; Yang, S.; Li, X.X.; Li, G.Q. Instar-dependent systemic RNA interference response in Leptinotarsa decemlineata larvae. Pestic Biochem. Physiol. 2015, 123, 64–73. [Google Scholar] [CrossRef]
- Ulvila, J.; Parikka, M.; Kleino, A.; Sormunen, R.; Ezekowitz, R.A.; Kocks, C.; Rämet, M. Double-stranded RNA is internalized by scavenger receptor-mediated endocytosis in Drosophila S2 cells. J. Biol. Chem. 2006, 281, 14370–14375. [Google Scholar] [CrossRef] [Green Version]
- Vélez, A.M.; Fishilevich, E. The mysteries of insect RNAi: A focus on dsRNA uptake and transport. Pestic. Biochem. Physiol. 2018, 151, 25–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khajuria, C.; Ivashuta, S.; Wiggins, E.; Flagel, L.; Moar, W.; Pleau, M.; Miller, K.; Zhang, Y.; Ramaseshadri, P.; Jiang, C.; et al. Development and characterization of the first dsRNA-resistant insect population from western corn rootworm, Diabrotica virgifera virgifera LeConte. PLoS ONE 2018, 13, e0197059. [Google Scholar] [CrossRef] [Green Version]
- Cooper, A.M.; Silver, K.; Zhang, J.; Park, Y.; Zhu, K.Y. Molecular mechanisms influencing efficiency of RNA interference in insects. Pest Manag. Sci. 2019, 75, 18–28. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grishok, A.; Pasquinelli, A.E.; Conte, D.; Li, N.; Parrish, S.; Ha, I.; Baillie, D.L.; Fire, A.; Ruvkun, G.; Mello, C.C. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 2001, 106, 23–34. [Google Scholar] [CrossRef] [Green Version]
- Bernstein, E.; Caudy, A.A.; Hammond, S.M.; Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001, 409, 363–366. [Google Scholar] [CrossRef]
- Lee, Y.S.; Nakahara, K.; Pham, J.W.; Kim, K.; He, Z.; Sontheimer, E.J.; Carthew, R.W. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 2004, 117, 69–81. [Google Scholar] [CrossRef] [Green Version]
- Liu, Q.; Rand, T.A.; Kalidas, S.; Du, F.; Kim, H.E.; Smith, D.P.; Wang, X. R2D2, a bridge between the initiation and effector steps of the Drosophila RNAi pathway. Science 2003, 301, 1921–1925. [Google Scholar] [CrossRef] [PubMed]
- Zamore, P.D.; Tuschl, T.; Sharp, P.A.; Bartel, D.P. RNAi: Double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 2000, 101, 25–33. [Google Scholar] [CrossRef] [Green Version]
- Elbashir, S.M.; Martinez, J.; Patkaniowska, A.; Lendeckel, W.; Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 2001, 20, 6877–6888. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elbashir, S.M.; Lendeckel, W.; Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 2001, 15, 188–200. [Google Scholar] [CrossRef] [Green Version]
- Vélez, A.M.; Khajuria, C.; Wang, H.; Narva, K.E.; Siegfried, B.D. Knockdown of RNA interference pathway genes in western corn rootworms (Diabrotica virgifera virgifera Le Conte) demonstrates a possible mechanism of resistance to lethal dsRNA. PLoS ONE 2016, 11, e0157520. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Camargo, C.; Fishilevich, E.; Narva, K.E.; Chen, X.; Taylor, C.E.; Siegfried, B.D. Distinct fitness costs associated with the knockdown of RNAi pathway genes in western corn rootworm adults. PLoS ONE 2017, 12, e0190208. [Google Scholar] [CrossRef] [Green Version]
- Davis-Vogel, C.; Ortiz, A.; Procyk, L.; Robeson, J.; Kassa, A.; Wang, Y.; Huang, E.; Walker, C.; Sethi, A.; Nelson, M.E.; et al. Knockdown of RNA interference pathway genes impacts the fitness of western corn rootworm. Sci. Rep. 2018, 8, 7858. [Google Scholar] [CrossRef] [PubMed]
- Camargo, C.; Wu, K.; Fishilevich, E.; Narva, K.E.; Siegfried, B.D. Knockdown of RNA interference pathway genes in western corn rootworm, Diabrotica virgifera virgifera, identifies no fitness costs associated with Argonaute 2 or Dicer-2. Pestic Biochem. Physiol. 2018, 148, 103–110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pham, J.W.; Pellino, J.L.; Lee, Y.S.; Carthew, R.W.; Sontheimer, E.J. A Dicer-2-dependent 80s complex cleaves targeted mRNAs during RNAi in Drosophila. Cell 2004, 117, 83–94. [Google Scholar] [CrossRef] [Green Version]
- Tomari, Y.; Du, T.; Haley, B.; Schwarz, D.S.; Bennett, R.; Cook, H.A.; Koppetsch, B.S.; Theurkauf, W.E.; Zamore, P.D. RISC assembly defects in the Drosophila RNAi mutant armitage. Cell 2004, 116, 831–841. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Jiang, F.; Kalidas, S.; Smith, D.; Liu, Q. Dicer-2 and R2D2 coordinately bind siRNA to promote assembly of the siRISC complexes. RNA 2006, 12, 1514–1520. [Google Scholar] [CrossRef] [Green Version]
- Pham, J.W.; Sontheimer, E.J. Molecular requirements for RNA-induced silencing complex assembly in the Drosophila RNA interference pathway. J. Biol. Chem. 2005, 280, 39278–39283. [Google Scholar] [CrossRef] [Green Version]
- Liang, C.; Wang, Y.; Murota, Y.; Liu, X.; Smith, D.; Siomi, M.C.; Liu, Q. TAF11 Assembles the RISC loading complex to enhance RNAi efficiency. Mol. Cell 2015, 59, 807–818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Okamura, K.; Ishizuka, A.; Siomi, H.; Siomi, M.C. Distinct roles for Argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev. 2004, 18, 1655–1666. [Google Scholar] [CrossRef] [Green Version]
- Hammond, S.M.; Boettcher, S.; Caudy, A.A.; Kobayashi, R.; Hannon, G.J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 2001, 293, 1146–1150. [Google Scholar] [CrossRef] [Green Version]
- Matranga, C.; Tomari, Y.; Shin, C.; Bartel, D.P.; Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 2005, 123, 607–620. [Google Scholar] [CrossRef] [Green Version]
- Hammond, S.M.; Bernstein, E.; Beach, D.; Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 2000, 404, 293–296. [Google Scholar] [CrossRef]
- Chen, J.; Peng, Y.; Zhang, H.; Wang, K.; Zhao, C.; Zhu, G.; Reddy Palli, S.; Han, Z. Off-target effects of RNAi correlate with the mismatch rate between dsRNA and non-target mRNA. RNA Biol. 2021, 18, 1747–1759. [Google Scholar] [CrossRef]
- Rivas, F.V.; Tolia, N.H.; Song, J.J.; Aragon, J.P.; Liu, J.; Hannon, G.J.; Joshua-Tor, L. Purified Argonaute2 and an siRNA form recombinant human RISC. Nat. Struct. Mol. Biol. 2005, 12, 340–349. [Google Scholar] [CrossRef]
- Kulkarni, M.M.; Booker, M.; Silver, S.J.; Friedman, A.; Hong, P.; Perrimon, N.; Mathey-Prevot, B. Evidence of off-target effects associated with long dsRNAs in Drosophila melanogaster cell-based assays. Nat. Methods 2006, 3, 833–838. [Google Scholar] [CrossRef]
- Pratt, A.J.; MacRae, I.J. The RNA-induced silencing complex: A versatile gene-silencing machine. J. Biol. Chem. 2009, 284, 17897–17901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iwasaki, S.; Kobayashi, M.; Yoda, M.; Sakaguchi, Y.; Katsuma, S.; Suzuki, T.; Tomari, Y. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol. Cell 2010, 39, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Ji, L.; Chen, X. Regulation of small RNA stability: Methylation and beyond. Cell Res. 2012, 22, 624–636. [Google Scholar] [CrossRef]
- Yoon, J.S.; Shukla, J.N.; Gong, Z.J.; Mogilicherla, K.; Palli, S.R. RNA interference in the Colorado potato beetle, Leptinotarsa decemlineata: Identification of key contributors. Insect Biochem. Mol. Biol. 2016, 78, 78–88. [Google Scholar] [CrossRef] [Green Version]
- Zhou, R.; Czech, B.; Brennecke, J.; Sachidanandam, R.; Wohlschlegel, J.A.; Perrimon, N.; Hannon, G.J. Processing of Drosophila endo-siRNAs depends on a specific Loquacious isoform. RNA 2009, 15, 1886–1895. [Google Scholar] [CrossRef] [Green Version]
- Yoon, J.S.; Mogilicherla, K.; Gurusamy, D.; Chen, X.; Chereddy, S.C.R.R.; Palli, S.R. Double-stranded RNA binding protein, Staufen, is required for the initiation of RNAi in coleopteran insects. Proc. Natl. Acad. Sci. USA 2018, 115, 8334–8339. [Google Scholar] [CrossRef] [Green Version]
- Tassetto, M.; Kunitomi, M.; Andino, R. Circulating immune cells mediate a systemic RNAi-based adaptive antiviral response in Drosophila. Cell 2017, 169, 314–325.e313. [Google Scholar] [CrossRef] [Green Version]
- Karlikow, M.; Goic, B.; Saleh, M.C. RNAi and antiviral defense in Drosophila: Setting up a systemic immune response. Dev. Comp. Immunol. 2014, 42, 85–92. [Google Scholar] [CrossRef] [Green Version]
- Pak, J.; Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 2007, 315, 241–244. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, W.; Shirayama, M.; Conte, D.; Vasale, J.; Batista, P.J.; Claycomb, J.M.; Moresco, J.J.; Youngman, E.M.; Keys, J.; Stoltz, M.J.; et al. Distinct argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans germline. Mol. Cell 2009, 36, 231–244. [Google Scholar] [CrossRef] [Green Version]
- Aoki, K.; Moriguchi, H.; Yoshioka, T.; Okawa, K.; Tabara, H. In vitro analyses of the production and activity of secondary small interfering RNAs in C. elegans. EMBO J. 2007, 26, 5007–5019. [Google Scholar] [CrossRef] [Green Version]
- Shih, J.D.; Hunter, C.P. SID-1 is a dsRNA-selective dsRNA-gated channel. RNA 2011, 17, 1057–1065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gordon, K.H.; Waterhouse, P.M. RNAi for insect-proof plants. Nat. Biotechnol. 2007, 25, 1231–1232. [Google Scholar] [CrossRef]
- Li, H.R.; Bowling, A.J.; Gandra, P.; Rangasamy, M.; Pence, H.E.; McEwan, R.E.; Khajuria, C.; Siegfried, B.D.; Narva, K.E. Systemic RNAi in western corn rootworm, Diabrotica virgifera virgifera, does not involve transitive pathways. Insect Sci. 2018, 25, 45–56. [Google Scholar] [CrossRef]
- Hu, X.; Richtman, N.M.; Zhao, J.Z.; Duncan, K.E.; Niu, X.; Procyk, L.A.; Oneal, M.A.; Kernodle, B.M.; Steimel, J.P.; Crane, V.C.; et al. Discovery of midgut genes for the RNA interference control of corn rootworm. Sci. Rep. 2016, 6, 30542. [Google Scholar] [CrossRef]
- Khajuria, C.; Velez, A.M.; Rangasamy, M.; Wang, H.; Fishilevich, E.; Frey, M.L.; Carneiro, N.P.; Gandra, P.; Narva, K.E.; Siegfried, B.D. Parental RNA interference of genes involved in embryonic development of the western corn rootworm, Diabrotica virgifera virgifera LeConte. Insect Biochem. Mol. Biol. 2015, 63, 54–62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mingels, L.; Wynant, N.; Santos, D.; Peeters, P.; Gansemans, Y.; Billen, J.; Van Nieuwerburgh, F.; Vanden Broeck, J. Extracellular vesicles spread the RNA interference signal of Tribolium castaneum TcA cells. Insect Biochem. Mol. Biol. 2020, 122, 103377. [Google Scholar] [CrossRef]
- Yoon, J.S.; Kim, K.; Palli, S.R. Double-stranded RNA in exosomes: Potential systemic RNA interference pathway in the Colorado potato beetle, Leptinotarsa decemlineata. J. Asia-Pac. Entomol. 2020, 23, 1160–1164. [Google Scholar] [CrossRef]
- Vaughn, T. A method of controlling corn rootworm feeding using a Bacillus thuringiensis protein expressed in transgenic maize. Crop. Sci. 2005, 45, 931–938. [Google Scholar] [CrossRef]
- Deitloff, J.; Dunbar, M.W.; Ingber, D.A.; Hibbard, B.E.; Gassmann, A.J. Effects of refuges on the evolution of resistance to transgenic corn by the western corn rootworm, Diabrotica virgifera virgifera LeConte. Pest Manag. Sci. 2016, 72, 190–198. [Google Scholar] [CrossRef] [Green Version]
- Schellenberger, U.; Oral, J.; Rosen, B.A.; Wei, J.Z.; Zhu, G.; Xie, W.; McDonald, M.J.; Cerf, D.C.; Diehn, S.H.; Crane, V.C.; et al. A selective insecticidal protein from Pseudomonas for controlling corn rootworms. Science 2016, 354, 634–637. [Google Scholar] [CrossRef]
- Domínguez-Arrizabalaga, M.; Villanueva, M.; Escriche, B.; Ancín-Azpilicueta, C.; Caballero, P. Insecticidal activity of Bacillus thuringiensis proteins against coleopteran pests. Toxins 2020, 12, 430. [Google Scholar] [CrossRef]
- Joga, M.R.; Zotti, M.J.; Smagghe, G.; Christiaens, O. RNAi Efficiency, systemic properties, and novel delivery methods for pest insect control: What we know so far. Front. Physiol. 2016, 7, 553. [Google Scholar] [CrossRef] [Green Version]
- Knorr, E.; Fishilevich, E.; Tenbusch, L.; Frey, M.L.F.; Rangasamy, M.; Billion, A.; Worden, S.E.; Gandra, P.; Arora, K.; Lo, W.; et al. Gene silencing in Tribolium castaneum as a tool for the targeted identification of candidate RNAi targets in crop pests. Sci. Rep. 2018, 8, 2061. [Google Scholar] [CrossRef]
- Koci, J.; Ramaseshadri, P.; Bolognesi, R.; Segers, G.; Flannagan, R.; Park, Y. Ultrastructural changes caused by Snf7 RNAi in larval enterocytes of western corn rootworm (Diabrotica virgifera virgifera Le Conte). PLoS ONE 2014, 9, e83985. [Google Scholar] [CrossRef] [Green Version]
- Ramaseshadri, P.; Segers, G.; Flannagan, R.; Wiggins, E.; Clinton, W.; Ilagan, O.; McNulty, B.; Clark, T.; Bolognesi, R. Physiological and cellular responses caused by RNAi- mediated suppression of Snf7 orthologue in western corn rootworm (Diabrotica virgifera virgifera) larvae. PLoS ONE 2013, 8, e54270. [Google Scholar] [CrossRef] [Green Version]
- Vélez, A.M.; Fishilevich, E.; Rangasamy, M.; Khajuria, C.; McCaskill, D.G.; Pereira, A.E.; Gandra, P.; Frey, M.L.; Worden, S.E.; Whitlock, S.L.; et al. Control of western corn rootworm via RNAi traits in maize: Lethal and sublethal effects of Sec23 dsRNA. Pest Manag. Sci. 2020, 76, 1500–1512. [Google Scholar] [CrossRef] [Green Version]
- Fishilevich, E.; Bowling, A.J.; Frey, M.L.F.; Wang, P.H.; Lo, W.; Rangasamy, M.; Worden, S.E.; Pence, H.E.; Gandra, P.; Whitlock, S.L.; et al. RNAi targeting of rootworm Troponin I transcripts confers root protection in maize. Insect Biochem. Mol. Biol. 2019, 104, 20–29. [Google Scholar] [CrossRef]
- Hu, X.; Steimel, J.P.; Kapka-Kitzman, D.M.; Davis-Vogel, C.; Richtman, N.M.; Mathis, J.P.; Nelson, M.E.; Lu, A.L.; Wu, G. Molecular characterization of the insecticidal activity of double-stranded RNA targeting the smooth septate junction of western corn rootworm (Diabrotica virgifera virgifera). PLoS ONE 2019, 14, e0210491. [Google Scholar] [CrossRef]
- Hu, X.; Boeckman, C.J.; Cong, B.; Steimel, J.P.; Richtman, N.M.; Sturtz, K.; Wang, Y.; Walker, C.L.; Yin, J.; Unger, A.; et al. Characterization of DvSSJ1 transcripts targeting the smooth septate junction (SSJ) of western corn rootworm (Diabrotica virgifera virgifera). Sci. Rep. 2020, 10, 11139. [Google Scholar] [CrossRef]
- Velez, A.M.; Fishilevich, E.; Matz, N.; Storer, N.P.; Narva, K.E.; Siegfried, B.D. Parameters for successful parental RNAi as an insect pest management tool in western corn rootworm, Diabrotica virgifera virgifera. Genes 2016, 8, 7. [Google Scholar] [CrossRef]
- Fishilevich, E.; Vélez, A.M.; Khajuria, C.; Frey, M.L.; Hamm, R.L.; Wang, H.; Schulenberg, G.A.; Bowling, A.J.; Pence, H.E.; Gandra, P.; et al. Use of chromatin remodeling ATPases as RNAi targets for parental control of western corn rootworm (Diabrotica virgifera virgifera) and Neotropical brown stink bug (Euschistus heros). Insect Biochem. Mol. Biol. 2016, 71, 58–71. [Google Scholar] [CrossRef] [Green Version]
- Niu, X.; Kassa, A.; Hu, X.; Robeson, J.; McMahon, M.; Richtman, N.M.; Steimel, J.P.; Kernodle, B.M.; Crane, V.C.; Sandahl, G.; et al. Control of western corn rootworm (Diabrotica virgifera virgifera) reproduction through plant-mediated RNA interference. Sci. Rep. 2017, 7, 12591. [Google Scholar] [CrossRef]
- Siomi, M.C.; Sato, K.; Pezic, D.; Aravin, A.A. PIWI-interacting small RNAs: The vanguard of genome defence. Nat. Rev. Mol. Cell Biol. 2011, 12, 246–258. [Google Scholar] [CrossRef]
- Fishilevich, E.; Vélez, A.M.; Storer, N.P.; Li, H.; Bowling, A.J.; Rangasamy, M.; Worden, S.E.; Narva, K.E.; Siegfried, B.D. RNAi as a management tool for the western corn rootworm, Diabrotica virgifera virgifera. Pest Manag. Sci. 2016, 72, 1652–1663. [Google Scholar] [CrossRef] [Green Version]
- Levine, S.L.; Tan, J.; Mueller, G.M.; Bachman, P.M.; Jensen, P.D.; Uffman, J.P. Independent action between DvSnf7 RNA and Cry3Bb1 protein in southern corn rootworm, Diabrotica undecimpunctata howardi and Colorado potato beetle, Leptinotarsa decemlineata. PLoS ONE 2015, 10, e0118622. [Google Scholar] [CrossRef] [Green Version]
- Vaccari, T.; Rusten, T.E.; Menut, L.; Nezis, I.P.; Brech, A.; Stenmark, H.; Bilder, D. Comparative analysis of ESCRT-I, ESCRT-II and ESCRT-III function in Drosophila by efficient isolation of ESCRT mutants. J. Cell Sci. 2009, 122, 2413–2423. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ulrich, J.; Dao, V.A.; Majumdar, U.; Schmitt-Engel, C.; Schwirz, J.; Schultheis, D.; Strohlein, N.; Troelenberg, N.; Grossmann, D.; Richter, T.; et al. Large scale RNAi screen in Tribolium reveals novel target genes for pest control and the proteasome as prime target. BMC Genom. 2015, 16, 674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lehman, W.; Galińska-Rakoczy, A.; Hatch, V.; Tobacman, L.S.; Craig, R. Structural basis for the activation of muscle contraction by troponin and tropomyosin. J. Mol. Biol. 2009, 388, 673–681. [Google Scholar] [CrossRef] [Green Version]
- Lord, C.; Ferro-Novick, S.; Miller, E.A. The highly conserved COPII coat complex sorts cargo from the endoplasmic reticulum and targets it to the golgi. Cold Spring Harb. Perspect Biol. 2013, 5. [Google Scholar] [CrossRef] [Green Version]
- Horn, T.; Narov, K.D.; Panfilio, K.A. Persistent parental RNAi in the beetle Tribolium castaneum involves maternal transmission of long double-stranded RNA. bioRxiv 2021. [Google Scholar] [CrossRef]
- Holoch, D.; Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 2015, 16, 71–84. [Google Scholar] [CrossRef] [PubMed]
- Ivashuta, S.; Zhang, Y.; Wiggins, B.E.; Ramaseshadri, P.; Segers, G.C.; Johnson, S.; Meyer, S.E.; Kerstetter, R.A.; McNulty, B.C.; Bolognesi, R.; et al. Environmental RNAi in herbivorous insects. RNA 2015, 21, 840–850. [Google Scholar] [CrossRef] [Green Version]
- Armstrong, T.A.; Chen, H.; Ziegler, T.E.; Iyadurai, K.R.; Gao, A.G.; Wang, Y.; Song, Z.; Tian, Q.; Zhang, Q.; Ward, J.M.; et al. Quantification of transgene-derived double-stranded RNA in plants using the QuantiGene nucleic acid detection platform. J. Agric. Food Chem. 2013, 61, 12557–12564. [Google Scholar] [CrossRef]
- Oleson, J.D.; Park, Y.-L.; Nowatzki, T.M.; Tollefson, J.J. Node-injury scale to evaluate root injury by corn rootworms (Coleoptera: Chrysomelidae). J. Econ. Entomol. 2005, 98, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Baum, J.A.; Roberts, J.K. Progress toward RNAi-mediated insect pest management. Adv. Insect Phys. 2014, 47, 249–295. [Google Scholar]
- USEPA [U.S. Environmental Protection Agency]. EPA Registers Innovative Tool to Control Corn Rootworm. 2017. Available online: https://archive.epa.gov/epa/newsreleases/epa-registers-innovative-tool-control-corn-rootworm.html (accessed on 27 December 2021).
- USEPA [U.S. Environmental Protection Agency]. Current and Previously Registered Section 3 Plant-Incorporated Protectant (PIP) Registrations. 2015. Available online: https://archive.epa.gov/pesticides/reregistration/web/html/current-previously-registered-section-3-plant-incorporated.html (accessed on 27 December 2021).
- Moar, W.; Khajuria, C.; Pleau, M.; Ilagan, O.; Chen, M.; Jiang, C.; Price, P.; McNulty, B.; Clark, T.; Head, G. Cry3Bb1-resistant western corn rootworm, Diabrotica virgifera virgifera (LeConte) does not exhibit cross-resistance to DvSnf7 dsRNA. PLoS ONE 2017, 12, e0169175. [Google Scholar] [CrossRef] [PubMed]
- USEPA [U.S. Environmental Protection Agency]. Pesticide Product Label, Bt11 × MIR162 × MIR604 × TC1507 × 5307 Corn. 2012. Available online: https://www3.epa.gov/pesticides/chem_search/ppls/067979-00023-20120731.pdf (accessed on 27 December 2021).
- Carrière, Y.; Fabrick, J.A.; Tabashnik, B.E. Can pyramids and seed mixtures delay resistance to Bt crops? Trends Biotechnol. 2016, 34, 291–302. [Google Scholar] [CrossRef] [Green Version]
- Pereira, A.E.; Carneiro, N.P.; Siegfried, B.D. Comparative susceptibility of southern and western corn rootworm adults and larvae to vATPase-A and Snf7 dsRNAs. J. RNAi Gene Silenc. 2016, 12, 528–535. [Google Scholar]
- Rangasamy, M.; Siegfried, B.D. Validation of RNA interference in western corn rootworm Diabrotica virgifera virgifera LeConte (Coleoptera: Chrysomelidae) adults. Pest Manag. Sci. 2012, 68, 587–591. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Taylor, C.E.; Fishilevich, E.; Narva, K.E.; Siegfried, B.D. Rapid and persistent RNAi response in western corn rootworm adults. Pestic. Biochem. Physiol. 2018, 150, 66–70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bachman, P.M.; Huizinga, K.M.; Jensen, P.D.; Mueller, G.; Tan, J.; Uffman, J.P.; Levine, S.L. Ecological risk assessment for DvSnf7 RNA: A plant-incorporated protectant with targeted activity against western corn rootworm. Regul. Toxicol. Pharmacol. 2016, 81, 77–88. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tabashnik, B.E.; Brévault, T.; Carrière, Y. Insect resistance to Bt crops: Lessons from the first billion acres. Nat. Biotechnol. 2013, 31, 510–521. [Google Scholar] [CrossRef] [PubMed]
- Tabashnik, B.E.; Gould, F.; Carrière, Y. Delaying evolution of insect resistance to transgenic crops by decreasing dominance and heritability. J. Evol. Biol. 2004, 17, 904–912. [Google Scholar] [CrossRef] [PubMed]
- Spencer, J.L.; Hibbard, B.E.; Moeser, J.; Onstad, D.W. Behaviour and ecology of the western corn rootworm (Diabrotica virgifera virgifera LeConte). Agric. For. Entomol. 2009, 11, 9–27. [Google Scholar] [CrossRef]
- USEPA [U.S. Environmental Protection Agency]. Framework to Delay Corn Rootworm Resistance. 2016. Available online: https://www.epa.gov/regulation-biotechnology-under-tsca-and-fifra/framework-delay-corn-rootworm-resistance (accessed on 27 December 2021).
- Martinez, J.C.; Caprio, M.A. IPM use with the deployment of a non-high dose Bt pyramid and mitigation of resistance for western corn rootworm (Diabrotica virgifera virgifera). Environ. Entomol. 2016, 45, 747–761. [Google Scholar] [CrossRef] [Green Version]
- Palli, S.R. RNA interference in Colorado potato beetle: Steps toward development of dsRNA as a commercial insecticide. Curr. Opin. Insect Sci. 2014, 6, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Flagel, L.E.; Bansal, R.; Kerstetter, R.A.; Chen, M.; Carroll, M.; Flannagan, R.; Clark, T.; Goldman, B.S.; Michel, A.P. Western corn rootworm (Diabrotica virgifera virgifera) transcriptome assembly and genomic analysis of population structure. BMC Genom. 2014, 15, 195. [Google Scholar] [CrossRef] [Green Version]
- Mishra, S.; Dee, J.; Moar, W.; Dufner-Beattie, J.; Baum, J.; Dias, N.P.; Alyokhin, A.; Buzza, A.; Rondon, S.I.; Clough, M.; et al. Selection for high levels of resistance to double-stranded RNA (dsRNA) in Colorado potato beetle (Leptinotarsa decemlineata Say) using non-transgenic foliar delivery. Sci. Rep. 2021, 11, 6523. [Google Scholar] [CrossRef] [PubMed]
- Carstens, K.L.; Hayter, K.; Layton, R.J. A perspective on problem formulation and exposure assessment of transgenic crops. In Proceedings of the Fourth Meeting on Ecological Impact of Genetically Modified Organisms, GMOs in Integrated Plant Production, Rostock, Germany, 14–16 May 2009; Volume 52, pp. 23–30. Available online: https://www.iobc-wprs.org/members/shop_en.cfm?mod_Shop_detail_produkte=59 (accessed on 27 December 2021).
- USEPA [U.S. Environmental Protection Agency]. Guidelines for Ecological Risk Assessment, EPA/630/R-95-002F. 1998. Available online: https://www.epa.gov/sites/default/files/2014-11/documents/eco_risk_assessment1998.pdf (accessed on 27 December 2021).
- Raybould, A. Problem formulation and hypothesis testing for environmental risk assessments of genetically modified crops. Environ. Biosaf. Res. 2006, 5, 119–125. [Google Scholar] [CrossRef]
- Wolt, J.D.; Peterson, R.K. Prospective formulation of environmental risk assessments: Probabilistic screening for Cry1A(b) maize risk to aquatic insects. Ecotoxicol. Environ. Saf. 2010, 73, 1182–1188. [Google Scholar] [CrossRef]
- Devos, Y.; Romeis, J.; Luttik, R.; Maggiore, A.; Perry, J.N.; Schoonjans, R.; Streissl, F.; Tarazona, J.V.; Brock, T.C. Optimising environmental risk assessments: Accounting for ecosystem services helps to translate broad policy protection goals into specific operational ones for environmental risk assessments. EMBO Rep. 2015, 16, 1060–1063. [Google Scholar] [CrossRef] [PubMed]
- Rose, R.I. White Paper on Tier-Based Testing for the Effects of Proteinaceous Insecticidal Plant-Incorporated Protectants on Non-Target Invertebrates for Regulatory Risk Assessment. 2007. Available online: https://www.epa.gov/sites/default/files/2015-09/documents/tier-based-testing.pdf (accessed on 27 December 2021).
- EFSA. Guidance on the environmental risk assessment of genetically modified plants. EFSA J. 2010, 8, 1879. [Google Scholar] [CrossRef]
- Romeis, J.; Hellmich, R.L.; Candolfi, M.P.; Carstens, K.; De Schrijver, A.; Gatehouse, A.M.; Herman, R.A.; Huesing, J.E.; McLean, M.A.; Raybould, A.; et al. Recommendations for the design of laboratory studies on non-target arthropods for risk assessment of genetically engineered plants. Transgenic Res. 2011, 20, 1–22. [Google Scholar] [CrossRef] [Green Version]
- Romeis, J.; Raybould, A.; Bigler, F.; Candolfi, M.P.; Hellmich, R.L.; Huesing, J.E.; Shelton, A.M. Deriving criteria to select arthropod species for laboratory tests to assess the ecological risks from cultivating arthropod-resistant genetically engineered crops. Chemosphere 2013, 90, 901–909. [Google Scholar] [CrossRef] [Green Version]
- Roberts, A.; Boeckman, C.J.; Mühl, M.; Romeis, J.; Teem, J.L.; Valicente, F.H.; Brown, J.K.; Edwards, M.G.; Levine, S.L.; Melnick, R.L.; et al. Sublethal endpoints in non-target organism testing for insect-active GE crops. Front. Bioeng. Biotechnol. 2020, 8, 556. [Google Scholar] [CrossRef]
- Garcia-Alonso, M.; Jacobs, E.; Raybould, A.; Nickson, T.E.; Sowig, P.; Willekens, H.; Van der Kouwe, P.; Layton, R.; Amijee, F.; Fuentes, A.M.; et al. A tiered system for assessing the risk of genetically modified plants to non-target organisms. Environ. Biosaf. Res. 2006, 5, 57–65. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.H.; Soumaila Issa, M.; Cooper, A.M.; Zhu, K.Y. RNA interference: Applications and advances in insect toxicology and insect pest management. Pestic. Biochem. Physiol. 2015, 120, 109–117. [Google Scholar] [CrossRef] [Green Version]
- Raybould, A.; Vlachos, D. Non-target organism effects tests on Vip3A and their application to the ecological risk assessment for cultivation of MIR162 maize. Transgenic Res. 2011, 20, 599–611. [Google Scholar] [CrossRef]
- Fischer, J.R.; MacQuarrie, G.R.; Malven, M.; Song, Z.; Rogan, G. Dissipation of DvSnf7 RNA from late-season maize tissue in aquatic microcosms. Environ. Toxicol. Chem. 2020, 39, 1032–1040. [Google Scholar] [CrossRef]
- Dubelman, S.; Fischer, J.; Zapata, F.; Huizinga, K.; Jiang, C.; Uffman, J.; Levine, S.; Carson, D. Environmental fate of double-stranded RNA in agricultural soils. PLoS ONE 2014, 9, e93155. [Google Scholar] [CrossRef]
- Parker, K.M.; Barragán Borrero, V.; van Leeuwen, D.M.; Lever, M.A.; Mateescu, B.; Sander, M. Environmental fate of RNA interference pesticides: Adsorption and degradation of double-stranded RNA molecules in agricultural soils. Environ. Sci. Technol. 2019, 53, 3027–3036. [Google Scholar] [CrossRef]
- Fischer, J.R.; Zapata, F.; Dubelman, S.; Mueller, G.M.; Uffman, J.P.; Jiang, C.; Jensen, P.D.; Levine, S.L. Aquatic fate of a double-stranded RNA in a sediment water system following an over-water application. Environ. Toxicol. Chem. 2017, 36, 727–734. [Google Scholar] [CrossRef]
- Albright, V.C.; Wong, C.R.; Hellmich, R.L.; Coats, J.R. Dissipation of double-stranded RNA in aquatic microcosms. Environ. Toxicol. Chem. 2017, 36, 1249–1253. [Google Scholar] [CrossRef] [Green Version]
- Fischer, J.R.; Zapata, F.; Dubelman, S.; Mueller, G.M.; Jensen, P.D.; Levine, S.L. Characterizing a novel and sensitive method to measure dsRNA in soil. Chemosphere 2016, 161, 319–324. [Google Scholar] [CrossRef]
- Bachman, P.; Fischer, J.; Song, Z.; Urbanczyk-Wochniak, E.; Watson, G. Environmental fate and dissipation of applied dsRNA in soil, aquatic systems, and plants. Front. Plant. Sci. 2020, 11, 21. [Google Scholar] [CrossRef]
- Li, H.; Khajuria, C.; Rangasamy, M.; Gandra, P.; Fitter, M.; Geng, C.; Woosely, A.; Hasler, J.; Schulenberg, G.; Worden, S.; et al. Long dsRNA but not siRNA initiates RNAi in western corn rootworm larvae and adults. J. Appl. Entomol. 2015, 139, 432–445. [Google Scholar] [CrossRef]
- Paces, J.; Nic, M.; Novotny, T.; Svoboda, P. Literature review of baseline information on RNAi to support the environmental risk assessment of RNAi-based GM plants. EFSA J. 2017, 15, 1424E. [Google Scholar] [CrossRef]
- Christiaens, O.; Dzhambazova, T.; Kostov, K.; Arpaia, S.; Joga, M.R.; Urru, I.; Sweet, J.; Smagghe, G. Literature review of baseline information on RNAi to support the environmental risk assessment of RNAi-based GM plants. EFSA J. 2018, 15, 1424E. [Google Scholar] [CrossRef]
- Romeis, J.; Bartsch, D.; Bigler, F.; Candolfi, M.P.; Gielkens, M.M.; Hartley, S.E.; Hellmich, R.L.; Huesing, J.E.; Jepson, P.C.; Layton, R.; et al. Assessment of risk of insect-resistant transgenic crops to nontarget arthropods. Nat. Biotechnol. 2008, 26, 203–208. [Google Scholar] [CrossRef] [Green Version]
- Vélez, A.M.; Jurzenski, J.; Matz, N.; Zhou, X.; Wang, H.; Ellis, M.; Siegfried, B.D. Developing an in vivo toxicity assay for RNAi risk assessment in honey bees, Apis mellifera L. Chemosphere 2016, 144, 1083–1090. [Google Scholar] [CrossRef] [Green Version]
- Pan, H.; Yang, X.; Bidne, K.; Hellmich, R.L.; Siegfried, B.D.; Zhou, X. Dietary risk assessment of v-ATPase A dsRNAs on monarch butterfly larvae. Front. Plant. Sci. 2017, 8, 242. [Google Scholar] [CrossRef] [Green Version]
- Haller, S.; Widmer, F.; Siegfried, B.D.; Zhuo, X.; Romeis, J. Responses of two ladybird beetle species (Coleoptera: Coccinellidae) to dietary RNAi. Pest Manag. Sci. 2019, 75, 2652–2662. [Google Scholar] [CrossRef]
- Pan, H.; Yang, X.; Romeis, J.; Siegfried, B.D.; Zhou, X. Dietary RNAi toxicity assay exhibits differential responses to ingested dsRNAs among lady beetles. Pest Manag. Sci. 2020, 76, 3606–3614. [Google Scholar] [CrossRef]
- Pan, H.; Xu, L.; Noland, J.E.; Li, H.; Siegfried, B.D.; Zhou, X. Assessment of potential risks of dietary RNAi to a soil micro-arthropod, Sinella curviseta Brook (Collembola: Entomobryidae). Front. Plant. Sci. 2016, 7, 1028. [Google Scholar] [CrossRef] [Green Version]
- Bachman, P.M.; Bolognesi, R.; Moar, W.J.; Mueller, G.M.; Paradise, M.S.; Ramaseshadri, P.; Tan, J.; Uffman, J.P.; Warren, J.; Wiggins, B.E.; et al. Characterization of the spectrum of insecticidal activity of a double-stranded RNA with targeted activity against western corn rootworm (Diabrotica virgifera virgifera LeConte). Transgenic Res. 2013, 22, 1207–1222. [Google Scholar] [CrossRef] [Green Version]
- Tan, J.; Levine, S.L.; Bachman, P.M.; Jensen, P.D.; Mueller, G.M.; Uffman, J.P.; Meng, C.; Song, Z.; Richards, K.B.; Beevers, M.H. No impact of DvSnf7 RNA on honey bee (Apis mellifera L.) adults and larvae in dietary feeding tests. Environ. Toxicol. Chem. 2016, 35, 287–294. [Google Scholar] [CrossRef] [Green Version]
- Raybould, A.; Burns, A. Problem formulation for off-target effects of externally applied double-stranded RNA-based products for pest control. Front. Plant. Sci. 2020, 11, 424. [Google Scholar] [CrossRef]
- Frizzi, A.; Zhang, Y.; Kao, J.; Hagen, C.; Huang, S. Small RNA profiles from virus-infected fresh market vegetables. J. Agric. Food Chem. 2014, 62, 12067–12074. [Google Scholar] [CrossRef]
- Ivashuta, S.I.; Petrick, J.S.; Heisel, S.E.; Zhang, Y.; Guo, L.; Reynolds, T.L.; Rice, J.F.; Allen, E.; Roberts, J.K. Endogenous small RNAs in grain: Semi-quantification and sequence homology to human and animal genes. Food Chem. Toxicol. 2009, 47, 353–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Petrick, J.S.; Frierdich, G.E.; Carleton, S.M.; Kessenich, C.R.; Silvanovich, A.; Zhang, Y.; Koch, M.S. Corn rootworm-active RNA DvSnf7: Repeat dose oral toxicology assessment in support of human and mammalian safety. Regul. Toxicol. Pharmacol. 2016, 81, 57–68, Erratum in Regul. Toxicol. Pharmacol. 2016, 82, 191. [Google Scholar] [CrossRef] [Green Version]
- Petrick, J.S.; Brower-Toland, B.; Jackson, A.L.; Kier, L.D. Safety assessment of food and feed from biotechnology-derived crops employing RNA-mediated gene regulation to achieve desired traits: A scientific review. Regul. Toxicol. Pharmacol. 2013, 66, 167–176. [Google Scholar] [CrossRef] [Green Version]
- Sherman, J.H.; Munyikwa, T.; Chan, S.Y.; Petrick, J.S.; Witwer, K.W.; Choudhuri, S. RNAi technologies in agricultural biotechnology: The Toxicology Forum 40th Annual Summer Meeting. Regul. Toxicol. Pharmacol. 2015, 73, 671–680. [Google Scholar] [CrossRef] [Green Version]
- Tripathi, S.; Suzuki, J.Y.; Ferreira, S.A.; Gonsalves, D. Papaya ringspot virus-P: Characteristics, pathogenicity, sequence variability and control. Mol. Plant. Pathol. 2008, 9, 269–280. [Google Scholar] [CrossRef]
- Wagner, N.; Mroczka, A.; Roberts, P.D.; Schreckengost, W.; Voelker, T. RNAi trigger fragment truncation attenuates soybean FAD2-1 transcript suppression and yields intermediate oil phenotypes. Plant. Biotechnol. J. 2011, 9, 723–728. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Xing, G.; Niu, L.; He, H.; Guo, D.; Du, Q.; Qian, X.; Yao, Y.; Li, H.; Zhong, X.; et al. Improved oil quality in transgenic soybean seeds by RNAi-mediated knockdown of GmFAD2-1B. Transgenic Res. 2018, 27, 155–166. [Google Scholar] [CrossRef]
- Waltz, E. Nonbrowning GM apple cleared for market. Nat. Biotechnol. 2015, 33, 326–327. [Google Scholar] [CrossRef]
- Pence, M.; Spence, R.; Rood, T.; Habig, J.; Collinge, S. Petition for Extension of Nonregulated Status for X17 Ranger Russet and Y9 Atlantic Potatoes with Late Blight Resistance, Low Acrylamide Potential, Lowered Reducing Sugars, and Reduced Black Spot, Petition No. 16-064-01P. 2016. Available online: https://www.aphis.usda.gov/brs/aphisdocs/16_06401p.pdf (accessed on 27 December 2021).
- Tran, N.L.; Barraj, L.M.; Collinge, S. Reduction in dietary acrylamide exposure-impact of potatoes with low acrylamide potential. Risk Anal. 2017, 37, 1754–1767. [Google Scholar] [CrossRef]
- ISAAA [International Service for the Acquisition of Agri-Biotech Applications]. GM Crop Events List. 2021. Available online: https://www.isaaa.org/gmapprovaldatabase/cropslist/default.asp (accessed on 27 December 2021).
- Heisel, S.E.; Zhang, Y.; Allen, E.; Guo, L.; Reynolds, T.L.; Yang, X.; Kovalic, D.; Roberts, J.K. Characterization of unique small RNA populations from rice grain. PLoS ONE 2008, 3, e2871. [Google Scholar] [CrossRef] [PubMed]
- Jensen, P.D.; Zhang, Y.; Wiggins, B.E.; Petrick, J.S.; Zhu, J.; Kerstetter, R.A.; Heck, G.R.; Ivashuta, S.I. Computational sequence analysis of predicted long dsRNA transcriptomes of major crops reveals sequence complementarity with human genes. GM Crops Food 2013, 4, 90–97. [Google Scholar] [CrossRef]
- O’Neill, M.J.; Bourre, L.; Melgar, S.; O’Driscoll, C.M. Intestinal delivery of non-viral gene therapeutics: Physiological barriers and preclinical models. Drug Discov. Today 2011, 16, 203–218. [Google Scholar] [CrossRef]
- Park, N.J.; Li, Y.; Yu, T.; Brinkman, B.M.; Wong, D.T. Characterization of RNA in saliva. Clin. Chem. 2006, 52, 988–994. [Google Scholar] [CrossRef] [Green Version]
- Huang, H.; Davis, C.D.; Wang, T.T.Y. Extensive degradation and low bioavailability of orally consumed corn miRNAs in mice. Nutrients 2018, 10, 215. [Google Scholar] [CrossRef] [Green Version]
- Dickinson, B.; Zhang, Y.; Petrick, J.S.; Heck, G.; Ivashuta, S.; Marshall, W.S. Lack of detectable oral bioavailability of plant microRNAs after feeding in mice. Nat. Biotechnol. 2013, 31, 965–967. [Google Scholar] [CrossRef]
- Witwer, K.W.; McAlexander, M.A.; Queen, S.E.; Adams, R.J. Real-time quantitative PCR and droplet digital PCR for plant miRNAs in mammalian blood provide little evidence for general uptake of dietary miRNAs: Limited evidence for general uptake of dietary plant xenomiRs. RNA Biol. 2013, 10, 1080–1086. [Google Scholar] [CrossRef] [Green Version]
- Snow, J.W.; Hale, A.E.; Isaacs, S.K.; Baggish, A.L.; Chan, S.Y. Ineffective delivery of diet-derived microRNAs to recipient animal organisms. RNA Biol. 2013, 10, 1107–1116. [Google Scholar] [CrossRef] [Green Version]
- Rodrigues, T.B.; Petrick, J.S. Safety considerations for humans and other vertebrates regarding agricultural uses of externally applied RNA molecules. Front. Plant. Sci. 2020, 11, 407. [Google Scholar] [CrossRef] [Green Version]
- Gilmore, I.R.; Fox, S.P.; Hollins, A.J.; Sohail, M.; Akhtar, S. The design and exogenous delivery of siRNA for post-transcriptional gene silencing. J. Drug Target. 2004, 12, 315–340. [Google Scholar] [CrossRef]
- Christensen, J.; Litherland, K.; Faller, T.; van de Kerkhof, E.; Natt, F.; Hunziker, J.; Krauser, J.; Swart, P. Metabolism studies of unformulated internally [3H]-labeled short interfering RNAs in mice. Drug Metab. Dispos. 2013, 41, 1211–1219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Layzer, J.M.; McCaffrey, A.P.; Tanner, A.K.; Huang, Z.; Kay, M.A.; Sullenger, B.A. In vivo activity of nuclease-resistant siRNAs. RNA 2004, 10, 766–771. [Google Scholar] [CrossRef] [Green Version]
- White, P.J. Barriers to successful delivery of short interfering RNA after systemic administration. Clin. Exp. Pharmacol. Physiol. 2008, 35, 1371–1376. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Kornbrust, D.J.; Foy, J.W.; Solano, E.C.; Schneider, D.J.; Feinstein, E.; Molitoris, B.A.; Erlich, S. Toxicological and pharmacokinetic properties of chemically modified siRNAs targeting p53 RNA following intravenous administration. Nucleic Acid Ther. 2012, 22, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Forbes, D.C.; Peppas, N.A. Oral delivery of small RNA and DNA. J. Control Release 2012, 162, 438–445. [Google Scholar] [CrossRef]
- Gilleron, J.; Querbes, W.; Zeigerer, A.; Borodovsky, A.; Marsico, G.; Schubert, U.; Manygoats, K.; Seifert, S.; Andree, C.; Stöter, M.; et al. Image-based analysis of lipid nanoparticle-mediated siRNA delivery, intracellular trafficking and endosomal escape. Nat. Biotechnol. 2013, 31, 638–646. [Google Scholar] [CrossRef]
- Juliano, R.L. The delivery of therapeutic oligonucleotides. Nucleic Acids Res. 2016, 44, 6518–6548. [Google Scholar] [CrossRef]
- Moroz, E.; Matoori, S.; Leroux, J.C. Oral delivery of macromolecular drugs: Where we are after almost 100 years of attempts. Adv. Drug Deliv. Rev. 2016, 101, 108–121. [Google Scholar] [CrossRef] [Green Version]
- Molitoris, B.A.; Dagher, P.C.; Sandoval, R.M.; Campos, S.B.; Ashush, H.; Fridman, E.; Brafman, A.; Faerman, A.; Atkinson, S.J.; Thompson, J.D.; et al. siRNA targeted to p53 attenuates ischemic and cisplatin-induced acute kidney injury. J. Am. Soc. Nephrol. 2009, 20, 1754–1764. [Google Scholar] [CrossRef] [Green Version]
- Ballarín-González, B.; Dagnaes-Hansen, F.; Fenton, R.A.; Gao, S.; Hein, S.; Dong, M.; Kjems, J.; Howard, K.A. Protection and systemic translocation of siRNA following oral administration of Chitosan/siRNA nanoparticles. Mol. Ther. Nucleic Acids 2013, 2, e76. [Google Scholar] [CrossRef]
- Tillman, L.G.; Geary, R.S.; Hardee, G.E. Oral delivery of antisense oligonucleotides in man. J. Pharm. Sci. 2008, 97, 225–236. [Google Scholar] [CrossRef]
- Witwer, K.W. Hypothetical plant-mammal RNA communication: Packaging and stoichiometry. In Non-Coding RNAs and Inter.-Kingdom Communication; Leitao, A.L., Enguita, F.J., Eds.; Springer International Publishing: Basel, Switzerland, 2016; pp. 161–176. [Google Scholar]
- Title, A.C.; Denzler, R.; Stoffel, M. Uptake and function studies of maternal milk-derived microRNAs. J. Biol. Chem. 2015, 290, 23680–23691. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Chen, T.; Wang, R.; Luo, J.Y.; He, J.J.; Ye, R.S.; Xie, M.Y.; Xi, Q.Y.; Jiang, Q.Y.; Sun, J.J.; et al. Plant MIR156 regulates intestinal growth in mammals by targeting the Wnt/β-catenin pathway. Am. J. Physiol. Cell Physiol. 2019, 317, C434–C448. [Google Scholar] [CrossRef] [PubMed]
- Mlotshwa, S.; Pruss, G.J.; MacArthur, J.L.; Endres, M.W.; Davis, C.; Hofseth, L.J.; Peña, M.M.; Vance, V. A novel chemopreventive strategy based on therapeutic microRNAs produced in plants. Cell Res. 2015, 25, 521–524. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Hou, D.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [CrossRef]
- Zhou, Z.; Li, X.; Liu, J.; Dong, L.; Chen, Q.; Kong, H.; Zhang, Q.; Qi, X.; Hou, D.; Zhang, L.; et al. Honeysuckle-encoded atypical microRNA2911 directly targets influenza A viruses. Cell Res. 2015, 25, 39–49. [Google Scholar] [CrossRef]
- Chan, S.Y.; Snow, J.W. Formidable challenges to the notion of biologically important roles for dietary small RNAs in ingesting mammals. Genes Nutr. 2017, 12, 13. [Google Scholar] [CrossRef]
- Kang, W.; Bang-Berthelsen, C.H.; Holm, A.; Houben, A.J.; Müller, A.H.; Thymann, T.; Pociot, F.; Estivill, X.; Friedländer, M.R. Survey of 800+ data sets from human tissue and body fluid reveals xenomiRs are likely artifacts. RNA 2017, 23, 433–445. [Google Scholar] [CrossRef] [Green Version]
- Witwer, K.W.; Halushka, M.K. Toward the promise of microRNAs—Enhancing reproducibility and rigor in microRNA research. RNA Biol. 2016, 13, 1103–1116. [Google Scholar] [CrossRef]
- Petrick, J.S.; Moore, W.M.; Heydens, W.F.; Koch, M.S.; Sherman, J.H.; Lemke, S.L. A 28-day oral toxicity evaluation of small interfering RNAs and a long double-stranded RNA targeting vacuolar ATPase in mice. Regul. Toxicol. Pharmacol. 2015, 71, 8–23. [Google Scholar] [CrossRef] [Green Version]
- Lusk, R.W. Diverse and widespread contamination evident in the unmapped depths of high throughput sequencing data. PLoS ONE 2014, 9, e110808. [Google Scholar] [CrossRef] [Green Version]
- Tosar, J.P.; Rovira, C.; Naya, H.; Cayota, A. Mining of public sequencing databases supports a non-dietary origin for putative foreign miRNAs: Underestimated effects of contamination in NGS. RNA 2014, 20, 754–757. [Google Scholar] [CrossRef] [Green Version]
| Name | NCBI Accession No | Type of Exposure | Reference |
|---|---|---|---|
| Larvicidal Genes | |||
| ESCRT III snf7 1 | XM_028287710.1 | Artificial diet, in planta | [31,55,124,125] |
| v-ATPase-A 1 | XM_028281990 | Artificial diet, in planta | [55] |
| Actin 2 | XM_028292745.1 | Artificial diet | [55] |
| apple ATPase 1 | XM_028281191.1 | Artificial diet | [55] |
| alpha tubulin 1 | XM_028282553.1 | Artificial diet | [55] |
| beta tubulin 1 | XM_028282553.1 | Artificial diet | [55] |
| COPI coatomer subunit β 1 | XM_028291201.1 | Artificial diet | [55] |
| ESCRT III_vps2 1 | XM_028296669.1 | Artificial diet | [55] |
| ESCRT I-Vps28 1 | XM_028283797.1 | Artificial diet | [55] |
| mov34 1 | XM_028287237.1 | Artificial diet | [55] |
| ribosomal protein L9 1 | XM_028294395.1 | Artificial diet | [55] |
| ribosomal protein L19 1 | XM_028289442.1 | Artificial diet | [55] |
| ribosomal protein S4 1 | XM_028298505.1 | Artificial diet | [55] |
| ribosomal protein rps-14 2 | XM_028275245.1 | Artificial diet | [55] |
| RNA polymerase II 1 | XM_028297193.1 | Artificial diet | [55] |
| vATPase-D 1 | XM_028287428.1 | Artificial diet | [55] |
| sec23 1 | MK474471 | Artificial diet, in planta | [126] |
| wupA/troponin I 1 | MH001576.1 | Artificial diet, in planta | [127] |
| rop 3 | XM_028277045.1 | Artificial diet, in planta | [123] |
| dre4 3 | XM_028288745.1 | Artificial diet, in planta | [123] |
| rpIII 140 3 | XM_028297193.1 | Artificial diet, in planta | [123] |
| ncm-1 | XM_028276581.1 | Artificial diet | [123] |
| Rpb7-1 | XM_028299763.1 | Artificial diet | [123] |
| smooth septate junction protein 1 (ssj1) | KU562965 | Artificial diet, in planta | [114,128,129] |
| smooth septate junction protein 1 (ssj2) | KU562966 | Artificial diet | [114] |
| proteasome subunit beta type-1-like (protb) | KU756279 | Artificial diet | [114] |
| proteasome subunit alpha type-3-like (pat3) | KU756280 | Artificial diet | [114] |
| ribosomal protein S10 (rps10) | KU756281 | Artificial diet | [114] |
| Parental and Reproductive Genes | |||
| hunchback 1 | XM_028272853.1 | Artificial diet, in planta | [115,130] |
| brahma 1 | KR152260.1 | Artificial diet, in planta | [115,130] |
| chd-1 1 | KT364642 | Artificial diet | [131] |
| iswi-1 1 | KT364640 | Artificial diet | [131] |
| mi-2 1 | KT364639 | Artificial diet | [131] |
| iswi-2 1 | KT364641 | Artificial diet | [131] |
| Vgr 1 | KY373243 | Artificial diet, in planta | [132] |
| bol 1 | KY373244 | Artificial diet, in planta | [132] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Darlington, M.; Reinders, J.D.; Sethi, A.; Lu, A.L.; Ramaseshadri, P.; Fischer, J.R.; Boeckman, C.J.; Petrick, J.S.; Roper, J.M.; Narva, K.E.; et al. RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions. Insects 2022, 13, 57. https://doi.org/10.3390/insects13010057
Darlington M, Reinders JD, Sethi A, Lu AL, Ramaseshadri P, Fischer JR, Boeckman CJ, Petrick JS, Roper JM, Narva KE, et al. RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions. Insects. 2022; 13(1):57. https://doi.org/10.3390/insects13010057
Chicago/Turabian StyleDarlington, Molly, Jordan D. Reinders, Amit Sethi, Albert L. Lu, Partha Ramaseshadri, Joshua R. Fischer, Chad J. Boeckman, Jay S. Petrick, Jason M. Roper, Kenneth E. Narva, and et al. 2022. "RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions" Insects 13, no. 1: 57. https://doi.org/10.3390/insects13010057
APA StyleDarlington, M., Reinders, J. D., Sethi, A., Lu, A. L., Ramaseshadri, P., Fischer, J. R., Boeckman, C. J., Petrick, J. S., Roper, J. M., Narva, K. E., & Vélez, A. M. (2022). RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions. Insects, 13(1), 57. https://doi.org/10.3390/insects13010057






