Microbial Ecology of European Foul Brood Disease in the Honey Bee (Apis mellifera): Towards a Microbiome Understanding of Disease Susceptibility
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Characterization of Disease Transmission Routes
2.1.1. PCR and MiSeq
2.1.2. MiSeq Sequence Analysis
2.1.3. Sequence Data Deposition
2.2. General Bacterial Culturing
2.3. Bacterial Inhibition Assay
2.4. Rearing of Host (A. mellifera) Larvae In Vitro
2.5. LD50 Determination M. plutonius
2.6. In Vitro Rescue Experiment
Statistical Analysis—Mortality
3. Results
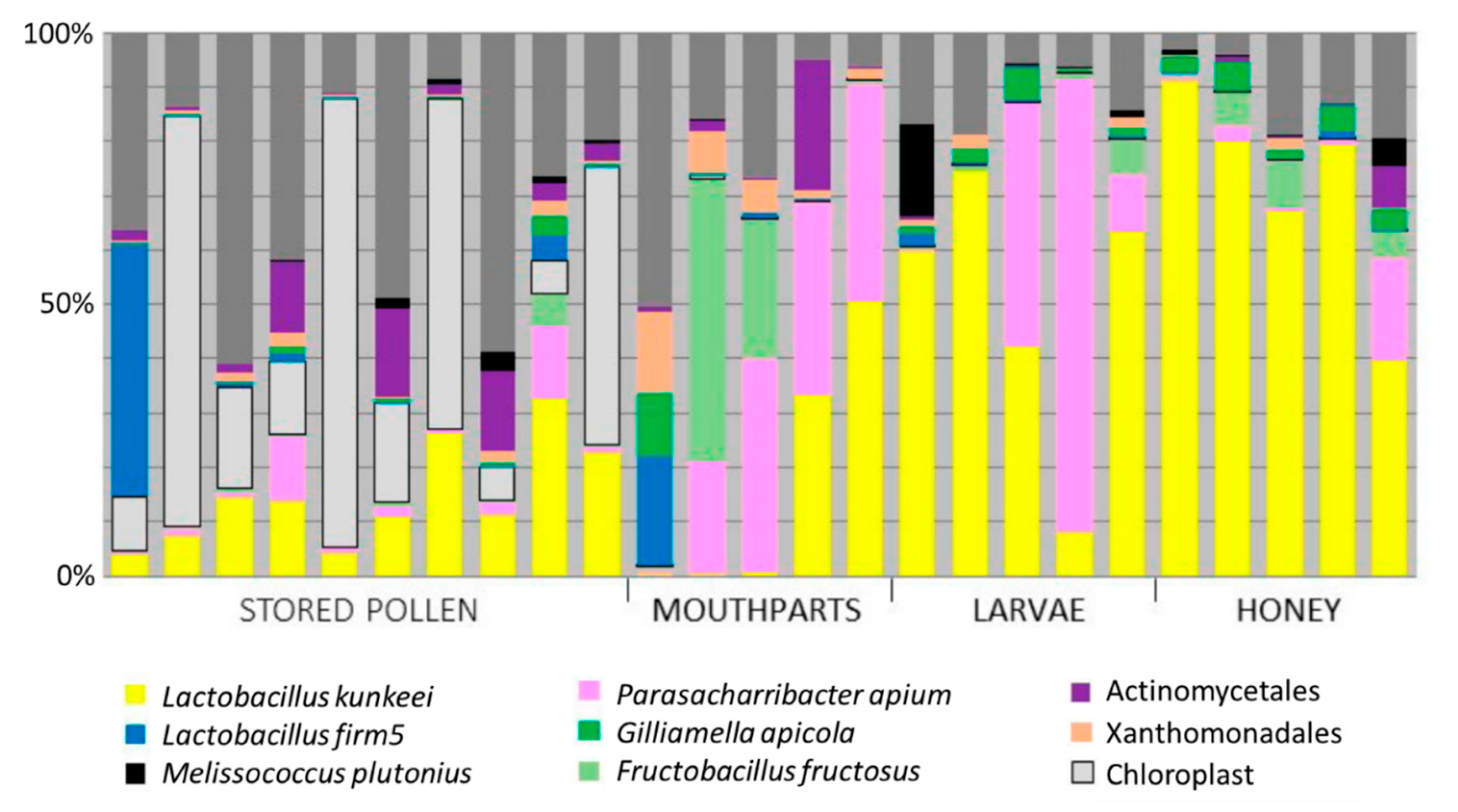
3.1. Niches Associated with Disease Transmission
3.2. Bacterial Culturing
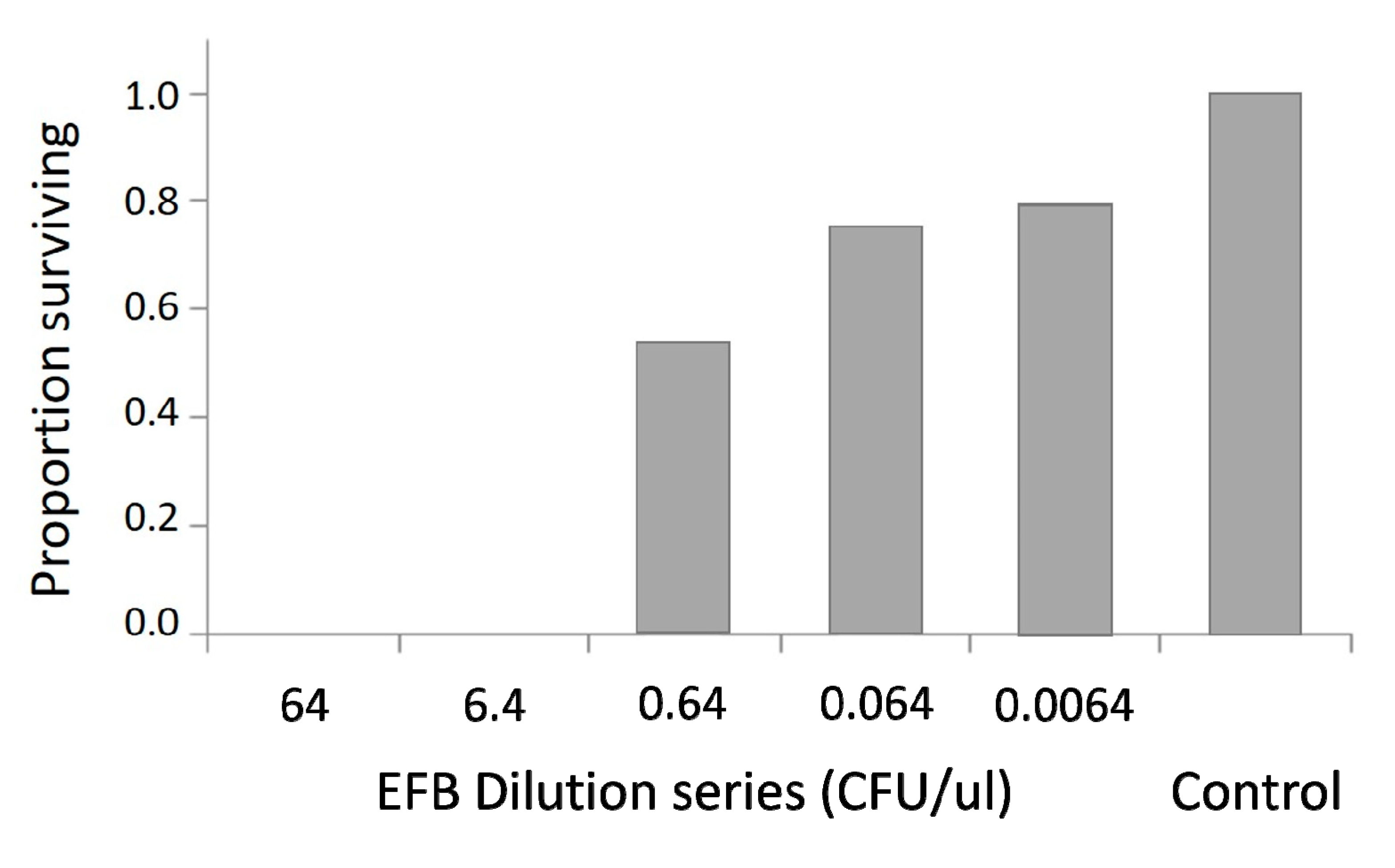
3.3. Bacterial Inhibition Assay
3.4. LD50 Determination M. plutonius
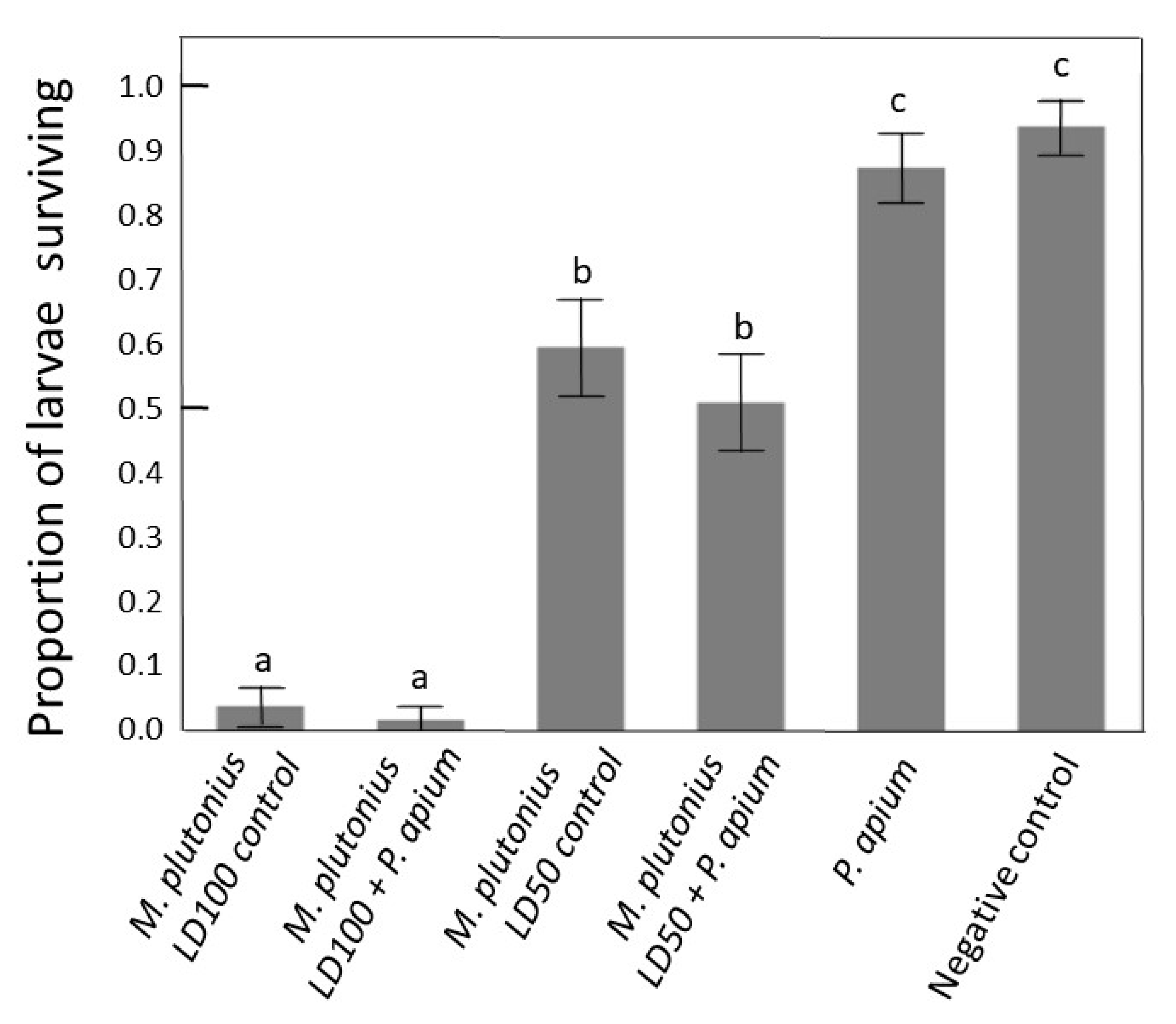
3.5. In Vitro Rescue Experiment
4. Discussion
4.1. Context of Disease Transmission
4.2. Bacterial Competition In Vitro
4.3. Larval Assay In Vitro
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chopra, S.S.; Bhavik, R.B.; Vikas, K. Economic Dependence of U.S. Industrial Sectors on Animal-Mediated Pollination Service. Environ. Sci. Technol. 2015, 49, 14441–14451. [Google Scholar] [CrossRef] [PubMed]
- Klein, A.M.; Vaissiere, B.E.; Cane, J.H.; Steffan-Dewenter, I.; Cunningham, S.A.; Kremen, C.; Tscharntke, T. Importance of pollinators in changing landscapes for world crops. Proc. Biol. Sci. 2007, 274, 303–313. [Google Scholar] [CrossRef] [PubMed]
- Ricigliano, V.A.; Mott, B.M.; Maes, P.W.; Floyd, A.S.; Fitz, W.; Copeland, D.C.; Meikle, W.G.; Anderson, K.E. Honey bee colony performance and health are enhanced by apiary proximity to US Conservation Reserve Program (CRP) lands. Sci. Rep. 2019, 9, 4894. [Google Scholar] [CrossRef] [PubMed]
- Genersch, E. American foulbrood in honey bees and its causative agent, Paenibacillus larvae. J. Invertebr. Pathol. 2010, 103, S10–S19. [Google Scholar] [CrossRef]
- Genersch, E. Honeybee pathology: Current threats to honeybees and beekeeping. Appl. Microbiol. Biotechnol. 2010, 87, 87–97. [Google Scholar] [CrossRef]
- Engelsdorp, D.V.; Lengerich, E.; Spleen, A.; Dainat, B.; Cresswell, J.; Baylis, K.; Nguyen, B.K.; Soroker, V.; Human, H.; Le Conte, Y.; et al. Standard epidemiological methods to understand and improve Apis mellifera health. J. Apicult. Res. 2015, 8839, 1–16. [Google Scholar] [CrossRef]
- Budge, G.E.; Shirley, M.D.F.; Jones, B.; Quill, E.; Tomkies, V.; Feil, E.J.; Brown, M.A.; Haynes, E.G. Molecular epidemiology and population structure of the honey bee brood pathogen Melissococcus plutonius. ISME J. 2014, 8, 1588–1597. [Google Scholar] [CrossRef]
- Lewkowski, O.; Erler, S. Virulence of Melissococcus plutonius and secondary invaders associated with European foulbrood disease of the honey bee. Microbiol. Open 2018, 8, e00649. [Google Scholar] [CrossRef]
- Maes, P.W.; Rodrigues, P.A.P.; Oliver, R.; Mott, B.M.; Anderson, K.E. Diet-related gut bacterial dysbiosis correlates with impaired development, increased mortality and Nosema disease in the honeybee (Apis mellifera). Mol. Ecol. 2016, 25, 5439–5450. [Google Scholar] [CrossRef]
- Anderson, K.E.; Ricigliano, V.A. Honey bee gut dysbiosis: A novel context of disease ecology. Curr. Opin. Insect Sci. 2017, 22, 125–132. [Google Scholar] [CrossRef]
- Erban, T.; Ledvinka, O.; Kamler, M.; Nesvorna, M.; Hortova, B.; Tyl, J.; Titera, D.; Markovic, M.; Hubert, J. Honeybee (Apis mellifera)-associated bacterial community affected by American foulbrood: Detection of Paenibacillus larvae via microbiome analysis. Sci. Rep. 2017, 7, 5084. [Google Scholar] [CrossRef] [PubMed]
- Hubert, J.; Bicianova, M.; Ledvinka, O.; Kamler, M.; Lester, P.J.; Nesvorna, M.; Kopecky, J.; Erban, T. Changes in the Bacteriome of Honey Bees Associated with the Parasite Varroa destructor, and Pathogens Nosema and Lotmaria passim. Microb. Ecol. 2017, 73, 685–698. [Google Scholar] [CrossRef] [PubMed]
- Sopko, B.; Zitek, J.; Nesvorna, M.; Markovic, M.; Kamler, M.; Titera, D.; Erban, T.; Hubert, J. Detection and quantification of Melissococcus plutonius in honey bee workers exposed to European foulbrood in Czechia through conventional PCR, qPCR, and barcode sequencing. J. Apic. Res. 2020, 59, 503–514. [Google Scholar] [CrossRef]
- Raymann, K.; Coon, K.L.; Shaffer, Z.; Salisbury, S.; Moran, N.A. Pathogenicity of Serratia marcescens Strains in Honey Bees. MBio 2018, 9, e01649-18. [Google Scholar] [CrossRef]
- Buford, T.W. (Dis)Trust your gut: The gut microbiome in age-related inflammation, health, and disease. Microbiome 2017, 5, 80. [Google Scholar] [CrossRef]
- Alberoni, D.; Gaggìa, F.; Baffoni, L.; Gioia, D.D. Beneficial microorganisms for honey bees: Problems and progresses. Appl. Microbiol. Biotechnol. 2016, 100, 9469–9482. [Google Scholar] [CrossRef]
- Killer, J.; Dubná, S.; Sedláček, I.; Švec, P. Lactobacillus apis sp. nov., from the stomach of honeybees (Apis mellifera), having an in vitro inhibitory effect on the causative agents of American and European foulbrood. Int. J. Syst. Evol. Microbiol. 2014, 64, 152–157. [Google Scholar] [CrossRef]
- Forsgren, E.; Olofsson, T.C.; Vasquez, A.; Fries, I. Novel lactic acid bacteria inhibiting Paenibacillus larvae in honey bee larvae. Apidologie 2010, 41, 99–108. [Google Scholar] [CrossRef]
- Wu, M.; Sugimura, Y.; Iwata, K.; Takaya, N.; Takamatsu, D.; Kobayashi, M.; Taylor, D.; Kimura, K.; Yoshiyama, M. Inhibitory effect of gut bacteria from the Japanese honey bee, Apis cerana japonica, against Melissococcus plutonius, the causal agent of European foulbrood disease. J. Insect. Sci. 2014, 14, 129. [Google Scholar] [CrossRef]
- Corby-Harris, V.; Snyder, L.A.; Schwan, M.R.; Maes, P.; McFrederick, Q.S.; Anderson, K.E. Origin and effect of Alpha 2.2 Acetobacteraceae in honey bee larvae and description of Parasaccharibacter apium gen. nov., sp. nov. Appl. Environ. Microbiol. 2014, 80, 7460–7472. [Google Scholar] [CrossRef]
- Stephan, J.G.; Lamei, S.; Pettis, J.S.; Riesbeck, K.; de Miranda, J.R.; Forsgren, E. Honeybee-Specific Lactic Acid Bacterium Supplements Have No Effect on American Foulbrood-Infected Honeybee Colonies. Appl. Environ. Microbiol. 2019, 85, e00606-19. [Google Scholar] [CrossRef] [PubMed]
- Rauch, S.; Ashiralieva, A.; Hedtke, K.; Genersch, E. Negative correlation between individual-insect-level virulence and colony-level virulence of Paenibacillus larvae, the etiological agent of American foulbrood of honeybees. Appl. Environ. Microbiol. 2009, 75, 3344–3347. [Google Scholar] [CrossRef] [PubMed]
- Vojvodic, S.; Jensen, A.B.; James, R.R.; Boomsma, J.J.; Eilenberg, J. Temperature dependent virulence of obligate and facultative fungal pathogens of honeybee brood. Vet. Microbiol. 2011, 149, 200–205. [Google Scholar] [CrossRef]
- Vojvodic, S.; Rehan, S.M.; Anderson, K.E. Microbial gut diversity of Africanized and European honey bee larval instars. PLoS ONE 2013, 8, e72106. [Google Scholar] [CrossRef]
- Anderson, K.E.; Sheehan, T.H.; Mott, B.M.; Maes, P.; Snyder, L.; Schwan, M.R.; Walton, A.; Jones, B.M.; Corby-Harris, V. Microbial ecology of the hive and pollination landscape: Bacterial associates from floral nectar, the alimentary tract and stored food of honey bees (Apis mellifera). PLoS ONE 2013, 8, e83125. [Google Scholar] [CrossRef] [PubMed]
- Hroncova, Z.; Havlik, J.; Killer, J.; Doskocil, I.; Tyl, J.; Kamler, M.; Titera, D.; Hakl, J.; Mrazek, J.; Bunesova, V.; et al. Variation in honey bee gut microbial diversity affected by ontogenetic stage, age and geographic location. PLoS ONE 2015, 10, e118707. [Google Scholar] [CrossRef]
- Seeley, T.D. Adaptive significance of the age polyeithism schedule in honeybee colonies. Behav. Ecol. Sociobiol. 1982, 11, 287–293. [Google Scholar] [CrossRef]
- Anderson, K.E.; Rodrigues, P.A.; Mott, B.M.; Maes, P.; Corby-Harris, V. Ecological succession in the honey bee gut: Shift in Lactobacillus strain dominance during early adult development. Microb. Ecol. 2016, 71, 1008–1019. [Google Scholar] [CrossRef]
- Forsgren, E. European foulbrood in honey bees. J. Invertebr. Pathol. 2010, 103, S5–S9. [Google Scholar] [CrossRef]
- Nakamura, K.; Yamazaki, Y.; Shiraishi, A.; Kobayashi, S.; Harada, M.; Yoshiyama, M.; Osaki, M.; Okura, M.; Takamatsu, D. Virulence Differences among Melissococcus plutonius Strains with Different Genetic Backgrounds in Apis mellifera Larvae under an Improved Experimental Condition. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Grossar, D.; Kilchenmann, V.; Forsgren, E.; Charrière, J.-D.; Gauthier, L.; Chapuisat, M.; Dietemann, V. Putative determinants of virulence in Melissococcus plutonius, the bacterial agent causing European foulbrood in honey bees. Virulence 2020, 11, 554–567. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Okumura, K.; Harada, M.; Okamota, M.; Okura, M.; Takamatsu, D. Different impacts of pMP19 on the virulence of Melissococcus plutonius strains with different genetic backgrounds. Environ. Microbiol. 2020, 22. [Google Scholar] [CrossRef] [PubMed]
- Bailey, L. The Pathogenicity for Honey-Bee Larvae of Microorganisms Associated with European Foulbrood. J. Insect Pathol. 1963, 5, 198–205. [Google Scholar]
- Tian, B.; Fadhil, N.H.; Powell, J.E.; Kwong, W.K.; Moran, N.A. Long-term exposure to antibiotics has caused accumulation of resistance determinants in the gut microbiota of honeybees. MBio 2012, 3, e00377-12. [Google Scholar] [CrossRef] [PubMed]
- Martinson, V.G.; Danforth, B.N.; Minckley, R.L.; Rueppell, O.; Tingek, S.; Moran, N.A. A simple and distinctive microbiota associated with honey bees and bumble bees. Mol. Ecol. 2011, 20, 619–628. [Google Scholar] [CrossRef] [PubMed]
- Yun, J.H.; Lee, J.Y.; Hyun, D.W.; Jung, M.J.; Bae, J.W. Bombella apis sp. nov., an acetic acid bacterium isolated from the midgut of a honey bee. Int. J. Syst. Evol. Microbiol. 2017, 67, 2184–2188. [Google Scholar] [CrossRef]
- McFrederick, Q.S.; Wcislo, W.T.; Taylor, D.R.; Ishak, H.D.; Dowd, S.E.; Mueller, U.G. Environment or kin: Whence do bees obtain acidophilic bacteria? Mol. Ecol. 2012, 21, 1754–1768. [Google Scholar] [CrossRef]
- Endo, A.; Salminen, S. Honeybees and beehives are rich sources for fructophilic lactic acid bacteria. Syst. Appl. Microbiol. 2013, 36, 444–448. [Google Scholar] [CrossRef]
- Anderson, K.E.; Ricigliano, V.A.; Mott, B.M.; Copeland, D.C.; Floyd, A.S.; Maes, P. The queen’s gut refines with age: Longevity phenotypes in a social insect model. Microbiome 2018, 6, 108. [Google Scholar] [CrossRef]
- Martinson, V.G.; Moy, J.; Moran, N.A. Establishment of characteristic gut bacteria during development of the honeybee worker. Appl. Environ. Microbiol. 2012, 78, 2830–2840. [Google Scholar] [CrossRef]
- Sabree, Z.; Hansen, A.; Moran, N.A. Independent studies using deep sequencing resolve the same set of core bacterial species dominating gut communities of honey bees. PLoS ONE 2012, 7, e41250. [Google Scholar] [CrossRef] [PubMed]
- Moran, N.A.; Hansen, A.K.; Powell, J.E.; Sabree, Z.L. Distinctive gut microbiota of honey bees assessed using deep sampling from individual worker bees. PLoS ONE 2012, 7, e36393. [Google Scholar] [CrossRef] [PubMed]
- Corby-Harris, V.; Maes, P.; Anderson, K.E. The bacterial communities associated with honey bee (Apis mellifera) foragers. PLoS ONE 2014, 9, e95056. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.E.; Carroll, M.J.; Sheehan, T.; Mott, B.M.; Maes, P.; Corby-Harris, V. Hive-stored pollen of honey bees: Many lines of evidence are consistent with pollen preservation, not nutrient conversion. Mol. Ecol. 2014, 23, 5904–5917. [Google Scholar] [CrossRef] [PubMed]
- Kapheim, K.M.; Rao, V.D.; Yeoman, C.J.; Wilson, B.A.; White, B.A.; Goldenfeld, N.; Robinson, G.E. Caste-specific differences in hindgut microbial communities of honey bees (Apis mellifera). PLoS ONE 2015, 10, e0123911. [Google Scholar] [CrossRef]
- Tarpy, D.R.; Mattila, H.R.; Newton, I.L.G. Development of the honey bee gut microbiome throughout the queen-rearing process. Appl. Environ. Microbiol. 2015, 81, 3182–3191. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.G.; Peplies, J.; Glockner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinform 2011, 27, 2194–2200. [Google Scholar] [CrossRef]
- Wang, Q.; Garrity, G.M.; Tiedje, J.M.; Cole, J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 2007, 73, 5261–5267. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Arai, R.; Tominaga, K.; Wu, M.; Okura, M.; Ito, K.; Okamura, N.; Onishi, H.; Osaki, M.; Sugimura, Y.; Yoshiyama, M.; et al. Diversity of Melissococcus plutonius from honeybee larvae in Japan and experimental reproduction of European foulbrood with cultured atypical isolates. PLoS ONE 2012, 7, e33708. [Google Scholar] [CrossRef] [PubMed]
- Chouaia, B.; Gaiarsa, S.; Crotti, E.; Comandatore, F.; Degli Esposti, M.; Ricci, I.; Alma, A.; Favia, G.; Bandi, C.; Daffonchio, D. Acetic acid bacteria genomes reveal functional traits for adaptation to life in insect guts. Genome Biol. Evol. 2014, 6, 912–920. [Google Scholar] [CrossRef] [PubMed]
- Kaftanoglu, O.; Linksvayer, T.A.; Page, R.E. Rearing honey bees (Apis mellifera L.) in vitro: Effects of feeding intervals on survival and development. J. Apic. Res. 2010, 49, 311–317. [Google Scholar] [CrossRef]
- McKee, B.A.; David Goodman, R.; Alan Hornitzky, M. The transmission of European foulbrood (Melissococcus plutonius) to artificially reared honey bee larvae (Apis mellifera). J. Apic. Res. 2004, 43, 93–100. [Google Scholar] [CrossRef]
- Garrido-Bailón, E.; Higes, M.; Martínez-Salvador, A.; Antúnez, K.; Botías, C.; Meana, A.; Prieto, L.; Martín-Hernández, R. The prevalence of the honeybee brood pathogens Ascosphaera apis, Paenibacillus larvae and Melissococcus plutonius in Spanish apiaries determined with a new multiplex PCR assay. Microb. biotechnol. 2013, 6, 731–739. [Google Scholar] [CrossRef]
- Fünfhaus, A.; Ebeling, J.; Genersch, E. Bacterial pathogens of bees. Curr. Opin. Insect Sci. 2018, 26, 89–96. [Google Scholar] [CrossRef]
- Govan, V.A.; Zel, V.B.; Allsopp, M.H.; Davison, A.S. A PCR Detection Method for Rapid Identification of Melissococcus pluton in Honeybee Larvae. Appl. Environ. Microbiol. 1998, 64. [Google Scholar] [CrossRef]
- Takamatsu, D.; Okumura, K.; Tabata, A.; Okamoto, M.; Okura, M. Transcriptional regulator SpxA1a controls the resistance of the honey bee pathogen Melissococcus plutonius to the antimicrobial activity of royal jelly. Environ. Microbiol. 2020, 22, 2736–2755. [Google Scholar] [CrossRef]
- Djukic, M.; Erler, S.; Leimbach, A.; Grossar, D.; Charrière, J.D.; Gauthier, L.; Hartken, D.; Dietrich, S.; Nacke, H.; Daniel, R.; et al. Comparative genomics and description of putative virulence factors of Melissococcus plutonius, the causative agent of European foulbrood disease in honey bees. Genes 2018, 9, 419. [Google Scholar] [CrossRef]
- Erban, T.; Ledvinka, O.; Kamler, M.; Hortova, B.; Nesvorna, M.; Tyl, J.; Titera, D.; Markovic, M.; Hubert, J. Bacterial community associated with worker honeybees (Apis mellifera) affected by European foulbrood. PeerJ 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Takamatsu, D.; Osawa, A.; Nakamura, K.; Yoshiyama, M.; Okura, M. High-level resistance of Melissococcus plutonius clonal complex 3 strains to antimicrobial activity of royal jelly. Environ. Microbiol. Rep. 2017, 9, 562–570. [Google Scholar] [CrossRef] [PubMed]
- Crailsheim, K.; Brodschneider, R.; Aupinel, P.; Behrens, D.; Genersch, E.; Vollmann, J.; Riessberger-Gallé, U. Standard methods for artificial rearing of Apis mellifera larvae. J. Apic. Res. 2013, 52, 1–16. [Google Scholar] [CrossRef]
- Schmehl, D.R.; Tomé, H.V.V.; Mortensen, A.N.; Martins, G.F.; Ellis, J.D. Protocol for the in vitro rearing of honey bee (Apis mellifera L.) workers. J. Apic. Res. 2016, 55, 113–129. [Google Scholar] [CrossRef]
- Mortensen, A.N.; Ellis, J.D. A honey bee (Apis mellifera) colony’s brood survival rate predicts its in vitro-reared brood survival rate. Apidologie 2018, 49, 573–580. [Google Scholar] [CrossRef]
| Study | Site | Tissue or Niche | Developmental Stage or Caste | Total Sequences | % P. apium |
|---|---|---|---|---|---|
| Relative abundance of P. apium from whole worker gut samples | |||||
| Martinson et al. 2012 [40] | AZ | whole guts | 9-day-old nurses (N = 3) | 78,595 | 0 |
| Martinson et al. 2012 [40] | AZ | whole guts | 30-day-old forager (N = 1) | 17,910 | 0 |
| Sabree et al. 2012 [41] | MA | whole guts | 12 days old | 106,344 | 1 |
| Moran et al. 2012 [42] | AZ, MD | whole guts | in hive, outer frames | 329,550 | 1 |
| Corby-Harris et al. 2014a [43] | AZ | whole guts | pollen foragers | 354,505 | 6 |
| Anderson et al. 2016 [28] | AZ | whole guts | 3-day-old bees A | 90,893 | 0 |
| Anderson et al. 2016 [28] | AZ | whole guts | 7-day-old bees A | 166,113 | 0 |
| Relative abundance of P. apium from honey-rich food storage environments | |||||
| Anderson et al. 2014 [44] | AZ | corbicular pollen | forager corbiculae | 85,043 | 6 |
| Anderson et al. 2014 [44] | AZ | stored pollen | hive environment | 116,593 | 3 |
| present study | AZ | ripening honey | hive environment | 784,635 | 4 |
| present study | AZ | stored pollen | hive environment | 314,705 | 6 |
| Relative abundance of P. apium from niches with >20% P. apium abundance | |||||
| Corby-Harris et al. 2014a [43] | AZ | crop (foregut) | pollen forager | 195,264 | 43 |
| Corby-Harris et al. 2014a [43] | AZ | crop (foregut) | nurse | 113,405 | 32 |
| Corby-Harris et al. 2014b [20] | AZ | nurse headglands | nurse | 127,157 | 34 |
| Corby-Harris et al. 2014b [20] | AZ | royal jelly | worker head glands | 105,820 | 40 |
| Kapheim et al. 2015 [45] | IL | dissected guts | queen (N = 4) | 78,355 | 38 |
| Tarpy et al. 2015 [46] | NC | dissected guts | queen B | 5000 | 24 |
| Anderson et al. 2018 [39] | AZ, CA | mouthparts | queen C | 1,305,332 | 89 |
| Anderson et al. 2018 [39] | AZ, CA | midgut | queen C | 1,572,752 | 86 |
| Anderson et al. 2018 [39] | AZ, CA | ileum | queen C | 1,419,416 | 38 |
| present study | AZ | mouthparts | nurse worker | 502,473 | 22 |
| present study | AZ | whole body | worker larvae 2nd instar | 330,556 | 25 |
| Time | * CFU 1 | CFU 2 | CFU 3 | Avg | CFU/µL in Diet |
|---|---|---|---|---|---|
| 0 h | 285 | 261 | 253 | 266 | 33.29 |
| 6 h | 114 | 142 | 152 | 136 | 13.6 |
| 24 h | 91 | 74 | 57 | 74 | 1.48 |
| 48 h | 59 | 85 | 53 | 66 | 1.31 |
| 72 h | 5 | 9 | 7 | 7 | 0.14 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Floyd, A.S.; Mott, B.M.; Maes, P.; Copeland, D.C.; McFrederick, Q.S.; Anderson, K.E. Microbial Ecology of European Foul Brood Disease in the Honey Bee (Apis mellifera): Towards a Microbiome Understanding of Disease Susceptibility. Insects 2020, 11, 555. https://doi.org/10.3390/insects11090555
Floyd AS, Mott BM, Maes P, Copeland DC, McFrederick QS, Anderson KE. Microbial Ecology of European Foul Brood Disease in the Honey Bee (Apis mellifera): Towards a Microbiome Understanding of Disease Susceptibility. Insects. 2020; 11(9):555. https://doi.org/10.3390/insects11090555
Chicago/Turabian StyleFloyd, Amy S., Brendon M. Mott, Patrick Maes, Duan C. Copeland, Quinn S. McFrederick, and Kirk E. Anderson. 2020. "Microbial Ecology of European Foul Brood Disease in the Honey Bee (Apis mellifera): Towards a Microbiome Understanding of Disease Susceptibility" Insects 11, no. 9: 555. https://doi.org/10.3390/insects11090555
APA StyleFloyd, A. S., Mott, B. M., Maes, P., Copeland, D. C., McFrederick, Q. S., & Anderson, K. E. (2020). Microbial Ecology of European Foul Brood Disease in the Honey Bee (Apis mellifera): Towards a Microbiome Understanding of Disease Susceptibility. Insects, 11(9), 555. https://doi.org/10.3390/insects11090555






