Probing the Honey Bee Diet-Microbiota-Host Axis Using Pollen Restriction and Organic Acid Feeding
Abstract
1. Introduction
2. Materials and Methods
2.1. Determination of Bacterial Abundance in the Honey Bee Anterior Rectum
2.2. Experimental Setup for Pollen and Organic Acid Feeding
2.3. RNA Extraction and qPCR
2.4. Comparison of Bacterial Abundance and Fermentative Enzyme Gene Expression
2.5. Comparison of Honey Bee Enteroendocrine Signaling Gene Expression
2.6. Statistical Analyses
3. Results
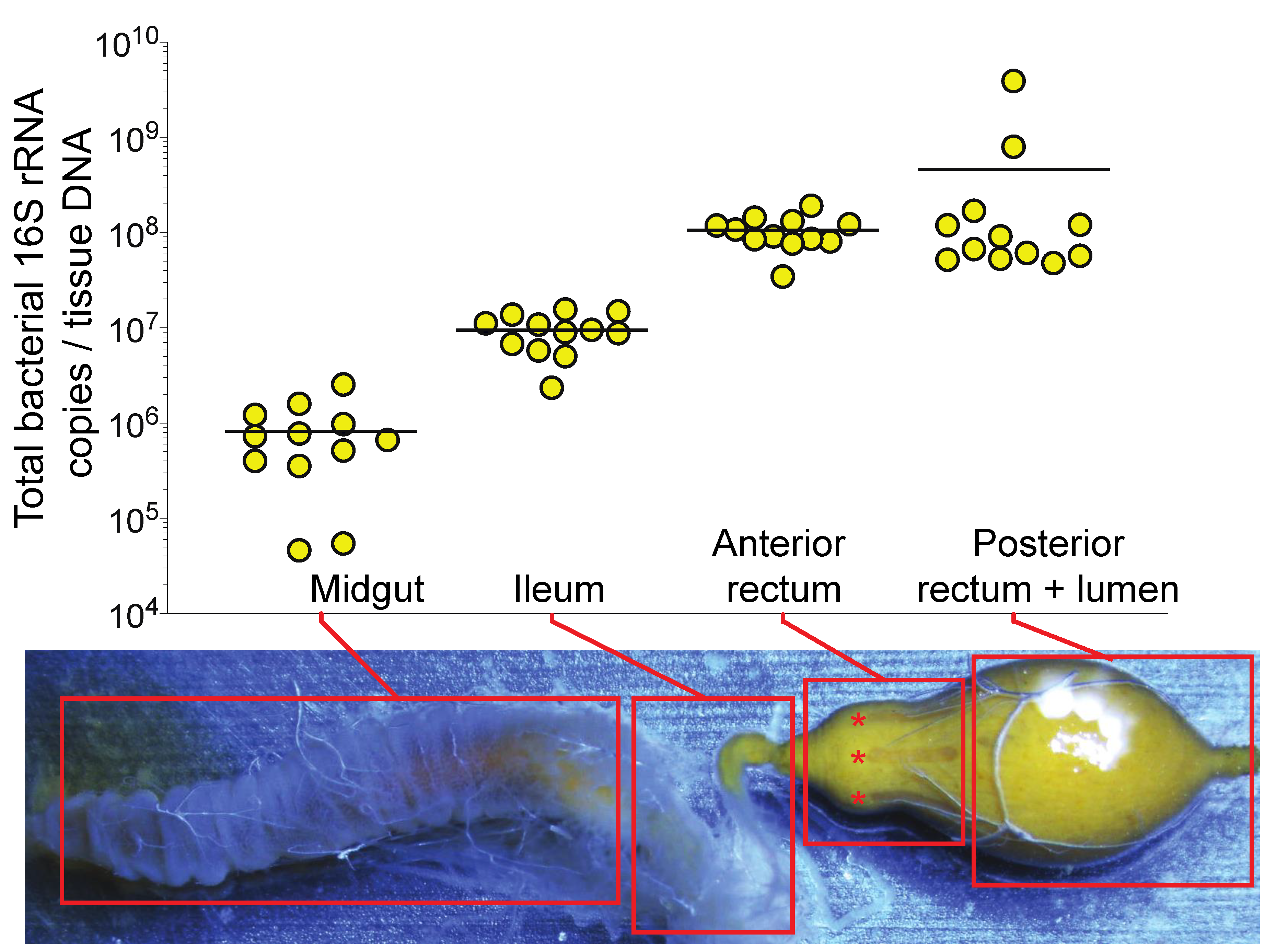
3.1. Bacterial Abundance in the Honey Bee Gut is Partially Associated with the Anterior Rectum Epithelium
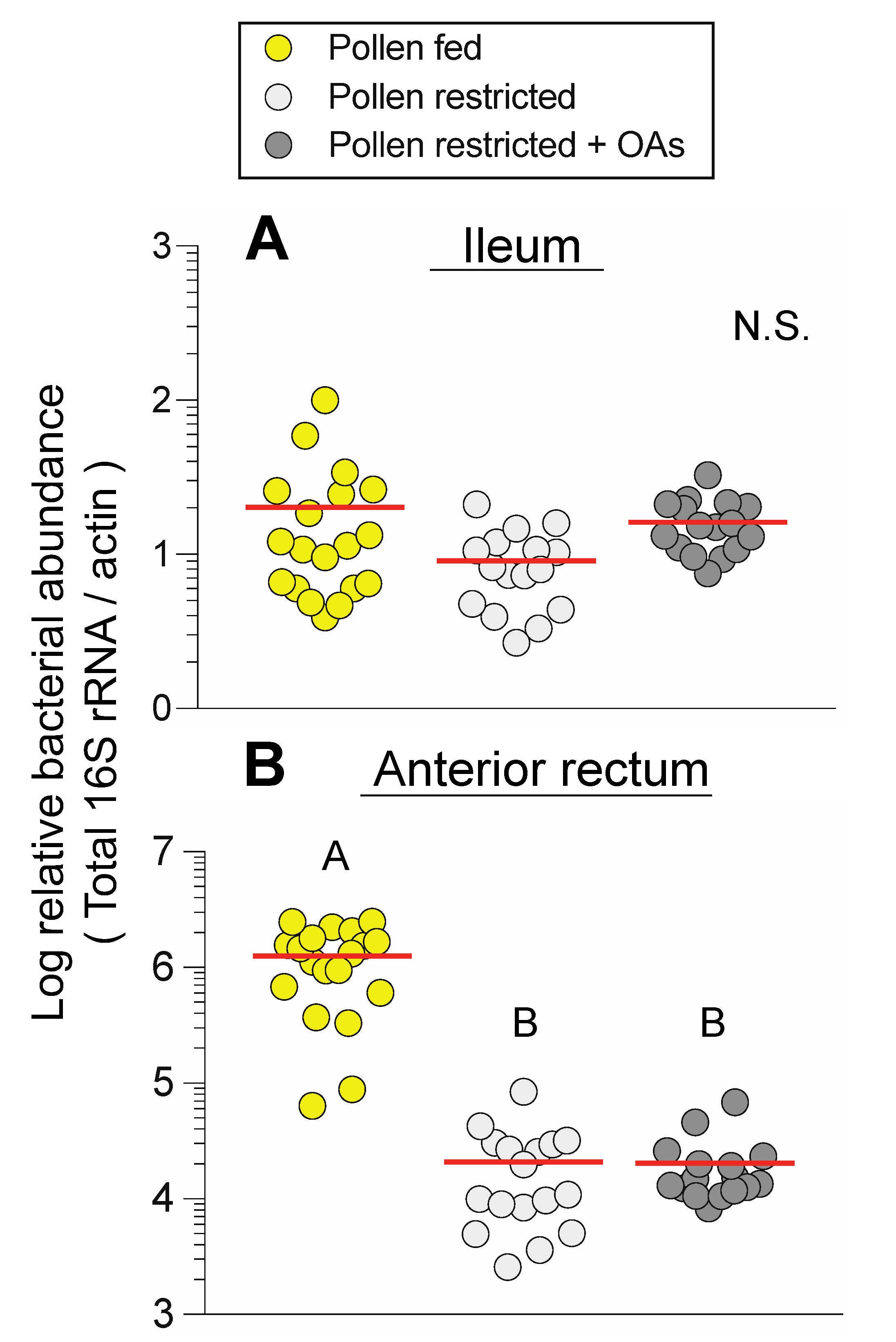
3.2. Dietary Pollen Restriction Markedly Reduces Total and Specific Bacterial Abundance in the Anterior Rectum
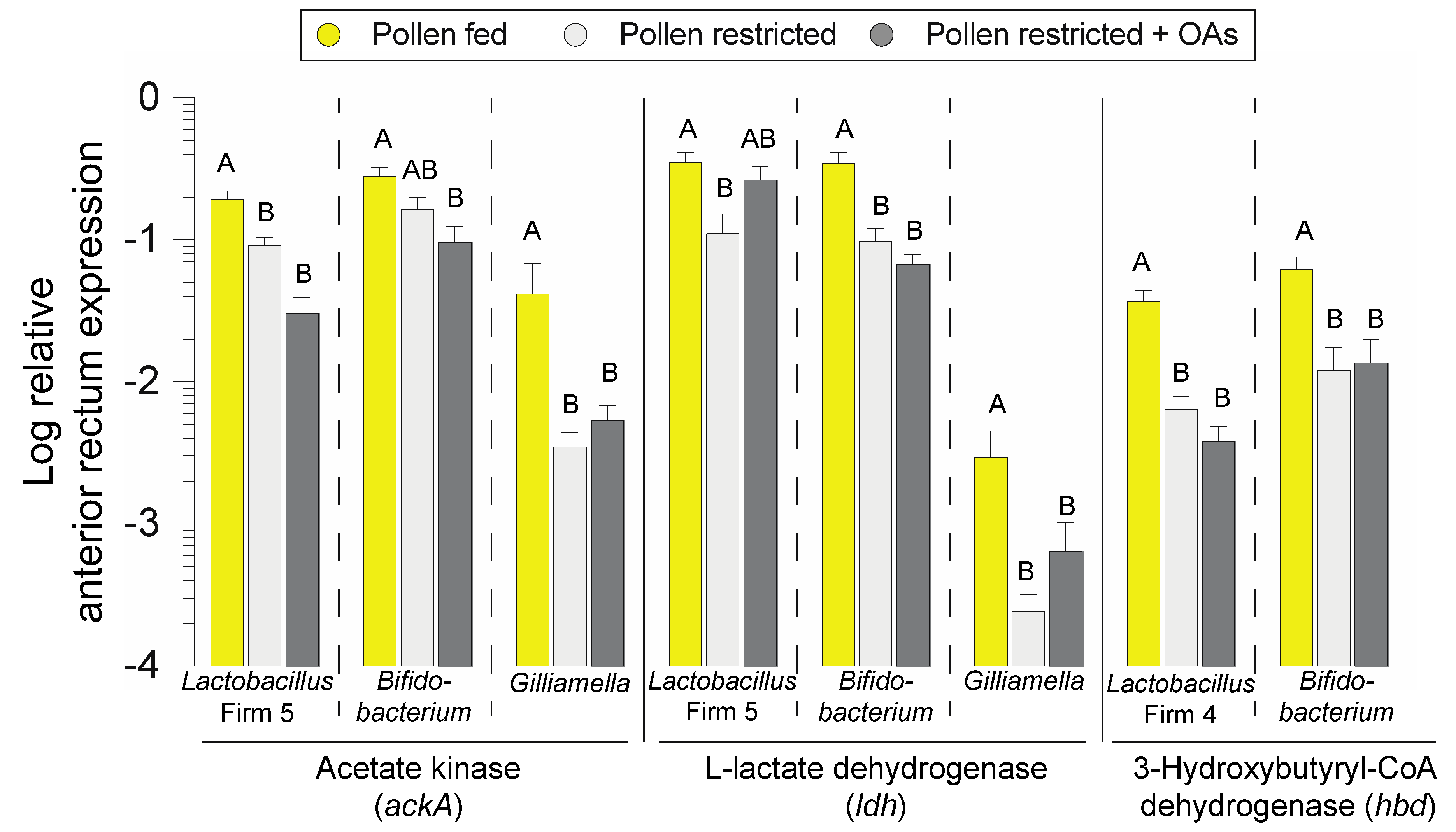
3.3. Bacterial Fermentative Enzyme Gene Transcripts in the Anterior Rectum are Reduced in Association with Diet-Induced Microbiota Shifts
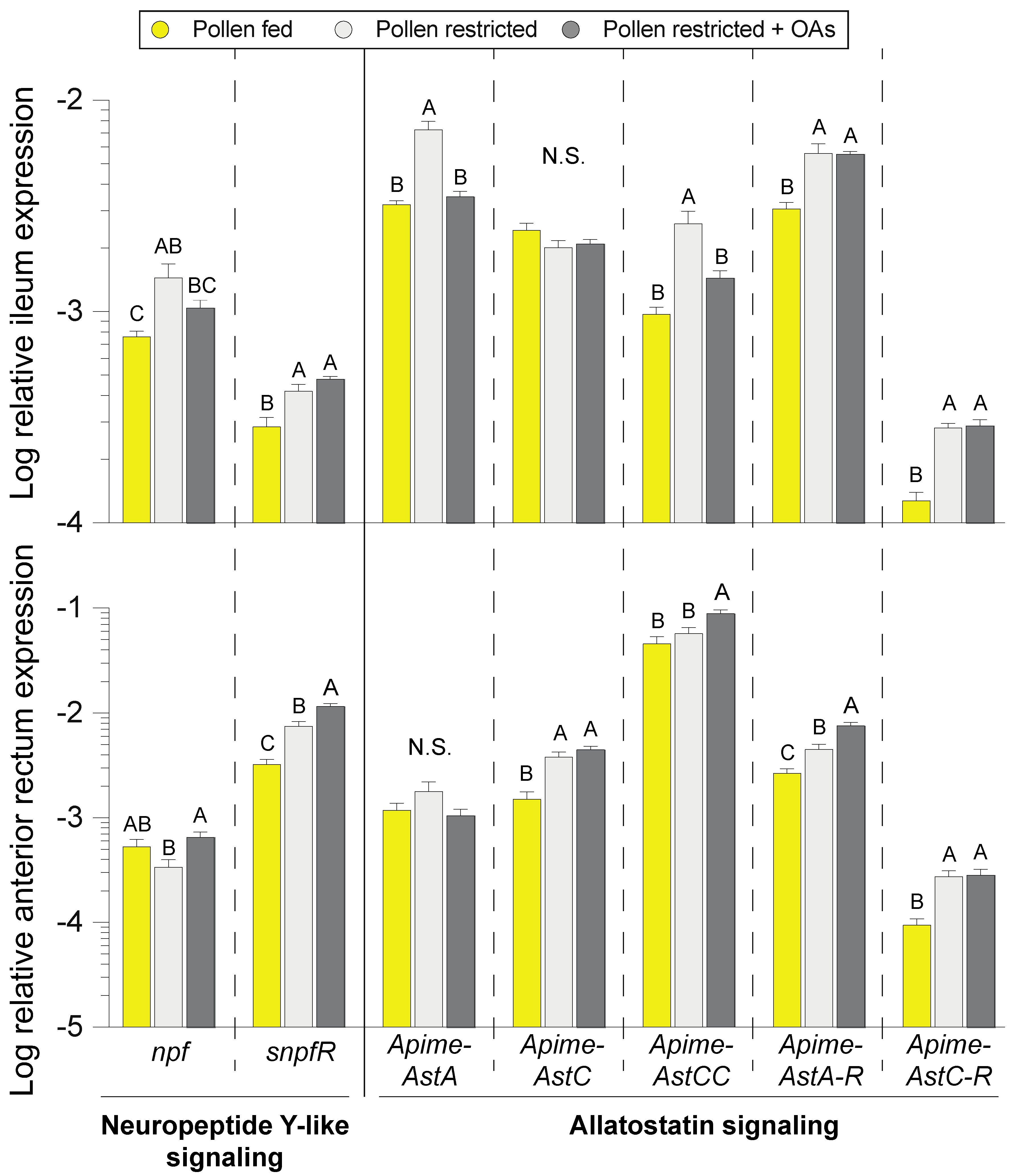
3.4. Modulation of Enteroendocrine Signaling by Pollen and Organic Acid Feeding
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kostic, A.D.; Howitt, M.R.; Garrett, W.S. Exploring host-microbiota interactions in animal models and humans. Genes Dev. 2013, 27, 701–718. [Google Scholar] [CrossRef] [PubMed]
- Pernice, M.; Simpson, S.J.; Ponton, F. Towards an integrated understanding of gut microbiota using insects as model systems. J. Insect Physiol. 2014, 69, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Steele, M.I.; Leonard, S.P.; Motta, E.V.S.; Moran, N.A. Honey bees as models for gut microbiota research. Lab Anim. 2018, 47, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Douglas, A.E. Multiorganismal Insects: Diversity and Function of Resident Microorganisms. Annu. Rev. Entomol. 2015, 60, 17–34. [Google Scholar] [CrossRef] [PubMed]
- Engel, P.; Moran, N.A. The gut microbiota of insects—Diversity in structure and function. FEMS Microbiol. Rev. 2013, 37, 699–735. [Google Scholar] [CrossRef] [PubMed]
- Ankrah, N.Y.D.; Douglas, A.E. Nutrient factories: Metabolic function of beneficial microorganisms associated with insects. Environ. Microbiol. 2018, 20, 2002–2011. [Google Scholar] [CrossRef] [PubMed]
- Kwong, W.K.; Mancenido, A.L.; Moran, N.A. Immune system stimulation by the native gut microbiota of honey bees. R. Soc. Open Sci. 2017, 4, 170003. [Google Scholar] [CrossRef]
- Emery, O.; Schmidt, K.; Engel, P. Immune system stimulation by the gut symbiont Frischella perrara in the honey bee (Apis mellifera). Mol. Ecol. 2017, 26, 2576–2590. [Google Scholar] [CrossRef]
- Zheng, H.; Powell, J.E.; Steele, M.I.; Dietrich, C.; Moran, N.A. Honeybee gut microbiota promotes host weight gain via bacterial metabolism and hormonal signaling. Proc. Natl. Acad. Sci. USA 2017, 114, 4775–4780. [Google Scholar] [CrossRef]
- Martinson, V.G.; Moy, J.; Moran, N.A. Establishment of Characteristic Gut Bacteria during Development of the Honeybee Worker. Appl. Environ. Microbiol. 2012, 78, 2830–2840. [Google Scholar] [CrossRef]
- Engel, P.; Moran, N.A. Functional and evolutionary insights into the simple yet specific gut microbiota of the honey bee from metagenomic analysis. Gut Microbes 2014, 4, 60–65. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.E.; Ricigliano, V.A. Honey bee gut dysbiosis: A novel context of disease ecology. Curr. Opin. Insect Sci. 2017, 22, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Kwong, W.K.; Moran, N.A. Cultivation and characterization of the gut symbionts of honey bees and bumble bees: Description of Snodgrassella alvi gen. nov., sp. nov., a member of the family Neisseriaceae of the Betaproteobacteria, and Gilliamella apicola gen. nov., sp. nov., a member of Orbaceae fam. nov., Orbales ord. nov., a sister taxon to the order “Enterobacteriales” of the Gammaproteobacteria. Int. J. Syst. Evol. Microbiol. 2013, 63, 2008–2018. [Google Scholar] [PubMed]
- Kwong, W.K.; Engel, P.; Koch, H.; Moran, N.A. Genomics and host specialization of honey bee and bumble bee gut symbionts. Proc. Natl. Acad. Sci. USA 2014, 111, 11509–11514. [Google Scholar] [CrossRef] [PubMed]
- Lee, F.J.; Rusch, D.B.; Stewart, F.J.; Mattila, H.R.; Newton, I.L.G. Saccharide breakdown and fermentation by the honey bee gut microbiome. Environ. Microbiol. 2014, 17, 796–815. [Google Scholar] [CrossRef]
- Zheng, H.; Nishida, A.; Kwong, W.K.; Koch, H.; Engel, P.; Steele, M.I.; Moran, N.A. Metabolism of Toxic Sugars by Strains of the Bee Gut Symbiont Gilliamella apicola. MBio 2016, 7, e01326-16. [Google Scholar] [CrossRef]
- Kešnerová, L.; Mars, R.A.T.; Ellegaard, K.M.; Troilo, M.; Sauer, U.; Engel, P. Disentangling metabolic functions of bacteria in the honey bee gut. PLoS Biol. 2017, 15, e2003467. [Google Scholar] [CrossRef]
- Gribble, F.M.; Reimann, F. Enteroendocrine Cells: Chemosensors in the Intestinal Epithelium. Annu. Rev. Physiol. 2016, 78, 277–299. [Google Scholar] [CrossRef]
- Gribble, F.M.; Reimann, F. Function and mechanisms of enteroendocrine cells and gut hormones in metabolism. Nat. Rev. Endocrinol. 2019, 15, 226–237. [Google Scholar] [CrossRef]
- Wegener, C.; Veenstra, J.A. Chemical identity, function and regulation of enteroendocrine peptides in insects. Curr. Opin. Insect Sci. 2015, 11, 8–13. [Google Scholar] [CrossRef]
- Cani, P.D.; Everard, A.; Duparc, T. Gut microbiota, enteroendocrine functions and metabolism. Curr. Opin. Pharmacol. 2013, 13, 935–940. [Google Scholar] [CrossRef] [PubMed]
- Davis, M.M.; Engström, Y. Immune response in the barrier epithelia: Lessons from the fruit fly Drosophila melanogaster. J. Innate Immun. 2012, 4, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Schoofs, L.; De Loof, A.; Van Hiel, M.B. Neuropeptides as Regulators of Behavior in Insects. Annu. Rev. Entomol. 2017, 62, 35–52. [Google Scholar] [CrossRef] [PubMed]
- Elphick, M.R.; Mirabeau, O.; Larhammar, D. Evolution of neuropeptide signalling systems. J. Exp. Biol. 2018, 221, jeb151092. [Google Scholar] [CrossRef] [PubMed]
- Christie, A.E. Assessment of midgut enteroendocrine peptide complement in the honey bee, Apis mellifera. Insect Biochem. Mol. Biol. 2020, 116, 103257. [Google Scholar] [CrossRef]
- Ament, S.A.; Velarde, R.A.; Kolodkin, M.H.; Moyse, D.; Robinson, G.E. Neuropeptide Y-like signalling and nutritionally mediated gene expression and behaviour in the honey bee. Insect Mol. Biol. 2011, 20, 335–345. [Google Scholar] [CrossRef]
- Nässel, D.R.; Wegener, C. A comparative review of short and long neuropeptide F signaling in invertebrates: Any similarities to vertebrate neuropeptide Y signaling? Peptides 2011, 32, 1335–1355. [Google Scholar] [CrossRef]
- Wu, Q.; Wen, T.; Lee, G.; Park, J.H.; Cai, H.N.; Shen, P. Developmental control of foraging and social behavior by the Drosophila neuropeptide Y-like system. Neuron 2003, 39, 147–161. [Google Scholar] [CrossRef]
- Ameku, T.; Yoshinari, Y.; Texada, M.J.; Kondo, S.; Amezawa, K.; Yoshizaki, G.; Shimada-Niwa, Y.; Niwa, R. Midgut-derived neuropeptide F controls germline stem cell proliferation in a mating-dependent manner. PLoS Biol. 2018, 16, e2005004. [Google Scholar] [CrossRef]
- Urlacher, E.; Soustelle, L.; Parmentier, M.-L.; Verlinden, H.; Gherardi, M.-J.; Fourmy, D.; Mercer, A.R.; Devaud, J.-M.; Massou, I. Honey Bee Allatostatins Target Galanin/Somatostatin-Like Receptors and Modulate Learning: A Conserved Function? PLoS ONE 2016, 11, e0146248-30. [Google Scholar] [CrossRef]
- Stay, B.; Tobe, S.S. The role of allatostatins in juvenile hormone synthesis in insects and crustaceans. Annu. Rev. Entomol. 2007, 52, 277–299. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Reiher, W.; Hermann-Luibl, C.; Sellami, A.; Cognigni, P.; Kondo, S.; Helfrich-Förster, C.; Veenstra, J.A.; Wegener, C. Allatostatin A Signalling in Drosophila Regulates Feeding and Sleep and Is Modulated by PDF. PLoS Genet 2016, 12, e1006346-33. [Google Scholar]
- Wang, C.; Chin-Sang, I.; Bendena, W.G. The FGLamide-Allatostatins Influence Foraging Behavior in Drosophila melanogaster. PLoS ONE 2012, 7, e36059. [Google Scholar] [CrossRef] [PubMed]
- McCauley, H.A. Enteroendocrine Regulation of Nutrient Absorption. J. Nutr. 2020, 150, 10–21. [Google Scholar] [CrossRef] [PubMed]
- Dalile, B.; Van Oudenhove, L.; Vervliet, B.; Verbeke, K. The role of short-chain fatty acids in microbiota-gut-brain communication. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 461–478. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Bayer, E.A.; Rincon, M.T.; Lamed, R.; White, B.A. Polysaccharide utilization by gut bacteria: Potential for new insights from genomic analysis. Nat. Rev. Microbiol. 2008, 6, 121–131. [Google Scholar] [CrossRef]
- den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Koh, A.; De Vadder, F.; Kovatcheva-Datchary, P.; Bäckhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016, 165, 1332–1345. [Google Scholar] [CrossRef]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef]
- Ricigliano, V.A.; Fitz, W.; Copeland, D.C.; Mott, B.M.; Maes, P.; Floyd, A.S.; Dockstader, A.; Anderson, K.E. The impact of pollen consumption on honey bee (Apis mellifera) digestive physiology and carbohydrate metabolism. Arch. Insect Biochem. Physiol. 2017, 96, e21406-14. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Ricigliano, V.A.; Mott, B.M.; Floyd, A.S.; Copeland, D.C.; Carroll, M.J.; Anderson, K.E. Honey bees overwintering in a southern climate: Longitudinal effects of nutrition and queen age on colony-level molecular physiology and performance. Sci. Rep. 2018, 8, 88. [Google Scholar] [CrossRef] [PubMed]
- Lee, F.J.; Miller, K.I.; McKinlay, J.B.; Newton, I.L.G. Differential carbohydrate utilization and organic acid production by honey bee symbionts. FEMS Microbiol. Ecol. 2018, 94, 3557. [Google Scholar] [CrossRef] [PubMed]
- Ellegaard, K.M.; Tamarit, D.; Javelind, E.; Olofsson, T.C.; Andersson, S.G.E.; Vásquez, A. Extensive intra-phylotype diversity in lactobacilli and bifidobacteria from the honeybee gut. BMC Genom. 2015, 16, 284. [Google Scholar] [CrossRef]
- Taylor, R.C.; Brown, A.K.; Singh, A.; Bhatt, A.; Besra, G.S. Characterization of a beta-hydroxybutyryl-CoA dehydrogenase from Mycobacterium tuberculosis. Microbiology 2010, 156, 1975–1982. [Google Scholar] [CrossRef]
- Cai, G.; Jin, B.; Saint, C.; Monis, P. Genetic manipulation of butyrate formation pathways in Clostridium butyricum. J. Biotechnol. 2011, 155, 269–274. [Google Scholar] [CrossRef]
- Engel, P.; Martinson, V.G. Functional diversity within the simple gut microbiota of the honey bee. Proc. Natl. Acad. Sci. USA 2012, 109, 11002–11007. [Google Scholar] [CrossRef]
- Phillips, J.E. Apparent transport of water by insect excretory systems. Am. Zool. 1970, 10, 413–436. [Google Scholar] [CrossRef]
- Andersen, M.K.; MacMillan, H.A.; Donini, A.; Overgaard, J. Cold tolerance of Drosophila species is tightly linked to the epithelial K+ transport capacity of the Malpighian tubules and rectal pads. J. Exp. Biol. 2017, 220, 4261–4269. [Google Scholar] [CrossRef]
- Ellegaard, K.M.; Engel, P. Genomic diversity landscape of the honey bee gut microbiota. Nat. Commun. 2019, 10, 446. [Google Scholar] [CrossRef]
- Kelly, C.J.; Zheng, L.; Campbell, E.L.; Saeedi, B.; Scholz, C.C.; Bayless, A.J.; Wilson, K.E.; Glover, L.E.; Kominsky, D.J.; Magnuson, A.; et al. Crosstalk between Microbiota-Derived Short-Chain Fatty Acids and Intestinal Epithelial HIF Augments Tissue Barrier Function. Cell Host Microbe 2015, 17, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Ricigliano, V.A. Microalgae as a promising and sustainable nutrition source for managed honey bees. Arch. Insect Biochem. Physiol. 2020, 27, 253. [Google Scholar] [CrossRef] [PubMed]
- Hartenstein, V. The neuroendocrine system of invertebrates: A developmental and evolutionary perspective. J. Endocrinol. 2006, 190, 555–570. [Google Scholar] [CrossRef] [PubMed]
- Anderson, K.E.; Ricigliano, V.A.; Mott, B.M.; Copeland, D.C.; Floyd, A.S.; Maes, P. The queen’s gut refines with age: Longevity phenotypes in a social insect model. Microbiome 2018, 6, 287. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ricigliano, V.A.; Anderson, K.E. Probing the Honey Bee Diet-Microbiota-Host Axis Using Pollen Restriction and Organic Acid Feeding. Insects 2020, 11, 291. https://doi.org/10.3390/insects11050291
Ricigliano VA, Anderson KE. Probing the Honey Bee Diet-Microbiota-Host Axis Using Pollen Restriction and Organic Acid Feeding. Insects. 2020; 11(5):291. https://doi.org/10.3390/insects11050291
Chicago/Turabian StyleRicigliano, Vincent A., and Kirk E. Anderson. 2020. "Probing the Honey Bee Diet-Microbiota-Host Axis Using Pollen Restriction and Organic Acid Feeding" Insects 11, no. 5: 291. https://doi.org/10.3390/insects11050291
APA StyleRicigliano, V. A., & Anderson, K. E. (2020). Probing the Honey Bee Diet-Microbiota-Host Axis Using Pollen Restriction and Organic Acid Feeding. Insects, 11(5), 291. https://doi.org/10.3390/insects11050291





