Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Some European Countries: Diffusion, Hosts, Molecular Characterization, and Natural Enemies
Abstract
1. Introduction
2. Materials and Methods
2.1. Monitoring Activities
2.2. Morpho-Molecular Characterization
3. Results
3.1. Monitoring Activities
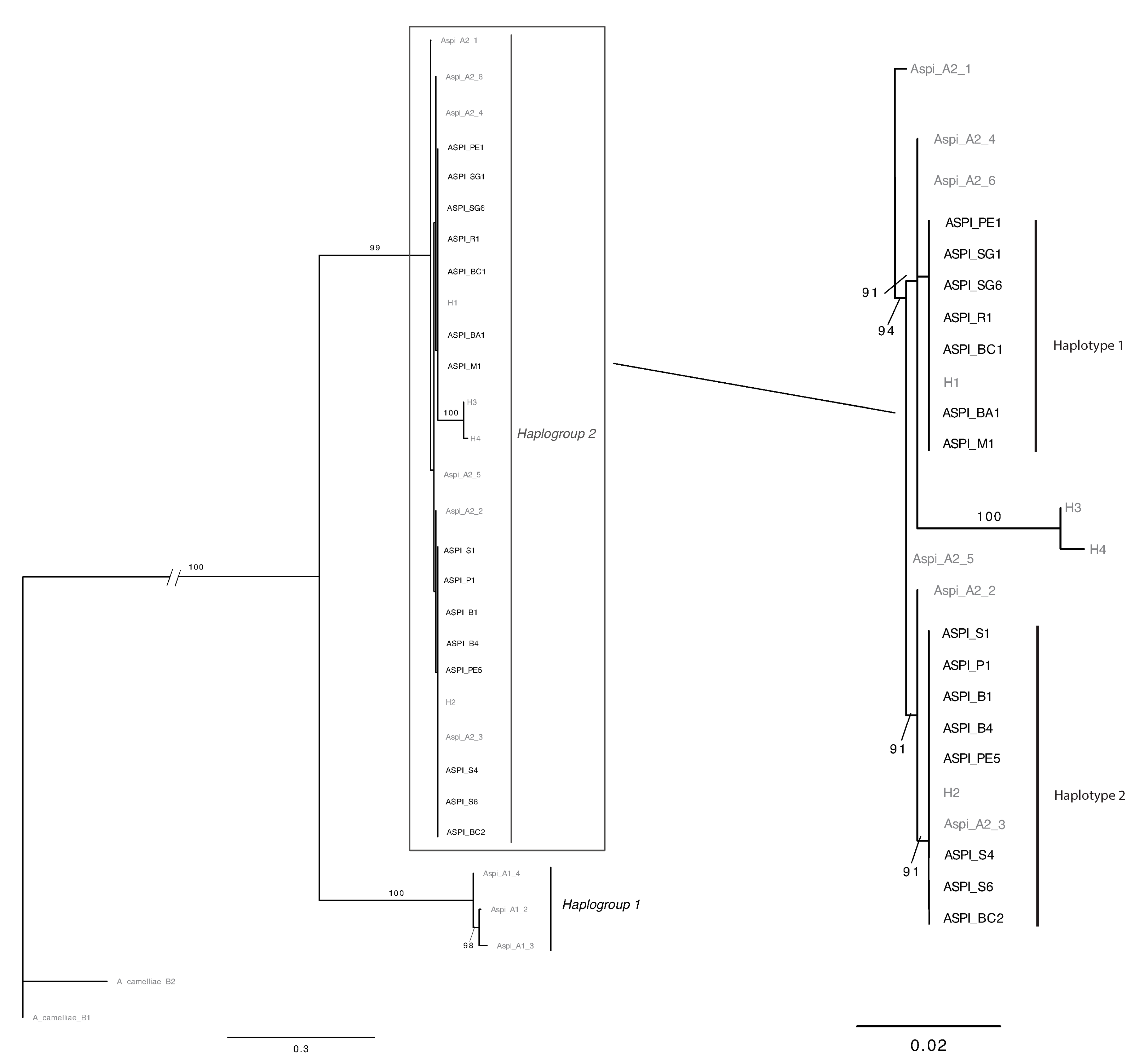
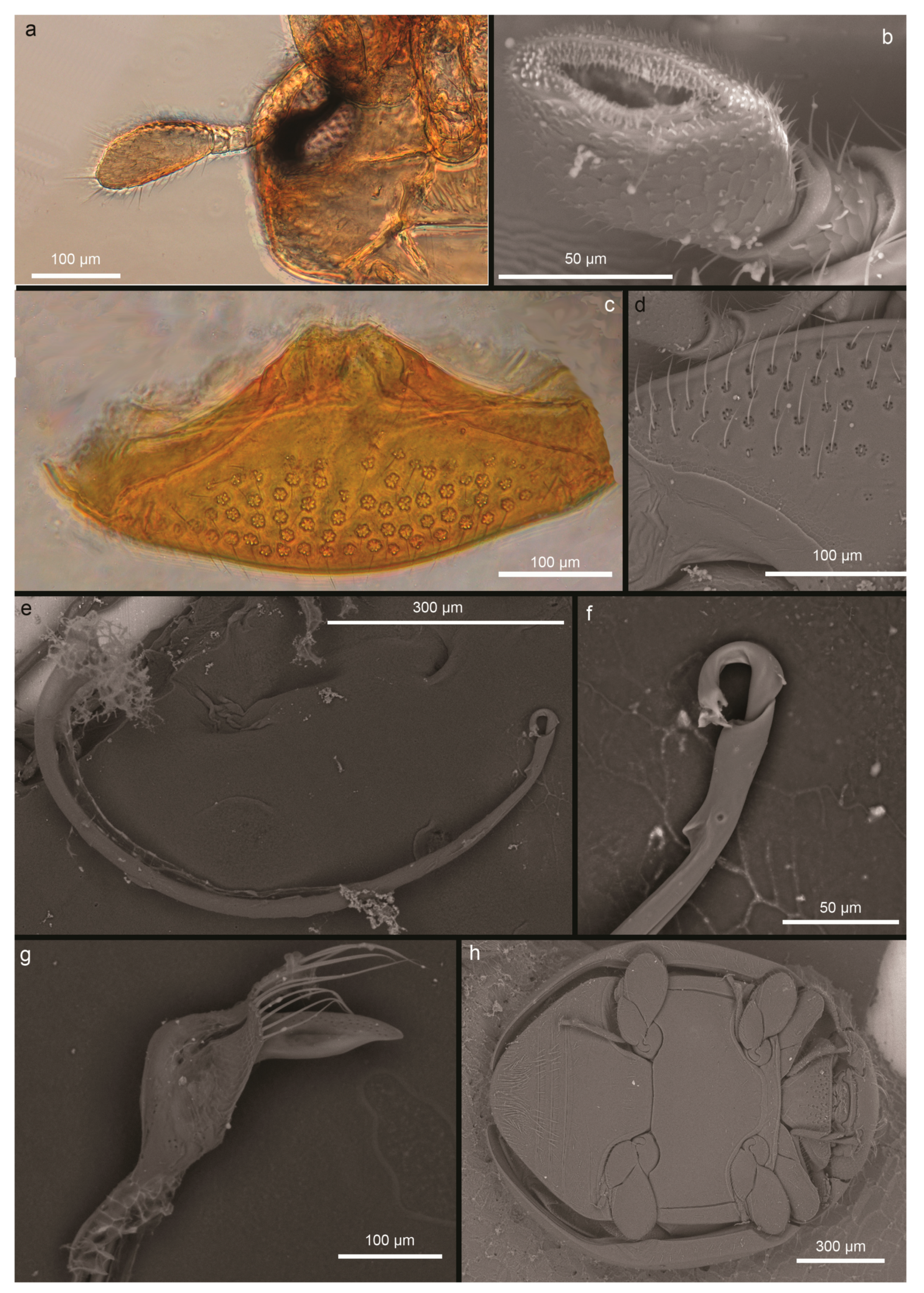
3.2. OSW Characterization
3.3. Natural Enemies
4. Discussion
- (1)
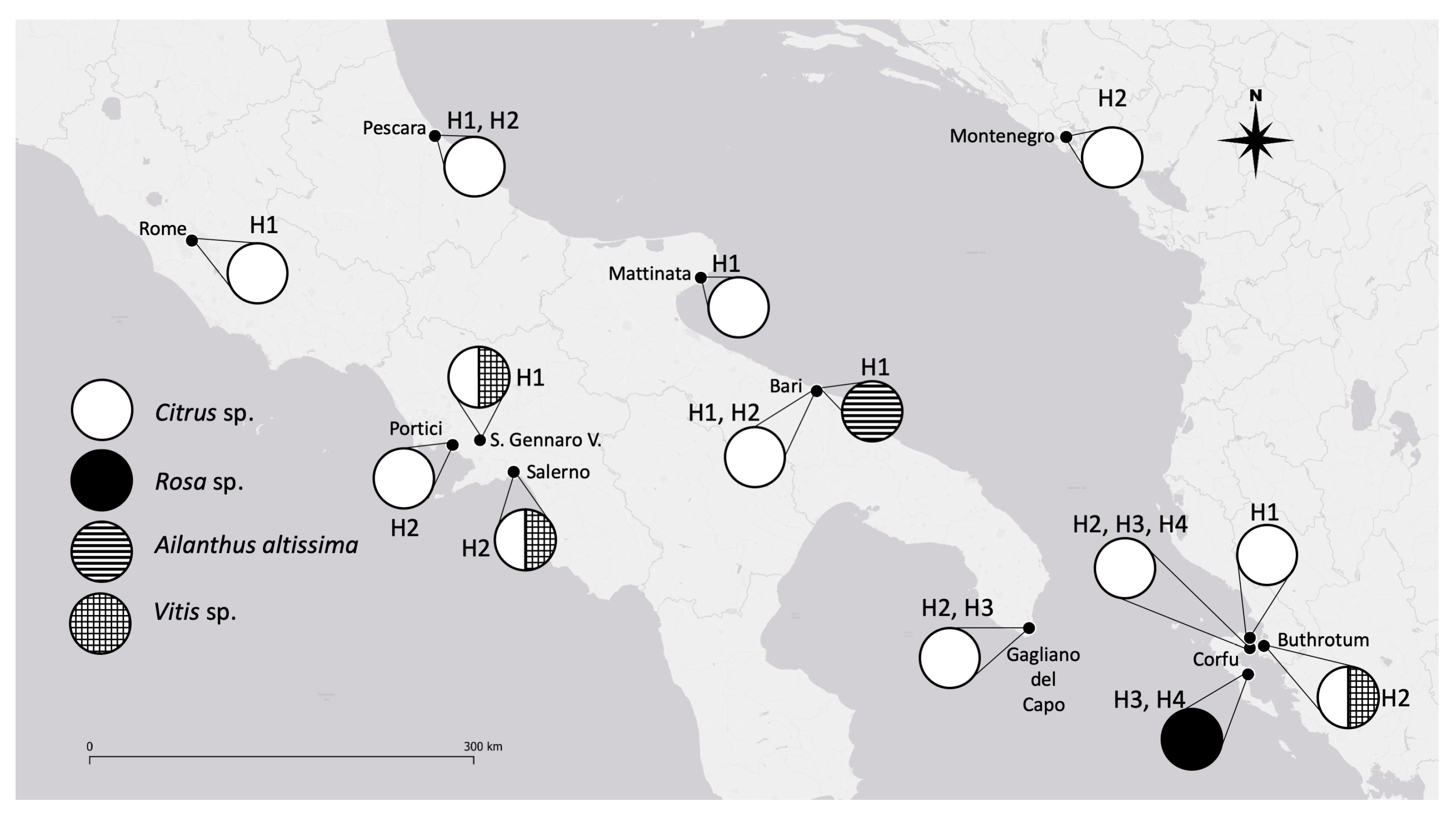
- The H4 haplotype seems to have a reduced diffusion (present only in Greece) [16] because it was not found in any other of the collection areas (Italy and Albania).
- (2)
- (3)
- The H1 haplotype was recovered in Apulia (both in Bari and in Mattinata) during our sampling but it was not found in the study performed by Kapantaidaki [16]. The different compositions of haplotypes in the samples collected by different authors could be linked both to different sampling methodologies and/or to a patch-like distribution of the different haplotypes.
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- EASIN—European Alien Species Information Network European Commission—Joint Research Centre 2019. Available online: https://easin.jrc.ec.europa.eu/ (accessed on 21 October 2019).
- Seebens, H.; Blackburn, T.M.; Dyer, E.E.; Genovesi, P.; Hulme, P.E.; Jeschke, J.M.; Essl, F. No saturation in the accumulation of alien species worldwide. Nat. Commun. 2017, 8, 14435. [Google Scholar] [CrossRef]
- Roques, A.; Auger-Rozenberg, A.M.; Blackburn, T.M.; Garnas, J.; Pyšek, P.; Rabitsch, W.; Richardson, D.M.; Wingfield, M.J.; Liebhold, A.M.; Duncan, R.P. Temporal and interspecific variation in rates of spread for insect species invading Europe during the last 200 years. Biol. Invasions 2016, 18, 907–920. [Google Scholar] [CrossRef]
- Bernardo, U.; van Nieukerken, E.J.; Sasso, R.; Gebiola, M.; Gualtieri, L.; Viggiani, G. Characterization, distribution, biology and impact on Italian walnut orchards of the invasive North-American leafminer Coptodisca lucifluella (Lepidoptera: Heliozelidae). Bull. Entomol. Res. 2015, 105, 210–224. [Google Scholar] [CrossRef]
- Garonna, A.P.; Scarpato, S.; Vicinanza, F.; Espinosa, B. First report of Toumeyella parvicornis (Cockerell) in Europe (Hemiptera: Coccidae). Zootaxa 2015, 3949, 142–146. [Google Scholar] [CrossRef]
- Lupi, D.; Bernardo, U.; Bonsignore, C.P.; Colombo, M.; Dindo, M.L.; Faccoli, M.; Ferracini, C.; Gualtieri, L.; Marullo, R.; Mazzon, L.; et al. Insects and globalization: Sustainable control of exotic species in Italian agro-forestry ecosystems. IOBC-WPRS Bull. 2014, 100, 87–90. [Google Scholar]
- Montecchio, L.; Faccoli, M. First record of thousand cankers disease Geosmithia morbida and walnut twig beetle Pityophthorus juglandis on Juglans nigra in Europe. Plant Dis. 2014, 98, 696. [Google Scholar] [CrossRef]
- Nugnes, F.; Russo, E.; Viggiani, G.; Bernardo, U. First record of an invasive fruit fly belonging to Bactrocera dorsalis complex (Diptera: Tephritidae) in Europe. Insects 2018, 9, 182. [Google Scholar] [CrossRef] [PubMed]
- Uesugi, R.; Sato, Y.; Han, B.Y.; Huang, Z.D.; Yara, K.; Furuhashi, K. Molecular evidence for multiple phylogenetic groups within two species of invasive spiny whiteflies and their parasitoid wasp. Bull. Entomol. Res. 2016, 10, 328–340. [Google Scholar] [CrossRef] [PubMed]
- Quaintance, A.L. New oriental Aleurodidae. Can. Entomol. 1903, 35, 61–64. [Google Scholar] [CrossRef]
- Van den Berg, M.A.; Greenland, J. Classical biological control of the spiny black fly, Aleurocanthus spiniferus (Hem.: Aleyrodidae), on citrus in Southern Africa. Entomophaga 1997, 42, 459–465. [Google Scholar] [CrossRef]
- Nguyen, R.; Sailer, R.I.; Hamon, A.B. Catalog of Aleyrodidae on Citrus and Their Natural Enemies (Homoptera—Aleyrodidae); Florida Department of Agriculture & Consumer Services, Division of Plant Industry, Bureau of Entomology: St. Gainesville, FL, USA; Centre for Agriculture and Bioscience International: Wallingford, UK, 1993; Volume 730.
- Muniappan, R.; Purea, M.; Sengebau, F.; Reddy, G.V.P. Orange spiny whitefly, Aleurocanthus spiniferus (Quaintance) (Homoptera: Aleyrodidae), and its parasitoids in the Republic of Palau. Proc. Hawaii. Entomol. Soc. 2006, 38, 21–25. [Google Scholar]
- Gillespie, P.S. A review of the whitefly genus Aleurocanthus Quaintance & Baker (Hemiptera: Aleyrodidae) in Australia. Zootaxa 2012, 3252, 1–42. [Google Scholar] [CrossRef]
- Cioffi, M.; Cornara, D.; Corrado, I.; Jansen, M.G.M.; Porcelli, F. The status of Aleurocanthus spiniferus from its unwanted introduction in Italy to date. Bull. Insectol. 2013, 66, 273–281. [Google Scholar]
- Kapantaidaki, D.E.; Antonatos, S.; Kontodimas, D.; Milonas, P.; Papachristos, D.P. Presence of the invasive whitefly Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Greece. OEPP/EPPO Bull. 2019, 49, 127–131. [Google Scholar] [CrossRef]
- Porcelli, F. First record of Aleurocanthus spiniferus (Homoptera: Aleyrodidae) in Apulia, Southern Italy. OEPP/EPPO Bull. 2008, 38, 516–518. [Google Scholar] [CrossRef]
- Nutricato, S.; D’onghia, A.M.; Fedele, A.; Filì, V.; Garonna, A.P.; Guario, A.; Guastamacchia, F.; Mazzeo, M.; Parisi, V.; Percoco, A.; et al. Nuovo aleirode da quarantena segnalato in Puglia. Inf. Agrar. 2009, 65, 57–59. [Google Scholar]
- El Kenawy, A.; Cornara, D.; Corrado, I.; El-Heneidy, A.; Rapisarda, C.; Porcelli, F. Aleurocanthus spiniferus (Quaintance) (Hemiptera Aleyrodidae) is spreading throughout the Italian region Apulia. In Proceedings of the 5th International Scientific Agricultural Symposium ‘Agrosym 2014’, Jahorina, Bosnia and Herzegovina, 23–26 October 2014. [Google Scholar] [CrossRef]
- EFSA—European Food Safety Authority. Scientific opinion on Pest categorisation of Aleurocanthus spp. EFSA J. 2018, 16, 5436. [Google Scholar]
- Šimala, M.; Masten, M.T. First record of the orange spiny whitefly, Aleurocanthus spiniferus Quaintance, 1903 (Hemiptera: Aleyrodidae), in Croatia. In Proceedings of the Conference ‘Zbornik Predavanj in Referatov, 11, Slovenskega Posvetovanja o Varstvu Rastlin Z Mednarodno Udelezbo, Bled, Slovenia, 5–6 March 2013; Available online: http://dvrs.bf.uni-lj.si/spvr/2013/60Simala.pdf (accessed on 3 August 2017).
- Radonjić, S.; Hrnčić, S.; Malumphy, C. First record of Aleurocanthus spiniferus (Quaintance) (Hemiptera Aleyrodidae) in Montenegro. Redia 2014, 77, 141–145. [Google Scholar]
- Jeffers, L. New Pest Advisory Group Report. Aleurocanthus Spiniferus Quaintance: Orange Spiny Whitefly, Hemiptera/Aleyrodidae. Available online: http://www.aphis.usda.gov/plant_health/cphst/npag/downloads/SampleReports/AleurocanthusspiniferusReport.pdf (accessed on 15 February 2018).
- Gyeltshen, J.; Hodges, A.; Hodges, G.S. Orange Spiny Whitefly, Aleurocanthus spiniferus (Quaintance) (Insecta: Hemiptera: Aleyrodidae). Available online: https://edis.ifas.ufl.edu/pdffiles/IN/IN61800.pdf (accessed on 12 August 2019).
- Zhang, Q.B. The reasons for rampage of citrus spiny white fly and its control. South China Fruits 2006, 2, 20–21. [Google Scholar]
- Clausen, C.P.; Bartlett, B.R.; Debach, P.; Goeden, R.D.; Legner, E.F.; Mcmurtry, J.A.; Oatman, E.R.; Bay, E.C.; Rosen, D. Introduced parasites and predators of arthropod pests and weeds: A world review. USDA Agric. Handb. 1978, 480, 1–545. [Google Scholar]
- Quezada, J.R. Biological control of Aleurocanthus woglumi (Homoptera: Aleyrodidae) in El Salvador. Entomophaga 1974, 19, 243–254. [Google Scholar] [CrossRef]
- Smith, H.D. Citrus blackfly on the west coast of Mexico and the importation and colonization of Eretmocerus serius Silv. for its control. Fitofolio 1945, 4, 67–103. [Google Scholar]
- Niu, J.Z.; Hull-Sanders, H.; Zhang, Y.X.; Lin, J.Z.; Dou, W.; Wang, J.J. Biological control of arthropod pests in citrus orchards in China. Biol. Control 2014, 68, 15–22. [Google Scholar] [CrossRef]
- Evans, G.A. Host Plant List of the Whiteflies (Aleyrodidae) of the World. 2007. Available online: http://entomofaune.qc.ca/entomofaune/aleurodes/references/Evans_2007_Hosts_whiteflies.pdf (accessed on 12 August 2019).
- Silvestri, F. Contribuzione alla conoscenza degli Aleurodidae (lnsecta: Hemiptera) viventi su Citrus in Estremo Oriente e dei loro parassiti. Bol. Lab. Zool. Gen. E Agric. R. Scuola Super. Agric. Portici 1928, 21, 1–60. [Google Scholar]
- Hodek, I.; Honěk, A. Ecology of Coccinellidae, 1st ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1966; p. 464. [Google Scholar]
- Mehrnejad, M.R.; Jalali, M.A. Life history parameters of the coccinellids beetle Oenopia conglobata contaminata, an important predator of the common pistachio psylla, Agonoscena pistaciae (Hemiptera: Psylloidea). Biocontrol Sci. Technol. 2004, 14, 701–711. [Google Scholar] [CrossRef]
- Han, B.Y.; Cui, L. The infection and prevalent of Aschersonia aleyrodis and Aegerita webberi against Aleurocanthus spiniferus. China Tea 2004, 2, 19. [Google Scholar]
- Zhang, Q.B.; Lei, H.D.; Li, H.J.; Liu, H.Q.; Yao, T.S.; Tian, W.H.; Qian, K.M. The damage and control of Aleurocanthus spiniferus. South China Fruits 2004, 33, 15. [Google Scholar]
- Guo, L.; Qiu, B.L.; Ren, S.X. Summary on the classify of natural enemy germplasm recourses of Aleurocanthus spiniferus (Homoptera: Aleyrodidae). Guangdong Agric. Sci. 2006, 2, 9–10. [Google Scholar]
- Campanian Executive Decree (Regione Campania) n. 2 June 22nd 2017—Misure Fitosanitarie Regionali Per il Controllo del Fitofago Aleurocanthus spiniferus. 2017. Available online: http://burc.regione.campania.it/eBurcWeb/publicContent/archivio/archivio.iface (accessed on 20 March 2019).
- Gebiola, M.; Bernardo, U.; Monti, M.M.; Navone, P.; Viggiani, G. Pnigalio agraules (Walker) and Pnigalio mediterraneus Ferrière and Delucchi (Hymenoptera: Eulophidae): Two closely related valid species. J. Nat. Hist. 2009, 43, 2465–2480. [Google Scholar] [CrossRef]
- Bink-Moenen, R.M. Revision of the African whiteflies (Aleyrodidae). Monogr. Ned. Entomol. Ver. 1983, 10, 1–211. [Google Scholar]
- Martin, J.H. An identification guide to the common whitefly pest species of the world (Homoptera, Aleyrodidae). Trop. Pest Manag. 1987, 33, 298–322. [Google Scholar] [CrossRef]
- Martin, J.H. The whitefly fauna of Australia (Sternorrhyncha: Aleyrodidae), a taxonomic account and identification guide. CSIRO Entomol. Tech. Pap. 1999, 38, 1–197. [Google Scholar]
- Kanmiya, K.; Ueda, S.; Kasai, A.; Yamashita, K.; Sato, Y.; Yoshiyasu, Y. Proposal of new specific status for tea-infesting populations of the nominal citrus spiny whitefly Aleurocanthus spiniferus (Homoptera: Aleyrodidae). Zootaxa 2011, 2797, 25–44. [Google Scholar] [CrossRef]
- Jansen, M.; Porcelli, F. Aleurocanthus camelliae (Hemiptera: Aleyrodidae), a species possibly new for the European fauna of a genus in great need of revision. Tijdschr. Entomol. 2018, 161, 63–78. [Google Scholar] [CrossRef]
- Uesugi, R.; Sato, Y. Differentiation of the tea-infesting population of citrus spiny whitefly Aleurocanthus spiniferus (Homoptera: Aleyrodidae) from the citrus-infesting population in Japan on the basis of differences in the mitochondrial cytochrome c oxidase subunit I gene. Jpn. J. Appl. Entomol. Zool. 2011, 55, 155–161. [Google Scholar] [CrossRef][Green Version]
- Simon, C.; Frati, F.; Beckenbach, A.T.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Clement, M.; Posada, D.; Crandall, K.A. TCS: A computer program to estimate gene genealogies. Mol. Ecol. 2001, 9, 1657–1659. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef]
- Posada, D. Jmodeltest: Phylogenetic model averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Dlugosch, K.M.; Parker, M. Founding events in species invasions: Genetic variation, adaptive evolution, and the role of multiple introductions. Mol. Ecol. 2008, 17, 431–449. [Google Scholar] [CrossRef] [PubMed]
- Gillespie, R.G.; Roderick, G.K. Evolution: Geology and climate drive diversification. Nature 2014, 509, 297–298. [Google Scholar] [CrossRef] [PubMed]
- Nugnes, F.; Gebiola, M.; Monti, M.M.; Gualtieri, L.; Giorgini, M.; Wang, J.; Bernardo, U. Genetic diversity of the invasive gall wasp Leptocybe invasa (Hymenoptera: Eulophidae) and of its Rickettsia endosymbiont, and associated sex-ratio differences. PLoS ONE 2015. [Google Scholar] [CrossRef] [PubMed]
- Dittrich-Schröder, G.; Hoareau, T.B.; Hurley, B.P.; Wingfield, M.J.; Lawson, S.; Nahrung, H.F.; Slippers, B. Population genetic analyses of complex global insect invasions in managed landscapes: A Leptocybe invasa (Hymenoptera) case study. Biol. Invasions 2018, 20, 2395–2420. [Google Scholar] [CrossRef]
- Mound, L.A.; Halsey, S.H. Whitefly of the World. A Systematic Catalogue of the Aleyrodidae (Homoptera) with Host Plant and Natural Enemy Data, 1st ed.; British Museum (Natural History): London, UK, 1978; p. 340. [Google Scholar]
- Evans, G. The Whiteflies (Hemiptera: Aleyrodidae) of the World and Their Host Plants and Natural Enemies. 2007. Available online: http://keys.lucidcentral.org/keys/v3/whitefly/PDF_PwP%20ETC/world-whitefly-catalog-Evans.pdf (accessed on 12 August 2019).
- Gebiola, M.; Gómez-Zurita, J.; Monti, M.M.; Navone, P.; Bernardo, U. Integration of molecular, ecological, morphological and endosymbiont data for species delimitation within the Pnigalio soemius complex (Hymenoptera: Eulophidae). Mol. Ecol. 2012, 21, 1190–1208. [Google Scholar] [CrossRef]
- Nugnes, F.; Bernardo, U.; Viggiani, G. An integrative approach to species discrimination in the Anagrus atomus group sensu stricto (Hymenoptera: Mymaridae), with a description of a new species. Syst. Biodivers. 2017, 15, 582–599. [Google Scholar] [CrossRef]
- Bernardo, U.; Nugnes, F.; Gualtieri, L.; Nicoletti, R.; Varricchio, P.; Sasso, R.; Viggiani, G. A new gall midge species of Asphondylia (Diptera: Cecidomyiidae) inducing flower galls on Clinopodium nepeta (Lamiaceae) from Europe, its phenology, and associated fungi. Environ. Entomol. 2018, 47, 609–622. [Google Scholar] [CrossRef]
- Lawson, S.; Griffiths, M.; Nahrung, H.; Noack, A.; Wingfield, M.; Wilcken, C.; Slippers, B.; Lo, N.; Thu, P.Q.; Lee, S.; et al. Biological Control of Eucalypt Pests Overseas and in Australia; Report Australian Centre for International Agricultural Research (ACIAR); ACIAR: Canberra, Australia, 2012; pp. 1–40. Available online: http://era.daf.qld.gov.au/id/eprint/2743/1/ACIAR_Final%20Report2012_26_biological_control_of_eucalypt_pests_Lawson.pdf (accessed on 12 November 2018).
- Muirhead, J.R.; Gray, D.K.; Kelly, D.W.; Ellis, S.M.; Heath, D.D.; Macisaac, H.J. Identifying the source of species invasions: Sampling intensity vs genetic diversity. Mol. Ecol. 2008, 17, 1020–1035. [Google Scholar] [CrossRef]
- Gordon, R.D. South American Coccinellidae (Coleoptera) part III: Taxonomic revision of the Western Hemisphere genus Delphastus Casey. Frustula Entomol. 1994, 17, 71–133. [Google Scholar]
- Booth, R.G.; Polaszek, A. The identities of ladybird beetle predators used for whitefly control, with notes on some whitefly parasitoids, in Europe. In Proceedings of the Brighton Crop Protection Conference—Pests and Diseases, Brighton, UK, 18–21 November 1996; pp. 69–74. [Google Scholar]
- Hoelmer, K.A.; Osborne, L.S.; Yokomi, R.K. Reproduction and feeding behavior of Delphastus catalinae (Coleoptera: Coccinellidae), a predator of Bemisia tabaci (Homoptera: Aleyrodidae). J. Econ. Entomol. 1993, 86, 322–329. [Google Scholar] [CrossRef]
- Hoelmer, K.A.; Osborne, L.S.; Yokomi, R.K. Interactions of the whitefly predator Delphastus catalinae (Coleoptera: Coccinellidae) with parasitized sweetpotato whitefly (Homoptera: Aleyrodidae). Environ. Entomol. 1994, 23, 136–139. [Google Scholar] [CrossRef]
- Pickett, C.; Casanave, K.; Schoenig, S.; Heinz, K. Rearing Delphastus catalinae (Coleoptera: Coccinellidae): Practical experience and a modelling analysis. Can. Entomol. 1999, 131, 115–129. [Google Scholar] [CrossRef]
- Heinz, K.M.; Parrella, M.P. Biological control of Bemisia argentifolii (Gennadius) (Homoptera: Aleyrodidae) infesting Euphorbia pulcherrima: Evaluations of releases of Encarsia luteola (Hymenoptera: Aphelinidae) and Delphastus pusillus (Coleoptera: Coccinellidae). Environ. Entomol. 1994, 23, 1346–1353. [Google Scholar] [CrossRef][Green Version]
- Heinz, K.M.; Brazzle, J.R.; Pickett, C.; Natwick, E.T.; Nelson, J.M.; Parrella, M.P. Predatory beetle may suppress silverleaf whitefly. Calif. Agric. 1994, 48, 35–40. [Google Scholar]
- Hoelmer, K.A.; Pickett, C.H. Geographic Origin and Taxonomic History of Delphastus spp. (Coleoptera: Coccinellidae) in Commercial Culture. Biocontrol Sci. Technol. 2003, 13, 529–535. [Google Scholar] [CrossRef]
- De Almeida, R.A.; Stouthamer, R. Molecular identification of Trichogramma cacoeciae Marchal (Hymenoptera: Trichogrammatidae): A new record for Peru. Neotrop. Entomol. 2003, 32, 269–272. [Google Scholar] [CrossRef]
| Specimen Code | Location | Coordinates | Host Plant | Date of Record | Haplotype | Accession Number |
|---|---|---|---|---|---|---|
| ASPI PE1 | Pescara | 42°27′ N 14°12′ E | Citrus sp. | 16 September 2019 | H1 | MN662884 |
| ASPI PE2 | H1 | MN662885 | ||||
| ASPI PE3 | H1 | MN662886 | ||||
| ASPI PE4 | H1 | MN662887 | ||||
| ASPI PE5 | H2 | MN662925 | ||||
| ASPI R1 | Rome | 41°54′ N 12°29′ E | Citrus sp. | 7 March 2019 | H1 | MN662888 |
| ASPI R2 | H1 | MN662889 | ||||
| ASPI M1 | Mattinata (FG) | 41°42′ N 16°04′ E | Citrus sp. | 16 August 2018 | H1 | MN662917 |
| ASPI M2 | H1 | MN662918 | ||||
| ASPI M3 | H1 | MN662919 | ||||
| ASPI BA1 | Bari | 41°06′ N 16°53′ E | Ailanthus altissima | 7 August 2017 | H1 | MN662912 |
| ASPI BA2 | H1 | MN662913 | ||||
| ASPI BA3 | H1 | MN662914 | ||||
| ASPI BA5 | H1 | MN662915 | ||||
| ASPI BC1 | 41°06′ N 16°52′ E | Citrus sp. | H1 | MN662916 | ||
| ASPI BC2 | H2 | MN662890 | ||||
| ASPI BC3 | H2 | MN662891 | ||||
| ASPI BC4 | H2 | MN662892 | ||||
| ASPI SG1 | San Gennaro Vesuviano (NA) | 40°51′ N 14°31′ E | Citrus nobilis | 13 September 2019 | H1 | MN662893 |
| ASPI SG2 | H1 | MN662894 | ||||
| ASPI SG3 | H1 | MN662895 | ||||
| ASPI SG4 | H1 | MN662896 | ||||
| ASPI SG5 | H1 | MN662897 | ||||
| ASPI SG6 | Vitis sp. | H1 | MN662898 | |||
| ASPI SG7 | H1 | MN662899 | ||||
| ASPI SG8 | H1 | MN662900 | ||||
| ASPI SG9 | H1 | MN662901 | ||||
| ASPI SG10 | H1 | MN662902 | ||||
| ASPI P1 | Portici (NA) | 40°49′ N 14°19′ E | Citrus limon | 14 February 2019 | H2 | MN662920 |
| ASPI P2 | H2 | MN662921 | ||||
| ASPI S1 | Salerno | 40°40′ N 14°45′ E | Citrus sinensis | 16 June 2018 | H2 | MN662922 |
| ASPI S2 | H2 | MN662903 | ||||
| ASPI S3 | H2 | MN662904 | ||||
| ASPI S4 | Citrus reticulata | 16 May 2019 | H2 | MN662905 | ||
| ASPI S5 | H2 | MN662906 | ||||
| ASPI S6 | Vitis sp. | H2 | MN662907 | |||
| ASPI S7 | H2 | MN662908 | ||||
| ASPI B1 | Buthrotum (Albania) | 39°44′ N 20°01′ E | Citrus sp. | 20 July 2018 | H2 | MN662923 |
| ASPI B2 | H2 | MN662924 | ||||
| ASPI B3 | H2 | MN662909 | ||||
| ASPI B4 | Vitis sp. | 21 July 2018 | H2 | MN662910 | ||
| ASPI B5 | H2 | MN662911 |
| Specimen Code | Preliminary Identification | Origin or Commercial Product | Date of Record | Host-Plant/Host | Molecular Identification | Morphological Re-Examination | Accession Number |
|---|---|---|---|---|---|---|---|
| DC1 | Delphastus catalinae | Salerno a | 21 October 2017 | Citrus limon/Aleurocanthus spiniferus | D. catalinae | D. catalinae | MN662936 |
| DC2 | MN662937 | ||||||
| DC3 | MN662938 | ||||||
| DC4 | MN662939 | ||||||
| DC-C1 | D. catalinae | Delphibug b | 22 August 2018 | MN662940 | |||
| DC-C2 | MN662941 | ||||||
| DP1 | D. pusillus | Delphastus-System c | 12 August 2018 | MN662942 | |||
| DP2 | MN662943 | ||||||
| DP NO1 | D. catalinae | Noto (Sicily) a | 27 August 2018 | C. limon/Aleurothrixus floccosus | MN662944 |
| Host Plant Family | Host Plant Species |
|---|---|
| Simaroubaceae | Ailanthus altissima (Mill.) Swingle |
| Ericaceae | Arbutus unedo L. |
| Rutaceae | Citrus medica L. |
| Citrus reticulata Blanco | |
| Ranunculaceae | Clematis vitalba L. |
| Anacardiaceae | Pistacia vera L. |
| Rosaceae | Prunus avium (L.) |
| P. cerasus L. | |
| P. domestica L. | |
| Rosa banksiae Aiton | |
| R. × damascena Herrm. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nugnes, F.; Laudonia, S.; Jesu, G.; Jansen, M.G.M.; Bernardo, U.; Porcelli, F. Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Some European Countries: Diffusion, Hosts, Molecular Characterization, and Natural Enemies. Insects 2020, 11, 42. https://doi.org/10.3390/insects11010042
Nugnes F, Laudonia S, Jesu G, Jansen MGM, Bernardo U, Porcelli F. Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Some European Countries: Diffusion, Hosts, Molecular Characterization, and Natural Enemies. Insects. 2020; 11(1):42. https://doi.org/10.3390/insects11010042
Chicago/Turabian StyleNugnes, Francesco, Stefania Laudonia, Giovanni Jesu, Maurice Gerardus Maria Jansen, Umberto Bernardo, and Francesco Porcelli. 2020. "Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Some European Countries: Diffusion, Hosts, Molecular Characterization, and Natural Enemies" Insects 11, no. 1: 42. https://doi.org/10.3390/insects11010042
APA StyleNugnes, F., Laudonia, S., Jesu, G., Jansen, M. G. M., Bernardo, U., & Porcelli, F. (2020). Aleurocanthus spiniferus (Hemiptera: Aleyrodidae) in Some European Countries: Diffusion, Hosts, Molecular Characterization, and Natural Enemies. Insects, 11(1), 42. https://doi.org/10.3390/insects11010042








