A Subset of Odorant Receptors from the Desert Locust Schistocerca gregaria Is Co-Expressed with the Sensory Neuron Membrane Protein 1
Abstract
:1. Introduction
2. Materials and Methods
2.1. Phylogenetic Analyses
2.2. Animals and Preparation of Antennae
2.3. Reverse Transcription PCR Amplification for Generating Riboprobes
2.4. Synthesis of Riboprobes for In Situ Hybridization
2.5. Fluorescence In Situ Hybridization (FISH)
2.6. RNA Extraction and Reverse Transcription PCR for Comparing SgreOR Expression in Males Versus Females
3. Results
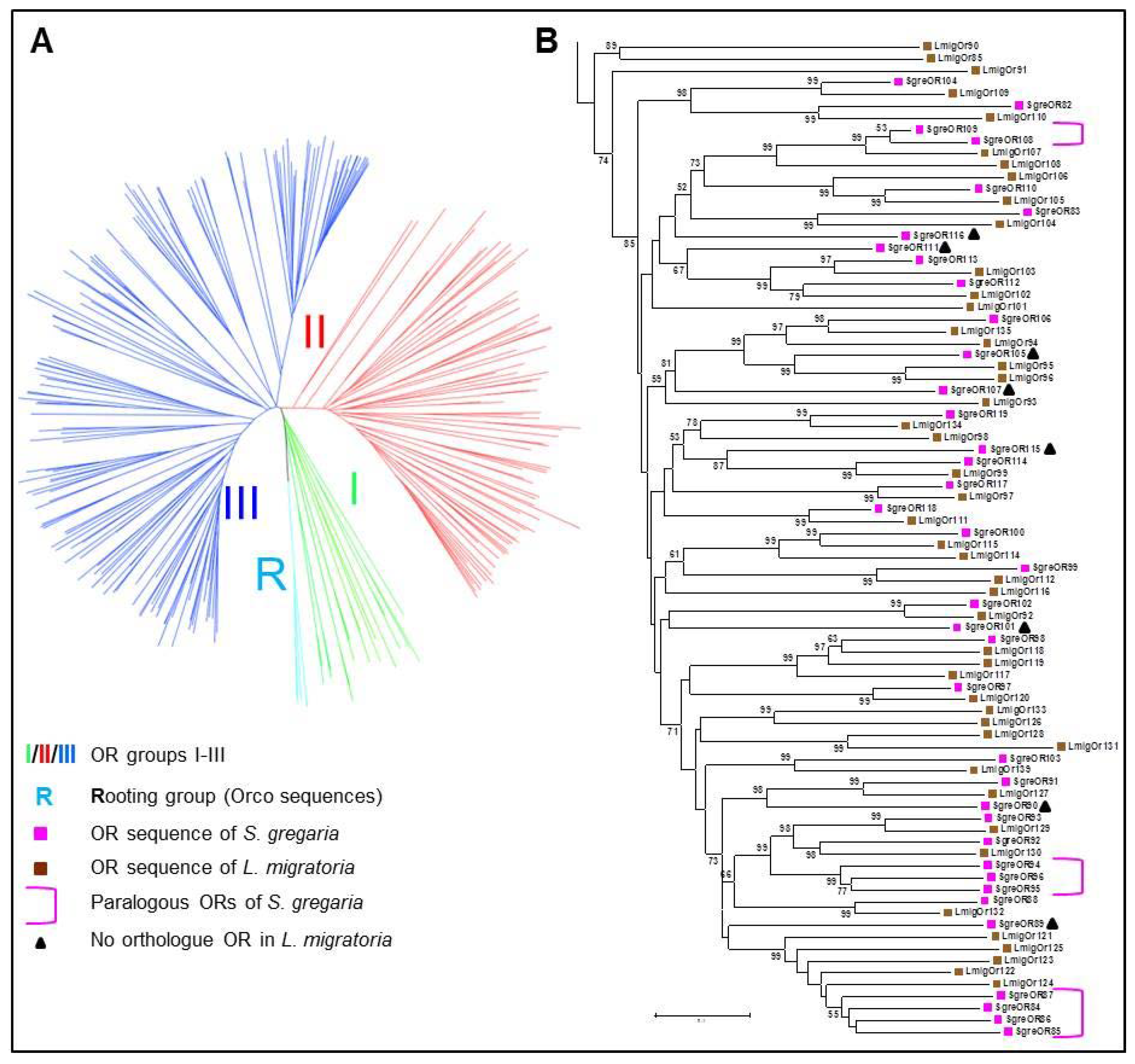
3.1. Categorization of SgreORs Based on Phylogenetic Analyses
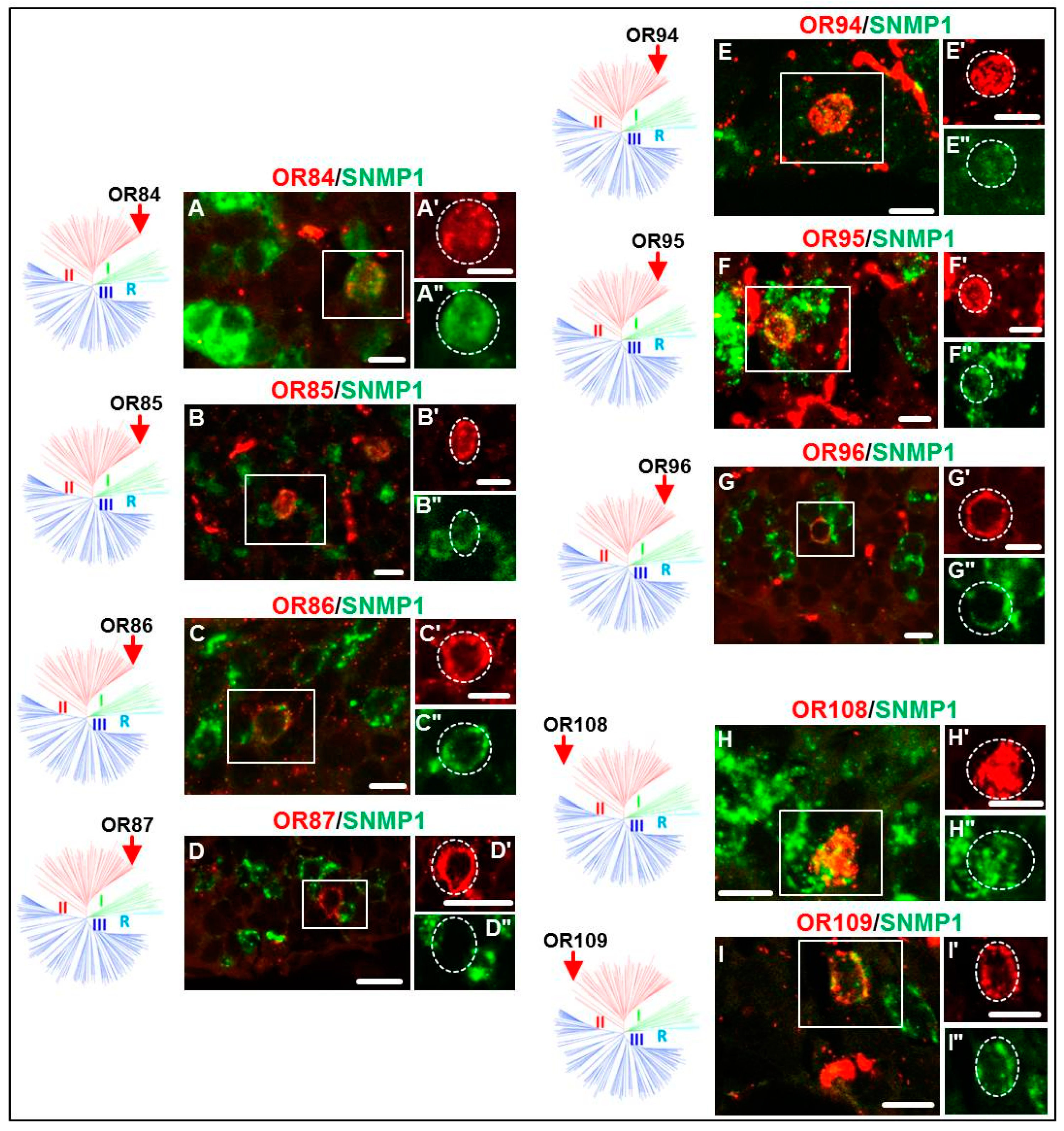
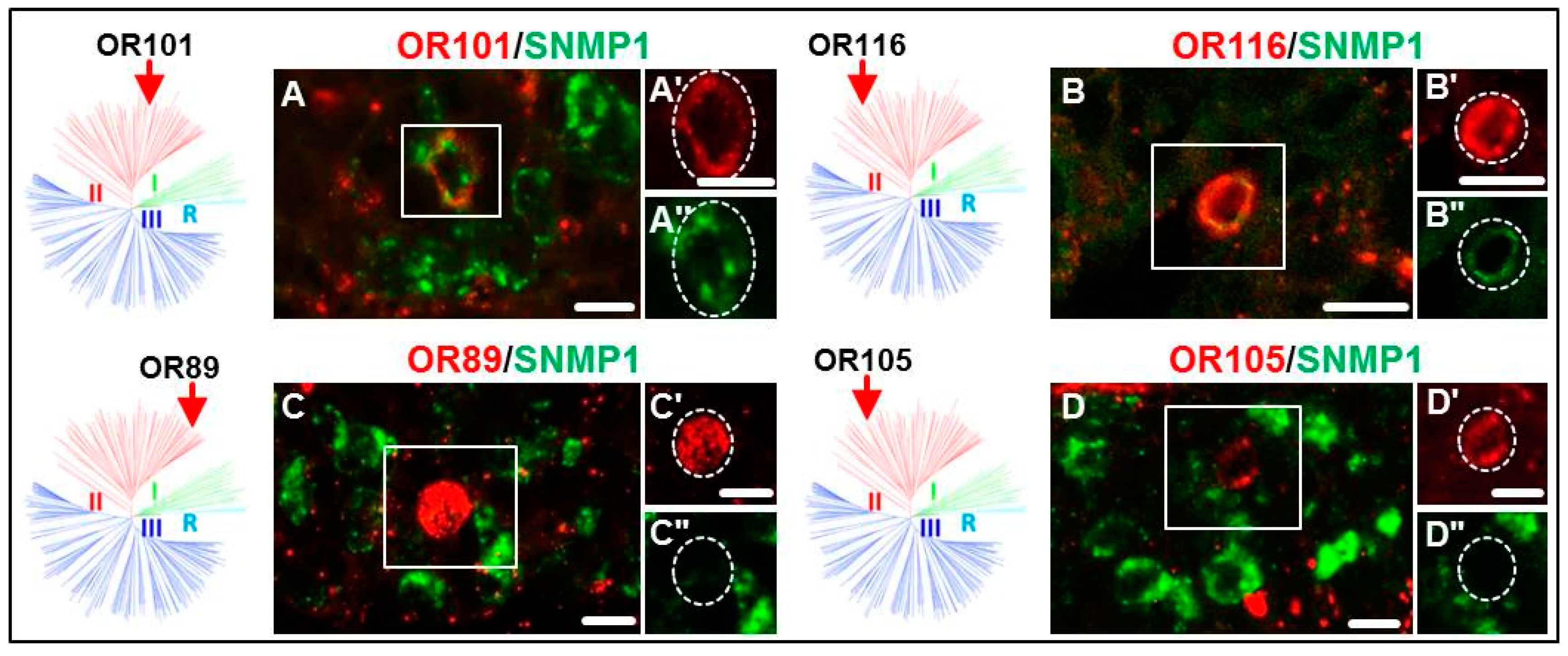
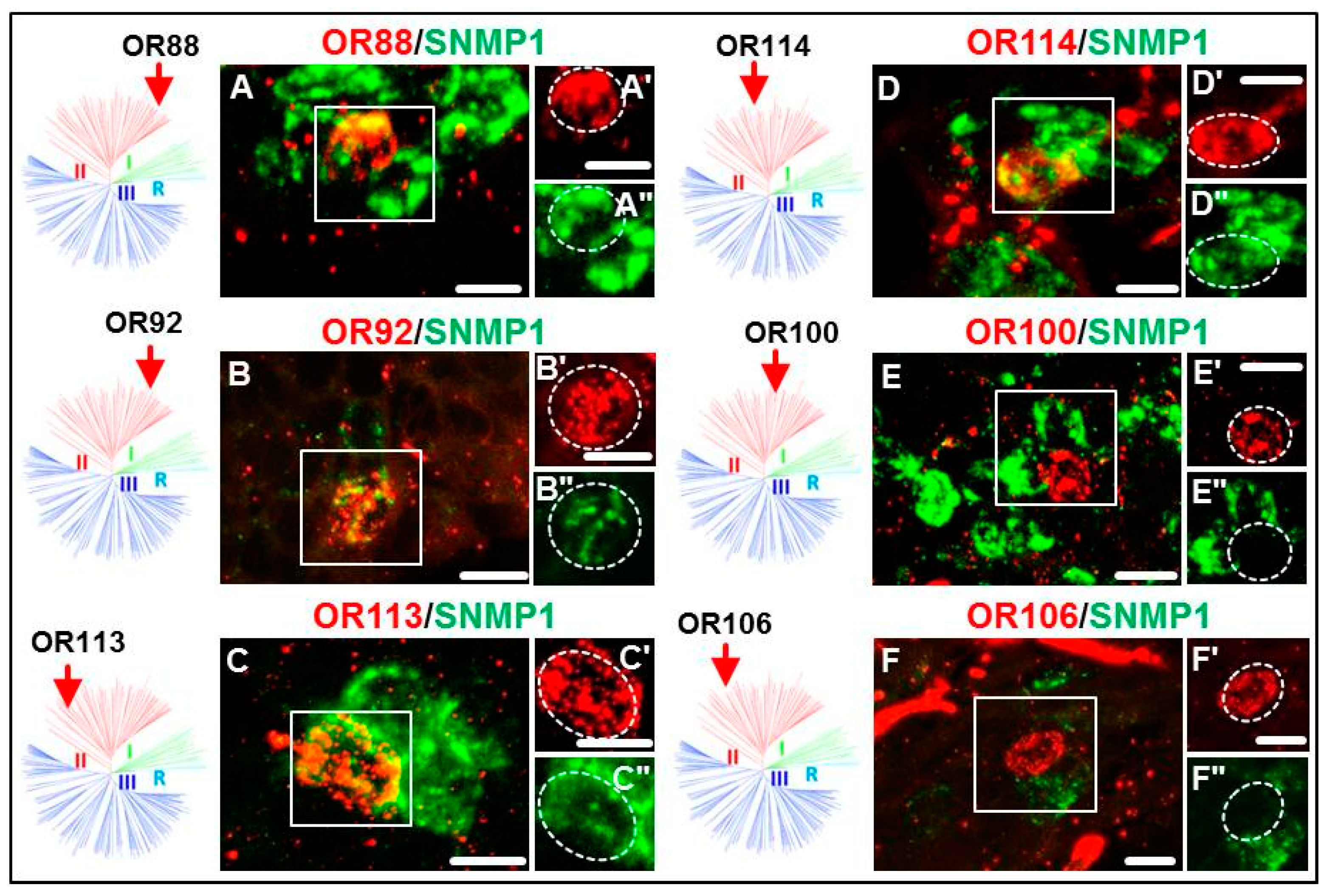
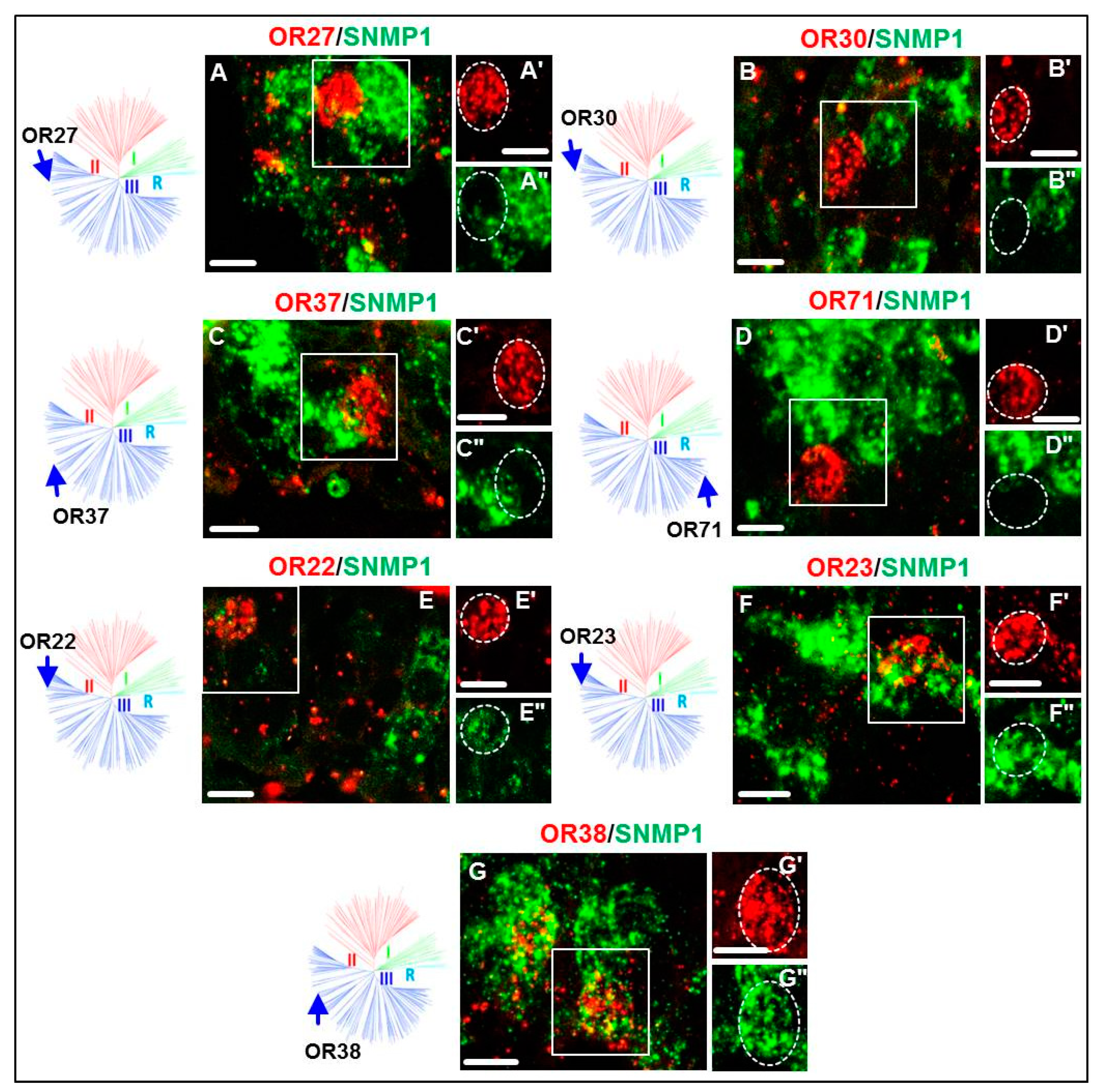
3.2. Expression of SgreORs and SNMP1
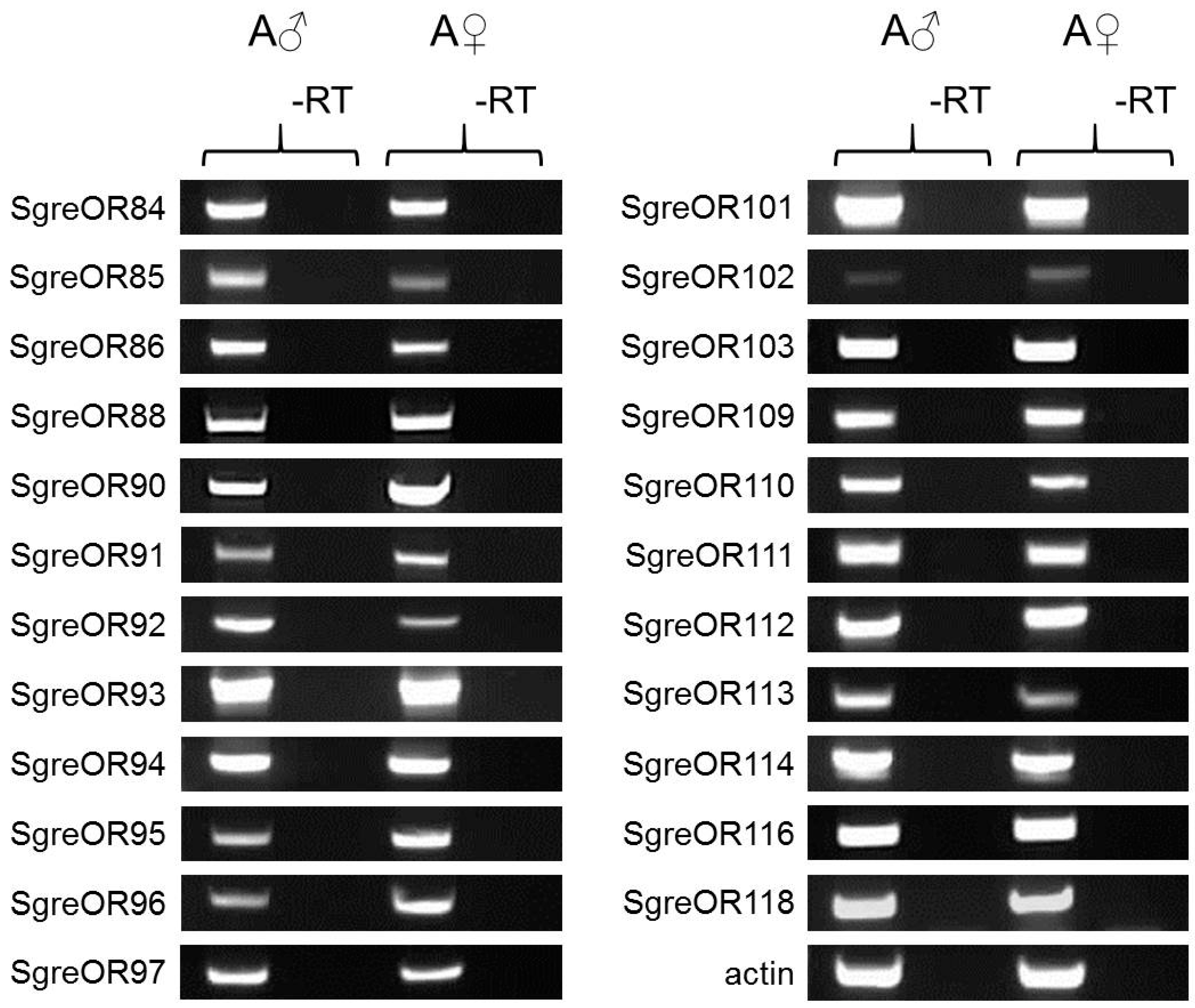
3.3. Expression of Group II ORs Co-Expressed with SNMP1 in Males and Females
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hansson, B.S.; Stensmyr, M.C. Evolution of insect olfaction. Neuron 2011, 72, 698–711. [Google Scholar] [CrossRef] [PubMed]
- Hassanali, A.; Njagi, P.G.N.; Bashir, M.O. Chemical Ecology of Locusts and Related Acridids. Annu. Rev. Entomol. 2005, 50, 223–245. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, J.; Pregitzer, P.; Breer, H.; Krieger, J. Access to the odor world: Olfactory receptors and their role for signal transduction in insects. Cell. Mol. Life Sci. 2018, 75, 485–508. [Google Scholar] [CrossRef] [PubMed]
- Benton, R.; Vannice, K.S.; Vosshall, L.B. An essential role for a CD36-related receptor in pheromone detection in Drosophila. Nature 2007, 450, 289–293. [Google Scholar] [CrossRef]
- Forstner, M.; Gohl, T.; Gondesen, I.; Raming, K.; Breer, H.; Krieger, J. Differential expression of SNMP-1 and SNMP-2 proteins in pheromone-sensitive hairs of moths. Chem. Senses 2008, 33, 291–299. [Google Scholar] [CrossRef]
- Krieger, J.; Klink, O.; Mohl, C.; Raming, K.; Breer, H. A candidate olfactory receptor subtype highly conserved across different insect orders. J. Comp. Physiol. A 2003, 189, 519–526. [Google Scholar] [CrossRef]
- Krieger, J.; Grosse-Wilde, E.; Gohl, T.; Dewer, Y.M.E.; Raming, K.; Breer, H. Genes encoding candidate pheromone receptors in a moth (Heliothis virescens). Proc. Natl. Acad. Sci. USA 2004, 101, 11845–11850. [Google Scholar] [CrossRef]
- Larsson, M.C.; Domingos, A.I.; Jones, W.D.; Chiappe, M.E.; Amrein, H.; Vosshall, L.B. Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 2004, 43, 703–714. [Google Scholar] [CrossRef]
- Sakurai, T.; Nakagawa, T.; Mitsuno, H.; Mori, H.; Endo, Y.; Tanoue, S.; Yasukochi, Y.; Touhara, K.; Nishioka, T. Identification and functional characterization of a sex pheromone receptor in the silkmoth Bombyx mori. Proc. Natl. Acad. Sci. USA 2004, 101, 16653–16658. [Google Scholar] [CrossRef]
- Rogers, M.E.; Krieger, J.; Vogt, R.G. Antennal SNMPS (sensory neuron membrane proteins) of lepidoptera define a unique family of invertebrate CD36-like proteins. J. Neurobiol. 2001, 49, 47–61. [Google Scholar] [CrossRef]
- Rogers, M.E.; Steinbrecht, R.A.; Vogt, R.G. Expression of SNMP-1 in olfactory neurons and sensilla of male and female antennae of the silkmoth Antheraea polyphemus. Cell Tissue Res. 2001, 303, 433–446. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.E.; Sun, M.; Lerner, M.R.; Vogt, R.G. Snmp-1, a novel membrane protein of olfactory neurons of the silk moth Antheraea polyphemus with homology to the CD36 family of membrane proteins. J. Biol. Chem. 1997, 272, 14792–14799. [Google Scholar] [CrossRef] [PubMed]
- German, P.F.; van der Poel, S.; Carraher, C.; Kralicek, A.V.; Newcomb, R.D. Insights into subunit interactions within the insect olfactory receptor complex using FRET. Insect Biochem. Mol. Biol. 2013, 43, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Pregitzer, P.; Greschista, M.; Breer, H.; Krieger, J. The sensory neurone membrane protein SNMP1 contributes to the sensitivity of a pheromone detection system. Insect Mol. Biol. 2014, 23, 733–742. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Diaz, C.; Bargeton, B.; Abuin, L.; Bukar, N.; Reina, J.H.; Bartoi, T.; Graf, M.; Ong, H.; Ulbrich, M.H.; Masson, J.F.; et al. A CD36 ectodomain mediates insect pheromone detection via a putative tunnelling mechanism. Nat. Commun. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Jin, X.; Ha, T.S.; Smith, D.P. SNMP is a signaling component required for pheromone sensitivity in Drosophila. Proc. Natl. Acad. Sci. USA 2008, 105, 10996–11001. [Google Scholar] [CrossRef] [PubMed]
- Showler, A.T. Locust 1 (Orthoptera: Acrididae) Outbreak in Africa and Asia, 1992–1994: An Overview. Am. Entomol. 1995, 41, 179–185. [Google Scholar] [CrossRef]
- Van Huis, A. Strategies to control the desert locust Schistocerca gregaria. In Area-Wide Control of Insect Pests; Vreysen, M.J.B., Robinson, A.S., Hendrichs, J., Eds.; Springer: Dordrecht, The Netherland, 2007; pp. 285–296. [Google Scholar]
- Simpson, S.J.; Sword, G.A. Locusts. Curr. Biol. 2008, 18, R364–R366. [Google Scholar] [CrossRef] [Green Version]
- Cui, X.; Wu, C.; Zhang, L. Electrophysiological response patterns of 16 olfactory neurons from the trichoid sensilla to odorant from fecal volatiles in the locust, Locusta migratoria manilensis. Arch. Insect Biochem. Physiol. 2011, 77, 45–57. [Google Scholar] [CrossRef]
- Hansson, B.; Ochieng’, S.; Grosmaitre, X.; Anton, S.; Njagi, P. Physiological responses and central nervous projections of antennal olfactory receptor neurons in the adult desert locust, Schistocerca gregaria (Orthoptera: Acrididae). J. Comp. Physiol. A 1996, 179, 157–167. [Google Scholar] [CrossRef]
- Ochieng’, S.A.; Hansson, B.S. Responses of olfactory receptor neurones to behaviourally important odours in gregarious and solitarious desert locust, Schistocerca gregaria. Physiol. Entomol. 1999, 24, 28–36. [Google Scholar] [CrossRef]
- Seidelmann, K.; Ferenz, H.J. Courtship inhibition pheromone in desert locusts, Schistocerca gregaria. J. Insect Physiol. 2002, 48, 991–996. [Google Scholar] [CrossRef]
- Pregitzer, P.; Jiang, X.; Grosse-Wilde, E.; Breer, H.; Krieger, J.; Fleischer, J. In search for pheromone receptors: Certain members of the odorant receptor family in the desert locust Schistocerca gregaria (orthoptera: Acrididae) are co-expressed with SNMP1. Int. J. Biol. Sci. 2017, 13, 911–922. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, J.; Krieger, J. Insect Pheromone Receptors—Key elements in sensing intraspecific chemical signals. Front. Cell. Neurosci. 2018, 12, 425. [Google Scholar] [CrossRef]
- Li, Z.; Ni, J.D.; Huang, J.; Montell, C. Requirement for Drosophila SNMP1 for rapid activation and termination of pheromone-induced activity. PLoS Genet. 2014, 10, e1004600. [Google Scholar] [CrossRef]
- Seidelmann, K.; Luber, K.; Ferenz, H.-J. Analysis of release and role of benzyl cyanide in male desert locusts, Schistocerca gregaria. J. Chem. Ecol. 2000, 26, 1897–1910. [Google Scholar] [CrossRef]
- Jiang, X.; Pregitzer, P.; Grosse-Wilde, E.; Breer, H.; Krieger, J. Identification and characterization of two “sensory neuron membrane proteins” (SNMPs) of the desert locust, Schistocerca gregaria (Orthoptera: Acrididae). J. Insect Sci. 2016, 16, 33. [Google Scholar] [CrossRef]
- Jiang, X.; Krieger, J.; Breer, H.; Pregitzer, P. Distinct subfamilies of odorant binding proteins in locust (Orthoptera, Acrididae): Molecular evolution, structural variation, and sensilla-specific expression. Front. Physiol. 2017, 8, 734. [Google Scholar] [CrossRef]
- Jiang, X.; Ryl, M.; Krieger, J.; Breer, H.; Pregitzer, P. Odorant binding proteins of the desert locust Schistocerca gregaria (Orthoptera, Acrididae): Topographic expression patterns in the antennae. Front. Physiol. 2018, 9, 417. [Google Scholar] [CrossRef]
- Wang, Z.; Yang, P.; Chen, D.; Jiang, F.; Li, Y.; Wang, X.; Kang, L. Identification and functional analysis of olfactory receptor family reveal unusual characteristics of the olfactory system in the migratory locust. Cell. Mol. Life Sci. 2015, 72, 4429–4443. [Google Scholar] [CrossRef] [Green Version]
- Jiang, X.; Breer, H.; Pregitzer, P. Sensilla-specific expression of odorant receptors in the desert locust Schistocerca gregaria. Front. Physiol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Lemke, R.S.; Pregitzer, P.; Eichhorn, A.S.; Breer, H.; Krieger, J.; Fleischer, J. SNMP1 and odorant receptors are co-expressed in olfactory neurons of the labial and maxillary palps from the desert locust Schistocerca gregaria (Orthoptera: Acrididae). Cell Tissue Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Couto, A.; Alenius, M.; Dickson, B.J. Molecular, anatomical, and functional organization of the Drosophila olfactory system. Curr. Biol. 2005, 15, 1535–1547. [Google Scholar] [CrossRef] [PubMed]
- Pener, M.P.; Simpson, S.J. Locust phase polyphenism: An update. Adv. Insect Physiol. 2009, 36, 1–272. [Google Scholar] [CrossRef]
- Seidelmann, K.; Weinert, H.; Ferenz, H.J. Wings and legs are production sites for the desert locust courtship-inhibition pheromone, phenylacetonitrile. J. Insect Physiol. 2003, 49, 1125–1133. [Google Scholar] [CrossRef]
- Wei, J.; Shao, W.; Wang, X.; Ge, J.; Chen, X.; Yu, D.; Kang, L. Composition and emission dynamics of migratory locust volatiles in response to changes in developmental stages and population density. Insect Sci. 2017, 24, 60–72. [Google Scholar] [CrossRef]
- Dweck, H.K.M.; Ebrahim, S.A.M.; Kromann, S.; Bown, D.; Hillbur, Y.; Sachse, S.; Hansson, B.S.; Stensmyr, M.C. Olfactory preference for egg laying on citrus substrates in Drosophila. Curr. Biol. 2013, 23, 2472–2480. [Google Scholar] [CrossRef]
- Ronderos, D.S.; Lin, C.-C.; Potter, C.J.; Smith, D.P. Farnesol-detecting olfactory neurons in Drosophila. J. Neurosci. 2014, 34, 3959–3968. [Google Scholar] [CrossRef]
- Krieger, J.; Große-Wilde, E.; Gohl, T.; Breer, H. Candidate pheromone receptors of the silkmoth Bombyx mori. Eur. J. Neurosci. 2005, 21, 2167–2176. [Google Scholar] [CrossRef]
- Legeai, F.; Malpel, S.; Montagné, N.; Monsempes, C.; Cousserans, F.; Merlin, C.; François, M.C.; Maïbèche-Coisné, M.; Gavory, F.; Poulain, J.; et al. An expressed sequence tag collection from the male antennae of the Noctuid moth Spodoptera littoralis: A resource for olfactory and pheromone detection research. BMC Genom. 2011, 12, 86. [Google Scholar] [CrossRef]
- Grosse-Wilde, E.; Kuebler, L.S.; Bucks, S.; Vogel, H.; Wicher, D.; Hansson, B.S. Antennal transcriptome of Manduca sexta. Proc. Natl. Acad. Sci. USA 2011, 108, 7449–7454. [Google Scholar] [CrossRef] [PubMed]
- Fuzeau-Braesch, S.; Genin, E.; Jullien, R.; Knowles, E.; Papin, C. Composition and role of volatile substances in atmosphere surrounding two gregarious locusts, Locusta migratoria and Schistocerca gregaria. J. Chem. Ecol. 1988, 14, 1023–1033. [Google Scholar] [CrossRef] [PubMed]
- Wei, J.; Shao, W.; Cao, M.; Ge, J.; Yang, P.; Chen, L.; Wang, X.; Kang, L. Phenylacetonitrile in locusts facilitates an antipredator defense by acting as an olfactory aposematic signal and cyanide precursor. Sci. Adv. 2019, 5, eaav5495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kurtovic, A.; Widmer, A.; Dickson, B.J. A single class of olfactory neurons mediates behavioural responses to a Drosophila sex pheromone. Nature 2007, 446, 542–546. [Google Scholar] [CrossRef] [PubMed]
- Steinbrecht, R.A. Structure and function of insect olfactory sensilla. Ciba Found. Symp. 1996, 200, 158–174. [Google Scholar] [PubMed]
- Stocker, R.F. Drosophila as a focus in olfactory research: Mapping of olfactory sensilla by fine structure, odor specificity, odorant receptor expression, and central connectivity. Microsc. Res. Technol. 2001, 55, 284–296. [Google Scholar] [CrossRef]
| SgreOR Group | Co-Expressed with SNMP1 | Not Co-Expressed with SNMP1 | ||||||
|---|---|---|---|---|---|---|---|---|
| Group I (9 ORs) 8 ORs examined | OR2 OR8 | OR3 OR9 | OR5 | OR6 | OR4 | OR7 | ||
| Group II (38 ORs) 33 ORs examined | OR84 OR90 OR94 OR101 OR109 OR113 | OR85 OR91 OR95 OR102 OR110 OR114 | OR86 OR92 OR96 OR103 OR111 OR116 | OR88 OR93 OR97 OR108 OR112 OR118 | OR82 OR98 OR106 | OR83 OR99 | OR87 OR100 | OR89 OR105 |
| Group III (70 ORs) 42 ORs examined | OR22 | OR23 | OR38 | OR11 OR16 OR27 OR31 OR35 OR41 OR49 OR54 OR66 OR71 | OR13 OR17 OR28 OR32 OR37 OR43 OR51 OR57 OR67 OR76 | OR14 OR25 OR29 OR33 OR39 OR45 OR52 OR61 OR68 OR80 | OR15 OR26 OR30 OR34 OR40 OR47 OR53 OR62 OR70 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pregitzer, P.; Jiang, X.; Lemke, R.-S.; Krieger, J.; Fleischer, J.; Breer, H. A Subset of Odorant Receptors from the Desert Locust Schistocerca gregaria Is Co-Expressed with the Sensory Neuron Membrane Protein 1. Insects 2019, 10, 350. https://doi.org/10.3390/insects10100350
Pregitzer P, Jiang X, Lemke R-S, Krieger J, Fleischer J, Breer H. A Subset of Odorant Receptors from the Desert Locust Schistocerca gregaria Is Co-Expressed with the Sensory Neuron Membrane Protein 1. Insects. 2019; 10(10):350. https://doi.org/10.3390/insects10100350
Chicago/Turabian StylePregitzer, Pablo, Xingcong Jiang, René-Sebastian Lemke, Jürgen Krieger, Jörg Fleischer, and Heinz Breer. 2019. "A Subset of Odorant Receptors from the Desert Locust Schistocerca gregaria Is Co-Expressed with the Sensory Neuron Membrane Protein 1" Insects 10, no. 10: 350. https://doi.org/10.3390/insects10100350







