Precision Medicine—Are We There Yet? A Narrative Review of Precision Medicine’s Applicability in Primary Care
Abstract
1. Introduction
2. The Emergence of -Omics
2.1. Genomics
2.1.1. Molecular Disease Definition
2.1.2. Polygenic Risk Scores
2.1.3. Pharmacogenomics
- Improving drug efficacy. Variation in the cytochrome P450 gene CYP2D6 affects the metabolism and elimination of more than 100 drugs [50]. One of these drugs is the analgesic codeine, which is metabolised to the bioactive form morphine. Patients can be classified by their rate of metabolism, with clinical implications for the ultrarapid metabolisers (UMs) and poor metabolisers [51], and pharmacogenomic guidance in the summary of product characteristics (SmPCs) [52]. Variants for another cytochrome P450 gene, CYP2C19, can also significantly impact clopidogrel metabolism and efficacy with an FDA “black box warning” for those carrying these variants [53].
- Testing at the time of prescribing. Typically, this is a single drug–gene test performed in advance of the prescription decision about to be made. For example, in oncology, DPYD testing in advance of initiating 5-FU, or in neonatal sepsis, testing the RNR1 gene for variants associated with aminoglycoside-induced hearing loss [57]. Opportunities for this approach, rapid testing for a single and significant gene–drug interaction, will broaden as molecular diagnostics continue to advance [58]. Although point-of-care testing (POCT) is already utilised in primary care [59], it is hard to see PGx POCT expand beyond a relatively limited number of indications. In primary care, given the range of clinical presentations and prescribing decisions, the feasibility of POCT, and the effect of even a modest delay on patient flow, an alternative approach is likely to be better suited.
- Testing patients in advance of prescribing decisions. This is a more distant prospect for primary care but would involve pre-emptively performing a PGx gene panel test for several key drug–gene interactions with the results captured to inform future prescribing decisions. This may involve testing at a separate time to the prescribing decision or be triggered by prescribing a single drug on the panel, the specific drug information available to guide that treatment, and the other PGx information available for future reference. An attractive approach is to use the existing sequencing data, captured for another indication to identify PGx variants, which is increasingly feasible as an increasing proportion of the population has genome or exome sequencing [47].
2.2. Transcriptomics, Epigenomics, Proteomics, Metabolomics, and Exposomics
3. Big Data, Data Analytics, and AI
3.1. Electronic Health Records
3.2. Digital Technologies including Wearables
3.3. Prediction Modelling
3.4. Artificial Intelligence (AI)
4. Discussion
- Co-development of technologies. To date, healthcare AI tools have often been driven by a focus on the technology and commercial need to find a marker rather than patient and clinician need [101]. The co-development of PM technologies with primary care clinicians, patients, and the public is key to ensure they address the needs and meet the standards and values of society and primary care. With a specific focus on ensuring an evidence base in the primary care setting they are to be deployed, consideration should be given to their impact on continuity of care and how they fit into the consultation and how they will avoid overmedicalisation and increasing health anxiety [101]. It will also be important to ensure that the outputs of PM technologies are delivered in such a way to have optimal impact, effectively inform clinical decision making and create meaningful change in people’s behaviour whilst not excessively burdening a healthcare system already under pressure.
- Real-world evidence in the population and setting it is to be deployed. Unrepresentative and biased datasets lead to PM tools that may exacerbate health inequalities. Before implementing PM at scale, health strategies need to ensure that the foundations upon which PM is built, datasets, genetic databases, cohort studies, and EHR datasets, are appropriately diverse and representative of their intended use population. Endeavours such as the STANDING Together initiative will be key to encourage representativeness in datasets and ensure transparency in how diversity is reported [120].Fundamentally, evidence needs to be gathered on PM technology in the setting that it is intended to be deployed. Environmental factors and where the technology is placed in the current workflow can significantly impact its performance and utility [76,121]. Before the widespread adoption of PM, implementation models need to be used suitably for these new technologies [122] and appropriate evaluation frameworks applied to ensure robust real-world evidence [109,123]. To minimise the workload impact of these technologies, care will be necessary to ensure they are implemented efficiently, but how we define and then measure efficiency is not clear, especially in the context of new technologies, and is an area for further research [124].Currently, in primary care, the EHR system plays a key role in the doctor–patient consultation, not only to inform the doctor and record clinical information but to facilitate patient involvement in the consultation by sharing the monitor screen [125]. When implementing PM technologies, not only should one consider how to use the contents of the EHR to develop the precision medicine insight but also how the EHR system interface will enable the clinician and patient to best understand and implement these PM insights meaningfully.
- Demonstrating the cost-effectiveness of PM. There is still much uncertainty about the affordability and health economic profile across the range of PM interventions [126]. PM enthusiasts, and frequently policy makers, highlight the potential cost savings of PM: avoiding ill health; promoting health prevention; streamlining diagnostic pathways with earlier diagnoses; making better therapeutic decisions, with less associated waste and adverse events; and decreasing the disease burden for the public at large [76,127,128]. However, the evidence for these health system efficiencies is hard to capture with the PM interventions adding an upfront cost, with the later benefit difficult to measure. For example, currently, drugs are prescribed without pharmacogenomic information. Will a net reduction in adverse events and inappropriate prescribing, and thus in health impact and cost, justify the expenditure of pharmacogenomic testing at a population level? Capturing sufficient information to understand cost-effectiveness, clinical impact, and the long-term viability of PM is likely to require an extended period of surveillance. Such ongoing surveillance of PM interventions should be ensured from the outset, adapting existing approaches of post-market surveillance for new drugs and medical devices.Whilst PM does not in itself seek to establish novel medications, it does stratify patients into subgroups who will respond to specific treatments regimes. Molecular disease definitions divide common conditions into multiple distinct subgroups, many of which will have their own treatment. In cancer, this has led to the development of drugs that have in many settings been prohibitively expensive [129]. Repositioning affordable licensed drugs based on specific molecular targets is an attractive proposition to reduce drug costs whilst maximising efficacy [121]. Although this has not been widely utilised to date, pharmaceutical companies and government research funders could use this opportunity to revisit “old drugs” for targeted personalised therapy in specific subgroups [130].
- Data collection sharing and transfer. Much of the potential of PM is dependent upon processing and analysing large amounts of data. Genomic sequencing has advanced at pace; however, the availability of information on diverse well-phenotyped individuals has not kept pace, hampering the ability to establish connections between disease and genomics [131]. Optimising the quality of data recorded is key. To achieve this, establishing standards for data recording, including clinical vocabulary that is used across care settings, and frameworks to share data, compliant with legal restrictions, that maintain patient privacy, and incorporating individual preferences for their data use need to be prioritised.In UK primary care, for example, better guidance regarding data sharing is needed to ensure practice is more uniform, with recent proposals that data controller responsibilities could be shared with national bodies [132]. In addition to ensuring the quality of the data recorded and standards for sharing, significant attention should be focussed on the storage and processing of the vast quantity of data that PM needs and will generate. This risks overwhelming an already stretched health system and workforce. Although plans are in place to advance NHS digital systems [133], this needs to be prioritised with suitable cautions given that previous large-scale IT infrastructure projects across the NHS have failed [134].
- Impact upon holistic care. There is concern that the advance of PM interventions may reduce the need for human interaction, with the virtual clinical assistant taking a greater role and affecting the patient–doctor relationship. As healthcare interventions become more personalised, derived from increasingly complex methodologies, the rational for the intervention may become more opaque [114,115]. Will this lack of transparency erode trust and further impact the doctor–patient relationship? Advocates of PM suggest that it will enable the better use or resources, bringing together disparate information to support the clinician to make the best decision, thus freeing time for human intelligence and restoring empathy [135]. However, evidence for or against this position needs to be established, with robust primary care-based qualitative research incorporating the views of clinicians and patients.
5. Conclusions
6. Take-Home Points
- Precision medicine (PM), a term often used interchangeably with targeted, stratified, individualised, and personalised medicine, is a rapidly developing area of research and practice.
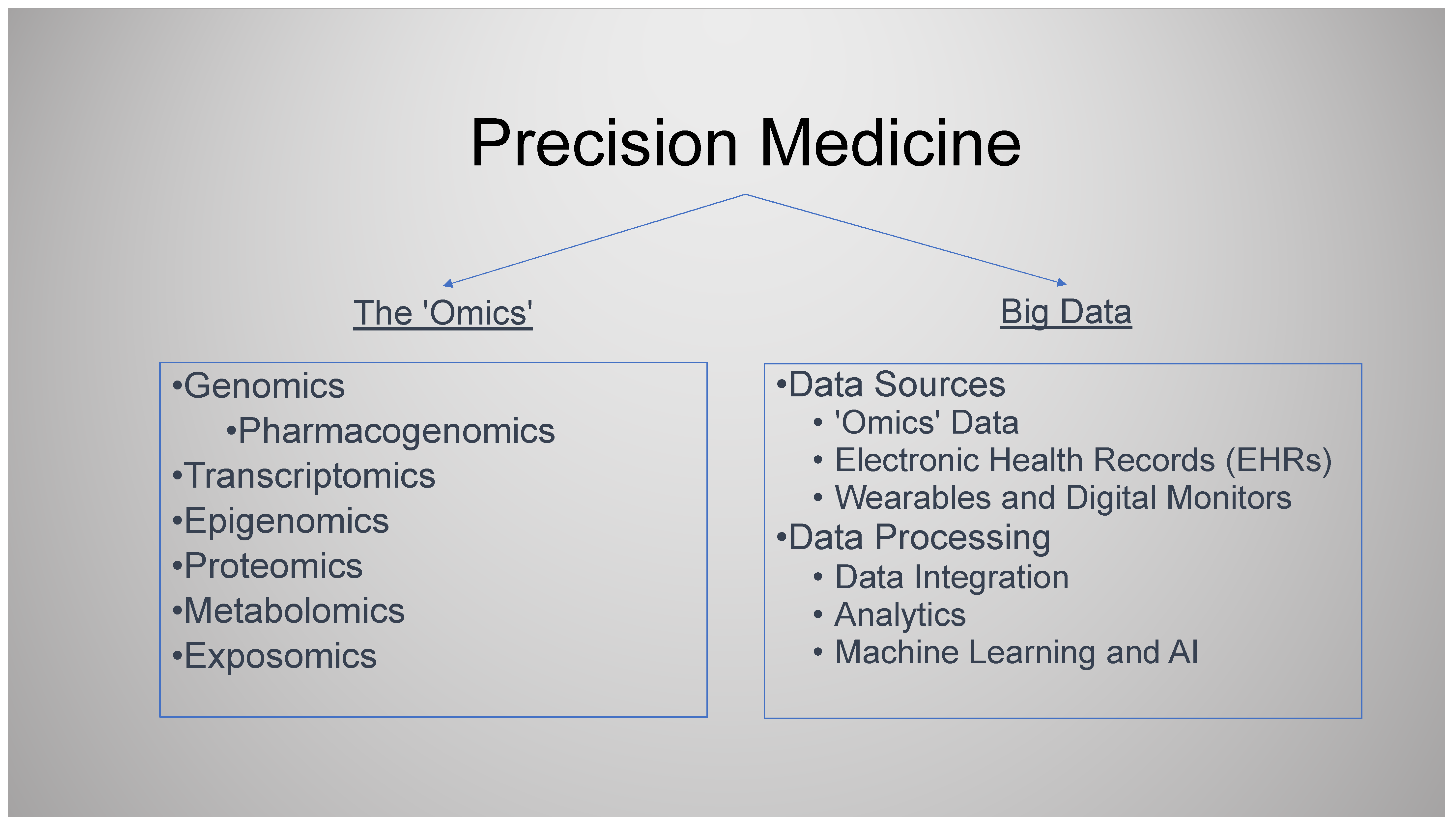
- PM can be considered to involve two broad areas, each with its own sub-domains. (1) The “-omics”, with a whole array of prefixes including but not limited to genomics, pharmacogenomics, transcriptomics, epigenomics, proteomics, metabolomics, and exposomics. (2) Big data, data analytics, and artificial intelligence (AI).
- PM is data science-driven and is built upon large volumes of biomedical data. AI is a key tool to manage, analyse, and communicate the insights from the multiple big data healthcare sources.
- The three -omics likely to impact primary care are polygenic risk scores to optimise patient risk stratification, pharmacogenomics to tailor treatment, and molecular genetic diagnostic testing. The impact of all will be optimised when integrated with the primary care electronic health record (EHR), which forms the key data resource for PM development and implementation in this setting.
- The evidence base for PM is still emerging. The pressures and incentives for the early adoption of PM technologies are multifactorial and complex. There is a need for PM initiatives that have real-world evidence of clinical utility in the context they are to be deployed. Particular attention to demonstrating cost-effectiveness and effect on health inequalities is required.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Osler, W. On the educational value of the medical society. In Equanimitas with Other Addresses to Medical Students, Nurses and Practitioners of Medicine; Blakiston: Philadelphia, PA, USA, 1904; pp. 343–362. Available online: https://onlinelibrary.wiley.com/doi/full/10.1046/j.1365-2141.2003.04615.x (accessed on 28 November 2023).
- Schleidgen, S.; Klingler, C.; Bertram, T.; Rogowski, W.H.; Marckmann, G. What is personalized medicine: Sharpening a vague term based on a systematic literature review. BMC Med. Ethics 2013, 14, 55. [Google Scholar] [CrossRef] [PubMed]
- Boyer, M.S.; Widmer, D.; Cohidon, C.; Desvergne, B.; Cornuz, J.; Guessous, I.; Cerqui, D. Representations of personalised medicine in family medicine: A qualitative analysis. BMC Prim. Care 2022, 23, 37. [Google Scholar] [CrossRef] [PubMed]
- Ginsburg, G.S.; Phillips, K.A. Precision Medicine: From Science to Value. Health Aff. Proj. Hope 2018, 37, 694–701. [Google Scholar] [CrossRef]
- National Research Council Committee on AF for D a NT of D. The National Academies Collection: Reports funded by National Institutes of Health. In Toward Precision Medicine: Building a Knowledge Network for Biomedical Research and a New Taxonomy of Disease; National Academies Press (US) National Academy of Sciences: Washington, DC, USA, 2011. [Google Scholar]
- Evans, R.S. Electronic Health Records: Then, Now, and in the Future. IMIA Yearbook. Med. Inform. 2016, 25 (Suppl. S1), S48–S61. [Google Scholar] [CrossRef]
- Abdelhalim, H.; Berber, A.; Lodi, M.; Jain, R.; Nair, A.; Pappu, A.; Patel, K.; Venkat, V.; Venkatesan, C.; Wable, R.; et al. Artificial Intelligence, Healthcare, Clinical Genomics, and Pharmacogenomics Approaches in Precision Medicine. Front. Genet. 2022, 13, 929736. [Google Scholar] [CrossRef]
- National Institutes of Health (NIH) All of Us. Available online: https://allofus.nih.gov/ (accessed on 13 June 2019).
- Stenzinger, A. Implementation of precision medicine in healthcare—A European perspective. In Journal of Internal Medicine; Wiley Online Library: Hoboken, NJ, USA, 2023; Available online: https://onlinelibrary.wiley.com/doi/10.1111/joim.13698 (accessed on 20 March 2024).
- International Consortium for Personalised Medicine—ICPerMed. Available online: https://www.icpermed.eu/ (accessed on 20 March 2024).
- Understanding the Global Landscape of Genomic Initiatives—IQVIA. Available online: https://www.iqvia.com/insights/the-iqvia-institute/reports-and-publications/reports/understanding-the-global-landscape-of-genomic-initiatives (accessed on 20 March 2024).
- Collins, F.S.; Varmus, H. A New Initiative on Precision Medicine. N. Engl. J. Med. 2015, 372, 793–795. [Google Scholar] [CrossRef] [PubMed]
- Joyner, M.J.; Paneth, N. Promises, promises, and precision medicine. J. Clin. Investig. 2019, 129, 946–948. [Google Scholar] [CrossRef] [PubMed]
- Evans, J.P.; Meslin, E.M.; Marteau, T.M.; Caulfield, T. Genomics. Deflating the genomic bubble. Science 2011, 331, 861–862. [Google Scholar] [CrossRef]
- Guttmacher, A.E.; Collins, F.S. (Eds.) Genomic Medicine—A Primer. N. Engl. J. Med. 2002, 347, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, N.; Humphries, S.E.; Gray, H. Personalised Medicine in General Practice: The Example of Raised Cholesterol. 2018. Available online: https://bjgp.org/content/68/667/68 (accessed on 20 February 2024).
- Davies, S.C. Annual Report of the Chief Medical Officer 2016, Generation Genome, 2016; Department of Health: London, UK, 2017. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/631043/CMO_annual_report_generation_genome.pdf (accessed on 20 February 2024).
- NHS England » Improving Outcomes through Personalised Medicine. Available online: https://www.england.nhs.uk/publication/improving-outcomes-through-personalised-medicine/ (accessed on 18 September 2023).
- Orrantia-Borunda, E.; Anchondo-Nuñez, P.; Acuña-Aguilar, L.E.; Gómez-Valles, F.O.; Ramírez-Valdespino, C.A. Subtypes of Breast Cancer. In Breast Cancer; Mayrovitz, H.N., Ed.; Exon Publications: Brisbane, AU, USA, 2022. Available online: http://www.ncbi.nlm.nih.gov/books/NBK583808/ (accessed on 13 October 2023).
- Wang, R.C.; Wang, Z. Precision Medicine: Disease Subtyping and Tailored Treatment. Cancers 2023, 15, 3837. [Google Scholar] [CrossRef]
- Borish, L.; Culp, J.A. Asthma: A syndrome composed of heterogeneous diseases. Ann. Allergy Asthma Immunol. Off. Publ. Am. Coll. Allergy Asthma Immunol. 2008, 101, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tsuo, K.; Zhou, W.; Wang, Y.; Kanai, M.; Namba, S.; Gupta, R.; Majara, L.; Nkambule, L.; Morisaki, T.; Okada, Y.; et al. Multi-ancestry meta-analysis of asthma identifies novel associations and highlights the value of increased power and diversity. Cell Genom. 2022, 2, 100212. [Google Scholar] [CrossRef] [PubMed]
- Colicino, S.; Munblit, D.; Minelli, C.; Custovic, A.; Cullinan, P. Validation of childhood asthma predictive tools: A systematic review. Clin. Exp. Allergy 2019, 49, 410–418. [Google Scholar] [CrossRef]
- Linder, J.E.; Allworth, A.; Bland, H.T.; Caraballo, P.J.; Chisholm, R.L.; Clayton, E.W.; Crosslin, D.; Dikilitas, O.; DiVietro, A.; Esplin, E.; et al. Returning integrated genomic risk and clinical recommendations: The eMERGE study. Genet. Med. Off. J. Am. Coll. Med. Genet. 2023, 25, 100006. [Google Scholar] [CrossRef]
- NICE. Overview|Familial Breast Cancer: Classification, Care and Managing Breast Cancer and Related Risks in People with a Family History of Breast Cancer|Guidance|NICE. NICE. 2017. Available online: https://www.nice.org.uk/guidance/cg164 (accessed on 23 May 2019).
- Evans, D.G.; Astley, S.; Stavrinos, P.; Harkness, E.; Donnelly, L.S.; Dawe, S.; Jacob, I.; Harvie, M.; Cuzick, J.; Brentall, C.; et al. Improvement in Risk Prediction, Early Detection and Prevention of Breast Cancer in the NHS Breast Screening Programme and Family History Clinics: A Dual Cohort Study; Programme Grants for Applied Research; NIHR Journals Library: Southampton, UK, 2016. Available online: http://www.ncbi.nlm.nih.gov/books/NBK379488/ (accessed on 24 September 2023).
- McHugh, J.K.; ni Raghallaigh, H.; Bancroft, E.; Kote-Jarai, Z.; Benafif, S.; Eeles, R.A. The BARCODE1 study in primary care: Early results targeting men with increased genetic risk of developing prostate cancer—Examining the interim data from a community-based screening program using polygenic risk score to target screening. J. Clin. Oncol. 2022, 40 (Suppl. S6), 231. [Google Scholar] [CrossRef]
- Fuat, A.; Adlen, E.; Monane, M.; Coll, R.; Groves, S.; Little, E.; Wild, J.; Kamali, F.; Soni, Y.; Haining, S.; et al. A polygenic risk score added to a QRISK®2 cardiovascular disease risk calculator demonstrated robust clinical acceptance and clinical utility in the primary care setting. Eur. J. Prev. Cardiol. 2024, zwae004. [Google Scholar] [CrossRef] [PubMed]
- Elliott, J.; Bodinier, B.; Bond, T.A.; Chadeau-Hyam, M.; Evangelou, E.; Moons, K.G.M.; Dehgan, A.; Muller, D.C.; Elliott, P.; Tzoulaki, I. Predictive Accuracy of a Polygenic Risk Score-Enhanced Prediction Model vs. a Clinical Risk Score for Coronary Artery Disease. JAMA 2020, 323, 636–645. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.D.; Hurson, A.N.; Zhang, H.; Choudhury, P.P.; Easton, D.F.; Milne, R.L.; Simard, J.; Hall, P.; Michailidou, K.; Dennis, J.; et al. Assessment of polygenic architecture and risk prediction based on common variants across fourteen cancers. Nat. Commun. 2020, 11, 3353. [Google Scholar] [CrossRef]
- Hingorani, A.; Gratton, J.; Finan, C.; Schmidt, A.; Patel, R.; Sofat, R.; Kuan, V.; Langenberg, C.; Hemingway, H.; Morris, J.K.; et al. Performance of polygenic risk scores in screening, prediction, and risk stratification: Secondary analysis of data in the Polygenic Score Catalog. BMJ Med. 2023, 2, e000554. [Google Scholar] [CrossRef]
- Kiflen, M.; Le, A.; Mao, S.; Lali, R.; Narula, S.; Xie, F.; Paré, G. Cost-Effectiveness of Polygenic Risk Scores to Guide Statin Therapy for Cardiovascular Disease Prevention. Circ. Genom. Precis. Med. 2022, 15, e003423. [Google Scholar] [CrossRef]
- Chung, R.; Xu, Z.; Arnold, M.; Ip, S.; Harrison, H.; Barrett, J.; Pennells, L.; Kim, L.G.; Di Angelantonio, E.; Paige, E.; et al. Using Polygenic Risk Scores for Prioritizing Individuals at Greatest Need of a Cardiovascular Disease Risk Assessment. J. Am. Heart Assoc. 2023, 12, e029296. [Google Scholar] [CrossRef]
- GWAS to the people. Nat. Med. 2018, 24, 1483. Available online: https://www.nature.com/articles/s41591-018-0231-3#citeas (accessed on 17 October 2023). [CrossRef]
- Martin, A.R.; Kanai, M.; Kamatani, Y.; Okada, Y.; Neale, B.M.; Daly, M.J. Current clinical use of polygenic scores will risk exacerbating health disparities. Nat. Genet. 2019, 51, 584–591. [Google Scholar] [CrossRef]
- Wang, Y.; Namba, S.; Lopera, E.; Kerminen, S.; Tsuo, K.; Läll, K.; Kanai, M.; Zhou, W.; Wu, K.; Favé, M.J.; et al. Global Biobank analyses provide lessons for developing polygenic risk scores across diverse cohorts. Cell Genom. 2023, 3, 100241. [Google Scholar] [CrossRef]
- Sud, A.; Horton, R.H.; Hingorani, A.D.; Tzoulaki, I.; Turnbull, C.; Houlston, R.S.; Lucassen, A. Realistic expectations are key to realising the benefits of polygenic scores. BMJ 2023, 380, e073149. [Google Scholar] [CrossRef]
- Lee, A.; Mavaddat, N.; Wilcox, A.N.; Cunningham, A.P.; Carver, T.; Hartley, S.; Babb de Villiers, C.; Izquierdo, A.; Simard, J.; Schmidy, M.K.; et al. BOADICEA: A comprehensive breast cancer risk prediction model incorporating genetic and nongenetic risk factors. Genet. Med. Off. J. Am. Coll. Med. Genet. 2019, 21, 1708–1718. [Google Scholar]
- Resnik, D.B.; Vorhaus, D.B. Genetic modification and genetic determinism. Philos. Ethics Humanit. Med. 2006, 1, 9. [Google Scholar] [CrossRef]
- Middlemass, J.B.; Yazdani, M.F.; Kai, J.; Standen, P.J.; Qureshi, N. Introducing genetic testing for cardiovascular disease in primary care: A qualitative study. Br. J. Gen. Pract. J. R Coll. Gen. Pract. 2014, 64, e282–e289. [Google Scholar] [CrossRef]
- Hollands, G.J.; French, D.P.; Griffin, S.J.; Prevost, A.T.; Sutton, S.; King, S.; Marteau, T.M. The impact of communicating genetic risks of disease on risk-reducing health behaviour: Systematic review with meta-analysis. BMJ 2016, 352, i1102. [Google Scholar] [CrossRef] [PubMed]
- Adminstration UF and DK: Paving the Way for Personalized Medicine: FDA’s ROle in a New Era of Medical Product Development. 2013. Available online: https://www.fdanews.com/ext/resources/files/10/10-28-13-Personalized-Medicine.pdf (accessed on 1 October 2023).
- Sadee, W.; Dai, Z. Pharmacogenetics/genomics and personalized medicine. Hum. Mol. Genet. 2005, 14, R207–R214. [Google Scholar] [CrossRef] [PubMed]
- Donaldson, L. An organisation with a memory. Clin. Med. 2002, 2, 452–457. Available online: https://pdfs.semanticscholar.org/a675/29ad58dbc031e82f33555d071a568a4e5c6c.pdf (accessed on 1 October 2023). [CrossRef]
- Overview|Medicines Optimisation: The Safe and Effective Use of Medicines to Enable the Best Possible Outcomes|Guidance|NICE. NICE. 2015. Available online: https://www.nice.org.uk/guidance/ng5 (accessed on 25 September 2023).
- Sultana, J.; Cutroneo, P.; Trifirò, G. Clinical and economic burden of adverse drug reactions. J. Pharmacol. Pharmacother. 2013, 4 (Suppl. S1), S73–S77. [Google Scholar] [CrossRef]
- RCP London. Personalised Prescribing: Using Pharmacogenomics to Improve Patient Outcomes. 2022. Available online: https://www.rcp.ac.uk/projects/outputs/personalised-prescribing-using-pharmacogenomics-improve-patient-outcomes (accessed on 25 September 2023).
- Tangamornsuksan, W.; Chaiyakunapruk, N.; Somkrua, R.; Lohitnavy, M.; Tassaneeyakul, W. Relationship between the HLA-B*1502 allele and carbamazepine-induced Stevens-Johnson syndrome and toxic epidermal necrolysis: A systematic review and meta-analysis. JAMA Dermatol. 2013, 149, 1025–1032. [Google Scholar] [CrossRef]
- BNF Content Published by NICE. 2023. Available online: https://bnf.nice.org.uk/ (accessed on 20 October 2023).
- Bertilsson, L.; Dahl, M.L.; Dalén, P.; Al-Shurbaji, A. Molecular genetics of CYP2D6: Clinical relevance with focus on psychotropic drugs. Br. J. Clin. Pharmacol. 2002, 53, 111–122. [Google Scholar] [CrossRef]
- Crews, K.R.; Gaedigk, A.; Dunnenberger, H.M.; Leeder, J.S.; Klein, T.E.; Caudle, K.E.; Haidar, C.E.; Shen, D.D.; Callaghan, J.T.; Sadhasivam, S.; et al. Clinical Pharmacogenetics Implementation Consortium guidelines for cytochrome P450 2D6 genotype and codeine therapy: 2014 update. Clin. Pharmacol. Ther. 2014, 95, 376–382. [Google Scholar] [CrossRef]
- Codeine Phosphate 30mg Tablets—Summary of Product Characteristics (SmPC)—(Emc). Available online: https://www.medicines.org.uk/emc/product/2375/smpc#gref (accessed on 26 September 2023).
- Administration UF and D. FDA Drug Safety Communication: Reduced Effectiveness of Plavix (Clopidogrel) in Patients Who Are Poor Metabolizers of the Drug|FDA. US_FDA. 2010. Available online: https://www.fda.gov/drugs/postmarket-drug-safety-information-patients-and-providers/fda-drug-safety-communication-reduced-effectiveness-plavix-clopidogrel-patients-who-are-poor (accessed on 1 October 2023).
- Martin, M.A.; Kroetz, D.L. Abacavir Pharmacogenetics—From Initial Reports to Standard of Care. Pharmacotherapy 2013, 33, 765–775. [Google Scholar] [CrossRef]
- Henricks, L.M.; Lunenburg, C.A.T.C.; Cats, A.; Mathijssen, R.H.J.; Guchelaar, H.J.; Schellens, J.H.M. DPYD genotype-guided dose individualisation of fluoropyrimidine therapy: Who and how?—Authors’ reply. Lancet Oncol. 2019, 20, e67. [Google Scholar] [CrossRef]
- PharmGKB. Available online: https://www.pharmgkb.org/ (accessed on 26 September 2023).
- McDermott, J.H.; Mahaveer, A.; James, R.A.; Booth, N.; Turner, M.; Harvey, K.E.; Miele, G.; Beaman, G.M.; Stoddard, D.C.; Tricker, K.; et al. Rapid Point-of-Care Genotyping to Avoid Aminoglycoside-Induced Ototoxicity in Neonatal Intensive Care. JAMA Pediatr. 2022, 176, 486–492. [Google Scholar] [CrossRef]
- McDermott, J.H.; Burn, J.; Donnai, D.; Newman, W.G. The rise of point-of-care genetics: How the SARS-CoV-2 pandemic will accelerate adoption of genetic testing in the acute setting. Eur. J. Hum. Genet. 2021, 29, 891–893. [Google Scholar] [CrossRef] [PubMed]
- Lingervelder, D.; Koffijberg, H.; Kusters, R.; IJzerman, M.J. Point-of-care testing in primary care: A systematic review on implementation aspects addressed in test evaluations. Int. J. Clin. Pract. 2019, 73, e13392. [Google Scholar] [CrossRef] [PubMed]
- Vermeulen, R.; Schymanski, E.L.; Barabási, A.L.; Miller, G.W. The exposome and health: Where chemistry meets biology. Science 2020, 367, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Topol, E.J. Individualized Medicine from Prewomb to Tomb. Cell 2014, 157, 241–253. [Google Scholar] [CrossRef] [PubMed]
- Baccarelli, A.; Dolinoy, D.C.; Walker, C.L. A precision environmental health approach to prevention of human disease. Nat. Commun. 2023, 14, 2449. [Google Scholar] [CrossRef] [PubMed]
- Pirola, L.; Balcerczyk, A.; Okabe, J.; El-Osta, A. Epigenetic phenomena linked to diabetic complications. Nat. Rev. Endocrinol. 2010, 6, 665–675. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Wang, L.; Chen, D.; Feng, M.; Lu, Y.; Chen, R.; Qiu, C.; Li, J. The application of proteomics in the diagnosis and treatment of bronchial asthma. Ann. Transl. Med. 2020, 8, 132. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, Z.; Mohamed, K.; Zeeshan, S.; Dong, X. Artificial intelligence with multi-functional machine learning platform development for better healthcare and precision medicine. Database J. Biol. Databases Curation 2020, 2020, baaa010. [Google Scholar] [CrossRef] [PubMed]
- Leon-Mimila, P.; Wang, J.; Huertas-Vazquez, A. Relevance of Multi-Omics Studies in Cardiovascular Diseases. Front. Cardiovasc. Med. 2019, 6, 91. Available online: https://www.frontiersin.org/articles/10.3389/fcvm.2019.00091 (accessed on 8 December 2023). [CrossRef] [PubMed]
- Wolf, A.; Dedman, D.; Campbell, J.; Booth, H.; Lunn, D.; Chapman, J.; Myles, P. Data resource profile: Clinical Practice Research Datalink (CPRD) Aurum. Int. J. Epidemiol. 2019, 48, 1740. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Yang, X.; Chen, A.; Smith, K.E.; PourNejatian, N.; Costa, A.B.; Martin, C.; Flores, M.G.; Zhang, Y.; Magoc, T.; et al. A Study of Generative Large Language Model for Medical Research and Healthcare. NPJ Digit. Med. 2023, 6, 210. [Google Scholar] [CrossRef]
- Dzau, V.J.; Ginsburg, G.S. Realizing the Full Potential of Precision Medicine in Health and Health CarePotential of Precision Medicine in Health and Health CarePotential of Precision Medicine in Health and Health Care. JAMA 2016, 316, 1659–1660. [Google Scholar] [CrossRef]
- Jones, L.A.; Nelder, J.R.; Fryer, J.M.; Alsop, P.H.; Geary, M.R.; Prince, M.; Cardinal, R.N. Public opinion on sharing data from health services for clinical and research purposes without explicit consent: An anonymous online survey in the, U.K. BMJ Open 2022, 12, e057579. [Google Scholar] [CrossRef]
- Armstrong, S. Pause plans for data sharing contract to ensure trust and patient consent, plead doctors’ leaders. BMJ 2023, 383, 2674. [Google Scholar] [CrossRef]
- Parr, E. UK Biobank Writes to All GP Practices Requesting They Share Patient Data. Pulse Today. 29 September 2023. Available online: https://www.pulsetoday.co.uk/news/technology/uk-biobank-writes-to-all-gp-practices-requesting-they-share-patient-data/ (accessed on 17 October 2023).
- Izmailova, E.S.; Wagner, J.A.; Perakslis, E.D. Wearable Devices in Clinical Trials: Hype and Hypothesis. Clin. Pharmacol. Ther. 2018, 104, 42–52. [Google Scholar] [CrossRef]
- The Topol Review. Available online: https://topol.hee.nhs.uk/the-topol-review/ (accessed on 18 September 2023).
- McGinnis, J.M.; Williams-Russo, P.; Knickman, J.R. The Case For More Active Policy Attention To Health Promotion. Health Aff. 2002, 21, 78–93. [Google Scholar] [CrossRef]
- Johnson, K.B.; Wei, W.; Weeraratne, D.; Frisse, M.E.; Misulis, K.; Rhee, K.; Zhao, J.; Snowdon, J.L. Precision Medicine, AI, and the Future of Personalized Health Care. Clin. Transl. Sci. 2021, 14, 86–93. [Google Scholar] [CrossRef]
- Hemingway, H.; Croft, P.; Perel, P.; Hayden, J.A.; Abrams, K.; Timmis, A.; Briggs, A.; Udumyan, R.; Moons, K.G.; Steyerberg, E.W.; et al. Prognosis research strategy (PROGRESS) 1: A framework for researching clinical outcomes. BMJ 2013, 346, e5595. [Google Scholar] [CrossRef]
- Cox, D.R. Regression Models and Life-Tables. J. R. Stat. Soc. Ser. B Methodol. 1972, 34, 187–220. [Google Scholar] [CrossRef]
- Vasan, R.S.; Beiser, A.; Seshadri, S.; Larson, M.G.; Kannel, W.B.; D’Agostino, R.B.; Levy, D. Residual Lifetime Risk for Developing Hypertension in Middle-aged Women and MenThe Framingham Heart Study. JAMA 2002, 287, 1003–1010. [Google Scholar] [CrossRef]
- Kanis, J.A.; Johnell, O.; Oden, A.; Johansson, H.; McCloskey, E. FRAX and the assessment of fracture probability in men and women from the UK. Osteoporos. Int. J. Establ. Result Coop. Eur. Found Osteoporos. Natl. Osteoporos. Found USA 2008, 19, 385–397. [Google Scholar] [CrossRef] [PubMed]
- Dent, E.; Kowal, P.; Hoogendijk, E.O. Frailty measurement in research and clinical practice: A review. Eur. J. Intern. Med. 2016, 31, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Hippisley-Cox, J.; Coupland, C.; Brindle, P. Development and validation of QRISK3 risk prediction algorithms to estimate future risk of cardiovascular disease: Prospective cohort study. BMJ 2017, 357, j2099. [Google Scholar] [CrossRef] [PubMed]
- Kingston, M.; Griffiths, R.; Hutchings, H.; Porter, A.; Russell, I.; Snooks, H. Emergency admission risk stratification tools in UK primary care: A cross-sectional survey of availability and use. Br. J. Gen. Pract. 2020, 70, e740–e748. [Google Scholar] [CrossRef]
- Papadakis, G.Z.; Karantanas, A.H.; Tsiknakis, M.; Tsatsakis, A.; Spandidos, D.A.; Marias, K. Deep learning opens new horizons in personalized medicine. Biomed. Rep. 2019, 10, 215–217. [Google Scholar] [CrossRef]
- Weng, S.F.; Reps, J.; Kai, J.; Garibaldi, J.M.; Qureshi, N. Can machine-learning improve cardiovascular risk prediction using routine clinical data? PLoS ONE 2017, 12, e0174944. [Google Scholar] [CrossRef]
- Callender, T.; Imrie, F.; Cebere, B.; Pashayan, N.; Navani, N.; van der Schaar, M.; Janes, S.M. Assessing eligibility for lung cancer screening using parsimonious ensemble machine learning models: A development and validation study. PLoS Med. 2023, 20, e1004287. [Google Scholar] [CrossRef] [PubMed]
- Montazeri, M.; Montazeri, M.; Bahaadinbeigy, K.; Montazeri, M.; Afraz, A. Application of machine learning methods in predicting schizophrenia and bipolar disorders: A systematic review. Health Sci. Rep. 2022, 6, e962. [Google Scholar] [CrossRef]
- Weng, S.F.; Vaz, L.; Qureshi, N.; Kai, J. Prediction of premature all-cause mortality: A prospective general population cohort study comparing machine-learning and standard epidemiological approaches. PLoS ONE 2019, 14, e0214365. [Google Scholar] [CrossRef]
- Topol, E.J. As artificial intelligence goes multimodal, medical applications multiply. Science 2023, 381, adk6139. [Google Scholar] [CrossRef]
- Archer-Williams, A. Microsoft and Epic Expand AI Collaboration to Tackle Current Healthcare Needs. htn. 2023. Available online: https://htn.co.uk/2023/08/23/microsoft-and-epic-expand-ai-collaboration-to-tackle-current-healthcare-needs/ (accessed on 29 August 2023).
- Lim, J.I.; Regillo, C.D.; Sadda, S.R.; Ipp, E.; Bhaskaranand, M.; Ramachandra, C.; Solanki, K. Artificial Intelligence Detection of Diabetic Retinopathy. Ophthalmol. Sci. 2022, 3, 100228. [Google Scholar] [CrossRef] [PubMed]
- Poplin, R.; Varadarajan, A.V.; Blumer, K.; Liu, Y.; McConnell, M.V.; Corrado, G.S.; Peng, L.; Webster, D.R. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat. Biomed. Eng. 2018, 2, 158–164. [Google Scholar] [CrossRef]
- Cheung, C.Y.; Ran, A.R.; Wang, S.; Chan, V.T.T.; Sham, K.; Hilal, S.; Venketasubramanian, N.; Cheng, C.; Sabanayagam, C.; Tham, Y.; et al. A deep learning model for detection of Alzheimer’s disease based on retinal photographs: A retrospective, multicentre case-control study. Lancet Digit. Health 2022, 4, e806–e815. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Pinto, A.; Ravikumar, N.; Attar, R.; Suinesiaputra, A.; Zhao, Y.; Levelt, E.; Dall’Armellina, E.; Lorenzi, M.; Chen, Q.; Kennan, T.D.L.; et al. Predicting myocardial infarction through retinal scans and minimal personal information. Nat. Mach. Intell. 2022, 4, 55–61. [Google Scholar] [CrossRef]
- Wagner, S.K.; Romero-Bascones, D.; Cortina-Borja, M.; Williamson, D.J.; Struyven, R.R.; Zhou, Y.; Patel, S.; Weil, R.S.; Antoniades, C.A.; Topol, E.J.; et al. Retinal Optical Coherence Tomography Features Associated With Incident and Prevalent Parkinson Disease. Neurology 2023, 101, e1581–e1593. Available online: https://n.neurology.org/content/early/2023/08/21/WNL.0000000000207727 (accessed on 31 August 2023). [CrossRef]
- Scheetz, J.; Rothschild, P.; McGuinness, M.; Hadoux, X.; Soyer, H.P.; Janda, M.; Condon, J.J.J.; Oakden-Rayner, L.; Palmer, L.J.; Keel, S.; et al. A survey of clinicians on the use of artificial intelligence in ophthalmology, dermatology, radiology and radiation oncology. Sci. Rep. 2021, 11, 5193. [Google Scholar] [CrossRef]
- Du-Harpur, X.; Watt, F.M.; Luscombe, N.M.; Lynch, M.D. What is AI? Applications of artificial intelligence to dermatology. Br. J. Dermatol. 2020, 183, 423–430. [Google Scholar] [CrossRef]
- Escalé-Besa, A.; Yélamos, O.; Vidal-Alaball, J.; Fuster-Casanovas, A.; Miró Catalina, Q.; Börve, A.; Ander-Egg Aguilar, R.; Fustà-Novell, X.; Cubiró, X.; Rafat, M.E.; et al. Exploring the potential of artificial intelligence in improving skin lesion diagnosis in primary care. Sci. Rep. 2023, 13, 4293. [Google Scholar] [CrossRef]
- Pan, K.; Hurault, G.; Arulkumaran, K.; Williams, H.C.; Tanaka, R.J. EczemaNet: Automating Detection and Severity Assessment of Atopic Dermatitis. In Machine Learning in Medical Imaging; Liu, M., Yan, P., Lian, C., Cao, X., Eds.; Lecture Notes in Computer Science; Springer International Publishing: Cham, Switzerland, 2020; pp. 220–230. [Google Scholar]
- Ambient Clinical Intelligence|Automatically Document Care|Nuance. Available online: https://www.nuance.com/healthcare/ambient-clinical-intelligence.html (accessed on 4 October 2023).
- Mistry, P. Artificial intelligence in primary care. Br. J. Gen. Pract. 2019, 69, 422–423. [Google Scholar] [CrossRef] [PubMed]
- ChatGPT. Available online: https://chat.openai.com (accessed on 5 October 2023).
- Bard—Chat Based AI Tool from Google, Powered by PaLM 2. Available online: https://bard.google.com (accessed on 20 October 2023).
- Yang, Z.; Li, L.; Lin, K.; Wang, J.; Lin, C.C.; Liu, Z.; Wang, L. The Dawn of LMMs: Preliminary Explorations with GPT-4V(ision). arXiv 2023, arXiv:2309.17421. Available online: http://arxiv.org/abs/2309.17421 (accessed on 16 October 2023).
- Magavern, E.F.; Smedley, D.; Caulfield, M.J. Factor V Leiden, oestrogen and multimorbidity association with venous thromboembolism in a British-South Asian Cohort. iScience 2023, 26, 107795. Available online: https://www.cell.com/iscience/abstract/S2589-0042(23)01872-2 (accessed on 20 September 2023). [CrossRef]
- Pandit, J.A.; Radin, J.M.; Quer, G.; Topol, E.J. Smartphone apps in the COVID-19 pandemic. Nat. Biotechnol. 2022, 40, 1013–1022. [Google Scholar] [CrossRef]
- Generative AI Country Automation. Available online: http://ceros.mckinsey.com/generative-ai-country-automation (accessed on 5 October 2023).
- Pause Giant AI Experiments: An Open Letter. Future of Life Institute. Available online: https://futureoflife.org/open-letter/pause-giant-ai-experiments/ (accessed on 20 October 2023).
- Sanderson, S.; Zimmern, R.; Kroese, M.; Higgins, J.; Patch, C.; Emery, J. How can the evaluation of genetic tests be enhanced? Lessons learned from the ACCE framework and evaluating genetic tests in the United Kingdom. Genet. Med. Off. J. Am. Coll. Med. Genet. 2005, 7, 495–500. [Google Scholar] [CrossRef] [PubMed]
- ACCE Model Process for Evaluating Genetic Tests|CDC. 2022. Available online: https://www.cdc.gov/genomics/gtesting/acce/index.htm (accessed on 20 October 2023).
- WHO Outlines Considerations for Regulation of Artificial Intelligence for Health. Available online: https://www.who.int/news/item/19-10-2023-who-outlines-considerations-for-regulation-of-artificial-intelligence-for-health (accessed on 25 October 2023).
- About the AI and Digital Regulations Service—AI Regulation Service—NHS. Available online: https://www.digitalregulations.innovation.nhs.uk/about-this-service/ (accessed on 25 October 2023).
- de Vries, A. The growing energy footprint of artificial intelligence. Joule 2023, 7, 2191–2194. Available online: https://www.cell.com/joule/abstract/S2542-4351(23)00365-3 (accessed on 12 October 2023). [CrossRef]
- Topol, E.J. High-performance medicine: The convergence of human and artificial intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, D.A.; Witkowski, E.; Gao, L.; Meireles, O.; Rosman, G. Artificial Intelligence in Anesthesiology: Current Techniques, Clinical Applications, and Limitations. Anesthesiology 2020, 132, 379–394. [Google Scholar] [CrossRef]
- Ibrahim, H.; Liu, X.; Zariffa, N.; Morris, A.D.; Denniston, A.K. Health data poverty: An assailable barrier to equitable digital health care. Lancet Digit. Health 2021, 3, e260–e265. [Google Scholar] [CrossRef]
- Fry, A.; Littlejohns, T.J.; Sudlow, C.; Doherty, N.; Adamska, L.; Sprosen, T.; Collins, R.; Allen, N.E. Comparison of Sociodemographic and Health-Related Characteristics of UK Biobank Participants With Those of the General Population. Am. J. Epidemiol. 2017, 186, 1026–1034. [Google Scholar] [CrossRef]
- Arora, A.; Alderman, J.E.; Palmer, J.; Ganapathi, S.; Laws, E.; McCradden, M.D.; Oakden-Rayner, L.; Pfohl, S.R.; Ghassemi, M.; McKay, F.; et al. The value of standards for health datasets in artificial intelligence-based applications. Nat. Med. 2023, 29, 2929–2938. [Google Scholar] [CrossRef] [PubMed]
- Tobias, D.K.; Merino, J.; Ahmad, A.; Aiken, C.; Benham, J.L.; Bodhini, D.; Clark, A.L.; Colclough, K.; Corcoy, R.; Cromer, S.J.; et al. Second international consensus report on gaps and opportunities for the clinical translation of precision diabetes medicine. Nat. Med. 2023, 29, 2438–2457. [Google Scholar] [CrossRef]
- Ganapathi, S.; Palmer, J.; Alderman, J.E.; Calvert, M.; Espinoza, C.; Gath, J.; Ghassemi, M.; Heller, K.; McKay, F.; Karthikesalingaam, A.; et al. Tackling bias in AI health datasets through the STANDING Together initiative. Nat. Med. 2022, 28, 2232–2233. [Google Scholar] [CrossRef]
- Jorgensen, A.L.; Prince, C.; Fitzgerald, G.; Hanson, A.; Downing, J.; Reynolds, J.; Zhang, E.J.; Alfirevic, A.; Pirmohamed, M. Implementation of genotype-guided dosing of warfarin with point-of-care genetic testing in three UK clinics: A matched cohort study. BMC Med. 2019, 17, 76. [Google Scholar] [CrossRef]
- Mogaka, J.J.O.; James, S.E.; Chimbari, M.J. Leveraging implementation science to improve implementation outcomes in precision medicine. Am. J. Transl. Res. 2020, 12, 4853–4872. [Google Scholar] [PubMed]
- Khoury, M.J. No Shortcuts on the Long Road to Evidence-Based Genomic Medicine. JAMA 2017, 318, 27–28. [Google Scholar] [CrossRef] [PubMed]
- Neri, M.; Cubi-Molla, P.; Cookson, G. Approaches to Measure Efficiency in Primary Care: A Systematic Literature Review. Appl. Health Econ. Health Policy 2022, 20, 19–33. [Google Scholar] [CrossRef] [PubMed]
- Milne, H.; Huby, G.; Buckingham, S.; Hayward, J.; Sheikh, A.; Cresswell, K.; Pinnock, H. Does sharing the electronic health record in the consultation enhance patient involvement? A mixed-methods study using multichannel video recording and in-depth interviews in primary care. Health Expect. Int. J. Public Particip. Health Care Health Policy 2016, 19, 602–616. [Google Scholar] [CrossRef]
- Chen, W.; Anothaisintawee, T.; Butani, D.; Wang, Y.; Zemlyanska, Y.; Wong, C.B.N.; Virabhak, S.; Hrishikesh, M.A.; Teerawattananon, Y. Assessing the cost-effectiveness of precision medicine: Protocol for a systematic review and meta-analysis. BMJ Open 2022, 12, e057537. [Google Scholar] [CrossRef] [PubMed]
- Mathur, S.; Sutton, J. Personalized medicine could transform healthcare. Biomed. Rep. 2017, 7, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Evans, B.J.; Flockhart, D.A.; Meslin, E.M. Creating incentives for genomic research to improve targeting of therapies. Nat. Med. 2020, 10, 1289–1291. [Google Scholar] [CrossRef] [PubMed]
- Prasad, V.; De Jesús, K.; Mailankody, S. The high price of anticancer drugs: Origins, implications, barriers, solutions. Nat. Rev. Clin. Oncol. 2017, 14, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Taylor, A.L.; Ziesche, S.; Yancy, C.; Carson, P.; D’Agostino, R.; Ferdinand, K.; Taylor, M.; Adams, K.; Sabolinski, M.; Worcel, M.; et al. Combination of Isosorbide Dinitrate and Hydralazine in Blacks with Heart Failure. N. Engl. J. Med. 2004, 351, 2049–2057. [Google Scholar] [CrossRef]
- Linder, J.E.; Bastarache, L.; Hughey, J.J.; Peterson, J.F. The Role of Electronic Health Records in Advancing Genomic Medicine. Annu Rev. Genom. Hum. Genet. 2021, 22, 219–238. [Google Scholar] [CrossRef]
- Lind, S. NHS England Could Take Co-Data Controller Responsibility for GP Patient Records. Pulse Today. 20 June 2023. Available online: https://www.pulsetoday.co.uk/news/breaking-news/nhs-england-could-take-co-data-controller-responsibility-for-gp-patient-records/ (accessed on 17 October 2023).
- NHS Long Term Plan » Overview and Summary. Available online: https://www.longtermplan.nhs.uk/online-version/overview-and-summary/ (accessed on 3 November 2023).
- Syal, R. Abandoned NHS IT System Has Cost £10bn So Far. The Guardian. 18 September 2013. Available online: https://www.theguardian.com/society/2013/sep/18/nhs-records-system-10bn (accessed on 3 November 2023).
- Topol, E.J. Machines and empathy in medicine. Lancet 2023, 402, 1411. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Evans, W.; Meslin, E.M.; Kai, J.; Qureshi, N. Precision Medicine—Are We There Yet? A Narrative Review of Precision Medicine’s Applicability in Primary Care. J. Pers. Med. 2024, 14, 418. https://doi.org/10.3390/jpm14040418
Evans W, Meslin EM, Kai J, Qureshi N. Precision Medicine—Are We There Yet? A Narrative Review of Precision Medicine’s Applicability in Primary Care. Journal of Personalized Medicine. 2024; 14(4):418. https://doi.org/10.3390/jpm14040418
Chicago/Turabian StyleEvans, William, Eric M. Meslin, Joe Kai, and Nadeem Qureshi. 2024. "Precision Medicine—Are We There Yet? A Narrative Review of Precision Medicine’s Applicability in Primary Care" Journal of Personalized Medicine 14, no. 4: 418. https://doi.org/10.3390/jpm14040418
APA StyleEvans, W., Meslin, E. M., Kai, J., & Qureshi, N. (2024). Precision Medicine—Are We There Yet? A Narrative Review of Precision Medicine’s Applicability in Primary Care. Journal of Personalized Medicine, 14(4), 418. https://doi.org/10.3390/jpm14040418






