Digital Validation in Breast Cancer Needle Biopsies: Comparison of Histological Grade and Biomarker Expression Assessment Using Conventional Light Microscopy, Whole Slide Imaging, and Digital Image Analysis
Abstract
1. Introduction
2. Materials and Methods
2.1. Case Selection and Immunohistochemistry
2.2. Conventional Light Microscope and Pathologist Visual Grading and Scoring
2.3. Slide Digitization, Re-Grading, and Scoring with WSI
2.4. Digital Image Analysis
2.5. Definition of Perfect Concordance, Minor Discordance, and Major Discordance
2.6. Statistical Analysis
3. Results
3.1. Patients and Clinicopathologic Characteristics
3.2. Intra-Observer Concordance and Agreement of Nottingham Grade and Its Components between CLM and WSI
3.3. Inter-Observer Agreement for Nottingham Grade and Its Components in CLM and WSI
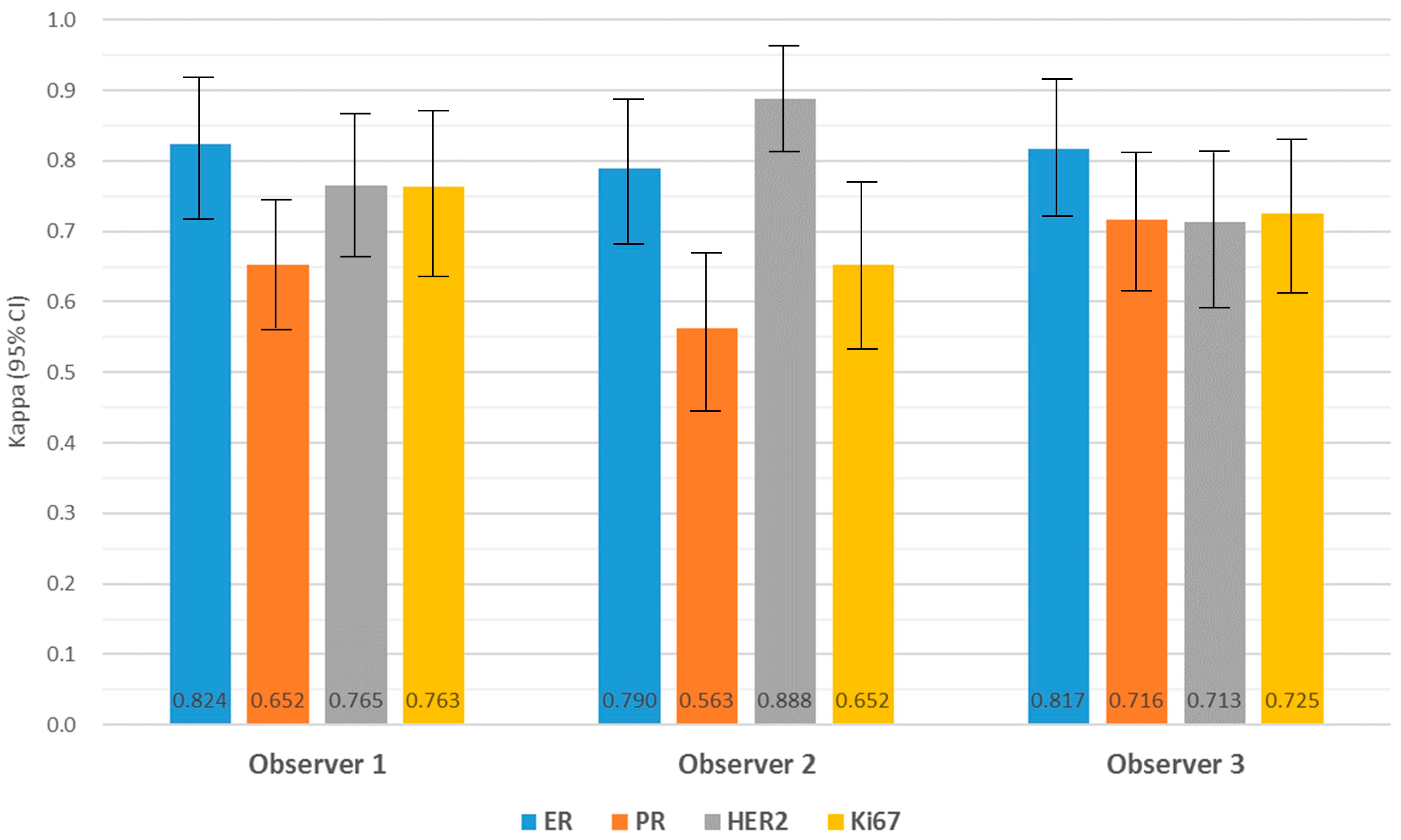
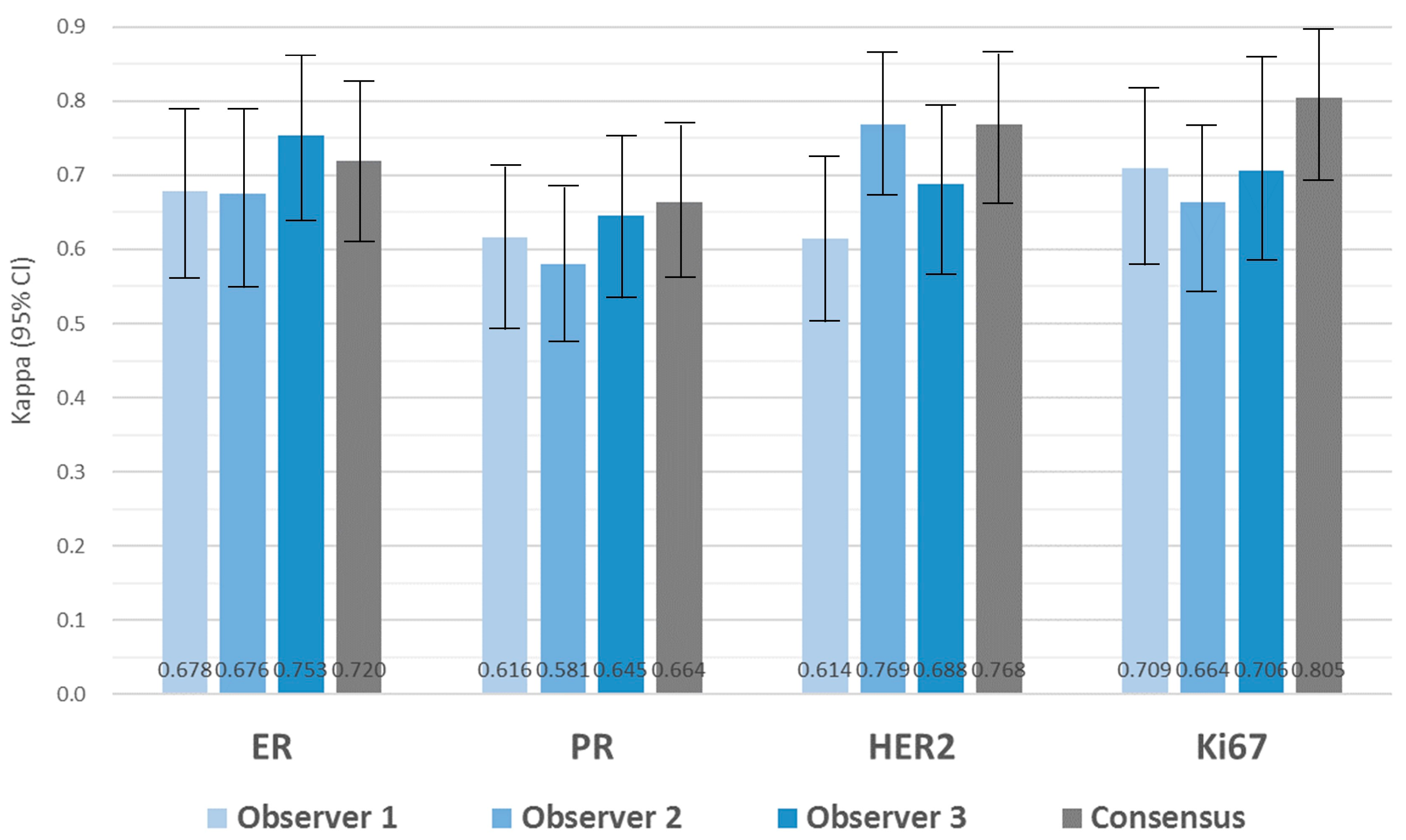
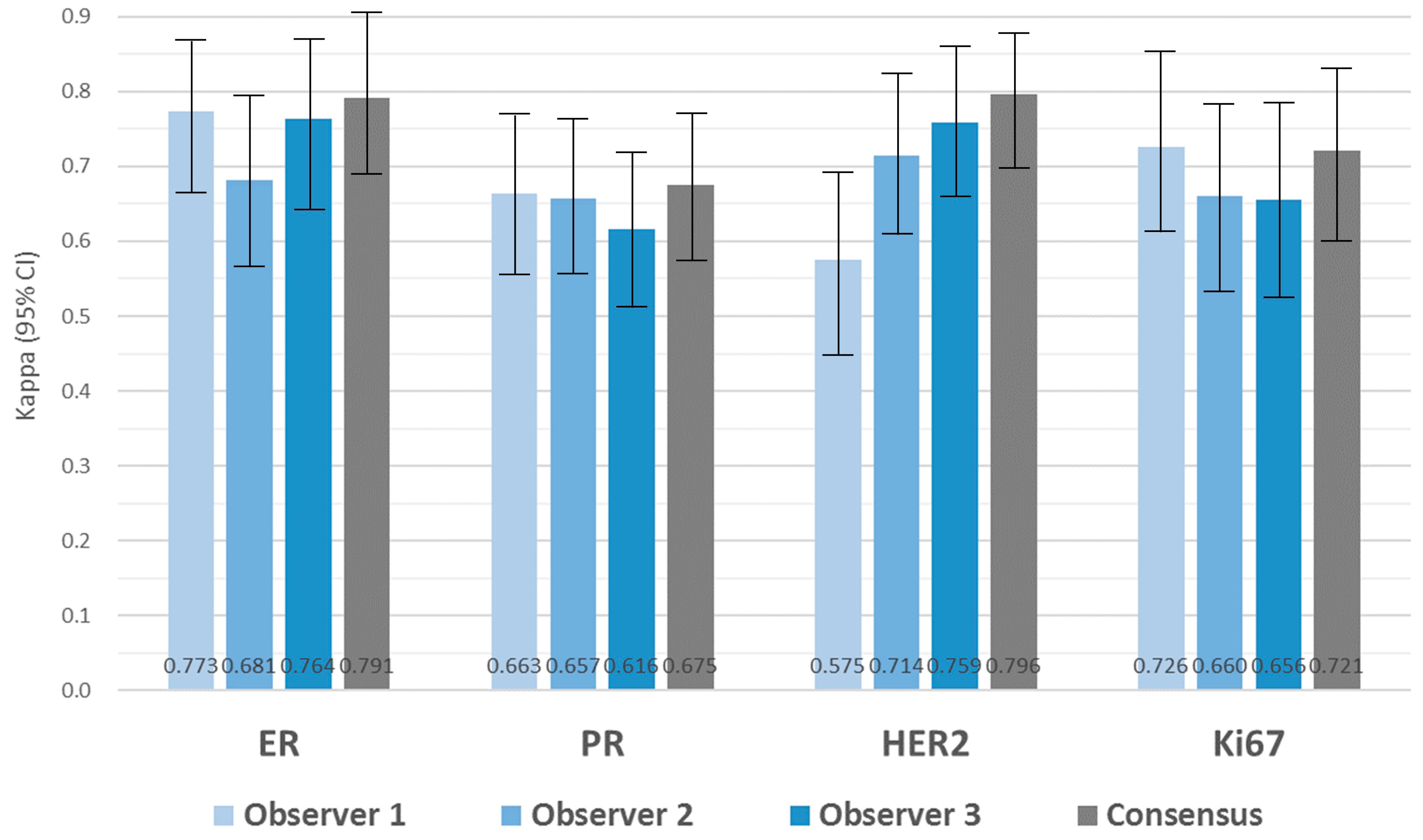
3.4. Agreement and Intra-Observer Variability in Biomarker Expression
3.5. Inter-Observer Variability in Biomarker Expression
3.6. Evaluation of Biomarker Expression with DIA
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Giuliano, A.E.; Connolly, J.L.; Edge, S.B.; Mittendorf, E.A.; Rugo, H.S.; Solin, L.J.; Weaver, D.L.; Winchester, D.J.; Hortobagyi, G.N. Breast Cancer-Major changes in the American Joint Committee on Cancer eighth edition cancer staging manual. CA Cancer J. Clin. 2017, 67, 290–303. [Google Scholar] [CrossRef]
- Allison, K.H. Ancillary Prognostic and Predictive Testing in Breast Cancer: Focus on Discordant, Unusual, and Borderline Results. Surg. Pathol. Clin. 2018, 11, 147–176. [Google Scholar] [CrossRef]
- Nielsen, T.O.; Leung, S.C.Y.; Rimm, D.L.; Dodson, A.; Acs, B.; Badve, S.; Denkert, C.; Ellis, M.J.; Fineberg, S.; Flowers, M.; et al. Assessment of Ki67 in Breast Cancer: Updated Recommendations From the International Ki67 in Breast Cancer Working Group. J. Natl. Cancer Inst. 2021, 113, 808–819. [Google Scholar] [CrossRef]
- Salama, A.M.; Hanna, M.G.; Giri, D.; Kezlarian, B.; Jean, M.H.; Lin, O.; Vallejo, C.; Brogi, E.; Edelweiss, M. Digital validation of breast biomarkers (ER, PR, AR, and HER2) in cytology specimens using three different scanners. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 2022, 35, 52–59. [Google Scholar] [CrossRef]
- Cuzick, J.; Dowsett, M.; Pineda, S.; Wale, C.; Salter, J.; Quinn, E.; Zabaglo, L.; Mallon, E.; Green, A.R.; Ellis, I.O.; et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breast cancer. J. Clin. Oncol. 2011, 29, 4273–4278. [Google Scholar] [CrossRef]
- Goldhirsch, A.; Wood, W.C.; Coates, A.S.; Gelber, R.D.; Thurlimann, B.; Senn, H.J.; Panel, m. Strategies for subtypes—Dealing with the diversity of breast cancer: Highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann. Oncol. 2011, 22, 1736–1747. [Google Scholar] [CrossRef] [PubMed]
- Rakha, E.A.; Aleskandarani, M.; Toss, M.S.; Green, A.R.; Ball, G.; Ellis, I.O.; Dalton, L.W. Breast cancer histologic grading using digital microscopy: Concordance and outcome association. J. Clin. Pathol. 2018, 71, 680–686. [Google Scholar] [CrossRef]
- Weinstein, R.S.; Holcomb, M.J.; Krupinski, E.A. Invention and Early History of Telepathology (1985–2000). J. Pathol. Inform. 2019, 10, 1. [Google Scholar] [CrossRef]
- Williams, B.J.; Lee, J.; Oien, K.A.; Treanor, D. Digital pathology access and usage in the UK: Results from a national survey on behalf of the National Cancer Research Institute’s CM-Path initiative. J. Clin. Pathol. 2018, 71, 463–466. [Google Scholar] [CrossRef]
- Pantanowitz, L.; Sinard, J.H.; Henricks, W.H.; Fatheree, L.A.; Carter, A.B.; Contis, L.; Beckwith, B.A.; Evans, A.J.; Lal, A.; Parwani, A.V. Validating whole slide imaging for diagnostic purposes in pathology: Guideline from the College of American Pathologists Pathology and Laboratory Quality Center. Arch. Pathol. Lab. Med. 2013, 137, 1710–1722. [Google Scholar] [CrossRef]
- Cornish, T.C.; Swapp, R.E.; Kaplan, K.J. Whole-slide imaging: Routine pathologic diagnosis. Adv. Anat. Pathol. 2012, 19, 152–159. [Google Scholar] [CrossRef]
- Lange, H. Digital Pathology: A Regulatory Overview. Lab. Med. 2011, 42, 587–591. [Google Scholar] [CrossRef][Green Version]
- Evans, A.J.; Brown, R.W.; Bui, M.M.; Chlipala, E.A.; Lacchetti, C.; Milner, D.A.; Pantanowitz, L.; Parwani, A.V.; Reid, K.; Riben, M.W.; et al. Validating Whole Slide Imaging Systems for Diagnostic Purposes in Pathology. Arch. Pathol. Lab. Med. 2022, 146, 440–450. [Google Scholar] [CrossRef]
- Reyes, C.; Ikpatt, O.F.; Nadji, M.; Cote, R.J. Intra-observer reproducibility of whole slide imaging for the primary diagnosis of breast needle biopsies. J. Pathol. Inform. 2014, 5, 5. [Google Scholar] [CrossRef]
- Krishnamurthy, S.; Mathews, K.; McClure, S.; Murray, M.; Gilcrease, M.; Albarracin, C.; Spinosa, J.; Chang, B.; Ho, J.; Holt, J.; et al. Multi-institutional comparison of whole slide digital imaging and optical microscopy for interpretation of hematoxylin-eosin-stained breast tissue sections. Arch. Pathol. Lab. Med. 2013, 137, 1733–1739. [Google Scholar] [CrossRef]
- Araújo, A.L.D.; Arboleda, L.P.A.; Palmier, N.R.; Fonsêca, J.M.; de Pauli Paglioni, M.; Gomes-Silva, W.; Ribeiro, A.C.P.; Brandão, T.B.; Simonato, L.E.; Speight, P.M.; et al. The performance of digital microscopy for primary diagnosis in human pathology: A systematic review. Virchows Arch. 2019, 474, 269–287. [Google Scholar] [CrossRef]
- Stålhammar, G.; Fuentes Martinez, N.; Lippert, M.; Tobin, N.P.; Mølholm, I.; Kis, L.; Rosin, G.; Rantalainen, M.; Pedersen, L.; Bergh, J.; et al. Digital image analysis outperforms manual biomarker assessment in breast cancer. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 2016, 29, 318–329. [Google Scholar] [CrossRef]
- Al-Janabi, S.; Huisman, A.; Van Diest, P.J. Digital pathology: Current status and future perspectives. Histopathology 2012, 61, 1–9. [Google Scholar] [CrossRef]
- Yousif, M.; Huang, Y.; Sciallis, A.; Kleer, C.G.; Pang, J.; Smola, B.; Naik, K.; McClintock, D.S.; Zhao, L.; Kunju, L.P.; et al. Quantitative Image Analysis as an Adjunct to Manual Scoring of ER, PgR, and HER2 in Invasive Breast Carcinoma. Am. J. Clin. Pathol. 2022, 157, 899–907. [Google Scholar] [CrossRef]
- Di Loreto, C.; Puglisi, F.; Rimondi, G.; Zuiani, C.; Anania, G.; Della Mea, V.; Beltrami, C.A. Large core biopsy for diagnostic and prognostic evaluation of invasive breast carcinomas. Eur. J. Cancer 1996, 32a, 1693–1700. [Google Scholar] [CrossRef]
- Rakha, E.A.; Allison, K.H.; Ellis, I.O.; Penault-Llorca, F.; Vincent-Salomon, A.; Masuda, S.; Tsuda, H.; Horii, R. Invasive breast carcinoma: General overview. In WHO Classification of Tumours. Breast Tumours, 5th ed.; Allison, K.H., Salgado, R., Eds.; International Agency for Research on Cancer: Lyon, France, 2019. [Google Scholar]
- Rossi, C.; Fraticelli, S.; Fanizza, M.; Ferrari, A.; Ferraris, E.; Messina, A.; Della Valle, A.; Anghelone, C.A.P.; Lasagna, A.; Rizzo, G.; et al. Concordance of immunohistochemistry for predictive and prognostic factors in breast cancer between biopsy and surgical excision: A single-centre experience and review of the literature. Breast Cancer Res. Treat. 2023, 198, 573–582. [Google Scholar] [CrossRef]
- Hammond, M.E.; Hayes, D.F.; Dowsett, M.; Allred, D.C.; Hagerty, K.L.; Badve, S.; Fitzgibbons, P.L.; Francis, G.; Goldstein, N.S.; Hayes, M.; et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J. Clin. Oncol. 2010, 28, 2784–2795. [Google Scholar] [CrossRef]
- Elston, C.W.; Ellis, I.O. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: Experience from a large study with long-term follow-up. Histopathology 1991, 19, 403–410. [Google Scholar] [CrossRef]
- Allred, D.C.; Harvey, J.M.; Berardo, M.; Clark, G.M. Prognostic and predictive factors in breast cancer by immunohistochemical analysis. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 1998, 11, 155–168. [Google Scholar]
- Wolff, A.C.; Hammond, M.E.H.; Allison, K.H.; Harvey, B.E.; Mangu, P.B.; Bartlett, J.M.S.; Bilous, M.; Ellis, I.O.; Fitzgibbons, P.; Hanna, W.; et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. Arch. Pathol. Lab. Med. 2018, 142, 1364–1382. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.; Kim, D.C.; Jung, C.K.; Kim, D.C.; Song, S.Y.; Joo, H.J.; Yi, S.Y. Recommendations for pathologic practice using digital pathology: Consensus report of the Korean Society of Pathologists. J. Pathol. Transl. Med. 2020, 54, 437–452. [Google Scholar] [CrossRef] [PubMed]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [PubMed]
- Hortobagyi, G.N.; Connolly, J.L.; D’Orsi, C.J.; Edge, S.B.; Mittendorf, E.A.; Rugo, H.S.; Solin, L.J.; Weaver, D.L.; Winchester, D.J.; Giuliano, A. AJCC Cancer Staging Manual 8th edition. In American Joint Committee on Cancer, 8th ed.; American Joint Committee on Cancer, Ed.; Springer: Chicago, IL, USA, 2017; pp. 589–628. [Google Scholar]
- Schwartz, A.M.; Henson, D.E.; Chen, D.; Rajamarthandan, S. Histologic grade remains a prognostic factor for breast cancer regardless of the number of positive lymph nodes and tumor size: A study of 161 708 cases of breast cancer from the SEER Program. Arch. Pathol. Lab. Med. 2014, 138, 1048–1052. [Google Scholar] [CrossRef] [PubMed]
- Meyer, J.S.; Alvarez, C.; Milikowski, C.; Olson, N.; Russo, I.; Russo, J.; Glass, A.; Zehnbauer, B.A.; Lister, K.; Parwaresch, R. Breast carcinoma malignancy grading by Bloom-Richardson system vs proliferation index: Reproducibility of grade and advantages of proliferation index. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 2005, 18, 1067–1078. [Google Scholar] [CrossRef]
- Longacre, T.A.; Ennis, M.; Quenneville, L.A.; Bane, A.L.; Bleiweiss, I.J.; Carter, B.A.; Catelano, E.; Hendrickson, M.R.; Hibshoosh, H.; Layfield, L.J.; et al. Interobserver agreement and reproducibility in classification of invasive breast carcinoma: An NCI breast cancer family registry study. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 2006, 19, 195–207. [Google Scholar] [CrossRef]
- Rakha, E.A.; Bennett, R.L.; Coleman, D.; Pinder, S.E.; Ellis, I.O. Review of the national external quality assessment (EQA) scheme for breast pathology in the UK. J. Clin. Pathol. 2017, 70, 51–57. [Google Scholar] [CrossRef]
- Davidson, T.M.; Rendi, M.H.; Frederick, P.D.; Onega, T.; Allison, K.H.; Mercan, E.; Brunyé, T.T.; Shapiro, L.G.; Weaver, D.L.; Elmore, J.G. Breast Cancer Prognostic Factors in the Digital Era: Comparison of Nottingham Grade using Whole Slide Images and Glass Slides. J. Pathol. Inform. 2019, 10, 11. [Google Scholar] [CrossRef]
- Ginter, P.S.; Idress, R.; D’Alfonso, T.M.; Fineberg, S.; Jaffer, S.; Sattar, A.K.; Chagpar, A.; Wilson, P.; Harigopal, M. Histologic grading of breast carcinoma: A multi-institution study of interobserver variation using virtual microscopy. Mod. Pathol. Off. J. United States Can. Acad. Pathol. Inc. 2021, 34, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Shaw, E.C.; Hanby, A.M.; Wheeler, K.; Shaaban, A.M.; Poller, D.; Barton, S.; Treanor, D.; Fulford, L.; Walker, R.A.; Ryan, D.; et al. Observer agreement comparing the use of virtual slides with glass slides in the pathology review component of the POSH breast cancer cohort study. J. Clin. Pathol. 2012, 65, 403–408. [Google Scholar] [CrossRef] [PubMed]
- Rakha, E.A.; Aleskandarany, M.A.; Toss, M.S.; Mongan, N.P.; ElSayed, M.E.; Green, A.R.; Ellis, I.O.; Dalton, L.W. Impact of breast cancer grade discordance on prediction of outcome. Histopathology 2018, 73, 904–915. [Google Scholar] [CrossRef] [PubMed]
- Kondo, Y.; Iijima, T.; Noguchi, M. Evaluation of immunohistochemical staining using whole-slide imaging for HER2 scoring of breast cancer in comparison with real glass slides. Pathol. Int. 2012, 62, 592–599. [Google Scholar] [CrossRef]
- Nassar, A.; Cohen, C.; Albitar, M.; Agersborg, S.S.; Zhou, W.; Lynch, K.A.; Heyman, E.R.; Lange, H.; Siddiqui, M.T. Reading immunohistochemical slides on a computer monitor—A multisite performance study using 180 HER2-stained breast carcinomas. Appl. Immunohistochem. Mol. Morphol. 2011, 19, 212–217. [Google Scholar] [CrossRef]
- Cserni, B.; Bori, R.; Csörgő, E.; Oláh-Németh, O.; Pancsa, T.; Sejben, A.; Sejben, I.; Vörös, A.; Zombori, T.; Nyári, T.; et al. The additional value of ONEST (Observers Needed to Evaluate Subjective Tests) in assessing reproducibility of oestrogen receptor, progesterone receptor, and Ki67 classification in breast cancer. Virchows Arch. 2021, 479, 1101–1109. [Google Scholar] [CrossRef]
- Regitnig, P.; Reiner, A.; Dinges, H.P.; Höfler, G.; Müller-Holzner, E.; Lax, S.F.; Obrist, P.; Rudas, M.; Quehenberger, F. Quality assurance for detection of estrogen and progesterone receptors by immunohistochemistry in Austrian pathology laboratories. Virchows Arch. 2002, 441, 328–334. [Google Scholar] [CrossRef]
- Baird, R.D.; Carroll, J.S. Understanding Oestrogen Receptor Function in Breast Cancer and its Interaction with the Progesterone Receptor. New Preclinical Findings and their Clinical Implications. Clin. Oncol. (R. Coll. Radiol.) 2016, 28, 1–3. [Google Scholar] [CrossRef][Green Version]
- Wells, C.A.; Sloane, J.P.; Coleman, D.; Munt, C.; Amendoeira, I.; Apostolikas, N.; Bellocq, J.P.; Bianchi, S.; Boecker, W.; Bussolati, G.; et al. Consistency of staining and reporting of oestrogen receptor immunocytochemistry within the European Union--an inter-laboratory study. Virchows Arch. 2004, 445, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Pu, T.; Shui, R.; Shi, J.; Liang, Z.; Yang, W.; Bu, H.; Li, Q.; Zhang, Z. External quality assessment (EQA) program for the immunohistochemical detection of ER, PR and Ki-67 in breast cancer: Results of an interlaboratory reproducibility ring study in China. BMC Cancer 2019, 19, 978. [Google Scholar] [CrossRef]
- Inamura, K. Update on Immunohistochemistry for the Diagnosis of Lung Cancer. Cancers 2018, 10, 72. [Google Scholar] [CrossRef] [PubMed]
- Stålhammar, G.; Rosin, G.; Fredriksson, I.; Bergh, J.; Hartman, J. Low concordance of biomarkers in histopathological and cytological material from breast cancer. Histopathology 2014, 64, 971–980. [Google Scholar] [CrossRef] [PubMed]
- Bera, K.; Schalper, K.A.; Rimm, D.L.; Velcheti, V.; Madabhushi, A. Artificial intelligence in digital pathology—New tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 2019, 16, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Bui, M.M.; Riben, M.W.; Allison, K.H.; Chlipala, E.; Colasacco, C.; Kahn, A.G.; Lacchetti, C.; Madabhushi, A.; Pantanowitz, L.; Salama, M.E.; et al. Quantitative Image Analysis of Human Epidermal Growth Factor Receptor 2 Immunohistochemistry for Breast Cancer: Guideline From the College of American Pathologists. Arch. Pathol. Lab. Med. 2019, 143, 1180–1195. [Google Scholar] [CrossRef]
- Skaland, I.; Ovestad, I.; Janssen, E.A.; Klos, J.; Kjellevold, K.H.; Helliesen, T.; Baak, J.P. Digital image analysis improves the quality of subjective HER-2 expression scoring in breast cancer. Appl. Immunohistochem. Mol. Morphol. 2008, 16, 185–190. [Google Scholar] [CrossRef]
- Mukhopadhyay, S.; Feldman, M.D.; Abels, E.; Ashfaq, R.; Beltaifa, S.; Cacciabeve, N.G.; Cathro, H.P.; Cheng, L.; Cooper, K.; Dickey, G.E.; et al. Whole Slide Imaging Versus Microscopy for Primary Diagnosis in Surgical Pathology: A Multicenter Blinded Randomized Noninferiority Study of 1992 Cases (Pivotal Study). Am. J. Surg. Pathol. 2018, 42, 39–52. [Google Scholar] [CrossRef]
- Pekmezci, M.; Uysal, S.P.; Orhan, Y.; Tihan, T.; Lee, H.S. Pitfalls in the use of whole slide imaging for the diagnosis of central nervous system tumors: A pilot study in surgical neuropathology. J. Pathol. Inform. 2016, 7, 25. [Google Scholar] [CrossRef]
| Perfect Concordance | Minor Discordance | Major Discordance | |
|---|---|---|---|
| Observer 1 | |||
| Nottingham grade | 78 (77.2%) | 23 (22.8%) | 0 (0.0%) |
| Tubule formation | 79 (78.2%) | 22 (21.8%) | 0 (0.0%) |
| Nuclear pleomorphism | 74 (73.3%) | 27 (27.7%) | 0 (0.0%) |
| Mitotic counts | 87 (86.1%) | 14 (13.9%) | 0 (0.0%) |
| Observer 2 | |||
| Nottingham grade | 81 (80.2%) | 20 (19.8%) | 0 (0.0%) |
| Tubule formation | 83 (82.2%) | 18 (17.8%) | 0 (0.0%) |
| Nuclear pleomorphism | 78 (77.2%) | 23 (22.8%) | 0 (0.0%) |
| Mitotic counts | 84 (83.2%) | 13 (12.9%) | 4 (3.9%) |
| Observer 3 | |||
| Nottingham grade | 77 (76.2%) | 24 (23.8%) | 0 (0.0%) |
| Tubule formation | 84 (83.2%) | 17 (16.8%) | 0 (0.0%) |
| Nuclear pleomorphism | 65 (64.4%) | 36 (35.6%) | 0 (0.0%) |
| Mitotic counts | 84 (83.2%) | 17 (16.8%) | 0 (0.0%) |
| Observer 1 | Observer 2 | Observer 3 | ||||
|---|---|---|---|---|---|---|
| Z-Score | p-Value | Z-Score | p | Z-Score | p-Value | |
| Nottingham grade | −2.294 | 0.022 | −1.342 | 0.180 | −0.816 | 0.541 |
| Tubule formation | −4.264 | <0.001 | −3.771 | <0.001 | −3.638 | <0.001 |
| Nuclear pleomorphism | −0.557 | 0.564 | −3.545 | <0.001 | −2.667 | 0.008 |
| Mitotic counts | 0.000 | 1.000 | −0.876 | 0.381 | −1.231 | 0.225 |
| CLM | WSI | |||
|---|---|---|---|---|
| Fleiss Kappa (95% CI) | p-Value | Fleiss Kappa (95% CI) | p-Value | |
| Nottingham grade | 0.630 (0.628–0.633) | <0.001 | 0.620 (0.618–0.623) | <0.001 |
| Tubule formation | 0.543 (0.540–0.546) | <0.001 | 0.523 (0.519–0.526) | <0.001 |
| Nuclear pleomorphism | 0.356 (0.353–0.359) | <0.001 | 0.394 (0.391–0.397) | <0.001 |
| Mitotic counts | 0.654 (0.651–0.657) | <0.001 | 0.720 (0.717–0.723) | <0.001 |
| Perfect Concordance | Minor Discordance | Major Discordance | |
|---|---|---|---|
| Observer 1 | |||
| ER | 92 (91.0%) | 5 (5.0%) | 4 (4.0%) |
| PR | 74 (73.2%) | 23 (22.8%) | 4 (4.0%) |
| HER2 | 85 (84.1%) | 5 (5.0%) | 11 (10.9%) |
| Ki67 | 86 (85.2%) | 15 (14.8%) | 0 (0.0%) |
| Observer 2 | |||
| ER | 89 (88.1%) | 9 (8.9%) | 3 (3.0%) |
| PR | 66 (65.3%) | 32 (31.7%) | 3 (3.0%) |
| HER2 | 93 (92.0%) | 6 (6.0%) | 2 (2.0%) |
| Ki67 | 78 (77.3%) | 23 (22.7%) | 0 (0.0%) |
| Observer 3 | |||
| ER | 92 (91.1%) | 8 (7.9%) | 1 (1.0%) |
| PR | 80 (79.2%) | 19 (18.8%) | 2 (2.0%) |
| HER2 | 80 (79.2%) | 11 (10.9%) | 10 (9.9%) |
| Ki67 | 84 (83.2%) | 17 (16.8%) | 0 (0.0%) |
| Observer 1 | Observer 2 | Observer 3 | ||||
|---|---|---|---|---|---|---|
| Z-Score | p-Value | Z-Score | p | Z-Score | p-Value | |
| ER | −0.420 | 0.674 | −0.243 | 0.808 | −0.490 | 0.624 |
| PR | −0.596 | 0.551 | −1.553 | 0.120 | −0.600 | 0.549 |
| HER2 | −0.688 | 0.491 | 0.000 | 1.000 | −1.528 | 0.127 |
| Ki67 | −0.258 | 0.796 | −1.877 | 0.061 | −1.698 | 0.090 |
| CLM | WSI | |||
|---|---|---|---|---|
| Fleiss Kappa (95% CI) | p-Value | Fleiss Kappa (95% CI) | p-Value | |
| ER | 0.792 (0.790–0.795) | <0.001 | 0.783 (0.781–0.786) | <0.001 |
| PR | 0.598 (0.596–0.600) | <0.001 | 0.648 (0.646–0.650) | <0.001 |
| HER2 | 0.680 (0.678–0.683) | <0.001 | 0.618 (0.615–0.620) | <0.001 |
| Ki67 | 0.577 (0.575–0.580) | <0.001 | 0.642 (0.639–0.644) | <0.001 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choi, J.E.; Kim, K.-H.; Lee, Y.; Kang, D.-W. Digital Validation in Breast Cancer Needle Biopsies: Comparison of Histological Grade and Biomarker Expression Assessment Using Conventional Light Microscopy, Whole Slide Imaging, and Digital Image Analysis. J. Pers. Med. 2024, 14, 312. https://doi.org/10.3390/jpm14030312
Choi JE, Kim K-H, Lee Y, Kang D-W. Digital Validation in Breast Cancer Needle Biopsies: Comparison of Histological Grade and Biomarker Expression Assessment Using Conventional Light Microscopy, Whole Slide Imaging, and Digital Image Analysis. Journal of Personalized Medicine. 2024; 14(3):312. https://doi.org/10.3390/jpm14030312
Chicago/Turabian StyleChoi, Ji Eun, Kyung-Hee Kim, Younju Lee, and Dong-Wook Kang. 2024. "Digital Validation in Breast Cancer Needle Biopsies: Comparison of Histological Grade and Biomarker Expression Assessment Using Conventional Light Microscopy, Whole Slide Imaging, and Digital Image Analysis" Journal of Personalized Medicine 14, no. 3: 312. https://doi.org/10.3390/jpm14030312
APA StyleChoi, J. E., Kim, K.-H., Lee, Y., & Kang, D.-W. (2024). Digital Validation in Breast Cancer Needle Biopsies: Comparison of Histological Grade and Biomarker Expression Assessment Using Conventional Light Microscopy, Whole Slide Imaging, and Digital Image Analysis. Journal of Personalized Medicine, 14(3), 312. https://doi.org/10.3390/jpm14030312








