Short- and Long-Term Effects of a Prebiotic Intervention with Polyphenols Extracted from European Black Elderberry—Sustained Expansion of Akkermansia spp.
Abstract
:1. Introduction
2. Materials and Methods
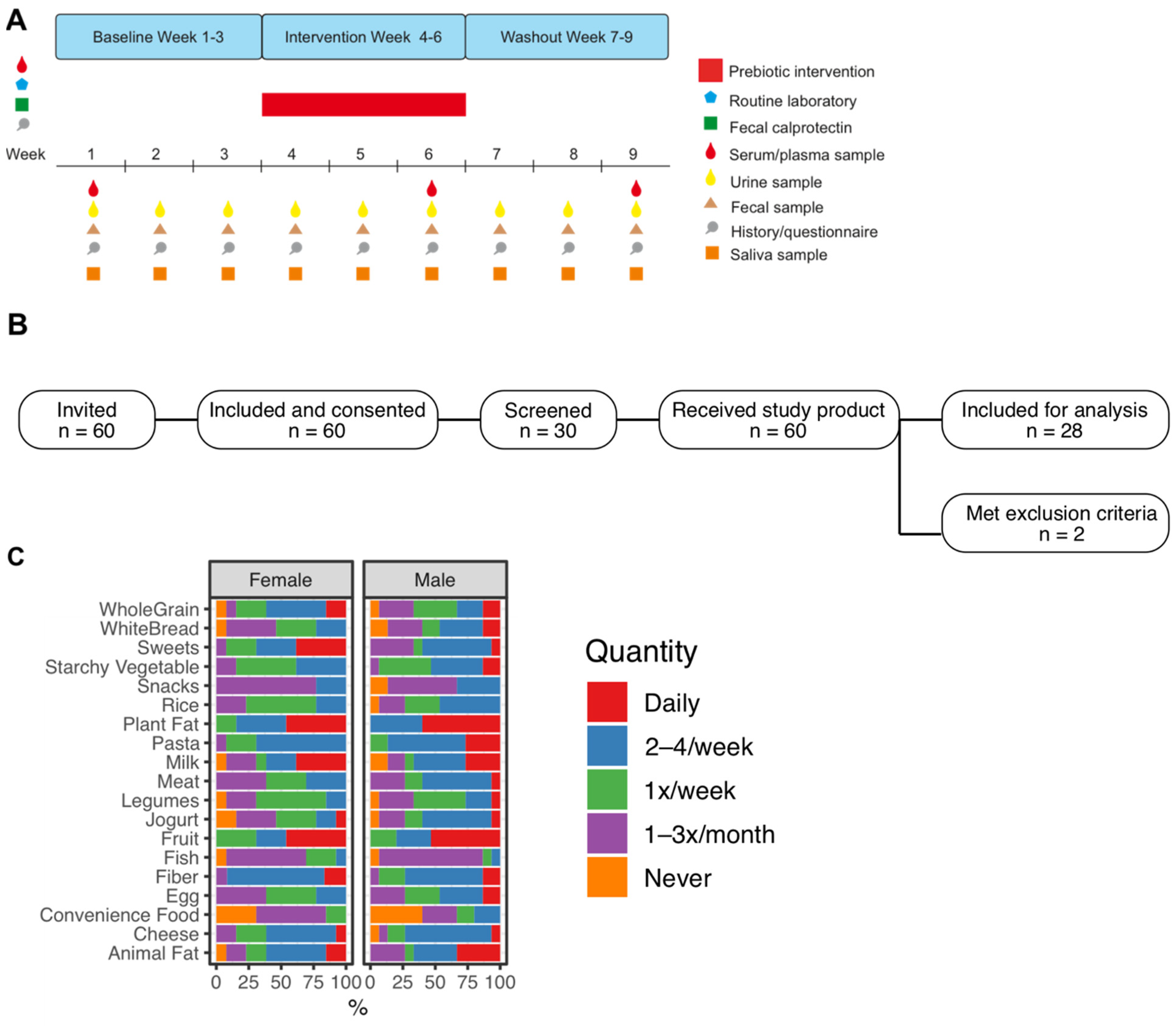
2.1. Trial Design and Study Cohort
2.2. Intervention
2.3. Biosample Acquisition and Biobanking
2.4. Fecal DNA Extraction
2.5. Metagenomic Analysis
2.6. Statistics
3. Results
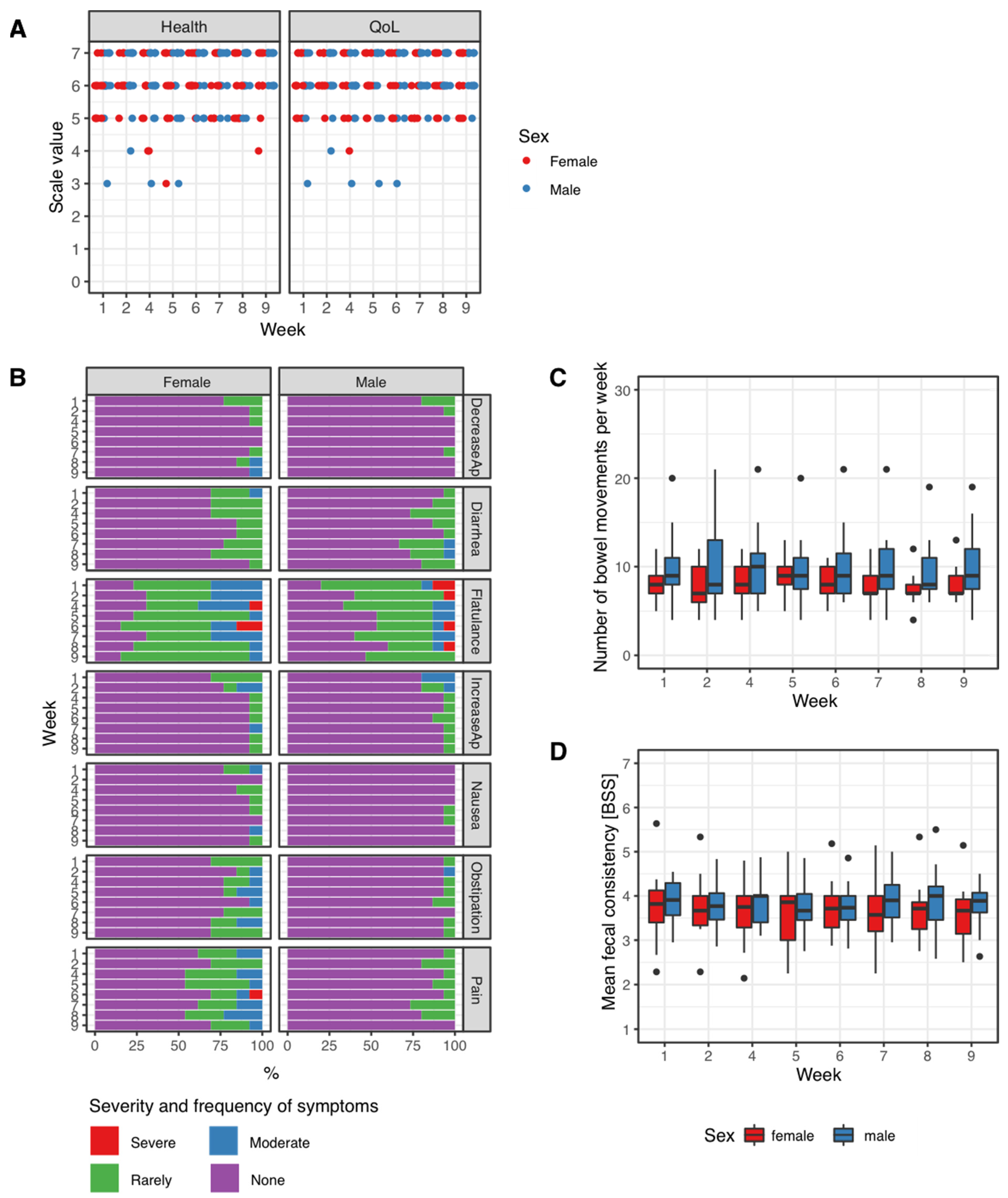
3.1. Patient-Reported Outcomes during the Trial Period
3.2. Alterations in 16S Amplicon-Based Metagenomic Profiles at the Beginning and after the End of the Prebiotic Intervention
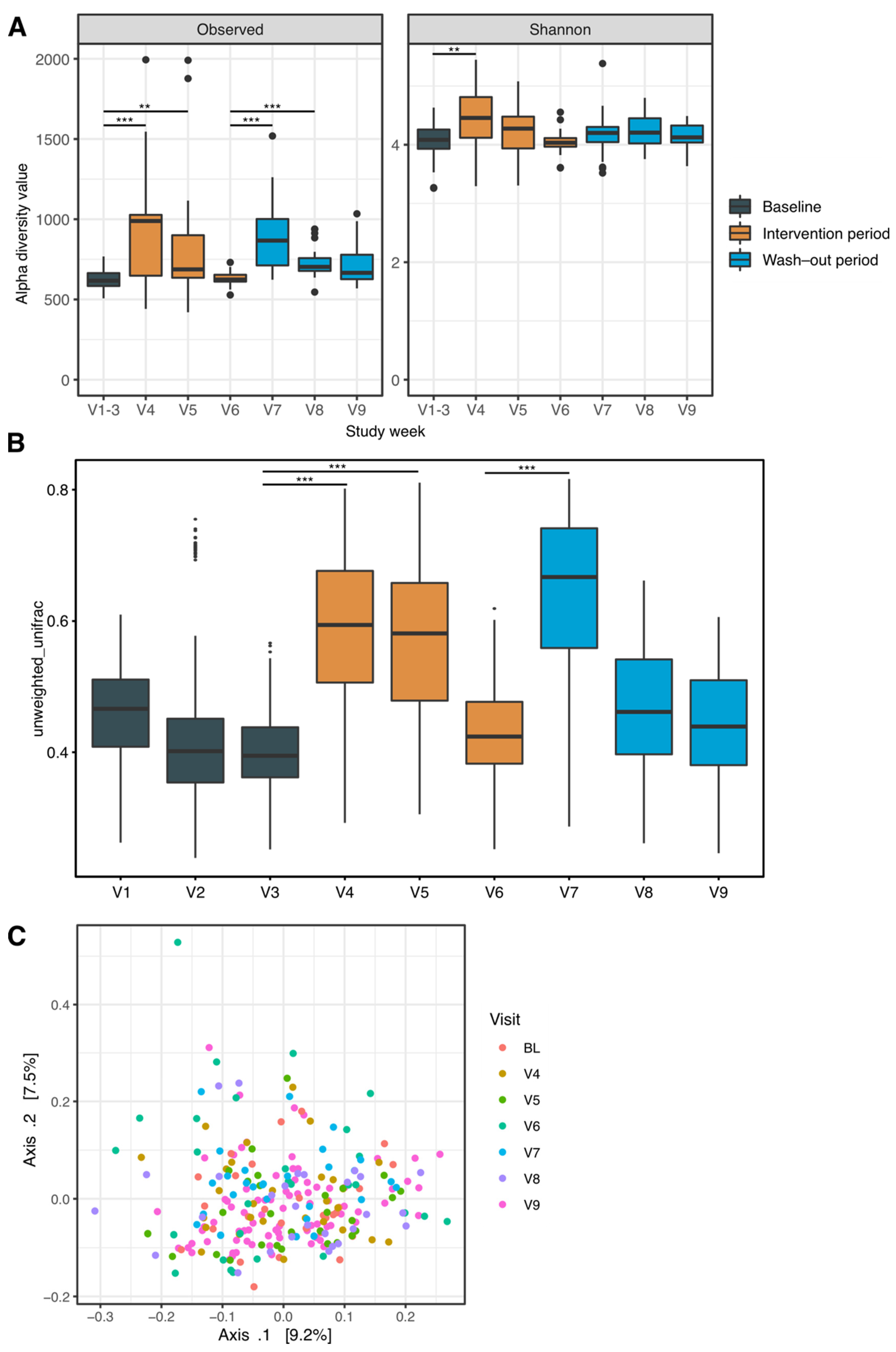
3.3. Changes in Intra-Sample Diversity
3.4. Compositional Differences over Time
3.5. Alterations on a Species Level Driving Overall Changes in Bacterial Ecology
3.6. Factors Determining Response to the Intervention
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brüssow, H. Problems with the concept of gut microbiota dysbiosis. Microb. Biotechnol. 2020, 13, 423–434. [Google Scholar] [CrossRef]
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Levy, M.; Kolodziejczyk, A.A.; Thaiss, C.A.; Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 2017, 17, 219–232. [Google Scholar] [CrossRef] [PubMed]
- Claesson, M.J.; Jeffery, I.B.; Conde, S.; Power, S.E.; O’Connor, E.M.; Cusack, S.; Harris, H.M.; Coakley, M.; Lakshminarayanan, B.; O’Sullivan, O.; et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 2012, 488, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, J.; Lange, B.; Frick, J.S.; Sauer, H.; Zimmermann, K.; Schwiertz, A.; Rusch, K.; Klosterhalfen, S.; Enck, P. A vegan or vegetarian diet substantially alters the human colonic faecal microbiota. Eur. J. Clin. Nutr. 2012, 66, 53–60. [Google Scholar] [CrossRef]
- Lozupone, C.A.; Stombaugh, J.I.; Gordon, J.I.; Jansson, J.K.; Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 2012, 489, 220–230. [Google Scholar] [CrossRef]
- Gibson, G.R.; Hutkins, R.; Sanders, M.E.; Prescott, S.L.; Reimer, R.A.; Salminen, S.J.; Scott, K.; Stanton, C.; Swanson, K.S.; Cani, P.D.; et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 491–502. [Google Scholar] [CrossRef]
- Plamada, D.; Vodnar, D.C. Polyphenols-Gut Microbiota Interrelationship: A Transition to a New Generation of Prebiotics. Nutrients 2022, 14, 137. [Google Scholar]
- Nazzaro, F.; Fratianni, F.; De Feo, V.; Battistelli, A.; Da Cruz, A.G.; Coppola, R. Chapter Two-Polyphenols, the new frontiers of prebiotics. In Advances in Food and Nutrition Research; da Cruz, A.G., Prudencio, E.S., Esmerino, E.A., da Silva, M.C., Eds.; Academic Press: Cambridge, MA, USA, 2020; Volume 94, pp. 35–89. [Google Scholar]
- Thilakarathna, W.P.D.W.; Langille, M.G.I.; Rupasinghe, H.P.V. Polyphenol-based prebiotics and synbiotics: Potential for cancer chemoprevention. Curr. Opin. Food Sci. 2018, 20, 51–57. [Google Scholar] [CrossRef]
- Dueñas, M.; Muñoz-González, I.; Cueva, C.; Jiménez-Girón, A.; Sánchez-Patán, F.; Santos-Buelga, C.; Moreno-Arribas, M.V.; Bartolomé, B. A survey of modulation of gut microbiota by dietary polyphenols. Biomed. Res. Int. 2015, 2015, 850902. [Google Scholar] [CrossRef] [Green Version]
- Netzel, M.; Strass, G.; Herbst, M.; Dietrich, H.; Bitsch, R.; Bitsch, I.; Frank, T. The excretion and biological antioxidant activity of elderberry antioxidants in healthy humans. Food Res. Int. 2005, 38, 905–910. [Google Scholar] [CrossRef]
- Bitsch, I.; Janßen, M.; Netzel, M.; Straß, G.; Frank, T. Bioavailability of anthocyanidin-3-glycosides following consumption of elderberry extract and blackcurrant juice. Int. J. Clin. Pharmacol. Ther. 2004, 42, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Frank, T.; Janßen, M.; Netzel, G.; Christian, B.; Bitsch, I.; Netzel, M. Absorption and excretion of elderberry (Sambucus nigra L.) anthocyanins in healthy humans. Methods Find. Exp. Clin. Pharmacol. 2007, 29, 525–533. [Google Scholar] [CrossRef]
- Frank, T.; Sonntag, S.; Strass, G.; Bitsch, I.; Bitsch, R.; Netzel, M. Urinary pharmacokinetics of cyanidin glycosides in healthy young men following consumption of elderberry juice. Int. J. Clin. Pharmacol. Res. 2005, 25, 47–56. [Google Scholar] [PubMed]
- Milbury, P.E.; Cao, G.; Prior, R.L.; Blumberg, J. Bioavailablility of elderberry anthocyanins. Mech. Ageing Dev. 2002, 123, 997–1006. [Google Scholar] [CrossRef]
- Czank, C.; Cassidy, A.; Zhang, Q.; Morrison, D.J.; Preston, T.; Kroon, P.A.; Botting, N.P.; Kay, C.D. Human metabolism and elimination of the anthocyanin, cyanidin-3-glucoside: A 13C-tracer study. Am. J. Clin. Nutr. 2013, 97, 995–1003. [Google Scholar] [CrossRef]
- Mülleder, U.; Murkovic, M.; Pfannhauser, W. Urinary excretion of cyanidin glycosides. J. Biochem. Biophys. Methods 2002, 53, 61–66. [Google Scholar] [CrossRef]
- Marín, L.; Miguélez, E.M.; Villar, C.J.; Lombó, F. Bioavailability of dietary polyphenols and gut microbiota metabolism: Antimicrobial properties. Biomed. Res. Int. 2015, 2015, 905215. [Google Scholar] [CrossRef]
- Aura, A.-M.; Mattila, I.; Seppänen-Laakso, T.; Miettinen, J.; Oksman-Caldentey, K.-M.; Orešič, M. Microbial metabolism of catechin stereoisomers by human faecal microbiota: Comparison of targeted analysis and a non-targeted metabolomics method. Phytochem. Lett. 2008, 1, 18–22. [Google Scholar] [CrossRef]
- Vlachojannis, J.E.; Cameron, M.; Chrubasik, S. A systematic review on the sambuci fructus effect and efficacy profiles. Phytother. Res. PTR 2010, 24, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Youdim, K.A.; Martin, A.; Joseph, J.A. Incorporation of the elderberry anthocyanins by endothelial cells increases protection against oxidative stress. Free. Radic. Biol. Med. 2000, 29, 51–60. [Google Scholar] [CrossRef]
- Gwiazdowska, D.; Juś, K.; Jasnowska-Małecka, J.; Kluczyńska, K. The impact of polyphenols on Bifidobacterium growth. Acta Biochim. Pol. 2015, 62, 895–901. [Google Scholar] [CrossRef] [PubMed]
- Roopchand, D.E.; Carmody, R.N.; Kuhn, P.; Moskal, K.; Rojas-Silva, P.; Turnbaugh, P.J.; Raskin, I. Dietary polyphenols promote growth of the gut bacterium akkermansia muciniphila and attenuate high-fat diet-induced metabolic syndrome. Front. Immunol. 2015, 64, 2847–2858. [Google Scholar] [CrossRef] [PubMed]
- Rastmanesh, R. High polyphenol, low probiotic diet for weight loss because of intestinal microbiota interaction. Chem. Biol. Interact. 2011, 189, 1–8. [Google Scholar] [CrossRef]
- Przybylska-Balcerek, A.; Szablewski, T.; Szwajkowska-Michałek, L.; Świerk, D.; Cegielska-Radziejewska, R.; Krejpcio, Z.; Suchowilska, E.; Tomczyk, Ł.; Stuper-Szablewska, K. Sambucus nigra extracts–natural antioxidants and antimicrobial compounds. Molecules 2021, 26, 2910. [Google Scholar] [CrossRef]
- Rodríguez-Daza, M.C.; Pulido-Mateos, E.C.; Lupien-Meilleur, J.; Guyonnet, D.; Desjardins, Y.; Roy, D. Polyphenol-Mediated Gut Microbiota Modulation: Toward Prebiotics and Further. Front. Nutr. 2021, 8, 689456. [Google Scholar] [CrossRef]
- Tihăuan, B.-M.; Axinie, M.; Marinaș, I.-C.; Avram, I.; Nicoară, A.-C.; Grădișteanu-Pîrcălăbioru, G.; Dolete, G.; Ivanof, A.-M.; Onisei, T.; Cășărică, A.; et al. Evaluation of the Putative Duplicity Effect of Novel Nutraceuticals Using Physico-Chemical and Biological in Vitro Models. Foods 2022, 11, 1636. [Google Scholar]
- de Ferrars, R.M.; Cassidy, A.; Curtis, P.; Kay, C.D. Phenolic metabolites of anthocyanins following a dietary intervention study in post-menopausal women. Mol. Nutr. Food Res. 2014, 58, 490–502. [Google Scholar] [CrossRef]
- Hidalgo, M.; Oruna-Concha, M.J.; Kolida, S.; Walton, G.E.; Kallithraka, S.; Spencer, J.P.E.; Gibson, G.R.; De Pascual-Teresa, S. Metabolism of anthocyanins by human gut microflora and their influence on gut bacterial growth. J. Agric. Food Chem. 2012, 60, 3882–3890. [Google Scholar] [CrossRef]
- Wan, M.L.Y.; Ling, K.H.; Wang, M.F.; El-Nezami, H. Green tea polyphenol epigallocatechin-3-gallate improves epithelial barrier function by inducing the production of antimicrobial peptide pBD-1 and pBD-2 in monolayers of porcine intestinal epithelial IPEC-J2 cells. Mol. Nutr. Food Res. 2016, 60, 1048–1058. [Google Scholar] [CrossRef] [PubMed]
- Chambers, K.F.; Day, P.E.; Aboufarrag, H.T.; Kroon, P.A. Polyphenol effects on cholesterol metabolism via bile acid biosynthesis, CYP7A1: A review. Nutrients 2019, 11, 2588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Masumoto, S.; Terao, A.; Yamamoto, Y.; Mukai, T.; Miura, T.; Shoji, T. Non-absorbable apple procyanidins prevent obesity associated with gut microbial and metabolomic changes. Sci. Rep. 2016, 6, 31208. [Google Scholar] [CrossRef] [PubMed]
- Alves-Santos, A.M.; Sugizaki, C.S.A.; Lima, G.C.; Naves, M.M.V. Prebiotic effect of dietary polyphenols: A systematic review. J. Funct. Foods 2020, 74, 104169. [Google Scholar] [CrossRef]
- Koh, A.; De Vadder, F.; Kovatcheva-Datchary, P.; Backhed, F. From Dietary Fiber to Host Physiology: Short-Chain Fatty Acids as Key Bacterial Metabolites. Cell 2016, 165, 1332–1345. [Google Scholar] [CrossRef]
- Tiralongo, E.; Wee, S.S.; Lea, R.A. Elderberry Supplementation Reduces Cold Duration and Symptoms in Air-Travellers: A Randomized, Double-Blind Placebo-Controlled Clinical Trial. Nutrients 2016, 8, 182. [Google Scholar] [CrossRef]
- Zakay-Rones, Z.; Thom, E.; Wollan, T.; Wadstein, J. Randomized study of the efficacy and safety of oral elderberry extract in the treatment of influenza A and B virus infections. J. Int. Med. Res. 2004, 32, 132–140. [Google Scholar] [CrossRef]
- Zakay-Rones, Z.; Varsano, N.; Zlotnik, M.; Manor, O.; Regev, L.; Schlesinger, M.; Mumcuoglu, M. Inhibition of Several Strains of Influenza Virus in Vitro and Reduction of Symptoms by an Elderberry Extract (Sambucus nigra L.) during an Outbreak of Influenza B Panama. J. Altern. Complement. Med. 1995, 1, 361–369. [Google Scholar] [CrossRef]
- Ho, G.T.T.; Ahmed, A.; Zou, Y.F.; Aslaksen, T.; Wangensteen, H.; Barsett, H. Structure-activity relationship of immunomodulating pectins from elderberries. Carbohydr. Polym. 2015, 125, 241–248. [Google Scholar] [CrossRef]
- Ho, G.T.T.; Wangensteen, H.; Barsett, H. Elderberry and elderflower extracts, phenolic compounds, and metabolites and their effect on complement, RAW 264.7 macrophages and dendritic cells. Int. J. Mol. Sci. 2017, 18, 584. [Google Scholar] [CrossRef]
- Lee, J.; Finn, C.E. Anthocyanins and other polyphenolics in American elderberry (Sambucus canadensis) and European elderberry (S. nigra) cultivars. J. Sci. Food Agric. 2007, 87, 2665–2675. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Haas, B.J.; Gevers, D.; Earl, A.M.; Feldgarden, M.; Ward, D.V.; Giannoukos, G.; Ciulla, D.; Tabbaa, D.; Highlander, S.K.; Sodergren, E.; et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 2011, 21, 494–504. [Google Scholar] [CrossRef]
- Edgar, R.C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Core Team: Vienna, Austria, 2020. [Google Scholar]
- McMurdie, P.J.; Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package; R Package Version 2.4-6. 2018. Available online: https://cran.r-project.org/web/packages/vegan/vegan.pdf (accessed on 1 September 2022).
- Mallick, H.; Rahnavard, A.; McIver, L.J.; Ma, S.; Zhang, Y.; Nguyen, L.H.; Tickle, T.L.; Weingart, G.; Ren, B.; Schwager, E.H.; et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 2021, 17, e1009442. [Google Scholar] [CrossRef]
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the Tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Kassambara, A. Rstatix: Pipe-Friendly Framework for Basic Statistical Tests; 0.6.0. 2020. Available online: https://rpkgs.datanovia.com/rstatix/ (accessed on 1 September 2022).
- Kassambara, A. Ggpubr: ‘Ggplot2’ Based Publication Ready Plots; 0.4.0. 2020. Available online: https://rpkgs.datanovia.com/ggpubr/ (accessed on 1 September 2022).
- Lewis, S.J.; Heaton, K.W. Stool form scale as a useful guide to intestinal transit time. Scand. J. Gastroenterol. 1997, 32, 920–924. [Google Scholar] [CrossRef] [PubMed]
- Blake, M.R.; Raker, J.M.; Whelan, K. Validity and reliability of the Bristol Stool Form Scale in healthy adults and patients with diarrhoea-predominant irritable bowel syndrome. Aliment. Pharmacol. Ther. 2016, 44, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Caruso, R.; Lo, B.C.; Núñez, G. Host-microbiota interactions in inflammatory bowel disease. Nat. Rev. Immunol. 2020, 20, 411–426. [Google Scholar] [CrossRef] [PubMed]
- Schirmer, M.; Franzosa, E.A.; Lloyd-Price, J.; McIver, L.J.; Schwager, R.; Poon, T.W.; Ananthakrishnan, A.N.; Andrews, E.; Barron, G.; Lake, K.; et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 2018, 3, 337–346. [Google Scholar] [CrossRef]
- Mehta, R.S.; Abu-Ali, G.S.; Drew, D.A.; Lloyd-Price, J.; Subramanian, A.; Lochhead, P.; Joshi, A.D.; Ivey, K.L.; Khalili, H.; Brown, G.T.; et al. Stability of the human faecal microbiome in a cohort of adult men. Nat. Microbiol. 2018, 3, 347–355. [Google Scholar] [CrossRef]
- Marcobal, A.; Barboza, M.; Froehlich, J.W.; Block, D.E.; German, J.B.; Lebrilla, C.B.; Mills, D.A. Consumption of human milk oligosaccharides by gut-related microbes. J. Agric. Food Chem. 2010, 58, 5334–5340. [Google Scholar] [CrossRef]
- Yasukawa, Z.; Inoue, R.; Ozeki, M.; Okubo, T.; Takagi, T.; Honda, A.; Naito, Y. Effect of Repeated Consumption of Partially Hydrolyzed Guar Gum on Fecal Characteristics and Gut Microbiota: A Randomized, Double-Blind, Placebo-Controlled, and Parallel-Group Clinical Trial. Nutrients 2019, 11, 2170. [Google Scholar] [CrossRef] [PubMed]
- Nearing, J.T.; Douglas, G.M.; Hayes, M.G.; MacDonald, J.; Desai, D.K.; Allward, N.; Jones, C.M.A.; Wright, R.J.; Dhanani, A.S.; Comeau, A.M.; et al. Microbiome differential abundance methods produce different results across 38 datasets. Nat. Commun. 2022, 13, 342. [Google Scholar] [CrossRef]
- Khomich, M.; Måge, I.; Rud, I.; Berget, I. Analysing microbiome intervention design studies: Comparison of alternative multivariate statistical methods. PLoS ONE 2021, 16, e0259973. [Google Scholar] [CrossRef]
- Anhê, F.F.; Roy, D.; Pilon, G.; Dudonné, S.; Matamoros, S.; Varin, T.V.; Garofalo, C.; Moine, Q.; Desjardins, Y.; Levy, E.; et al. A polyphenol-rich cranberry extract protects from diet-induced obesity, insulin resistance and intestinal inflammation in association with increased Akkermansia spp. population in the gut microbiota of mice. Gut 2015, 64, 872–883. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Depommier, C.; Derrien, M.; Everard, A.; de Vos, W.M. Akkermansia muciniphila: Paradigm for next-generation beneficial microorganisms. Nat. Rev. Gastroenterol. Hepatol. 2022. epub ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Grander, C.; Adolph, T.E.; Wieser, V.; Lowe, P.; Wrzosek, L.; Gyongyosi, B.; Ward, D.V.; Grabherr, F.; Gerner, R.R.; Pfister, A.; et al. Recovery of ethanol-induced Akkermansia muciniphila depletion ameliorates alcoholic liver disease. Gut 2018, 67, 891–901. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Zhang, H.; Liu, H.; Zhang, H.; Bao, Y.; Di, J.; Hu, C. The genus Sutterella is a potential contributor to glucose metabolism improvement after Roux-en-Y gastric bypass surgery in T2D. Diabetes Res. Clin. Pract. 2020, 162, 108116. [Google Scholar] [CrossRef] [PubMed]
- Hiippala, K.; Kainulainen, V.; Kalliomäki, M.; Arkkila, P.; Satokari, R. Mucosal Prevalence and Interactions with the Epithelium Indicate Commensalism of Sutterella spp. Front. Microbiol. 2016, 7, 1706. [Google Scholar] [CrossRef]
- Paramsothy, S.; Nielsen, S.; Kamm, M.A.; Deshpande, N.P.; Faith, J.J.; Clemente, J.C.; Paramsothy, R.; Walsh, A.J.; van den Bogaerde, J.; Samuel, D.; et al. Specific Bacteria and Metabolites Associated with Response to Fecal Microbiota Transplantation in Patients with Ulcerative Colitis. Gastroenterology 2019, 156, 1440–1454. [Google Scholar] [CrossRef]
- Moon, C.; Baldridge, M.T.; Wallace, M.A.; Burnham, C.-A.D.; Virgin, H.W.; Stappenbeck, T.S. Vertically transmitted faecal IgA levels determine extra-chromosomal phenotypic variation. Nature 2015, 521, 90–93. [Google Scholar] [CrossRef]
- Robert, C.; Chassard, C.; Lawson, P.A.; Bernalier-Donadille, A. Bacteroides cellulosilyticus sp. nov., a cellulolytic bacterium from the human gut microbial community. Int. J. Syst. Evol. Microbiol. 2007, 57, 1516–1520. [Google Scholar] [CrossRef]
- McNulty, N.P.; Wu, M.; Erickson, A.R.; Pan, C.; Erickson, B.K.; Martens, E.C.; Pudlo, N.A.; Muegge, B.D.; Henrissat, B.; Hettich, R.L.; et al. Effects of diet on resource utilization by a model human gut microbiota containing Bacteroides cellulosilyticus WH2, a symbiont with an extensive glycobiome. PLoS Biol. 2013, 11, e1001637. [Google Scholar] [CrossRef]
- Dees, K.J.; Koo, H.; Humphreys, J.F.; Hakim, J.A.; Crossman, D.K.; Crowley, M.R.; Nabors, L.B.; Benveniste, E.N.; Morrow, C.D.; McFarland, B.C. Human gut microbial communities dictate efficacy of anti-PD-1 therapy in a humanized microbiome mouse model of glioma. Neurooncol. Adv. 2021, 3, vdab023. [Google Scholar] [CrossRef]
- Xu, X.; Zhang, W.; Guo, M.; Xiao, C.; Fu, Z.; Yu, S.; Jiang, L.; Wang, S.; Ling, Y.; Liu, F.; et al. Integrated analysis of gut microbiome and host immune responses in COVID-19. Front. Med. 2022, 16, 263–275. [Google Scholar] [CrossRef] [PubMed]
- Porter, N.T.; Luis, A.S.; Martens, E.C. Bacteroides thetaiotaomicron. Trends Microbiol. 2018, 26, 966–967. [Google Scholar] [CrossRef]
- Delday, M.; Mulder, I.; Logan, E.T.; Grant, G. Bacteroides thetaiotaomicron Ameliorates Colon Inflammation in Preclinical Models of Crohn’s Disease. Inflamm. Bowel. Dis. 2019, 25, 85–96. [Google Scholar] [CrossRef]
- Li, K.; Hao, Z.; Du, J.; Gao, Y.; Yang, S.; Zhou, Y. Bacteroides thetaiotaomicron relieves colon inflammation by activating aryl hydrocarbon receptor and modulating CD4(+)T cell homeostasis. Int. Immunopharmacol. 2021, 90, 107183. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.; Hong, J.; Xu, X.; Feng, Q.; Zhang, D.; Gu, Y.; Shi, J.; Zhao, S.; Liu, W.; Wang, X.; et al. Gut microbiome and serum metabolome alterations in obesity and after weight-loss intervention. Nat. Med. 2017, 23, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Cullen, T.W.; Schofield, W.B.; Barry, N.A.; Putnam, E.E.; Rundell, E.A.; Trent, M.S.; Degnan, P.H.; Booth, C.J.; Yu, H.; Goodman, A.L. Gut microbiota. Antimicrobial peptide resistance mediates resilience of prominent gut commensals during inflammation. Science 2015, 347, 170–175. [Google Scholar] [CrossRef] [Green Version]

| Mean Number of Bowel Movements/Week | Mean BSS Value/Week | |||
|---|---|---|---|---|
| Week | Mean | std. Deviation | Mean | std. Deviation |
| 1 | 9.1 | 3.2 | 3.8 | 0.6 |
| 2 | 9.1 | 4 | 3.7 | 0.6 |
| 4 | 9.1 | 3.4 | 3.7 | 0.6 |
| 5 | 9.4 | 3 | 3.7 | 0.6 |
| 6 | 9.1 | 3.3 | 3.7 | 0.6 |
| 7 | 8.9 | 3.5 | 3.8 | 0.7 |
| 8 | 8.7 | 2.9 | 3.8 | 0.7 |
| 9 | 9 | 3.3 | 3.7 | 0.6 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reider, S.; Watschinger, C.; Längle, J.; Pachmann, U.; Przysiecki, N.; Pfister, A.; Zollner, A.; Tilg, H.; Plattner, S.; Moschen, A.R. Short- and Long-Term Effects of a Prebiotic Intervention with Polyphenols Extracted from European Black Elderberry—Sustained Expansion of Akkermansia spp. J. Pers. Med. 2022, 12, 1479. https://doi.org/10.3390/jpm12091479
Reider S, Watschinger C, Längle J, Pachmann U, Przysiecki N, Pfister A, Zollner A, Tilg H, Plattner S, Moschen AR. Short- and Long-Term Effects of a Prebiotic Intervention with Polyphenols Extracted from European Black Elderberry—Sustained Expansion of Akkermansia spp. Journal of Personalized Medicine. 2022; 12(9):1479. https://doi.org/10.3390/jpm12091479
Chicago/Turabian StyleReider, Simon, Christina Watschinger, Julia Längle, Ulrike Pachmann, Nicole Przysiecki, Alexandra Pfister, Andreas Zollner, Herbert Tilg, Stephan Plattner, and Alexander R. Moschen. 2022. "Short- and Long-Term Effects of a Prebiotic Intervention with Polyphenols Extracted from European Black Elderberry—Sustained Expansion of Akkermansia spp." Journal of Personalized Medicine 12, no. 9: 1479. https://doi.org/10.3390/jpm12091479
APA StyleReider, S., Watschinger, C., Längle, J., Pachmann, U., Przysiecki, N., Pfister, A., Zollner, A., Tilg, H., Plattner, S., & Moschen, A. R. (2022). Short- and Long-Term Effects of a Prebiotic Intervention with Polyphenols Extracted from European Black Elderberry—Sustained Expansion of Akkermansia spp. Journal of Personalized Medicine, 12(9), 1479. https://doi.org/10.3390/jpm12091479






