An Idealized Clinicogenomic Registry to Engage Underrepresented Populations Using Innovative Technology
Abstract
1. Introduction
2. Barriers and Benefits of an Idealized Clinicogenomic Registry
2.1. Underlying Cultural and Social Determinants of Health (SDOH)
2.2. Institutional Review Board (IRB) and Protocol Adherence
2.3. Informed Consent
2.4. Big Data
- To inform clinical trial design;
- To support clinical decision support, clinical guidelines, and policy; and
- To address post-market safety, adverse events, and regulatory decision making.
2.5. Data Standards
2.6. Boundary Problems
2.7. Protocols
2.8. Technology
3. Vision for an Idealized Clinicogenomic Registry
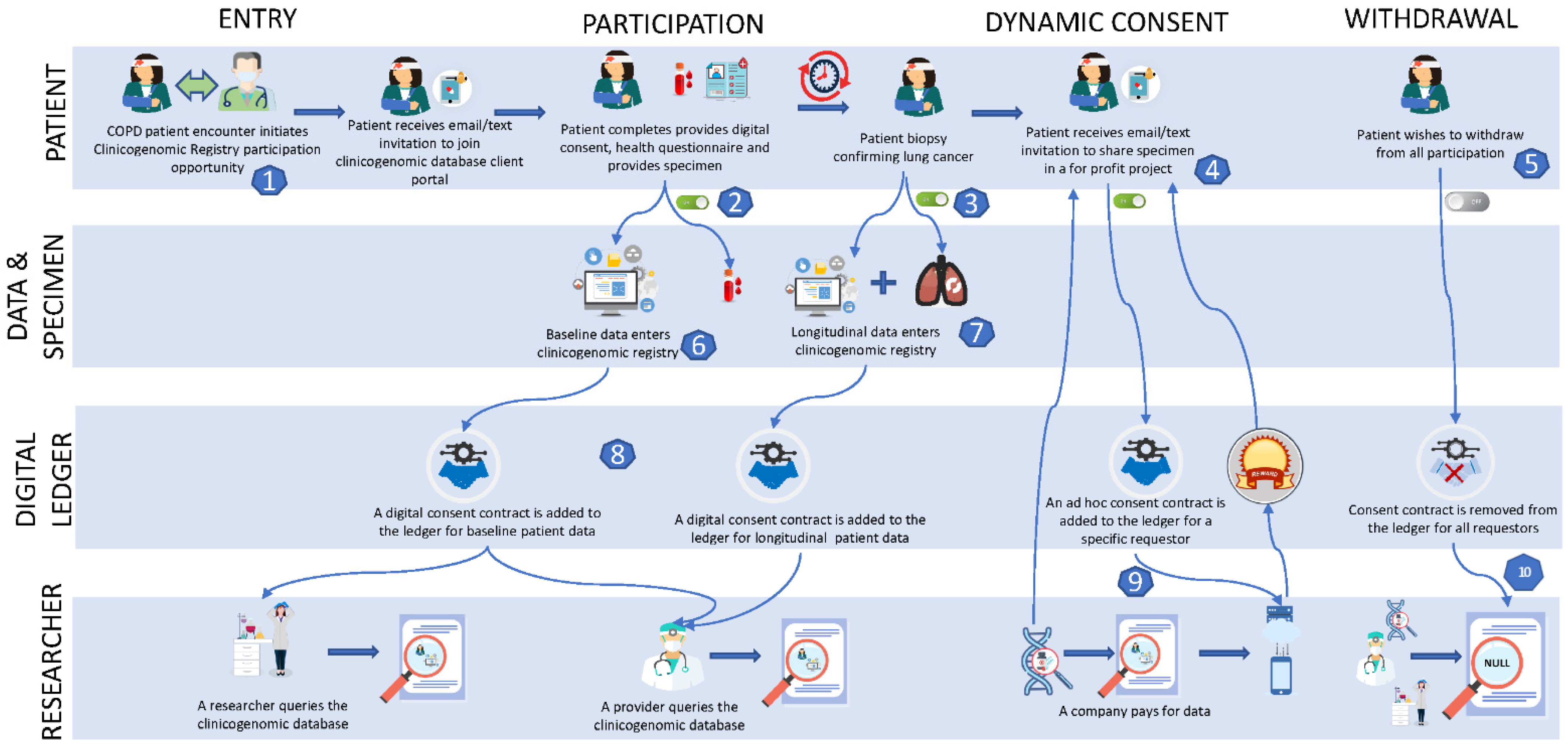
4. Hypothetical Case Study
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Oh, S.S.; White, M.J.; Gignoux, C.R.; Burchard, E.G. Making Precision Medicine Socially Precise. Take a Deep Breath. Am. J. Respir. Crit. Care Med. 2016, 193, 348–350. [Google Scholar] [CrossRef] [PubMed]
- Levy, D.E.; Byfield, S.D.; Comstock, C.B. Underutilization of BRCA1/2 Testing to Guide Breast Cancer Treatment: Black and Hispanic Women Particularly at Risk. Genet. Med. 2011, 13, 349–355. [Google Scholar] [CrossRef] [PubMed]
- Zavala, V.A.; Bracci, P.M.; Carethers, J.M.; Carvajal-Carmona, L.; Coggins, N.B.; Cruz-Correa, M.R.; Davis, M.; de Smith, A.J.; Dutil, J.; Figueiredo, J.C.; et al. Cancer health disparities in racial/ethnic minorities in the United States. Br. J. Cancer 2021, 124, 315–332. [Google Scholar] [CrossRef]
- Wheeler, D.A.; Wang, L. From human genome to cancer genome: The first decade. Genome Res. 2013, 23, 1054–1062. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.C.; Mensah, G.A.; Khoury, M.J. Leveraging Implementation Science to Address Health Disparities in Genomic Medicine: Examples from the Field. Ethn. Dis. 2019, 21, 187–192. [Google Scholar] [CrossRef]
- El-Deiry, W.S.; Goldberg, R.M.; Lenz, H.-J.; Shields, A.F.; Gibney, G.T.; Tan, A.R.; Brown, J.; Eisenberg, B.; Heath, E.I.; Phuphanich, S.; et al. The current state of molecular testing in the treatment of patients with solid tumors, 2019. CA Cancer J. Clin. 2019, 69, 305–343. [Google Scholar] [CrossRef] [PubMed]
- Bentley, A.R.; Callier, S.; Rotimi, C.N. Diversity and inclusion in genomic research: Why the uneven progress? J. Community Genet. 2017, 8, 255–266. [Google Scholar] [CrossRef]
- Landry, L.G.; Ali, N.; Williams, D.R.; Rehm, H.L.; Bonham, V.L. Lack of Diversity In Genomic Databases Is A Barrier To Translating Precision Medicine Research Into Practice. Health Aff. 2018, 37, 780–785. [Google Scholar] [CrossRef]
- Taube, S.E.; Clark, G.M.; Dancey, J.E.; McShane, L.M.; Sigman, C.C.; Gutman, S.I. A Perspective on Challenges and Issues in Biomarker Development and Drug and Biomarker Codevelopment. J. Natl. Cancer Inst. 2009, 101, 1453–1463. [Google Scholar] [CrossRef]
- Flaherty, K.T.; Gray, R.J.; Chen, A.P.; Li, S.; McShane, L.M.; Patton, D.; Hamilton, S.R.; Williams, P.M.; Iafrate, A.J.; Sklar, J.; et al. Molecular Landscape and Actionable Alterations in a Genomically Guided Cancer Clinical Trial: National Cancer Institute Molecular Analysis for Therapy Choice (NCI-MATCH). J. Clin. Oncol. 2020, 38, 3883–3894. [Google Scholar] [CrossRef]
- Manolio, T.A.; Chisholm, R.L.; Ozenberger, B.; Roden, D.M.; Williams, M.S.; Wilson, R.; Bick, D.; Bottinger, E.P.; Brilliant, M.H.; Eng, C.; et al. Implementing genomic medicine in the clinic: The future is here. Genet. Med. 2013, 15, 258–267. [Google Scholar] [CrossRef] [PubMed]
- Deverka, P.A.; Douglas, M.P.; Phillips, K.A. Use of Real-World Evidence in US Payer Coverage Decision-Making for Next-Generation Sequencing–Based Tests: Challenges, Opportunities, and Potential Solutions. Value Health 2020, 23, 540–550. [Google Scholar] [CrossRef] [PubMed]
- Reid, S.; Cadiz, S.; Pal, T. Disparities in Genetic Testing and Care Among Black Women with Hereditary Breast Cancer. Curr. Breast Cancer Rep. 2020, 12, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.; Jacobs, D.; Kriak, J.; Abu-Baker, A.; Udeani, G.; Neal, G.; Ramos, K. Implementation of Pharmacogenomics and Artificial Intelligence Tools for Chronic Disease Management in Primary Care Setting. J. Pers. Med. 2021, 11, 443. [Google Scholar] [CrossRef] [PubMed]
- Mapes, B.; Charif, O.E.; Al-Sawwaf, S.; Dolan, E.M. Genome-Wide Association Studies of Chemotherapeutic Toxicities: Ge-nomics of Inequality. Clin. Cancer Res. 2017, 23, 4010–4019. [Google Scholar] [CrossRef]
- Nevins, J.R.; Huang, E.S.; Dressman, H.; Pittman, J.; Huang, A.T.; West, M. Towards integrated Clinico-Genomic models for personalized medicine: Combining gene expression signatures and clinical factors in breast cancer outcomes prediction. Hum. Mol. Genet. 2003, 12, R153–R157. [Google Scholar] [CrossRef]
- Agarwala, V.; Khozin, S.; Singal, G.; O’Connell, C.; Kuk, D.; Li, G.; Gossai, A.; Miller, V.; Abernethy, A.P. Real-World Evidence in Support of Precision Medicine: Clinico-Genomic Cancer Data as a Case Study. Health Aff. 2018, 37, 765–772. [Google Scholar] [CrossRef]
- Thorlund, K.; Dron, L.; Park, J.J.; Mills, E.J. Synthetic and External Controls in Clinical Trials—A Primer for Researchers. Clin. Epidemiol. 2020, 12, 457–467. [Google Scholar] [CrossRef]
- Boulos, M.K.; Zhang, P. Digital Twins: From Personalised Medicine to Precision Public Health. J. Pers. Med. 2021, 11, 745. [Google Scholar] [CrossRef]
- Fisher, C.K.; Smith, A.M.; Walsh, J.R. Machine learning for comprehensive forecasting of Alzheimer’s Disease progression. Sci. Rep. 2019, 9, 13622. [Google Scholar] [CrossRef]
- Kolonel, L.N.; Henderson, B.E.; Hankin, J.H.; Nomura, A.M.; Wilkens, L.R.; Pike, M.C.; Stram, D.O.; Monroe, K.R.; Earle, M.E.; Nagamine, F.S. A Multiethnic Cohort in Hawaii and Los Angeles: Baseline Characteristics. Am. J. Epidemiol. 2000, 151, 346–357. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Guan, S.; Xu, Y.; Chen, W.; Huang, X.; Wang, X.; Zhang, M. Effects of the Two-Dimensional Structure of Trust on Patient Adherence to Medication and Non-pharmaceutical Treatment: A Cross-Sectional Study of Rural Patients with Essen-tial Hypertension in China. Front. Public Health 2022, 4, 818426. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Feng, Y.; Fang, J.; Wu, J.; Huang, X.; Wang, X.; Zhang, M. Effect of trust in primary care physicians on patient satisfaction: A cross-sectional study among patients with hypertension in rural China. BMC Prim. Care 2020, 21, 196. [Google Scholar] [CrossRef]
- Wang, M.; Li, M.; Dong, X. The Associations Between Sociodemographic Characteristics and Trust in Physician with Immunization Service Use in U.S. Chin. Older Adults. Res. Aging 2022, 44, 164–173. [Google Scholar]
- Cao, Q.; Krok-Schoen, J.L.; Guo, M.; Dong, X. Trust in physicians, health insurance, and health care utilization among Chinese older immigrants. Ethn. Health 2022, 18, 1–18. [Google Scholar] [CrossRef]
- Bakhireva, L.N.; Nebeker, C.; Ossorio, P.; Angal, J.; Thomason, M.E.; Croff, J.M. Inclusion of American Indians and Alaskan Natives in Large National Studies: Ethical Considerations and Implications for Biospecimen Collection in the HEALthy Brain and Child Development Study. Advers. Resil. Sci. 2020, 1, 285–294. [Google Scholar] [CrossRef]
- Desmond, H. Precision Medicine, Data, and the Anthropology of Social Status. Am. J. Bioeth. 2021, 21, 80–83. [Google Scholar] [CrossRef]
- Mikk, K.A.; Sleeper, H.A.; Topol, E.J. The Pathway to Patient Data Ownership and Better Health. J. Am. Med. Assoc. 2017, 318, 1433–1434. [Google Scholar] [CrossRef]
- Contreras, J.L.; Rumbold, J.; Pierscionek, B. Patient Data Ownership. J. Am. Med. Assoc. 2018, 319, 935. [Google Scholar] [CrossRef]
- Contreras, J. Genetic Property, 2016. University of Utah College of Law Research Paper No. 171. Georget. Law J. 2016, 105, 1. [Google Scholar]
- Mirchev, M.; Mircheva, I.; Kerekovska, A. The Academic Viewpoint on Patient Data Ownership in the Context of Big Data: Scoping Review. J. Med. Internet Res. 2020, 22, e22214. [Google Scholar] [CrossRef] [PubMed]
- Charles, W.M.; Delgado, B.M. Health Datasets as Assets: Blockchain-Based Valuation and Transaction Methods. Blockchain Healthc. Today 2022, 5, 185. [Google Scholar] [CrossRef]
- Press, W. What’s So Special About Science (And How Much Should We Spend on It?). Science 2013, 342, 817–822. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.; Ramos, K. Academic Medical Centers as Innovation Ecosystems: Evolution of Industry Partnership Models Beyond Bayh-Dole. Act. Acad. Med. 2018, 93, 1135–1141. [Google Scholar] [CrossRef]
- Wilbanks, J. Design Issues in E-Consent. J. Law Med. Ethics 2018, 46, 110–118. [Google Scholar] [CrossRef]
- Kaye, J.; Whitley, E.A.; Lund, D.; Morrison, M.; Teare, H.; Melham, K. Dynamic Consent: A Patient Interface for Twenty-First Century Research Networks. Eur. J. Hum. Genet. 2015, 23, 141–146. [Google Scholar] [CrossRef]
- Privacy and Health Research in a Data Driven World: An Exploratory Workshop. September 2019. Available online: https://www.hhs.gov/ohrp/sites/default/files/report-privacy-and-health-10-31-19.pdf (accessed on 12 April 2022).
- Liddell, K.; Simon, D.A.; Lucassen, A. Patient Data Ownership: Who Owns Your Health? J. Law Biosci. 2021, 8, 23. [Google Scholar] [CrossRef]
- Adashi, E.Y.; Walters, L.B.; Menikoff, J.A. The Belmont Report at 40: Reckoning with Time. Am. J. Public Health 2018, 108, 1345–1348. [Google Scholar] [CrossRef]
- Kayaalp, M. Patient Privacy in the Era of Big Data. Balk. Med. J. 2018, 35, 8–17. [Google Scholar] [CrossRef]
- Mainous, A.G.; Kern, D.; Hainer, B.; Kneuper-Hall, R.; Stephens, J.; Geesey, M.E. The relationship between continuity of care and trust with stage of cancer at diagnosis. Fam. Med. 2004, 36, 35–44. [Google Scholar]
- Stange, K.C. The Problem of Fragmentation and the Need for Integrative Solutions. Ann. Fam. Med. 2009, 7, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Musmade, P.B.; Nijhawan, L.P.; Udupa, N.; Bairy, K.L.; Bhat, K.; Janodia, M.; Muddukrishna, B.S. Informed consent: Issues and challenges. J. Adv. Pharm. Technol. Res. 2013, 4, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Maloy, J.W.; Bass, P.F. Understanding Broad Consent. Ochsner J. 2020, 20, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Fisher, C.B.; Layman, D.M. Genomics, Big Data, and Broad Consent: A New Ethics Frontier for Prevention Science. Prev. Sci. 2018, 19, 871–879. [Google Scholar] [CrossRef]
- Grady, C.; Eckstein, L.; Berkman, B.; Brock, D.; Cook-Deegan, R.; Fullerton, S.M.; Greely, H.; Hansson, M.G.; Hull, S.; Kim, S.; et al. Broad Consent for Research With Biological Samples: Workshop Conclusions. Am. J. Bioeth. 2015, 15, 34–42. [Google Scholar] [CrossRef]
- Chandler, R.; Brady, K.T.; Jerome, R.N.; Eder, M.; Rothwell, E.; Brownley, K.A.; Harris, P.A. Broad-scale informed consent: A survey of the CTSA landscape. J. Clin. Transl. Sci. 2019, 3, 253–260. [Google Scholar] [CrossRef]
- Vries, D.; Tomlinson, R.G.; Kim, H.M.; Krenz, C.; Haggerty, D.; Ryan, K.A.; Kim, S.Y. Understanding the Public’s Reservations about Broad Consent and Study-By-Study Consent for Donations to a Biobank: Results of a National Survey. PLoS ONE 2016, 11, 159113. [Google Scholar] [CrossRef]
- Dickert, N.W.; Eyal, N.; Goldkind, S.F.; Grady, C.; Joffe, S.; Lo, B.; Miller, F.G.; Pentz, R.D.; Silbergleit, R.; Weinfurt, K.P.; et al. Reframing Consent for Clinical Research: A Function-Based Approach. Am. J. Bioeth. 2017, 17, 3–11. [Google Scholar] [CrossRef]
- Smith, R.G. Clinical data to be used as a foundation to combat COVID-19 vaccine hesitancy. J. Interprof. Educ. Pract. 2021, 26, 100483. [Google Scholar] [CrossRef]
- Suter, S.M. GINA at 10 Years: The Battle Over ‘Genetic Information’ Continues in Court. J. Law Biosci. 2018, 5, 495–526. [Google Scholar] [CrossRef]
- Matsui, K.; Kita, Y.; Ueshima, H. Informed consent, participation in, and withdrawal from a population based cohort study involving genetic analysis. J. Med. Ethic 2005, 31, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Doerr, M.; Moore, S.; Barone, V.; Sutherland, S.; Bot, B.M.; Suver, C.; Wilbanks, J. Assessment of the All of Us research program’s informed consent process. AJOB Empir. Bioeth. 2021, 12, 72–83. [Google Scholar] [CrossRef] [PubMed]
- Byrd, T.F.; Ahmad, F.S.; Liebovitz, D.M.; Kho, A.N. Defragmenting Heart Failure Care: Medical Records Integration. Heart Fail Clin. 2020, 16, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Mulberg, A.E.; Bucci-Rechtweg, C.; Giuliano, J.; Jacoby, D.; Johnson, F.K.; Liu, Q.; Marsden, D.; McGoohan, S.; Nelson, R.; Patel, N.; et al. Regulatory strategies for rare diseases under current global regulatory statutes: A discussion with stakeholders. Orphanet J. Rare Dis. 2019, 14, 36. [Google Scholar] [CrossRef] [PubMed]
- Corral-Acero, J.; Margara, F.; Marciniak, M. The ’Digital Twin’ to Enable the Vision of Precision Cardiology. Eur. Heart J. 2020, 41, 4556–4564. [Google Scholar] [CrossRef]
- Lal, A.; Li, G.; Cubro, E. Development and Verification of a Digital Twin Patient Model to Predict Specific Treatment Response During the First 24 Hours of Sepsis. Crit. Care Explor. 2020, 2, 249. [Google Scholar] [CrossRef]
- Andre, E.B.; Reynolds, R.; Caubel, P.; Azoulay, L.; Dreyer, N.A. Trial Designs Using Real-World Data: The Changing Landscape of the Regulatory Approval Process. Pharmacoepidemiol. Drug Saf. 2020, 29, 1201–1212. [Google Scholar] [CrossRef] [PubMed]
- Goemans, N.; Signorovitch, J.; Sajeev, G. Suitability of External Controls for Drug Evaluation in Duchenne Muscular Dystrophy. Neurology 2020, 95, 1381. [Google Scholar] [CrossRef]
- Schulz, A.; Ajayi, T.; Specchio, N.; Reyes, E.D.L.; Gissen, P.; Ballon, D.; Dyke, J.P.; Cahan, H.; Slasor, P.; Jacoby, D.; et al. Study of Intraventricular Cerliponase Alfa for CLN2 Disease. N. Engl. J. Med. 2018, 378, 1898–1907. [Google Scholar] [CrossRef]
- Johnson, T.B.; Cain, J.T.; White, K.A.; Ramirez-Montealegre, D.; Pearce, D.A.; Weimer, J.M. Therapeutic Landscape for Batten Disease: Current Treatments and Future Prospects. Nat. Rev. Neurol. 2019, 15, 161–178. [Google Scholar] [CrossRef]
- Hawk, E.T.; Habermann, E.B.; Ford, J.G. Five National Cancer Institute-Designated Cancer Centers’ Data Collection on Ra-cial/Ethnic Minority Participation in Therapeutic Trials: A Current View and Opportunities for Improvement. Cancer 2014, 120, 1113–1121. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.H.; Cook, E.D. A primer for cancer research programs on defining and evaluating the catchment area and evaluating minority clinical trials recruitment. Adv. Cancer Res. 2020, 146, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, C.; Wang, Q.; Li, Z.; Lin, J.; Wang, H. Differences in Stage of Cancer at Diagnosis, Treatment, and Survival by Race and Ethnicity Among Leading Cancer Types. JAMA Netw. Open 2020, 3, e202950. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Finkelstein, J. Inconsistency in Race and Ethnic Classification in Pharmacogenetics Studies and its Potential Clinical Implications. Pharmgenom. Pers. Med. 2019, 12, 107–123. [Google Scholar] [CrossRef]
- Rogers, S.L.; Keeling, N.J.; Giri, J.; Gonzaludo, N.; Jones, J.S.; Glogowski, E.; Formea, C.M. PARC report: A health-systems focus on reimbursement and patient access to pharmacogenomics testing. Pharmacogenomics 2020, 21, 785–796. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, S.J.; Sireci, A.; Pendrick, D. Clinical Utility and Reimbursement for Expanded Genomic Panel Testing in Adult Oncology. J. Clin. Oncol. 2019, 37, 6593. [Google Scholar] [CrossRef]
- Hsiao, S.J.; Sireci, A.N.; Pendrick, D.; Freeman, C.; Fernandes, H.; Schwartz, G.K.; Henick, B.S.; Mansukhani, M.M.; Roth, K.A.; Carvajal, R.D.; et al. Clinical Utilization, Utility, and Reimbursement for Expanded Genomic Panel Testing in Adult Oncology. JCO Precis. Oncol. 2020, 4, 1038–1048. [Google Scholar] [CrossRef]
- Perdrizet, K.; Stockley, T.L.; Law, J.H.; Smith, A.; Zhang, T.; Fernandes, R.; Shabir, M.; Sabatini, P.; Al Youssef, N.; Ishu, C.; et al. Integrating comprehensive genomic sequencing of non-small cell lung cancer into a public healthcare system. Cancer Treat. Res. Commun. 2022, 31, 100534. [Google Scholar] [CrossRef]
- Horgan, D.; Curigliano, G.; Rieß, O.; Hofman, P.; Büttner, R.; Conte, P.; Cufer, T.; Gallagher, W.M.; Georges, N.; Kerr, K.; et al. Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe. J. Pers. Med. 2022, 12, 72. [Google Scholar] [CrossRef]
- Data Standards Catalog V8.0—Supported and Required Standards. Federal Drug Administration, Data Standards Catalog; 2022. Available online: https://www.fda.gov/industry/fda-data-standards-advisory-board/study-data-standards-resources (accessed on 12 April 2022).
- Ross, S.; Grant, A.; Counsell, C.; Gillespie, W.; Russell, I.; Prescott, R. Barriers to Participation in Randomised Controlled Trials: A Systematic Review. J. Clin. Epidemiol. 1999, 52, 141–150. [Google Scholar] [CrossRef]
- Mahajan, A.P. Health Information Exchange—Obvious Choice or Pipe Dream? JAMA Intern. Med. 2016, 176, 429–430. [Google Scholar] [CrossRef] [PubMed]
- Budin-Ljøsne, I.; Bentzen, H.B.; Solbakk, J.H.; Myklebost, O. Genome Sequencing in Research Requires a New Approach to Consent. Tidsskr. Nor. Laegeforen. 2015, 135, 2031–2033. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Martinez-Martin, N.; Luo, Z.; Kaushal, A.; Adeli, E.; Haque, A.; Kelly, S.S.; Wieten, S.; Cho, M.K.; Magnus, D.; Fei-Fei, L.; et al. Ethical issues in using ambient intelligence in health-care settings. Lancet Digit. Health 2021, 3, e115–e123. [Google Scholar] [CrossRef]
- Braunstein, M.L. Health care in the age of interoperability part 5: The personal health record. IEEE Pulse 2019, 10, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Total Cancer Care Protocol: A Lifetime Partnership with Patients Who Have or May be at Risk of Having Cancer (TCCP). Clinical Trial: NCT03977402; 2021. Available online: https://clinicaltrials.gov/ (accessed on 12 April 2022).
- Haddad, D.N.; Sandler, K.L.; Henderson, L.M.; Rivera, M.P.; Aldrich, M.C. Disparities in Lung Cancer Screening: A Review. Ann. Am. Thorac. Soc. 2020, 17, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Wong, R.J.; Ahmed, A. Understanding Gaps in the Hepatocellular Carcinoma Cascade of Care: Opportunities to Improve Hepatocellular Carcinoma Outcomes. J. Clin. Gastroenterol. 2020, 54, 850–856. [Google Scholar] [CrossRef]
- Popejoy, A.B.; Crooks, K.R.; Fullerton, S.M. Clinical Genetics Lacks Standard Definitions and Protocols for the Collection and Use of Diversity Measures. Am. J. Hum. Genet. 2020, 107, 72–82. [Google Scholar] [CrossRef]
- Rotimi, C.N.; Jorde, L.B. Ancestry and Disease in the Age of Genomic Medicine. N. Engl. J. Med. 2010, 363, 1551–1558. [Google Scholar] [CrossRef]
- Clayton, E. Ethical, Legal, and Social Implications of Genomic Medicine. N. Engl. J. Med. 2003, 349, 562–569. [Google Scholar] [CrossRef]
- Beskow, L.M. Lessons from HeLa Cells: The Ethics and Policy of Biospecimens. Annu. Rev. Genom. Hum. Genet. 2016, 17, 395–417. [Google Scholar] [CrossRef]
- Mortimer, D.; Peacock, S. Social welfare and the Affordable Care Act: Is it ever optimal to set aside comparative cost? Soc. Sci. Med. 2012, 75, 1156–1162. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Y.; Sheets, L.; Gao, X.; Shen, Y.; Shae, Z.-Y.; Tsai, J.J.P.; Shyu, C.-R. Development of A Blockchain Framework for Virtual Clinical Trials. AMIA Annu. Symp. Proc. 2021, 2020, 1412–1420. [Google Scholar] [PubMed]
- Roehr, B. Pfizer launches virtual clinical trial. BMJ 2011, 342, d3722. [Google Scholar] [CrossRef] [PubMed]
- TriCare Select Navigator Pilot Program. Available online: https://www.tricare.mil/Plans/SpecialPrograms/SelectNavigator (accessed on 12 April 2022).
- 21st Century Cures Act: Interoperability, Information Blocking, and the ONC Health IT Certification Program.
- Pollard, C.; Bailey, K.A.; Petitte, T.; Baus, A.; Swim, M.; Hendryx, M. Electronic Patient Registries Improve Diabetes Care and Clinical Outcomes in Rural Community Health Centers. J. Rural Health 2009, 25, 77–84. [Google Scholar] [CrossRef]
- Calman, N.S.; Hauser, D.; Weiss, L. Becoming a Patient-Centered Medical Home: A 9-Year Transition for a Network of Fed-erally Qualified Health Centers. Ann. Fam. Med. 2013, 11 (Suppl. S1), 68. [Google Scholar] [CrossRef]
- Kuo, T.T.; Kim, H.E.; Ohno-Machado, L. Blockchain Distributed Ledger Technologies for Biomedical and Health Care Ap-plications. J. Am. Med. Inform. Assoc. 2017, 24, 1211–1220. [Google Scholar] [CrossRef]
- Abu-Elezz, I.; Hassan, A.; Nazeemudeen, A.; Househ, M.; Abd-alrazaq, A. The Benefits and Threats of Blockchain Technol-ogy in Healthcare: A Scoping Review. Int. J. Med. Inform. 2020, 142, 104246. [Google Scholar] [CrossRef]
- Mackey, T.K.; Kuo, T.T.; Gummadi, B. Fit-for-Purpose? Challenges and Opportunities for Applications of Blockchain Tech-nology in the Future of Healthcare. BMC Med. 2019, 17, 68. [Google Scholar] [CrossRef]
- Singal, G.; Miller, P.; Agarwala, V.; Li, G.; Kaushik, G.; Backenroth, D.; Gossai, A.; Frampton, G.M.; Torres, A.Z.; Lehnert, E.M.; et al. Association of Patient Characteristics and Tumor Genomics with Clinical Outcomes Among Patients with Non–Small Cell Lung Cancer Using a Clinicogenomic Database. J. Am. Med. Assoc. 2019, 321, 1391–1399. [Google Scholar] [CrossRef]
- García-Barriocanal, E.; Sánchez-Alonso, S.; Sicilia, M.-A. Deploying Metadata on Blockchain Technologies. In Proceedings of the Communications in Computer and Information Science, Babahoyo, Ecuador, 8–10 November 2017; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2017; Volume 2017, pp. 38–49. [Google Scholar]
- Booth, G.S.; Gehrie, E.A. Non-fungible tokens: Stem cell transplantation in the blockchain. Transfus. Apher. Sci. 2021, 60, 103196. [Google Scholar] [CrossRef] [PubMed]
- Sazandrishvili, G. Asset tokenization in plain English. J. Corp. Account. Financ. 2020, 31, 68–73. [Google Scholar] [CrossRef]
| Ideal Registry | Challenge | Blockchain Feature to Solve It |
|---|---|---|
| Patients provide consent for a wide swath of research activities | Patient control of future use of data | Governance; smart contracts |
| Incentives for health systems and patients to share data | The chain of custody of fungible data makes attribution of bulk data impractical | Monetization; smart contracts; digital ledger |
| Users and providers have comfort with provenance of data in a collaboration | Third-party obligations and lack of granularity of data and specimens shared | Digital ledger; governance; smart contracts |
| Assembling a cohort involves minimal institutional touchpoints and bypasses cumbersome processes | Transactional frictions of health data sharing | Governance layers; smart contracts |
| Complete control of who uses data and for what purpose | Unauthorized use or replication of fungible data | Smart contracts; digital ledger |
| Low-friction methods to define the rules of engagement for compliance and legal constraints for data recipients | Operational costs to administer data governance and administer legal contracts | Governance layers; smart contracts |
| Facile HIPAA and compliance reporting | Lack of granularity of bulk data and lack of visibility to compliance administrators | Smart contracts; digital ledger |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, P.; Dahlke, D.V.; Smith, M.L.; Charles, W.; Gomez, J.; Ory, M.G.; Ramos, K.S. An Idealized Clinicogenomic Registry to Engage Underrepresented Populations Using Innovative Technology. J. Pers. Med. 2022, 12, 713. https://doi.org/10.3390/jpm12050713
Silva P, Dahlke DV, Smith ML, Charles W, Gomez J, Ory MG, Ramos KS. An Idealized Clinicogenomic Registry to Engage Underrepresented Populations Using Innovative Technology. Journal of Personalized Medicine. 2022; 12(5):713. https://doi.org/10.3390/jpm12050713
Chicago/Turabian StyleSilva, Patrick, Deborah Vollmer Dahlke, Matthew Lee Smith, Wendy Charles, Jorge Gomez, Marcia G. Ory, and Kenneth S. Ramos. 2022. "An Idealized Clinicogenomic Registry to Engage Underrepresented Populations Using Innovative Technology" Journal of Personalized Medicine 12, no. 5: 713. https://doi.org/10.3390/jpm12050713
APA StyleSilva, P., Dahlke, D. V., Smith, M. L., Charles, W., Gomez, J., Ory, M. G., & Ramos, K. S. (2022). An Idealized Clinicogenomic Registry to Engage Underrepresented Populations Using Innovative Technology. Journal of Personalized Medicine, 12(5), 713. https://doi.org/10.3390/jpm12050713








