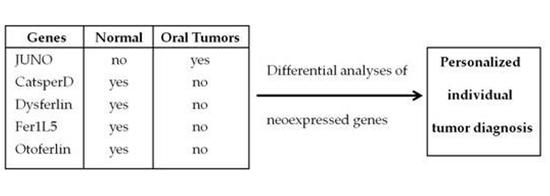
Neoexpression of JUNO in Oral Tumors Is Accompanied with the Complete Suppression of Four Other Genes and Suggests the Application of New Biomarker Tools
Abstract
:1. Introduction
2. Materials and Methods
2.1. Tissue Sampling
2.2. Reverse Transcription Real-Time PCR
2.3. Statistical Analysis
3. Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Monk, M.; Holding, C. Human embryonic genes re-expressed in cancer cells. Oncogene 2001, 20, 8085–8091. [Google Scholar] [CrossRef] [Green Version]
- Yu, X.; Xu, J. A ‘Goldmine’ for digging cancer-specific targets: The genes essential for embryo development but non-essential for adult life. J. Mol. Cell Biol. 2020, 12, 669–673. [Google Scholar] [CrossRef]
- Bruggeman, J.W.; Irie, N.; Lodder, P.; Van Pelt, A.M.M.; Koster, J.; Hamer, G. Tumors Widely Express Hundreds of Embryonic Germline Genes. Cancers 2020, 12, 3812. [Google Scholar] [CrossRef]
- Kraus, D.; Glassmann, A.; Golletz, C.; Kristiansen, G.; Winter, J.; Probstmeier, R. Zona Pellucida Protein 2 (ZP2) Is Expressed in Colon Cancer and Promotes Cell Proliferation. Cancers 2021, 13, 1759. [Google Scholar] [CrossRef]
- Kraus, D.; Reckenbeil, J.; Perner, S.; Winter, J.; Probstmeier, R. Expression Pattern of Matrix Metalloproteinase 20 (MMP20) in Human Tumors. Anticancer Res. 2016, 36, 2713–2718. [Google Scholar]
- Bianchi, E.; Doe, B.; Goulding, D.; Wright, G. Juno is the egg Izumo receptor and is essential for mammalian fertilization. Nature 2014, 508, 483–487. [Google Scholar] [CrossRef] [PubMed]
- National Library of Medicine. Available online: https://pubmed.ncbi.nlm.nih.gov (accessed on 1 December 2021).
- Sun, X.-H.; Zhu, Y.-Y.; Wang, L.; Liu, H.-L.; Ling, Y.; Li, Z.-L.; Sun, L.-B. The Catsper channel and its roles in male fertility: A systematic review. Reprod. Biol. Endocrinol. 2017, 15, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Brown, S.G.; Publicover, S.J.; Barratt, C.L.R.; da Silva, S.J.M. Human sperm ion channel (dys)function: Implications for fer-zilization. Hum. Reprod. Update 2019, 25, 758–776. [Google Scholar] [CrossRef]
- Posey, A.D., Jr.; Pytel, P.; Gardikiotes, K.; Demonbreun, A.R.; Rainey, M.; George, M.; Band, H.; McNally, E.M. Endocytic Recycling Proteins EHD1 and EHD2 Interact with Fer-1-like-5 (Fer1L5) and Mediate Myoblast Fusion. J. Biol. Chem. 2011, 286, 7379–7388. [Google Scholar] [CrossRef] [Green Version]
- Peulen, O.; Rademaker, G.; Anania, S.; Turtoi, A.; Bellahcène, A.; Castronovo, V. Ferlin Overview: From Membrane to Cancer Biology. Cells 2019, 8, 954. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.K.; Bansal, P.; Ganguly, A.; Bhandari, B.; Chakrabarti, K. Human zona pellucida glycoproteins: Functional relevance during fertilization. J. Reprod. Immunol. 2009, 83, 50–55. [Google Scholar] [CrossRef]
- Bansal, P.; Gupta, S.K. Binding characteristics of sperm with recombinant human zona pellucida glycoprotein-3 coated beads. Indian J. Med. Res. 2009, 130, 37–43. [Google Scholar] [PubMed]
- Costa, J.; Pereira, R.; Oliveira, J.; Alves, Â.; Marques-Magalhães, Â.; Frutuoso, A.; Leal, C.; Barros, N.; Fernandes, R.; Queiroz Almeida, D.; et al. Structural and molecular analysis of the cancer prostate cell line PC3: Oocyte zona pellucida glycoproteins. Tissue Cell 2018, 55, 91–106. [Google Scholar] [CrossRef]
- Rahman, N.A.; Bennink, H.J.; Chrusciel, M.; Sharp, V.; Zimmerman, Y.; Dina, R.; Li, X.; Ellonen, A.; Rivero-Müller, A.; Dilworth, S.; et al. A novel treatment strategy for ovarian cancer based on immunization against zona pellucida protein (ZP) 3. FASEB J. 2012, 26, 324–333. [Google Scholar] [CrossRef]
- Reckenbeil, J.; Kraus, D.; Probstmeier, R.; Allam, J.-P.; Novak, N.; Frentzen, M.; Martini, M.; Wenghoefer, M.; Winter, J. Cellular Distribution and Gene Expression Pattern of Metastasin (S100A4), Calgranulin A (S100A8), and Calgranulin B (S100A9) in Oral Lesions as Markers for Molecular Pathology. Cancer Investig. 2016, 34, 246–254. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [Green Version]
- Brierley, J.D.; Gospodarowicz, M.K.; Wittekind, C. TNM Classification of Malignant Tumours, 8th ed.; John Wiley & Sons: Hoboken, NJ, USA, 2017. [Google Scholar]
- Nebion.com. Available online: https://genevisible.com/search (accessed on 1 December 2021).
- Horpaopan, S.; Kirfel, J.; Peters, S.; Kloth, M.; Hüneburg, R.; Altmüller, J.; Drichel, D.; Odenthal, M.; Kristiansen, G.; Strassburg, C.; et al. Exome sequencing characterizes the somatic mutation spectrum of early serrated lesions in a patient with serrated polyposis syndrome (SPS). Hered. Cancer Clin. Pract. 2017, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qureshi, M.A.; Khan, S.; Tauheed, M.S.; Syed, S.A.; Ujjan, I.D.; Lail, A.; Sharafat, S. Pan-Cancer Multiomics Analysis of TC2N Gene Suggests its Important Role(s) in Tumourigenesis of Many Cancers. Asian Pac. J. Cancer Prev. APJCP 2020, 21, 3199–3209. [Google Scholar] [CrossRef]
- Turtoi, A.; Blomme, A.; Bellahcène, A.; Gilles, C.; Hennequière, V.; Peixoto, P.; Bianchi, E.; Noel, A.; De Pauw, E.; Lifrange, E.; et al. Myoferlin is a key regulator of EGFR activity in breast cancer. Cancer Res. 2013, 73, 5438–5448. [Google Scholar] [CrossRef] [Green Version]
- Volakis, L.I.; Li, R.; Ackerman, W.E.; Mihai, C.; Bechel, M.; Summerfield, T.L.; Ahn, C.S.; Powell, H.M.; Zielinski, R.; Rosol, T.J.; et al. Loss of Myoferlin Redirects Breast Cancer Cell Motility towards Collective Migration. PLoS ONE 2014, 9, e86110. [Google Scholar] [CrossRef] [Green Version]
- Kumar, B.; Brown, N.V.; Swanson, B.J.; Schmitt, A.C.; Old, M.; Ozer, E.; Agrawal, A.; Schuller, D.E.; Teknos, T.N.; Kumar, P. High expression of myoferlin is associated with poor outcome in oropharyngeal squamous cell carcinoma patients and is inversely associated with HPV-status. Oncotarget 2016, 7, 18665–18677. [Google Scholar] [CrossRef] [Green Version]
- Fahmy, K.; Gonzalez, A.; Arafa, M.; Peixoto, P.; Bellahcène, A.; Turtoi, A.; Delvenne, P.; Thiry, M.; Castronovo, V.; Peulen, O. Myoferlin plays a key role in VEGFA secretion and impacts tumor-associated angiogenesis in human pancreas cancer. Int. J. Cancer 2015, 138, 652–663. [Google Scholar] [CrossRef]
- Song, D.H.; Ko, G.H.; Lee, J.H.; Lee, J.S.; Yang, J.W.; Kim, M.H.; An, H.J.; Kang, M.H.; Jeon, K.N.; Kim, D.C. Prognostic role of myoferlin expression in patients with clear cell renal cell carcinoma. Oncotarget 2017, 8, 89033–89039. [Google Scholar] [CrossRef] [Green Version]
- Rademaker, G.; Costanza, B.; Bellier, J.; Herfs, M.; Peiffer, R.; Agirman, F.; Maloujahmoum, N.; Habraken, Y.; Delvenne, P.; Bellahcène, A.; et al. Human colon cancer cells highly express myoferlin to maintain a fit mitochondrial network and escape p53-driven apoptosis. Oncogenesis 2019, 8, 21. [Google Scholar] [CrossRef] [Green Version]
- Kim, M.H.; Song, D.H.; Ko, G.H.; Lee, J.H.; Kim, D.C.; Yang, J.W.; Lee, H.I.; An, H.J.; Lee, J.S. Myoferlin Expression and Its Correlation with FIGO Histologic Grading in Early-Stage Endometrioid Carcinoma. J. Pathol. Transl. Med. 2018, 52, 93–97. [Google Scholar] [CrossRef] [Green Version]
- Cox, A.; Tolkach, Y.; Stein, J.; Kristiansen, G.; Ritter, M.; Ellinger, J. Otoferlin is a prognostic biomarker in patients with clear cell renal cell carcinoma: A systematic expression analysis. Int. J. Urol. 2021, 28, 424–431. [Google Scholar] [CrossRef]
- Ahluwalia, P.; Ahluwalia, M.; Mondal, A.K.; Sahajpal, N.; Kota, V.; Rojiani, M.V.; Rojiani, A.M.; Kolhe, R. Prognostic and therapeutic implications of extracellular matrix associated gene signature in renal clear cell carcinoma. Sci. Rep. 2021, 11, 1–14. [Google Scholar] [CrossRef]
- Ohto, U.; Ishida, H.; Krayukhina, E.; Uchiyama, S.; Inoue, N.; Shimizu, T. Structure of IZUMO1–JUNO reveals sperm–oocyte recognition during mammalian fertilization. Nature 2016, 534, 566–569. [Google Scholar] [CrossRef]
| Gene | Primer Sequences (Sense/Antisense) | Efficiency | Annealing Temperature (°C) |
|---|---|---|---|
| β-actin | 5′-CATGGATGATGATATCGCCGCG-3′ 5′-ACATGATCTGGGTCATCTTCTCG-3′ | 1.84 | 69 |
| B2M | 5′-GCCTTAGCTGTGCTCGCGCT-3′ 5′-TGCTGCTTACATGTCTCGATCCCA-3′ | 2.04 | 64 |
| GAPDH | 5′- TGGTATCGTGGAAGGACTCA-3′ 5′-CCAGTAGAGGCAGGGATGAT-3′ | 1.93 | 67 |
| RPO | 5′-GCCTTGACCTTTTCAGCAAG-3′ 5′-GCAGCATCTACAACCCTGAAG-3′ | 1.97 | 62 |
| CatsperB | 5′-TCTTTTTGGACAGCCTCCAGATATGGG-3′ 5′-ACAAGGCCTGAACAACTTGTGAATCA-3′ | 2.02 | 62 |
| CatsperD | 5′-GGTGGAGCTGTGGCGAAAAGAC-3′ 5′-CTCCAGTTGACAGTAGCTGTAGTACGG-3′ | 2.08 | 57 |
| Dysferlin | 5′-TGGTGGTCAAAGACCATGAG-3′ 5′-ACATCCAGGTCAGGCAGAGT-3′ | 1.89 | 55 |
| Fer1L5 | 5′-CGCTTGTCGGGCAAGGTGAAG-3′ 5′-GCCAAATAGTGCGGAGCTGAATAGATG-3′ | 1.94 | 56 |
| Juno | 5′-CTCTATGAGGAGTGCATCCCCTG-3′ 5′-CGGTTCTTCCCCTGACTCCAG-3′ | 2.06 | 60 |
| Myoferlin | 5′-TGTGGAATCTGCCAGCAATA-3′ 5′-CAGGTCCTTCAGGGCTACAG-3′ | 1.88 | 62 |
| Otoferlin | 5′-ATGGCCACCGGGGAGGTGGA-3′ 5′-AGCTCGTGTCGGGCCGGTTG-3′ | 2.01 | 63 |
| ZP2 | 5′-GCTCTCTAGCCTGGTCTACTTCCACT -3′ 5′-GTCCATAGCACCTCGTGAGCCA-3′ | 2.08 | 69 |
| ZP3 | 5′GGATGTGTCCGGTGCCATAG-3′ 5′-CACTCGTGGAGTCCAACCTC-3′ | 2.05 | 61 |
| Oral Mucosa | OSCC | |
|---|---|---|
| Juno | n.d. | 0.0003 (0.00007) |
| CatsperB | 0.001 (0.0002) | 0.0005 (0.0001) |
| CatsperD | 0.0015 (0.0004) | n.d. |
| Dysferlin | 0.0005 (0.00009) | n.d. |
| Fer1L5 | 0.00085 (0.00014) | n.d. |
| Myoferlin | 0.022 (0.006) | 0.0078 (0.002) |
| Otoferlin | 0.0036 (0.0007) | n.d. |
| ZP2 | 0.0000047 (0.00000073) | 0.000001 (0.00000004) |
| ZP3 | 0.000024 (0.000007) | 0.000081 (0.000023) |
| Oral Mucosa | OSCC | |
|---|---|---|
| Juno | n.d. | 10.0 * |
| CatsperB | 1.0 | 0.5 |
| CatsperD | 50.0 * | n.d. |
| Dysferlin | 16.7 * | n.d. |
| Fer1L5 | 25.0 * | n.d. |
| Myoferlin | 1.0 | 0.35 * |
| Otoferlin | 111.0 * | n.d. |
| ZP2 | 1.0 | 0.21 * |
| ZP3 | 1.0 | 0.34 * |
| Oral Mucosa | OSCC | |||
|---|---|---|---|---|
| + | − | + | − | |
| Juno | 0 | 100 | 100 | 0 |
| CatsperB | 100 | 0 | 100 | 0 |
| CatsperD | 100 | 0 | 0 | 100 |
| Dysferlin | 100 | 0 | 0 | 100 |
| Fer1L5 | 100 | 0 | 0 | 100 |
| Myoferlin | 100 | 0 | 100 | 0 |
| Otoferlin | 100 | 0 | 0 | 100 |
| ZP2 | 45 | 55 | 38 | 62 |
| ZP3 | 100 | 0 | 64 | 36 |
| Patient | Sex | Age | Staging/Grading | Tumor Site |
|---|---|---|---|---|
| 1 | m | 70 | pT2, N0, M0, L0, V0, Pn0, R0, G1 | jaw |
| 2 | f | 69 | pT2, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 3 | m | 67 | pT2, N2b, M0, L0, V0, Pn1, R0, G1 | jaw |
| 4 (†) | m | 64 | pT2, N2c, M0, L1, V0, Pn0, R0, G2 | tongue |
| 5 | m | 72 | pT2, N0, M0, L1, V0, Pn0, R0, G2 | jaw |
| 6 (†) | m | 69 | pT2, N2c, M0, L1, V0, Pn1, R0, G3 | jaw |
| 7 | f | 72 | pT1, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 8 | m | 63 | pT1, N0, M0, L0, V0, Pn0, R0, G1 | tongue |
| 9 | m | 59 | pT2, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 10 (†) | m | 60 | pT2, N0, M0, L0, V0, Pn0, R1, G3-4 | jaw |
| 11 | m | 73 | pT2, N0, M0, L0, V0, Pn1, R0, G2 | tongue |
| 12 | m | 78 | pT2b, N2b, M0, L0, V0, Pn0, R0, G2 | tongue |
| 13 | f | 60 | pT3, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 14 (†) | f | 82 | pT4a, N2c, M0, L0, V0, Pn0, R0, G2-3 | jaw |
| 15 | f | 62 | pT1, N0, M0, L0, V0, Pn0, R0, G1 | tongue |
| 16 (†) | m | 96 | pT3, Nx, M0, L0, V0, Pn1, R0, G2 | jaw |
| 17 | f | 60 | pT2, N1, M0, L0, V0, Pn0, R0, G2 | tongue |
| 18 | m | 65 | pT2, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 19 | m | 47 | pT1, N0, M0, L0, V0, Pn0, R0, G2 | tongue |
| 20 | m | 70 | pT2, N1, M0, L0, V0, Pn0, R0, G2 | tongue |
| 21 | f | 69 | pT2, N0, M0, L0, V0, Pn0, R0, G2 | jaw |
| 22 | m | 62 | pT3, N0, M0, L0, V0, Pn0, R0, G3 | jaw |
| 23 (†) | f | 98 | pT4a, N2b, M0, L0, V1, Pn1, R1, G2 | jaw |
| 24 (†) | m | 62 | pT4a, N0, M0, L0, V0, Pn0, R0, G3 | jaw |
| 25 | m | 78 | pT1, N0, M0, L0, V0, Pn0, R0, G1 | tongue |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kraus, D.; Weider, S.; Probstmeier, R.; Winter, J. Neoexpression of JUNO in Oral Tumors Is Accompanied with the Complete Suppression of Four Other Genes and Suggests the Application of New Biomarker Tools. J. Pers. Med. 2022, 12, 494. https://doi.org/10.3390/jpm12030494
Kraus D, Weider S, Probstmeier R, Winter J. Neoexpression of JUNO in Oral Tumors Is Accompanied with the Complete Suppression of Four Other Genes and Suggests the Application of New Biomarker Tools. Journal of Personalized Medicine. 2022; 12(3):494. https://doi.org/10.3390/jpm12030494
Chicago/Turabian StyleKraus, Dominik, Simone Weider, Rainer Probstmeier, and Jochen Winter. 2022. "Neoexpression of JUNO in Oral Tumors Is Accompanied with the Complete Suppression of Four Other Genes and Suggests the Application of New Biomarker Tools" Journal of Personalized Medicine 12, no. 3: 494. https://doi.org/10.3390/jpm12030494
APA StyleKraus, D., Weider, S., Probstmeier, R., & Winter, J. (2022). Neoexpression of JUNO in Oral Tumors Is Accompanied with the Complete Suppression of Four Other Genes and Suggests the Application of New Biomarker Tools. Journal of Personalized Medicine, 12(3), 494. https://doi.org/10.3390/jpm12030494








