MicroRNA Alterations Induced in Human Skin by Diesel Fumes, Ozone, and UV Radiation
Abstract
1. Introduction
2. Materials and Methods
2.1. Ex Vivo Human Skin Explants Preparation
2.2. Ex Vivo Human Skin Explants Ozone (O3) Exposure
2.3. Ex Vivo Human Skin Explants Diesel Engine Exhaust (Diesel) Exposure
2.4. Ex Vivo Human Skin Explants Ultraviolet Light (UV) Exposure
2.5. Total RNA Extraction and Lyophilization
2.6. miRNA-Microarray and Bioinformatic Analyses
3. Results
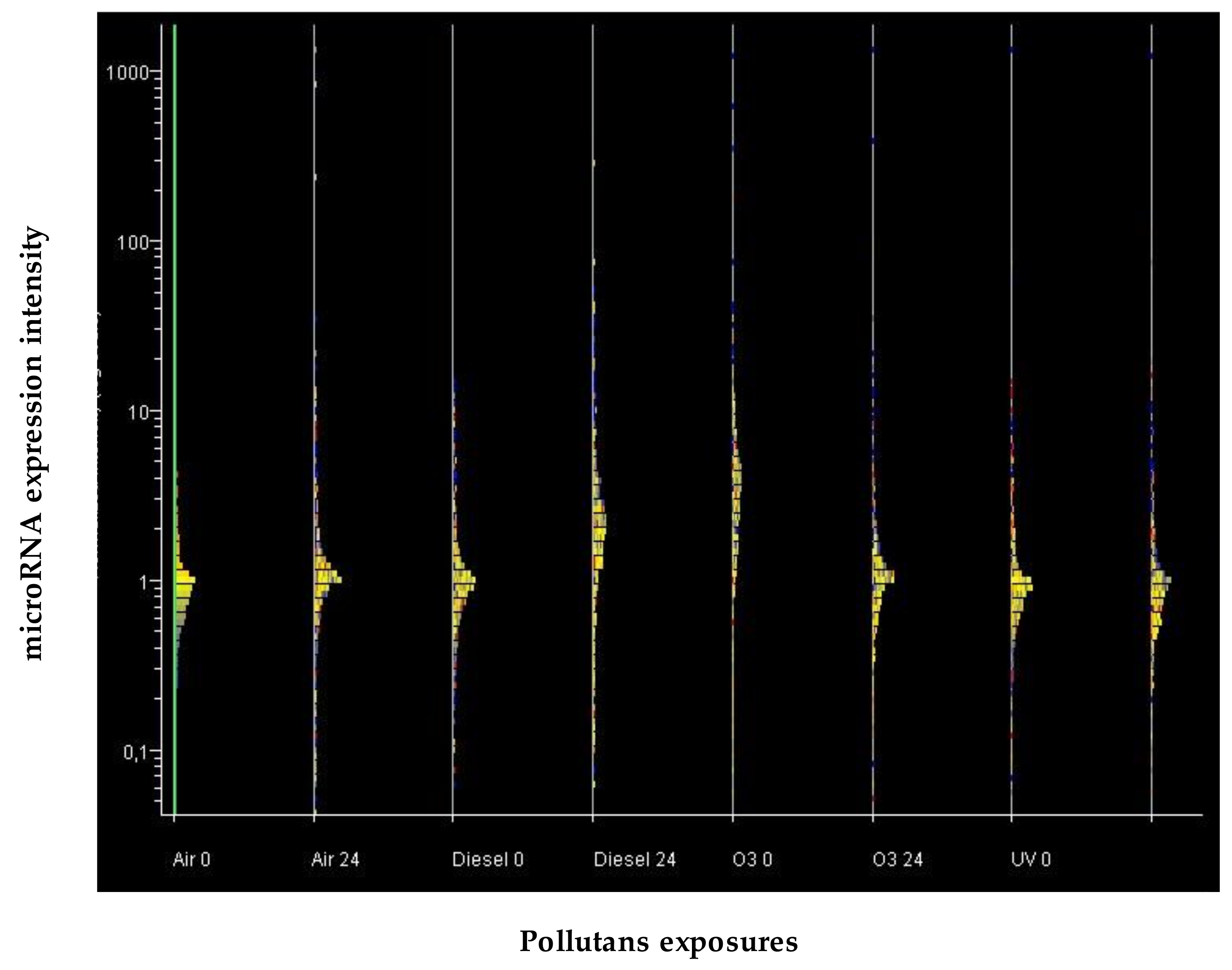
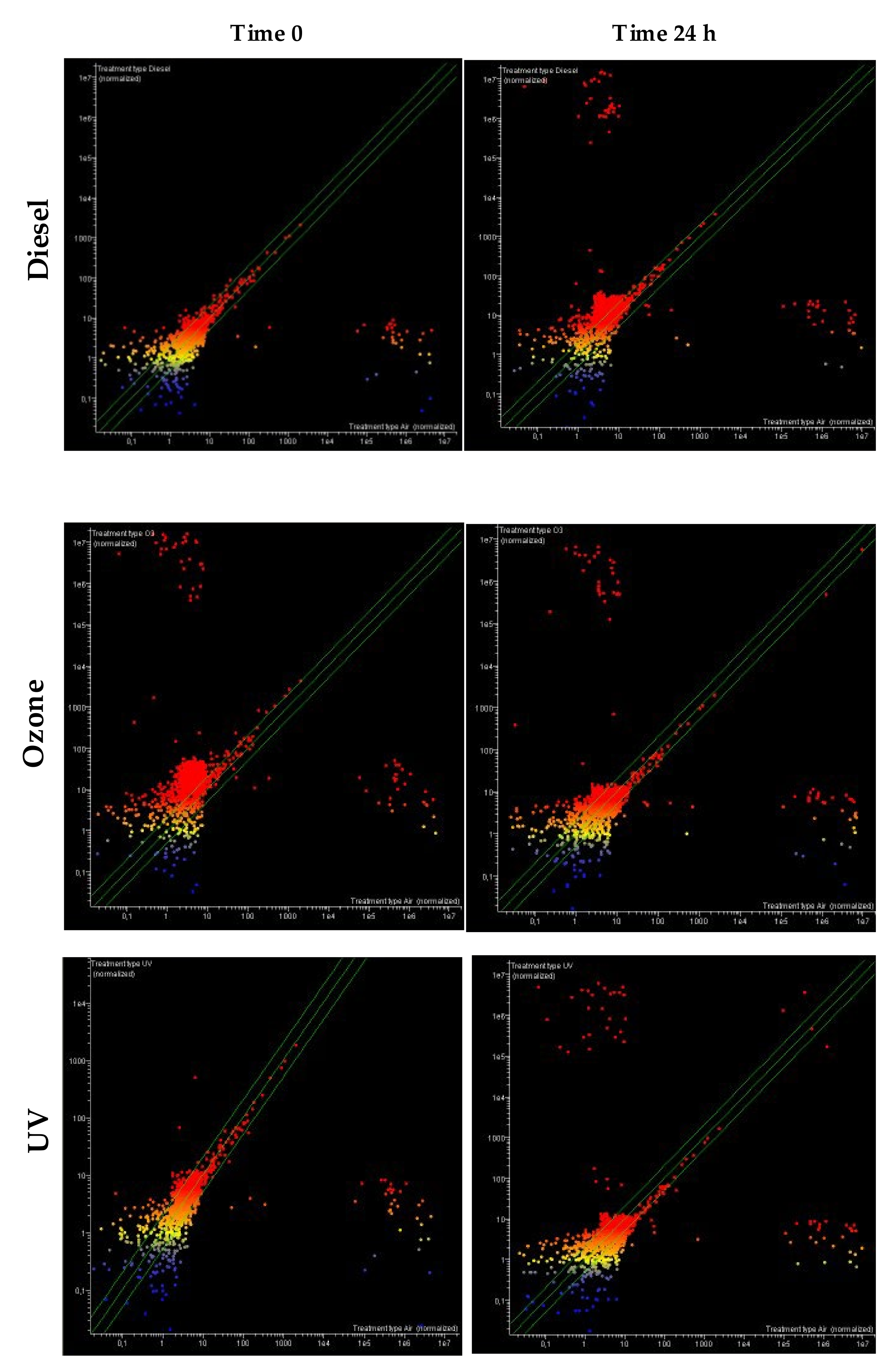
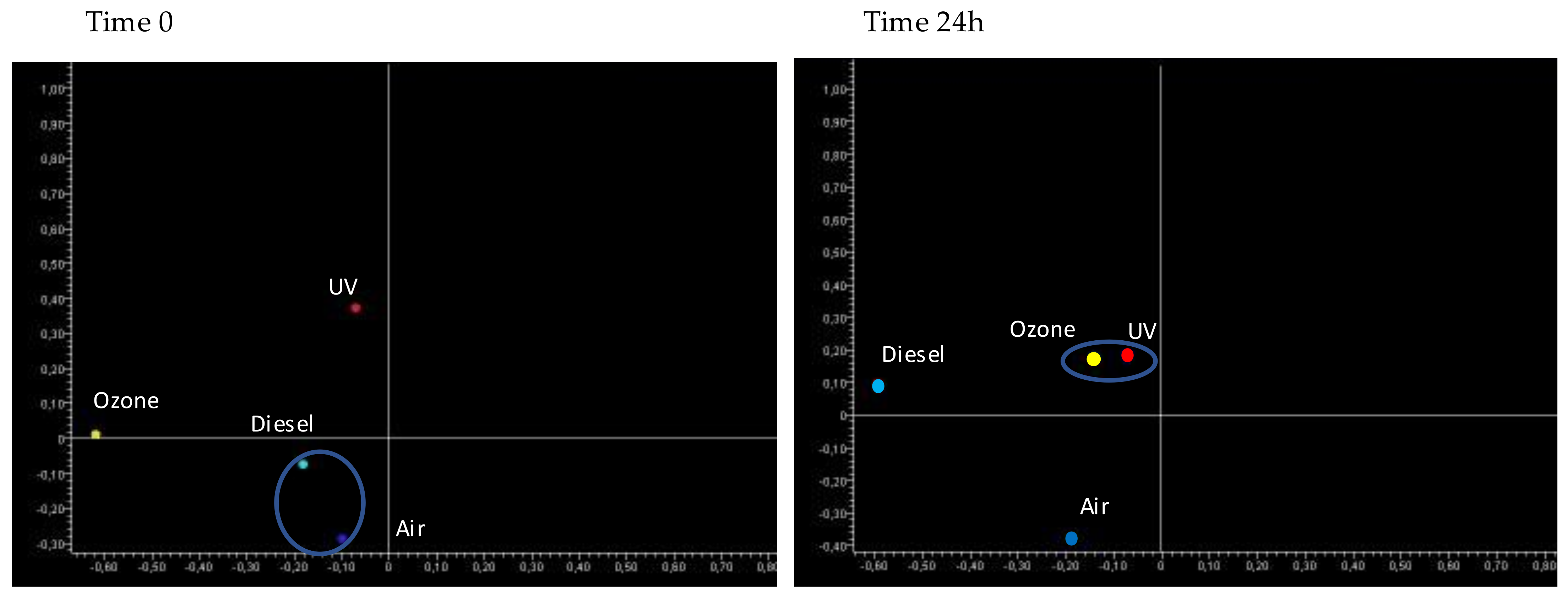
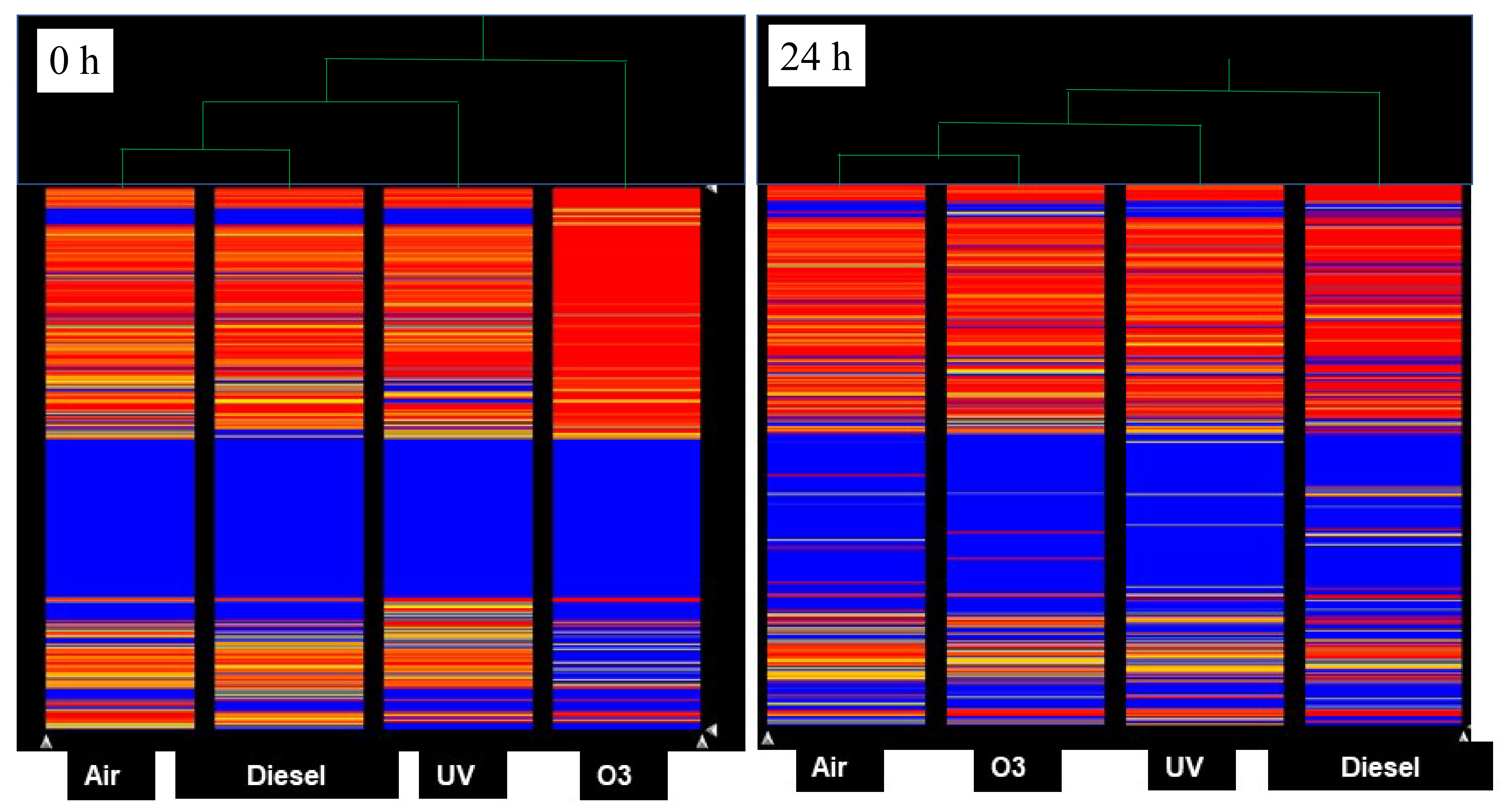
Comparison of miRNA Expression Profile between Pollutants
4. Discussion and Conclusions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lelieveld, J.; Pozzer, A.; Pöschl, U.; Fnais, M.; Haines, A.; Münzel, T. Loss of life expectancy from air pollution compared to other risk factors: A worldwide perspective. Cardiovasc. Res. 2020, 116, 1910–1917. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). WHO’s Urban Ambient Air Pollution Database—Update 2016; WHO: Geneva, Switzerland, 2016. [Google Scholar]
- Fuchs, E. Epithelial Skin Biology. Three Decades of Developmental Biology, a Hundred Questions Answered and a Thousand New Ones to Address. Curr. Top. Dev. Biol. 2016, 116, 357–374. [Google Scholar] [CrossRef] [PubMed]
- Feingold, K.R.; Schmuth, M.; Elias, P.M. The regulation of permeability barrier homeostasis. J. Investig. Dermatol. 2007, 127, 1574–1576. [Google Scholar] [CrossRef]
- Valacchi, G.; Magnani, N.; Woodby, B.; Ferreira, S.M.; Evelson, P. Particulate Matter Induces Tissue OxInflammation: From Mechanism to Damage. Antioxid. Redox Signal. 2020, 33, 308–326. [Google Scholar] [CrossRef] [PubMed]
- Criteria Air Pollutants. Available online: https://www.epa.gov/criteria-air-pollutants (accessed on 30 September 2021).
- Fuks, K.B.; Hüls, A.; Sugiri, D.; Altug, H.; Vierkötter, A.; Abramson, M.J.; Goebel, J.; Wagner, G.G.; Demuth, I.; Krutmann, J.; et al. Tropospheric ozone and skin aging: Results from two German cohort studies. Environ. Int. 2019, 124, 139–144. [Google Scholar] [CrossRef]
- Xu, F.; Yan, S.; Wu, M.; Li, F.; Xu, X.; Song, W.; Zhao, J.; Xu, J.; Kan, H. Ambient ozone pollution as a risk factor for skin disorders. Br. J. Dermatol. 2011, 165, 224–225. [Google Scholar] [CrossRef]
- Kousha, T.; Valacchi, G. The air quality health index and emergency department visits for urticaria in Windsor, Canada. J. Toxicol. Environ. Health Part A Curr. Issues 2015, 78, 524–533. [Google Scholar] [CrossRef]
- Valacchi, G.; Porada, E.; Rowe, B.H. Ambient ozone and bacterium streptococcus: A link between cellulitis and pharyngitis. Int. J. Occup. Med. Environ. Health 2015, 28, 771–774. [Google Scholar] [CrossRef]
- Lademann, J.; Otberg, N.; Jacobi, U.; Hoffman, R.M.; Blume-Peytavi, U. Follicular penetration and targeting. J. Investig. Dermatol. Symp. Proc. 2005, 10, 301–303. [Google Scholar] [CrossRef]
- Dijkhoff, I.M.; Drasler, B.; Karakocak, B.B.; Petri-Fink, A.; Valacchi, G.; Eeman, M.; Rothen-Rutishauser, B. Impact of airborne particulate matter on skin: A systematic review from epidemiology to in vitro studies. Part. Fibre Toxicol. 2020, 17, 35. [Google Scholar] [CrossRef]
- Ferrara, F.; Woodby, B.; Pecorelli, A.; Schiavone, M.L.; Pambianchi, E.; Messano, N.; Therrien, J.P.; Choudhary, H.; Valacchi, G. Additive effect of combined pollutants to UV induced skin OxInflammation damage. Evaluating the protective topical application of a cosmeceutical mixture formulation. Redox Biol. 2020, 34, 1014481. [Google Scholar] [CrossRef] [PubMed]
- Plusquin, M.; Guida, F.; Polidoro, S.; Vermeulen, R.; Raaschou-Nielsen, O.; Campanella, G.; Hoek, G.; Kyrtopoulos, S.A.; Georgiadis, P.; Naccarati, A.; et al. DNA methylation and exposure to ambient air pollution in two prospective cohorts. Environ. Int. 2017, 108, 127–136. [Google Scholar] [CrossRef] [PubMed]
- Ding, R.; Jin, Y.; Liu, X.; Zhu, Z.; Zhang, Y.; Wang, T.; Xu, Y. Characteristics of DNA methylation changes induced by traffic-related air pollution. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2016, 15, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.J.; Söderhäll, C.; Bustamante, M.; Baïz, N.; Gruzieva, O.; Gehring, U.; Mason, D.; Chatzi, L.; Basterrechea, M.; Llop, S.; et al. DNA methylation in childhood asthma: An epigenome-wide meta-analysis. Lancet Respir. Med. 2018, 6, 379–388. [Google Scholar] [CrossRef]
- Alfano, R.; Herceg, Z.; Nawrot, T.S.; Chadeau-Hyam, M.; Ghantous, A.; Plusquin, M. The Impact of Air Pollution on Our Epigenome: How Far Is the Evidence? (A Systematic Review). Curr. Environ. Health Rep. 2018, 5, 544–578. [Google Scholar] [CrossRef]
- Weinhold, B. Epigenetics: The science of change. Environ. Health Perspect. 2006, 114, 160–167. [Google Scholar] [CrossRef]
- Jin, B.; Li, Y.; Robertson, K.D. DNA methylation: Superior or subordinate in the epigenetic hierarchy? Genes Cancer 2011, 2, 607–617. [Google Scholar] [CrossRef]
- Breton, C.V.; Marutani, A.N. Air Pollution and Epigenetics: Recent Findings. Curr. Environ. Health Rep. 2014, 1, 35–45. [Google Scholar] [CrossRef]
- Chuang, J.C.; Jones, P.A. Epigenetics and microRNAs. Pediatr. Res. 2007, 61, 24–29. [Google Scholar] [CrossRef]
- Cancer, I.A. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans, No. 100D.; World Health Organization: Geneva, Switzerland, 2012; Volume 100D, p. 341. ISBN 978-9283213215. [Google Scholar]
- Rooks, M.G.; Veiga, P.; Wardwell-Scott, L.H.; Tickle, T.; Segata, N.; Michaud, M.; Gallini, C.A.; Beal, C.; Van Hylckama-Vlieg, J.E.T.; Ballal, S.A.; et al. Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME J. 2014, 8, 1403–1417. [Google Scholar] [CrossRef]
- Guo, B.; Hui, Q.; Xu, Z.; Chang, P.; Tao, K. miR-495 inhibits the growth of fibroblasts in hypertrophic scars. Aging 2019, 11, 2898–2910. [Google Scholar] [CrossRef]
- Jun, H.; Ying, H.; Daiwen, C.; Bing, Y.; Xiangbing, M.; Ping, Z.; Jie, Y.; Zhiqing, H.; Junqiu, L. MIR-628, a microRNA that is induced by Toll-like receptor stimulation, regulates porcine innate immune responses. Sci. Rep. 2015, 5, 1–10. [Google Scholar] [CrossRef]
- Chen, H.; Ji, X.; She, F.; Gao, Y.; Tang, P. MIR-628-3p regulates osteoblast differentiation by targeting RUNX2: Possible role in atrophic non-union. Int. J. Mol. Med. 2017, 39, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Dong, P.; Xiong, Y.; Yue, J.; Xu, D.; Ihira, K.; Konno, Y.; Kobayashi, N.; Todo, Y.; Watari, H. Long noncoding RNA NEAT1 drives aggressive endometrial cancer progression via miR-361-regulated networks involving STAT3 and tumor microenvironment-related genes. J. Exp. Clin. Cancer Res. 2019, 8, 38–295. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Cai, X.; Li, Q.; Xue, P.; Chen, Z.; Dong, X.; Xue, Y. Hsa-miR-875-5p exerts tumor suppressor function through down-regulation of EGFR in colorectal carcinoma (CRC). Oncotarget 2016, 7, 42225–42240. [Google Scholar] [CrossRef]
- Kang, N.; Ou, Y.; Wang, G.; Chen, J.; Li, D.; Zhan, Q. miR-875-5p exerts tumor-promoting function via down-regulation of CAPZA1 in esophageal squamous cell carcinoma. PeerJ 2021, 9, e10020. [Google Scholar] [CrossRef]
- Liang, J.-J.; Wang, J.-Y.; Zhang, T.-J.; An, G.-S.; Ni, J.-H.; Li, S.-Y.; Jia, H.-T. MiR-509-3-5p-NONHSAT112228.2 Axis Regulates p21 and Suppresses Proliferation and Migration of Lung Cancer Cells. Curr. Top. Med. Chem. 2020, 20, 835–846. [Google Scholar] [CrossRef]
- Ram Kumar, R.M.; Boro, A.; Fuchs, B. Involvement and Clinical Aspects of MicroRNA in Osteosarcoma. Int. J. Mol. Sci. 2016, 17, 877. [Google Scholar] [CrossRef]
- Yang, H.; Ren, J.; Bai, Y.; Jiang, J.; Xiao, S. Microrna-518-3p suppresses cell proliferation, invasiveness, and migration in colorectal cancer via targeting trip4. Biochem. Cell Biol. 2020, 98, 575–582. [Google Scholar] [CrossRef]
- Zhao, Y.; Lukiw, W.J. Bacteroidetes Neurotoxins and Inflammatory Neurodegeneration. Mol. Neurobiol. 2018, 55, 9100–9107. [Google Scholar] [CrossRef]
- Lu, L.; Zhang, H.; Dong, W.; Peng, W.; Yang, J. MiR-381 negatively regulates cardiomyocyte survival by suppressing Notch signaling. Vitr. Cell. Dev. Biol. Anim. 2018, 54, 610–619. [Google Scholar] [CrossRef]
- Reddy, S.D.N.; Pakala, S.B.; Ohshiro, K.; Rayala, S.K.; Kumar, R. MicroRNA-661, a c/EBPα target, inhibits metastatic tumor antigen 1 and regulates its functions. Cancer Res. 2009, 69, 5639–5642. [Google Scholar] [CrossRef] [PubMed]
- Yonemori, K.; Seki, N.; Idichi, T.; Kurahara, H.; Osako, Y.; Koshizuka, K.; Arai, T.; Okato, A.; Kita, Y.; Arigami, T.; et al. The microRNA expression signature of pancreatic ductal adenocarcinoma by RNA sequencing: Anti-tumour functions of the microRNA-216 cluster. Oncotarget 2017, 8, 70097–70115. [Google Scholar] [CrossRef] [PubMed]
- Son, G.H.; Kim, Y.; Lee, J.J.; Lee, K.Y.; Ham, H.; Song, J.E.; Park, S.T.; Kim, Y.H. MicroRNA-548 regulates high mobility group box 1 expression in patients with preterm birth and chorioamnionitis. Sci. Rep. 2019, 9, 19746. [Google Scholar] [CrossRef] [PubMed]
- Treiber, T.; Treiber, N.; Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 2019, 20, 5–20. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.H.; Huang, K.H.; Liu, K.; Wang, G.T.; Sun, Q. DGCR5 induces osteogenic differentiation by up-regulating Runx2 through miR-30d-5p. Biochem. Biophys. Res. Commun. 2018, 505, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wu, D.; Jiang, Z.; Zhang, Y.; Wang, S.; Ma, Z.; Hui, B.; Wang, J.; Qian, W.; Ge, Z.; et al. MiR-616-3p modulates cell proliferation and migration through targeting tissue factor pathway inhibitor 2 in preeclampsia. Cell Prolif. 2018, 51, e12490. [Google Scholar] [CrossRef]
- Ntoumou, E.; Tzetis, M.; Braoudaki, M.; Lambrou, G.; Poulou, M.; Malizos, K.; Stefanou, N.; Anastasopoulou, L.; Tsezou, A. Serum microRNA array analysis identifies miR-140-3p, miR-33b-3p and miR-671-3p as potential osteoarthritis biomarkers involved in metabolic processes. Clin. Epigenetics 2017, 12, 9–127. [Google Scholar] [CrossRef]
- Sun, S.; Su, C.; Zhu, Y.; Li, H.; Liu, N.; Xu, T.; Sun, C.; Lv, Y. MicroRNA-544a Regulates Migration and Invasion in Colorectal Cancer Cells via Regulation of Homeobox A10. Dig. Dis. Sci. 2016, 61, 2535–2544. [Google Scholar] [CrossRef]
- Shin, C.-H.; Byun, J.; Lee, K.; Kim, B.; Noh, Y.K.; Tran, N.L.; Park, K.; Kim, S.-H.; Kim, T.H.; Oh, S.J. Exosomal miRNA-19a and miRNA-614 Induced by Air Pollutants Promote Proinflammatory M1 Macrophage Polarization via Regulation of RORα Expression in Human Respiratory Mucosal Microenvironment. J. Immunol. 2020, 205, 3179–3190. [Google Scholar] [CrossRef]
- Xiong, G.; Zhang, J.; Zhang, Y.; Pang, X.; Wang, B.; Zhang, Y. Circular RNA_0074027 participates in cell proliferation, apoptosis and metastasis of colorectal cancer cells through regulation of miR-525-3p. Mol. Med. Rep. 2021, 23, 324. [Google Scholar] [CrossRef] [PubMed]
- Templin, C.; Volkmann, J.; Emmert, M.Y.; Mocharla, P.; Müller, M.; Kraenkel, N.; Ghadri, J.R.; Meyer, M.; Styp-Rekowska, B.; Briand, S.; et al. Increased proangiogenic activity of mobilized CD34 + progenitor cells of patients with acute ST-segment-elevation myocardial infarction: Role of differential MicroRNA-378 expression. Arterioscler. Thromb. Vasc. Biol. 2017, 37, 341–349. [Google Scholar] [CrossRef]
- Wang, H.; Chen, X.; Yang, B.; Xia, Z.; Chen, Q. MiR-924 as a tumor suppressor inhibits non-small cell lung cancer by inhibiting RHBDD1/Wnt/β-catenin signaling pathway. Cancer Cell Int. 2020, 8, 20–49. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Tan, Y.; Xu, B.; Lu, L.; Zhao, M.; Ma, J.; Liang, H.; Liu, J.; Yu, S. GPR30 Attenuates Myocardial Fibrosis in Diabetic Ovariectomized Female Rats: Role of iNOS Signaling. DNA Cell Biol. 2018, 37, 821–830. [Google Scholar] [CrossRef] [PubMed]
- Shuai, F.; Wang, B.; Dong, S. MiR-522-3p promotes tumorigenesis in human colorectal cancer via targeting bloom syndrome protein. Oncol. Res. 2018, 26, 1113–1121. [Google Scholar] [CrossRef]
- Vettori, A.; Pompucci, G.; Paolini, B.; Del Ciondolo, I.; Bressan, S.; Dundar, M.; Kenanoğlu, S.; Unfer, V.; Bertelli, M. Genetic background, nutrition and obesity: A review. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 1751–1761. [Google Scholar] [CrossRef]
- Wang, L.; Sun, M.; Cao, Y.; Ma, L.; Shen, Y.; Velikanova, A.A.; Li, X.; Sun, C.; Zhao, Y. miR-34a regulates lipid metabolism by targeting SIRT1 in non-alcoholic fatty liver disease with iron overload. Arch. Biochem. Biophys. 2020, 695, 108642. [Google Scholar] [CrossRef]
- Xiao, K.; Luo, X.; Wang, X.; Gao, Z. MicroRNA-185 regulates transforming growth factor-β1 and collagen-1 in hypertrophic scar fibroblasts. Mol. Med. Rep. 2017, 15, 1489–1496. [Google Scholar] [CrossRef]
- Li, K.P.; Fang, Y.P.; Liao, J.Q.; Duan, J.D.; Feng, L.G.; Luo, X.Z.; Liang, Z.J. Upregulation of miR-598 promotes cell proliferation and cell cycle progression in human colorectal carcinoma by suppressing INPP5E expression. Mol. Med. Rep. 2018, 17, 2991–2997. [Google Scholar] [CrossRef]
- Xiao, Y. MiR-486-5p inhibits the hyperproliferation and production of collagen in hypertrophic scar fibroblasts via IGF1/PI3K/AKT pathway. J. Dermatolog. Treat. 2020, 32, 973–982. [Google Scholar] [CrossRef]
- Pedersen, I.M.; Cheng, G.; Wieland, S.; Volinia, S.; Croce, C.M.; Chisari, F.V.; David, M. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 2007, 449, 919–922. [Google Scholar] [CrossRef] [PubMed]
- Feng, H.J.; Ouyang, W.; Liu, J.H.; Sun, Y.G.; Hu, R.; Huang, L.H.; Xian, J.L.; Jing, C.F.; Zhou, M.J. Global microRNA profiles and signaling pathways in the development of cardiac hypertrophy. Braz. J. Med. Biol. Res. 2014, 47, 361–368. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chaudhari, U.; Nemade, H.; Gaspar, J.A.; Hescheler, J.; Hengstler, J.G.; Sachinidis, A. MicroRNAs as early toxicity signatures of doxorubicin in human-induced pluripotent stem cell-derived cardiomyocytes. Arch. Toxicol. 2016, 90, 3087–3098. [Google Scholar] [CrossRef]
- Xia, X.; Wang, S.; Ni, B.; Xing, S.; Cao, H.; Zhang, Z.; Yu, F.; Zhao, E.; Zhao, G. Hypoxic gastric cancer-derived exosomes promote progression and metastasis via MiR-301a-3p/PHD3/HIF-1α positive feedback loop. Oncogene 2020, 39, 6231–6244. [Google Scholar] [CrossRef]
- Zhang, L.; Li, J.; Cui, L.; Shang, J.; Tian, F.; Wang, R.; Xing, G. Retraction notice to MicroRNA-30b promotes lipopolysaccharide-induced in fl ammatory injury and alleviates autophagy through JNK and NF- κ B. Biomed. Pharmacother. 2020, 128, 266021. [Google Scholar]
- Guan, C.; Wang, Y. LncRNA CASC9 attenuates lactate dehydrogenase-mediated oxidative stress and inflammation in spinal cord injury via sponging miR-383-5p. Inflammation 2021, 44, 923–933. [Google Scholar] [CrossRef]
- Sun, R.; Ge, L.; Cao, Y.; Wu, W.; Wu, Y.; Zhu, H.; Li, J.; Yu, D. Corrigendum to “MiR-429 regulates blood-spinal cord barrier permeability by targeting Krüppel-like factor 6”. Biochem. Biophys. Res. Commun. 2020, 525, 740–746. [Google Scholar] [CrossRef]
- Chen, X.F.; Zhang, L.J.; Zhang, J.; Dou, X.; Shao, Y.; Jia, X.J.; Zhang, W.; Yu, B. MiR-151a is involved in the pathogenesis of atopic dermatitis by regulating interleukin-12 receptor β2. Exp. Dermatol. 2018, 27, 427–432. [Google Scholar] [CrossRef]
- Xiao, S.; Zhang, M.; Liu, C.; Wang, D. MiR-514 attenuates proliferation and increases chemoresistance by targeting ATP binding cassette subfamily in ovarian cancer. Mol. Genet. Genom. 2018, 293, 1159–1167. [Google Scholar] [CrossRef]
- Chu, H.T.; Li, L.; Jia, M.; Diao, L.L.; Li, Z.B. Correlation between serum microRNA-136 levels and RAAS biochemical markers in patients with essential hypertension. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 11761–11767. [Google Scholar] [CrossRef]
- Mishima, Y.; Stahlhut, C.; Giraldez, A.J. miR-1-2 Gets to the Heart of the Matter. Cell 2007, 20, 247–249. [Google Scholar] [CrossRef] [PubMed]
- Qi, B.; Dong, Y.; Qiao, X.L. Effects of miR-18a on proliferation and apoptosis of gastric cancer cells by regulating RUNX1. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 9957–9964. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Yu, M.; Lin, Y.; Hou, B.; Wu, Z.; Li, Z.; Sun, J. MiR-502-3P suppresses cell proliferation, migration, and invasion in hepatocellular carcinoma by targeting SET. Onco. Targets. Ther. 2016, 9, 3281–3289. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chu, X.; Wei, Q. Mir-451 promotes cell apoptosis and inhibits autophagy in pediatric acute myeloid leukemia by targeting hmgb1. J. Environ. Pathol. Toxicol. Oncol. 2021, 40, 45–53. [Google Scholar] [CrossRef]
- Rodosthenous, R.S.; Coull, B.A.; Lu, Q.; Vokonas, P.S.; Schwartz, J.D.; Baccarelli, A.A. Ambient particulate matter and microRNAs in extracellular vesicles: A pilot study of older individuals. Part. Fibre Toxicol. 2016, 13, 13. [Google Scholar] [CrossRef]
- Wang, Z.; Liu, W.; Wang, C.; Ai, Z. Mir-873-5p inhibits cell migration and invasion of papillary thyroid cancer via regulation of CXCL16. Onco. Targets. Ther. 2020, 13, 1037–1046. [Google Scholar] [CrossRef]
- Tu, H.; Wei, G.; Cai, Q.; Chen, X.X.; Sun, Z.; Cheng, C.; Zhang, L.; Feng, Y.; Zhou, H.; Zhou, B.; et al. MicroRNA-212 inhibits hepatocellular carcinoma cell proliferation and induces apoptosis by targeting FOXAI. Onco. Targets. Ther. 2015, 24, 2227–2235. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, Y.; Dou, L. MiR-552 promotes the proliferation and metastasis of cervical cancer cells through targeting MUC15 pathway. J. Cancer 2021, 12, 6094–6104. [Google Scholar] [CrossRef]
- Fang, L.; Xu, X.; Lu, Y.; Wu, Y.Y.; Li, J.J. MicroRNA-495 attenuates proliferation and inflammatory response in rheumatoid arthritis fibroblast-like synoviocytes through attenuating β-catenin pathway. J. Biol. Regul. Homeost. Agents 2020, 34, 837–844. [Google Scholar] [CrossRef]
- Zhang, K.; Guo, L. MiR-767 promoted cell proliferation in human melanoma by suppressing CYLD expression. Gene 2018, 641, 272–278. [Google Scholar] [CrossRef]
- Sun, L.; Trajkovski, M. MiR-27 orchestrates the transcriptional regulation of brown adipogenesis. Metabolism 2014, 63, 272–282. [Google Scholar] [CrossRef] [PubMed]
- Si, Z.; Yu, L.; Jing, H.; Wu, L.; Wang, X. Oncogenic lncRNA ZNF561-AS1 is essential for colorectal cancer proliferation and survival through regulation of miR-26a-3p/miR-128-5p-SRSF6 axis. J. Exp. Clin. Cancer Res. 2021, 40, 78. [Google Scholar] [CrossRef] [PubMed]
- Tao, X.; Yang, X.; Wu, K.; Yang, L.; Huang, Y.; Jin, Q.; Chen, S. miR-629–5p promotes growth and metastasis of hepatocellular carcinoma by activating β-catenin. Exp. Cell Res. 2019, 15, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Gan, L.; Zhang, J. miR-543 inhibites cervical cancer growth and metastasis by targeting TRPM7. Chem. Biol. Interact. 2019, 302, 83–92. [Google Scholar] [CrossRef]
- Zhang, M.; Muralimanoharan, S.; Wortman, A.C.; Mendelson, C.R. Primate-specific miR-515 family members inhibit key genes in human trophoblast differentiation and are upregulated in preeclampsia. Proc. Natl. Acad. Sci. USA 2016, 113, 7069–7076. [Google Scholar] [CrossRef]
- Witwer, K.W.; Sarbanes, S.L.; Liu, J.; Clements, J.E. A plasma microRNA signature of acute lentiviral infection: Biomarkers of central nervous system disease. AIDS 2011, 25, 2057–2067. [Google Scholar] [CrossRef]
- Li, B.; Wang, Y.; Li, S.; He, H.; Sun, F.; Wang, C.; Lu, Y.; Wang, X.; Tao, B. Decreased expression of miR-378 correlates with tumor invasiveness and poor prognosis of patients with glioma. Int. J. Clin. Exp. Pathol. 2015, 8, 7016–7021. [Google Scholar]
- Li, Q.; Tian, Y.; Liang, Y.; Li, C. CircHIPK3/miR-876-5p/PIK3R1 axis regulates regulation proliferation, migration, invasion, and glutaminolysis in gastric cancer cells. Cancer Cell Int. 2020, 13, 391. [Google Scholar] [CrossRef]
- Guo, M.; Jiang, Z.; Zhang, X.; Lu, D.; Ha, A.D.; Sun, J.; Du, W.; Wu, Z.; Hu, L.; Khadarian, K.; et al. miR-656 inhibits glioma tumorigenesis through repression of BMPR1A. Carcinogenesis 2014, 35, 1698–1706. [Google Scholar] [CrossRef]
- Lv, L.; Wang, X.; Ma, T. MicroRNA-944 inhibits the malignancy of hepatocellular carcinoma by directly targeting IGF-1R and deactivating the PI3K/Akt signaling pathway. Cancer Manag. Res. 2019, 11, 2531–2543. [Google Scholar] [CrossRef]
- Zhang, W.; Zhong, B.; Zhang, C.; Luo, C.; Zhan, Y. miR-373 regulates inflammatory cytokine-mediated chondrocyte proliferation in osteoarthritis by targeting the P2X7 receptor. FEBS Open Bio 2018, 8, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Aqeilan, R.I.; Calin, G.A.; Croce, C.M. MiR-15a and miR-16-1 in cancer: Discovery, function and future perspectives. Cell Death Differ. 2010, 17, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wei, Z.; Wu, X.; Yang, H. Screening of exosomal miRNAs derived from subcutaneous and visceral adipose tissues: Determination of targets for the treatment of obesity and associated metabolic disorders. Mol. Med. Rep. 2018, 18, 3314–3324. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wang, L.; Jiang, L.; Zhang, X. Novel MicroRNA Biomarkers, miR-142-5p, miR-550a, miR-1826, and miR-1201, Were Identified for Primary Melanoma. J. Comput. Biol. 2020, 27, 815–824. [Google Scholar] [CrossRef]
- Jiang, X.; Huang, H.; Li, Z.; He, C.; Li, Y.; Chen, P.; Gurbuxani, S.; Arnovitz, S.; Hong, G.M.; Price, C.; et al. miR-495 is a tumor-suppressor microRNA down-regulated in MLL-rearranged leukemia. Proc. Natl. Acad. Sci. USA 2012, 109, 19397–19402. [Google Scholar] [CrossRef]
- Wei, Q.; Tu, Y.; Zuo, L.; Zhao, J.; Chang, Z.; Zou, Y.; Qiu, J. MiR-345-3p attenuates apoptosis and inflammation caused by oxidized low-density lipoprotein by targeting TRAF6 via TAK1/p38/NF-kB signaling in endothelial cells. Life Sci. 2020, 241, 117142. [Google Scholar] [CrossRef]
- Xu, X.; Zheng, S. MiR-887-3p negatively regulates StARD13 and promotes pancreatic cancer progression. Cancer Manag. Res. 2020, 12, 6137–6147. [Google Scholar] [CrossRef]
- Wu, K.; Yu, Z.; Tang, Z.; Wei, W.; Xie, D.; Xie, Y.; Xiao, Q. MiR-877-5p suppresses gastric cancer cell proliferation through targeting FOXM1. Onco. Targets. Ther. 2020, 13, 4731–4742. [Google Scholar] [CrossRef]
- Muti, P.; Donzelli, S.; Sacconi, A.; Hossain, A.; Ganci, F.; Frixa, T.; Sieri, S.; Krogh, V.; Berrino, F.; Biagioni, F.; et al. MiRNA-513a-5p inhibits progesterone receptor expression and constitutes a risk factor for breast cancer: The hOrmone and Diet in the ETiology of breast cancer prospective study. Carcinogenesis 2018, 39, 98–108. [Google Scholar] [CrossRef]
- Zhu, F.; Li, H.; Ding, F.; Guo, H.; Mou, H.; Ma, J. MiR-422a in gastric cancer cells directly targets CDC40 and modulates cell proliferation. Am. J. Transl. Res. 2020, 12, 4693. [Google Scholar]
- Wang, Y.; Yang, L.; Chen, T.; Liu, X.; Guo, Y.; Zhu, Q.; Tong, X.; Yang, W.; Xu, Q.; Huang, D.; et al. A novel lncRNA MCM3AP-AS1 promotes the growth of hepatocellular carcinoma by targeting miR-194-5p/FOXA1 axis. Mol. Cancer 2019, 18, 28. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Hu, J.; Wang, X.; Zhao, X.; Li, Z.; Niu, J.; Steer, C.J.; Zheng, G.; Song, G. MicroRNA-378 promotes hepatic inflammation and fibrosis via modulation of the NF-κB-TNFα pathway. J. Hepatol. 2019, 70, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.P.; Chen, L.L. Circular rna mat2b induces colorectal cancer proliferation via sponging mir-610, resulting in an increased e2f1 expression. Cancer Manag. Res. 2020, 12, 7107. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Long, G.; Zhao, C.; Li, H.; Chaugai, S.; Wang, Y.; Chen, C.; Wang, D.W. Atherosclerosis-related circulating miRNAs as novel and sensitive predictors for acute myocardial infarction. PLoS ONE 2014, 9, e105734. [Google Scholar] [CrossRef]
- Chen, Z.; Li, Y.; Tan, B.; Li, F.; Zhao, Q.; Fan, L.; Zhang, Z.; Zhao, X.; Liu, Y.; Wang, D. Long Non-coding RNA ASNR Targeting miR-519e-5p Promotes Gastric Cancer Development by Regulating FGFR2. Front. Cell Dev. Biol. 2021, 9, 679176. [Google Scholar] [CrossRef]
- Chen, F.; Liu, M.; Yu, Y.; Sun, Y.; Li, J.; Hu, W.; Wang, X.; Tong, D. LINC00958 regulated miR-627-5p/YBX2 axis to facilitate cell proliferation and migration in oral squamous cell carcinoma. Cancer Biol. Ther. 2019, 20, 1270–1280. [Google Scholar] [CrossRef]
- Liang, T.; Guo, L.; Liu, C. Genome-wide analysis of mir-548 gene family reveals evolutionary and functional implications. J. Biomed. Biotechnol. 2012, 2012, 679563. [Google Scholar] [CrossRef]
- Wang, L.; Li, H. MiR-770-5p facilitates podocyte apoptosis and inflammation in diabetic nephropathy by targeting TIMP3. Biosci. Rep. 2020, 40, BSR20193653. [Google Scholar] [CrossRef]
- Li, J.; Wang, L.; He, F.; Li, B.; Han, R. Long noncoding RNA LINC00629 restrains the progression of gastric cancer by upregulating AQP4 through competitively binding to miR-196b-5p. J. Cell. Physiol. 2020, 235, 2973–2985. [Google Scholar] [CrossRef]
- Zhu, H.; Hu, Y.; Wang, C.; Zhang, X.; He, D. CircGCN1L1 promotes synoviocyte proliferation and chondrocyte apoptosis by targeting miR-330-3p and TNF-α in TMJ osteoarthritis. Cell Death Dis. 2020, 11, 284. [Google Scholar] [CrossRef]
- Liu, T.; Feng, X.; Liao, Y. miR-617 Promotes the Growth of IL-22-Stimulated Keratinocytes Through Regulating FOXO4 Expression. Biochem. Genet. 2021, 59, 547–559. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Tang, Y.; Liu, Y.; Zhang, P.; Lv, L.; Zhang, X.; Jia, L.; Zhou, Y. Exosomes derived from miR-375-overexpressing human adipose mesenchymal stem cells promote bone regeneration. Cell Prolif. 2019, 52, e12669. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Gong, Y.; Xu, X.D.; Shen, H.; Gao, S.; Bao, H.D.; Guo, S.B.; Yu, X.F.; Gong, J. MicroRNA-936/ERBB4/Akt axis exhibits anticancer properties of gastric cancer through inhibition of cell proliferation, migration, and invasion. Kaohsiung J. Med. Sci. 2021, 37, 111–120. [Google Scholar] [CrossRef]
- Wang, P.; Wang, Z.; Liu, G.; Jin, C.; Zhang, Q.; Man, S.; Wang, Z. MiR-657 promotes macrophage polarization toward M1 by targeting FAM46C in gestational diabetes mellitus. Mediat. Inflamm. 2019, 2019, 4851214. [Google Scholar] [CrossRef] [PubMed]
- Garros, R.F.; Paul, R.; Connolly, M.; Lewis, A.; Garfield, B.E.; Natanek, S.A.; Bloch, S.; Mouly, V.; Griffiths, M.J.; Polkey, M.I.; et al. MicroRNA-542 promotes mitochondrial dysfunction and SMAD activity and is elevated in intensive care unit–acquired weakness. Am. J. Respir. Crit. Care Med. 2017, 196, 1422–1433. [Google Scholar] [CrossRef]
- Chen, Y.; Yu, H.; Zhu, D.; Liu, P.; Yin, J.; Liu, D.; Zheng, M.; Gao, J.; Zhang, C.; Gao, Y. miR-136-3p targets PTEN to regulate vascularization and bone formation and ameliorates alcohol-induced osteopenia. FASEB J. 2020, 34, 5348–5362. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y. MicroRNA-136-3p inhibits glioma tumorigenesis in vitro and in vivo by targeting KLF7. World J. Surg. Oncol. 2020, 18, 1–10. [Google Scholar] [CrossRef]
- Xue, Q.; Yang, D.; Zhang, J.; Gan, P.; Lin, C.; Lu, Y.; Zhang, W.; Zhang, L.; Guang, X. USP7, negatively regulated by miR-409-5p, aggravates hypoxia-induced cardiomyocyte injury. APMIS 2021, 129, 152–162. [Google Scholar] [CrossRef]
- Wang, Y.; Lin, W.; Ju, J. MicroRNA-409-5p promotes retinal neovascularization in diabetic retinopathy. Cell Cycle 2020, 19, 1314–1325. [Google Scholar] [CrossRef]
- Fan, X.D.; Luo, Y.; Wang, J.; An, N. MiR-154-3p and miR-487-3p synergistically modulate RHOA signaling in the carcinogenesis of thyroid cancer. Biosci. Rep. 2020, 40, BSR20193158. [Google Scholar] [CrossRef]
- Ma, J.; Wu, D.; Yi, J.; Yi, Y.; Zhu, X.; Qiu, H.; Kong, R.; Lin, J.; Qian, J.; Deng, Z. MiR-378 promoted cell proliferation and inhibited apoptosis by enhanced stem cell properties in chronic myeloid leukemia K562 cells. Biomed. Pharmacother. 2019, 112, 108623. [Google Scholar] [CrossRef] [PubMed]
- Tang, B.; Xuan, L.; Tang, M.; Wang, H.; Zhou, J.; Liu, J.; Wu, S.; Li, M.; Wang, X.; Zhang, H. miR-93-3p alleviates lipopolysaccharide-induced inflammation and apoptosis in H9c2 cardiomyocytes by inhibiting toll-like receptor 4. Pathol. Res. Pract. 2018, 214, 1686–1693. [Google Scholar] [CrossRef] [PubMed]
- Jin, W.; Chen, L.; Gao, F.; Yang, M.; Liu, Y.; Wang, B. Down-regulation of miR-556-3p inhibits hemangioma cell proliferation and promotes apoptosis by targeting VEGFC. Cell. Mol. Biol. 2020, 66, 204–207. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yang, S.; Song, J.; Yang, W.; Ji, Y.; Zhang, F.; Rao, J. Dysregulation of miR-23b-5p promotes cell proliferation via targeting FOXM1 in hepatocellular carcinoma. Cell Death Discov. 2021, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.; Mao, Y.; Xiong, L.; Li, L. Integrated Transcriptome Analysis of microRNA and mRNA in Mouse Skin Derived Precursors (SKPs) and SKP Derived Fibroblast (SFBs) by RNA-Seq. Curr. Genomics 2018, 20, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Tang, P. Clinical diagnostic value of circulating serum miR-509-3p in pulmonary arterial hypertension with congenital heart disease. Hell. J. Cardiol. 2020, 61, 26–30. [Google Scholar] [CrossRef]
- Stark, M.S.; Bonazzi, V.F.; Boyle, G.M.; Palmer, J.M.; Symmons, J.; Lanagan, C.M.; Schmidt, C.W.; Herington, A.C.; Ballotti, R.; Pollock, P.M.; et al. miR-514a regulates the tumour suppressor NF1 and modulates BRAFi sensitivity in melanoma. Oncotarget 2015, 6, 17753. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, K.; Yuan, X.; Xu, N.; Zhao, S.; Hou, L.; Yang, L.; Zhang, N. MiR-431-5p regulates cell proliferation and apoptosis in fibroblast-like synoviocytes in rheumatoid arthritis by targeting XIAP. Arthritis Res. Ther. 2020, 22, 231. [Google Scholar] [CrossRef]
- Wang, Y.; Lei, X.; Gao, C.; Xue, Y.; Li, X.; Wang, H.; Feng, Y. MiR-506-3p suppresses the proliferation of ovarian cancer cells by negatively regulating the expression of MTMR6. J. Biosci. 2019, 44, 126. [Google Scholar] [CrossRef]
- Zhu, B.; Tian, T.; Zhao, M. MiR-645 promotes proliferation and migration of non-small cell lung cancer cells by targeting TP53I11. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 6150–6156. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, Y.; Wei, Y.; Li, M.; Yu, S.; Ye, M.; Zhang, H.; Chen, S.; Liu, W.; Zhang, J. MiR-129-3p promotes docetaxel resistance of breast cancer cells via CP110 inhibition. Sci. Rep. 2015, 5, 15424. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Jiang, L.; Wei, G. Circ_0006168 promotes the migration, invasion and proliferation of esophageal squamous cell carcinoma cells via mir-516b-5p-dependent regulation of xbp1. Onco. Targets. Ther. 2021, 14, 475–2488. [Google Scholar] [CrossRef] [PubMed]
- Kong, M.; Han, Y.; Zhao, Y.; Zhang, H. miR-512-3p Overcomes Resistance to Cisplatin in Retinoblastoma by Promoting Apoptosis Induced by Endoplasmic Reticulum Stress. Med. Sci. Monit. 2020, 26, 923817. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Guo, Y.; Mi, N.; Zhou, L. miR-101-3p and miR-199b-5p promote cell apoptosis in oral cancer by targeting BICC1. Mol. Cell. Probes 2020, 52, 101567. [Google Scholar] [CrossRef]
- Liao, Z.J.; Zheng, Q.; Wei, T.; Zhang, Y.B.; Ma, J.Q.; Zhao, Z.; Sun, H.F.; Nan, K.J. MicroRNA-561 Affects Proliferation and Cell Cycle Transition through PTEN/AKT Signaling Pathway by Targeting P-REX2a in NSCLC. Oncol. Res. 2020, 28, 147–159. [Google Scholar] [CrossRef]
- He, Q.; Liu, N.; Hu, F.; Shi, Q.; Pi, X.; Chen, H.; Li, J.; Zhang, B. Circ_0061012 contributes to IL-22-induced proliferation, migration and invasion in keratinocytes through miR-194-5p/GAB1 axis in psoriasis. Biosci. Rep. 2021, 4, BSR20203130. [Google Scholar] [CrossRef]
- Wang, Y.P.; Li, H.Q.; Chen, J.X.; Kong, F.G.; Mo, Z.H.; Wang, J.Z.; Huang, K.M.; Li, X.N.; Yan, Y. Overexpression of XIST facilitates cell proliferation, invasion and suppresses cell apoptosis by reducing radio-sensitivity of glioma cells via miR-329-3p/CREB1 axis. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 3190–3203. [Google Scholar] [CrossRef]
- Wang, Y.F.; Lian, X.L.; Zhong, J.Y.; Su, S.X.; Xu, Y.F.; Xie, X.F.; Wang, Z.P.; Li, W.; Zhang, L.; Che, D.; et al. Serum exosomal microRNA let-7i-3p as candidate diagnostic biomarker for Kawasaki disease patients with coronary artery aneurysm. IUBMB Life 2019, 71, 891–900. [Google Scholar] [CrossRef]
- Liu, Z.; Dou, C.; Yao, B.; Xu, M.; Ding, L.; Wang, Y.; Jia, Y.; Li, Q.; Zhang, H.; Tu, K.; et al. Methylation-mediated repression of microRNA-129-2 suppresses cell aggressiveness by inhibiting high mobility group box 1 in human hepatocellular carcinoma. Oncotarget 2016, 7, 36909–36923. [Google Scholar] [CrossRef]
- Zhao, G.; Wang, T.; Huang, Q.K.; Pu, M.; Sun, W.; Zhang, Z.C.; Ling, R.; Tao, K.S. MicroRNA-548a-5p promotes proliferation and inhibits apoptosis in hepatocellular carcinoma cells by targeting Tg737. World J. Gastroenterol. 2016, 22, 5364–5373. [Google Scholar] [CrossRef]
- Jiang, Y.; Wang, N.; Yin, D.; Li, Y.K.; Guo, L.; Shi, L.P.; Huang, X. Changes in the Expression of Serum MiR-887-5p in Patients with Endometrial Cancer. Int. J. Gynecol. Cancer 2016, 26, 1143–1147. [Google Scholar] [CrossRef] [PubMed]
- Moura, S.R.; Bras, J.P.; Freitas, J.; Osório, H.; Barbosa, M.A.; Santos, S.G.; Almeida, M.I. miR-99a in bone homeostasis: Regulating osteogenic lineage commitment and osteoclast differentiation. Bone 2020, 134, 115303. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Hao, J.; Cen, X.; Song, K.; Yang, C.; Xiao, S.; Cheng, S. Downregulation of miR-487a-3p suppresses the progression of non-small cell lung cancer via targeting Smad7. Drug Dev. Res. 2021. [Google Scholar] [CrossRef]
- Liu, W.; Yang, Y.J.; An, Q. LINC00963 promotes ovarian cancer proliferation, migration and EMT via the miR-378g/CHI3L1 axis. Cancer Manag. Res. 2020, 12, 463–473. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Liu, H.; Cheng, Y.; Yang, J.; Liu, Y.; Wang, Y.; Yang, Z.; Shi, C.; Xu, Y. Association of variants in microRNA with Parkinson’s disease in Chinese Han population. Neurol. Sci. 2018, 39, 353–357. [Google Scholar] [CrossRef]
- Dong, L.; Zheng, Y.; Gao, L.; Luo, X. lncRNA NEAT1 prompts autophagy and apoptosis in MPTP-induced Parkinson’s disease by impairing miR-374c-5p. Acta Biochim. Biophys. Sin. 2021, 53, 870–882. [Google Scholar] [CrossRef]
- Zhu, M.; Zhang, N.; He, S.; Yan, R.; Zhang, J. MicroRNA-106a functions as an oncogene in human gastric cancer and contributes to proliferation and metastasis in vitro and in vivo. Clin. Exp. Metastasis 2016, 3, 509–519. [Google Scholar] [CrossRef]
- Long, C.Y.; Xiao, Y.X.; Li, S.Y.; Tang, X.B.; Yuan, Z.W.; Bai, Y.Z. Upregulation of miR-92a-2-5p potentially contribute to anorectal malformations by inhibiting proliferation and enhancing apoptosis via PRKCA/β-catenin. Biomed. Pharmacother. 2020, 127, 110117. [Google Scholar] [CrossRef]
- Zhu, L.M.; Li, N. Downregulation of long noncoding RNA TUSC7 promoted cell growth, invasion and migration through sponging with miR-616-5p/GSK3β pathway in ovarian cancer. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 7253–7265. [Google Scholar] [CrossRef]
- Fu, L.; Li, Z.; Zhu, J.; Wang, P.; Fan, G.; Dai, Y.; Zheng, Z.; Liu, Y. Serum expression levels of microRNA-382-3p, -598-3p, -1246 and -184 in breast cancer patients. Oncol. Lett. 2016, 12, 269–274. [Google Scholar] [CrossRef]
- Li, G.; Xu, Y.; Wang, S.; Yan, W.; Zhao, Q.; Guo, J. MiR-873-5p inhibits cell migration, invasion and epithelial-mesenchymal transition in colorectal cancer via targeting ZEB1. Pathol. Res. Pract. 2019, 215, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Su, D.; Qi, X.; Ma, J. MiR-500a-5p promotes glioblastoma cell proliferation, migration and invasion by targeting chromodomain helicase DNA binding protein 5. Mol. Med. Rep. 2018, 18, 2689–2696. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; He, C.; Qu, Y.; Chen, X.; Zhu, H.; Xiang, B. MiR-659-3p regulates the progression of chronic myeloid leukemia by targeting SPHK1. Int. J. Clin. Exp. Pathol. 2018, 11, 2470. [Google Scholar]
- Lin, Q.; Jia, Y.; Zhang, D.; Jin, H. NCK1-AS1 promotes the progression of melanoma by accelerating cell proliferation and migration via targeting miR-526b-5p/ADAM15 axis. Cancer Cell Int. 2021, 21, 1–10. [Google Scholar] [CrossRef]
- Zhu, F.; Li, Q.; Li, J.; Li, B.; Li, D. Long noncoding Mirt2 reduces apoptosis to alleviate myocardial infarction through regulation of the miR-764/PDK1 axis. Lab. Investig. 2021, 101, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Mi, Y.; Guan, B.; Zheng, B.; Wei, P.; Gu, Y.; Zhang, Z.; Cai, S.; Xu, Y.; Li, X.; et al. Tumor-derived exosomal miR-934 induces macrophage M2 polarization to promote liver metastasis of colorectal cancer. J. Hematol. Oncol. 2020, 13, 156. [Google Scholar] [CrossRef]
- Ye, X.Y.; Xu, L.; Lu, S.; Chen, Z.W. Mir-516a-5p inhibits the proliferation of non-small cell lung cancer by targeting hist3h2a. Int. J. Immunopathol. Pharmacol. 2019, 33, 2058738419841481. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X.; Ma, R.; Yang, S.; Jiang, L.; Wang, Z.; Zhu, Z.; Li, H. miR-520g and miR-520h overcome bortezomib resistance in multiple myeloma via suppressing APE1. Cell Cycle 2019, 18, 1660–1669. [Google Scholar] [CrossRef]
- Wang, J.; Wang, H.; Liu, A.; Fang, C.; Hao, J.; Wang, Z. Lactate dehydrogenase A negatively regulated by miRNAs promotes aerobic glycolysis and is increased in colorectal cancer. Oncotarget 2015, 6, 19465. [Google Scholar] [CrossRef]
- Su, X.; Gao, C.; Feng, X.; Jiang, M. miR-613 suppresses migration and invasion in esophageal squamous cell carcinoma via the targeting of G6PD. Exp. Ther. Med. 2020, 19, 3081–3089. [Google Scholar] [CrossRef]
- Wang, M.; Zhao, H.Y.; Zhang, J.L.; Wan, D.M.; Li, Y.M.; Jiang, Z.X. Dysregulation of LncRNA ANRIL mediated by miR-411–3p inhibits the malignant proliferation and tumor stem cell like property of multiple myeloma via hypoxia-inducible factor 1α. Exp. Cell Res. 2020, 396, 112280. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Li, A.L.; Xiong, W.Q.; Chen, R. Hsa Circ 001839 Promoted Inflammation in Renal Ischemia-Reperfusion Injury through NLRP3 by miR-432-3p. Nephron 2021, 145, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Fan, R.F.; Cao, C.Y.; Chen, M.H.; Shi, Q.X.; Xu, S.W. Gga-let-7f-3p promotes apoptosis in selenium deficiency-induced skeletal muscle by targeting selenoprotein K. Metallomics 2018, 10, 941–952. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B. Guizhi Fuling pills inhibit the proliferation, migration and invasion of human cutaneous malignant melanoma cells by regulating the molecular axis of LncRNA TPT1-AS1/miR-671-5p. Cell. Mol. Biol. 2020, 66, 148–154. [Google Scholar] [CrossRef]
- Li, Y.; Kuscu, C.; Banach, A.; Zhang, Q.; Pulkoski-Gross, A.; Kim, D.; Liu, J.; Roth, E.; Li, E.; Shroyer, K.R.; et al. miR-181a-5p inhibits cancer cell migration and angiogenesis via downregulation of matrix metalloproteinase-14. Cancer Res. 2015, 75, 2674–2685. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.A.; Wang, F.; Liu, Y.; Kriegel, A.J.; Geurts, A.M.; Usa, K.; Xue, H.; Wang, D.; Kong, Y.; Liang, M. MiR-192-5p in the Kidney Protects Against the Development of Hypertension. Hypertension 2019, 73, 399–406. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Z.; Gao, L.; Qiao, Z.; Yu, M.; Yu, B.; Yang, T. miR-1-3p suppresses proliferation of hepatocellular carcinoma through targeting SOX9. Onco. Targets. Ther. 2019, 12, 2149–2157. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Q. Long Noncoding RNA MALAT1 Knockdown Inhibits Proliferation, Migration, and Invasion and Promotes Apoptosis in Non-Small-Cell Lung Cancer Cells Through Regulating miR-515-3p/TRIM65 Axis. Cancer Biother. Radiopharm. 2020. [Google Scholar] [CrossRef]
- Liu, L.; Zhang, H.; Mao, H.; Li, X.; Hu, Y. Exosomal miR-320d derived from adipose tissue-derived MSCs inhibits apoptosis in cardiomyocytes with atrial fibrillation (AF). Artif. Cells Nanomed. Biotechnol. 2019, 47, 3976–3984. [Google Scholar] [CrossRef]
- Peng, X.; Wu, M.; Liu, W.; Guo, C.; Zhan, L.; Zhan, X. MiR-502-5p inhibits the proliferation, migration and invasion of gastric cancer cells by targeting SP1. Oncol. Lett. 2020, 20, 2757–2762. [Google Scholar] [CrossRef]
- Wu, X.; Chen, B.; Shi, H.; Zhou, J.; Zhou, F.; Cao, J.; Sun, X. miR-758-3p suppresses human bladder cancer cell proliferation, migration and invasion by targeting NOTCH2. Exp. Ther. Med. 2019, 17, 4273–4278. [Google Scholar] [CrossRef]
- Pourhanifeh, M.H.; Mahjoubin-Tehran, M.; Karimzadeh, M.R.; Mirzaei, H.R.; Razavi, Z.S.; Sahebkar, A.; Hosseini, N.; Mirzaei, H.; Hamblin, M.R. Autophagy in cancers including brain tumors: Role of MicroRNAs. Cell Commun. Signal. 2020, 18, 88. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Lei, S.; Sun, S.; Zhang, W.; Zhu, F.; Yang, H.; Xu, Q.; Zhang, B.; Li, H.; Zhu, M.; et al. Mir-324-3p regulates fibroblast proliferation via targeting tgf-β1 in atrial fibrillation. Int. Heart J. 2020, 61, 20–423. [Google Scholar] [CrossRef] [PubMed]
- Kordaß, T.; Weber, C.E.M.; Eisel, D.; Pane, A.A.; Osen, W.; Eichmüller, S.B. miR-193b and miR-30c-1* inhibit, whereas miR-576-5p enhances melanoma cell invasion in vitro. Oncotarget 2018, 9, 32507–32522. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Shao, W.; Zhao, J. MiR-520a-3p inhibits malignant progression of epithelial ovarian cancer by targeting SUV39H1 expression. Hum. Cell 2021, 34, 570–578. [Google Scholar] [CrossRef]
- Fang, Y.; Gu, X.; Li, Z.; Xiang, J.; Chen, Z. miR-449b inhibits the proliferation of SW1116 colon cancer stem cells through downregulation of CCND1 and E2F3 expression. Oncol. Rep. 2013, 30, 399–506. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Martínez, M.; Benito-Jardon, L.; Alonso, L.; Koetz-Ploch, L.; Hernando, E.; Teixido, J. miR-204-5p and miR-211-5p contribute to BRAF inhibitor resistance in melanoma. Cancer Res. 2018, 78, 1017–1030. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Xu, Z.; Wang, X.; Liu, F. Integrated bioinformatics analysis identifies microRNA-376a-3p as a new microRNA biomarker in patient with coronary artery disease. Am. J. Transl. Res. 2020, 12, 633–648. [Google Scholar]
- Han, X.; Du, C.; Chen, Y.; Zhong, X.; Wang, F.; Wang, J.; Liu, C.; Li, M.; Chen, S.; Li, B. Overexpression of miR-939-3p predicts poor prognosis and promotes progression in lung cancer. Cancer Biomark. 2019, 25, 335–342. [Google Scholar] [CrossRef]
- Xu, M.; Sun, J.; Yu, Y.; Pang, Q.; Lin, X.; Barakat, M.; Lei, R.; Xu, J. TM4SF1 involves in miR-1-3p/miR-214-5p-mediated inhibition of the migration and proliferation in keloid by regulating AKT/ERK signaling. Life Sci. 2020, 254, 117746. [Google Scholar] [CrossRef]
- Van Solingen, C.; Oldebeken, S.R.; Salerno, A.G.; Wanschel, A.C.B.A.; Moore, K.J. High-Throughput Screening Identifies MicroRNAs Regulating Human PCSK9 and Hepatic Low-Density Lipoprotein Receptor Expression. Front. Cardiovasc. Med. 2021, 8, 701. [Google Scholar] [CrossRef] [PubMed]
- Sassi, Y.; Avramopoulos, P.; Ramanujam, D.; Grüter, L.; Werfel, S.; Giosele, S.; Brunner, A.D.; Esfandyari, D.; Papadopoulou, A.S.; De Strooper, B.; et al. Cardiac myocyte miR-29 promotes pathological remodeling of the heart by activating Wnt signaling. Nat. Commun. 2017, 8, 1614. [Google Scholar] [CrossRef] [PubMed]
- Miao, L.J.; Huang, S.F.; Sun, Z.T.; Gao, Z.Y.; Zhang, R.X.; Liu, Y.; Wang, J. MiR-449c targets c-Myc and inhibits NSCLC cell progression. FEBS Lett. 2013, 587, 1359–1365. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.H.; Yu, G.S.; Deng, K.L.; Feng, Z.H.; Huang, Q.; Pan, B.; Deng, J.Z. Hsa_circ_0088233 Alleviates Proliferation, Migration, and Invasion of Prostate Cancer by Targeting hsa-miR-185-3p. Front. Cell Dev. Biol. 2020, 8, 528155. [Google Scholar] [CrossRef]
- Ye, W.; Ma, J.; Wang, F.; Wu, T.; He, M.; Li, J.; Pei, R.; Zhang, L.; Wang, Y.; Zhou, J. LncRNA MALAT 1 Regulates miR-144-3p to Facilitate Epithelial-Mesenchymal Transition of Lens Epithelial Cells via the ROS/NRF2/Notch1/Snail Pathway. Oxid. Med. Cell. Longev. 2020, 2020, 8184314. [Google Scholar] [CrossRef]
- Watanabe, K.; Yamaji, R.; Ohtsuki, T. MicroRNA-664a-5p promotes neuronal differentiation of SH-SY5Y cells. Genes Cells 2018, 23, 225–233. [Google Scholar] [CrossRef]
- Huang, D.; Liu, Y.; Gao, L.; Wei, X.; Xu, Y.; Cai, R.; Su, Q. MiR-32-3p Regulates Myocardial Injury Induced by Microembolism and Microvascular Obstruction by Targeting RNF13 to Regulate the Stability of Atherosclerotic Plaques. J. Cardiovasc. Transl. Res. 2021, 1–24. [Google Scholar] [CrossRef]
- Wang, G.; Liu, L.; Zhang, J.; Huang, C.; Chen, Y.; Bai, W.; Wang, Y.; Zhao, K.; Li, S. Lncrna hcg11 suppresses cell proliferation and promotes apoptosis via sponging mir-224-3p in non-small-cell lung cancer cells. Onco. Targets. Ther. 2020, 3, 6553–6563. [Google Scholar] [CrossRef]
- Pan, X.; He, Y.; Chen, Z.; Yan, G.; Ma, G. Circulating miR-130 is a potential bio signature for early prognosis of acute myocardial infarction. J. Thorac. Dis. 2020, 12, 7320–7325. [Google Scholar] [CrossRef]
- Krist, B.; Florczyk, U.; Pietraszek-Gremplewicz, K.; Józkowicz, A.; Dulak, J. The role of miR-378a in metabolism, angiogenesis, and muscle biology. Int. J. Endocrinol. 2015, 2015, 281756. [Google Scholar] [CrossRef]
- Kurowska, W.; Kuca-Warnawin, E.; Radzikowska, A.; Jakubaszek, M.; Maślińska, M.; Kwiatkowska, B.; Maśliński, W. Monocyte-related biomarkers of rheumatoid arthritis development in undifferentiated arthritis patients—A pilot study. Reumatologia 2018, 56, 10–16. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ma, H.; Huang, C.; Huang, Q.; Li, G.; Li, J.; Huang, B.; Zhong, Q.; Cao, C. Circular RNA circ_0014717 Suppresses Hepatocellular Carcinoma Tumorigenesis Through Regulating miR-668-3p/BTG2 Axis. Front. Oncol. 2021, 10, 3013. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Yin, Y.; Xie, L.; He, H. Plasma miR-18 screens acute myocardial infarction from healthy controls by targeting hypoxia inducible factor 1α. Clin. Lab. 2018, 64, 1207–1212. [Google Scholar] [CrossRef]
- Lupini, L.; Pepe, F.; Ferracin, M.; Braconi, C.; Callegari, E.; Pagotto, S.; Spizzo, R.; Zagatti, B.; Lanuti, P.; Fornari, F.; et al. Over-expression of the miR-483-3p overcomes the miR-145/TP53 pro-apoptotic loop in hepatocellular carcinoma. Oncotarget 2016, 7, 31361. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Qin, J.; Sun, H.; Xu, T.; Wang, S.; He, B. MiR-485-5p as a potential biomarker and tumor suppressor in human colorectal cancer. Biomark. Med. 2020, 14, 239–248. [Google Scholar] [CrossRef]
- Carlomosti, F.; D’Agostino, M.; Beji, S.; Torcinaro, A.; Rizzi, R.; Zaccagnini, G.; Maimone, B.; Di Stefano, V.; De Santa, F.; Cordisco, S.; et al. Oxidative Stress-Induced miR-200c Disrupts the Regulatory Loop among SIRT1, FOXO1, and eNOS. Antioxid. Redox Signal. 2017, 27, 328–344. [Google Scholar] [CrossRef]
- Li, S.N.; Li, P.; Liu, W.H.; Shang, J.J.; Qiu, S.L.; Zhou, M.X.; Liu, H.X. Danhong injection enhances angiogenesis after myocardial infarction by activating MiR-126/ERK/VEGF pathway. Biomed. Pharmacother. 2019, 120, 109538. [Google Scholar] [CrossRef]
- Tsai, M.M.; Huang, H.W.; Wang, C.S.; Lee, K.F.; Tsai, C.Y.; Lu, P.H.; Chi, H.C.; Lin, Y.H.; Kuo, L.M.; Lin, K.H. MicroRNA-26b inhibits tumor metastasis by targeting the KPNA2/c-jun pathway in human gastric cancer. Oncotarget 2016, 7, 39511–39526. [Google Scholar] [CrossRef]
- Skrzypek, K.; Tertil, M.; Golda, S.; Ciesla, M.; Weglarczyk, K.; Collet, G.; Guichard, A.; Kozakowska, M.; Boczkowski, J.; Was, H.; et al. Interplay between heme oxygenase-1 and miR-378 affects non-small cell lung carcinoma growth, vascularization, and metastasis. Antioxid. Redox Signal. 2013, 19, 644–660. [Google Scholar] [CrossRef]
- Wu, M.; Li, X.; Liu, Q.; Xie, Y.; Yuan, J.; Wanggou, S. miR-526b-3p serves as a prognostic factor and regulates the proliferation, invasion, and migration of glioma through targeting WEEI. Cancer Manag. Res. 2019, 11, 3099–3110. [Google Scholar] [CrossRef]
- Wang, Y.N.; Xu, F.; Zhang, P.; Wang, P.; Wei, Y.N.; Wu, C.; Cheng, S.J. MicroRNA-575 regulates development of gastric cancer by targeting PTEN. Biomed. Pharmacother. 2019, 113, 108716. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Xu, Y.; Ge, H.; Xing, B.; Li, G.; Li, G.; Wu, J. MiR-564 inhibits hepatocellular carcinoma cell proliferation and invasion by targeting the GRB2-ERK1/2-AKT axis. Oncotarget 2017, 8, 107543–107557. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, S.; Xu, Y.N.; Niu, X.; Li, Z.; Wang, J.F. miR-513a-5p targets Bcl-2 to promote dichlorvos induced apoptosis in HK-2 cells. Biomed. Pharmacother. 2018, 108, 876–882. [Google Scholar] [CrossRef] [PubMed]
- Shu, W.; Zhang, Y.; Zhang, C.; You, Q.; Zhou, H.; Wen, S. Triclosan inhibits the activation of human periodontal ligament fibroblasts induced by lipopolysaccharide from Porphyromonas gingivalis. J. Biomed. Res. 2021, 35, 206. [Google Scholar] [CrossRef]
- Wang, M.; Qiu, R.; Gong, Z.; Zhao, X.; Wang, T.; Zhou, L.; Lu, W.; Shen, B.; Zhu, W.; Xu, W. miR-188-5p emerges as an oncomiRNA to promote gastric cancer cell proliferation and migration via upregulation of SALL4. J. Cell. Biochem. 2019, 120, 15027–15037. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, M.; Sun, G.; Bai, Y.; Lv, D.; Liu, C. MiR-563 restrains cell proliferation via targeting LIN28B in human lung cancer. Thorac. Cancer 2020, 11, 55–61. [Google Scholar] [CrossRef]
- Zhang, W.; Xu, J.; Wang, K.; Tang, X.; He, J. MIR-139-3p suppresses the invasion and migration properties of breast cancer cells by targeting RAB1A. Oncol. Rep. 2019, 42, 1699–1708. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Hui, X.; Chong Hoo, R.L.; Ye, D.; Cheung Chan, C.Y.; Feng, T.; Wang, Y.; Ling Lam, K.S.; Xu, A. Adipocyte-secreted exosomal microRNA-34a inhibits M2 macrophage polarization to promote obesity-induced adipose inflammation. J. Clin. Investig. 2019, 129, 834–849. [Google Scholar] [CrossRef]
- Liu, X.; Feng, J.; Tang, L.; Liao, L.; Xu, Q.; Zhu, S. The regulation and function of mir-21-foxo3a-mir-34b/c signaling in breast cancer. Int. J. Mol. Sci. 2015, 16, 3148–3162. [Google Scholar] [CrossRef]
- Luo, X.; Zhang, X.; Peng, J.; Chen, Y.; Zhao, W.; Jiang, X.; Su, L.; Xie, M.; Lin, B. miR-371b-5p promotes cell proliferation, migration and invasion in non-small cell lung cancer via SCAI. Biosci. Rep. 2020, 40, BSR290200163. [Google Scholar] [CrossRef]
- Zhu, G.; Zhang, W.; Liu, Y.; Wang, S. MiR-371b-5p inhibits endothelial cell apoptosis in monocrotaline-induced pulmonary arterial hypertension via PTEN/PI3K/Akt signaling pathways. Mol. Med. Rep. 2018, 8, 5489–5501. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Chen, L.; Zhao, W.; Hao, J.; An, R. MicroRNA-let-7f-1 is induced by lycopene and inhibits cell proliferation and triggers apoptosis in prostate cancer. Mol. Med. Rep. 2016, 13, 2708–2714. [Google Scholar] [CrossRef] [PubMed]
- Qiu, J.; Hao, Y.; Huang, S.; Ma, Y.; Li, X.; Li, D.; Mao, Y. MIR-557 works as a tumor suppressor in human lung cancers by negatively regulating LEF1 expression. Tumor Biol. 2017, 39, 1010428317709467. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Qi, S.; Liao, R.; Su, D.; Wang, Y.; Xue, S. MiR-574–5p promotes the differentiation of human cardiac fibroblasts via regulating ARID3A. Biochem. Biophys. Res. Commun. 2020, 521, 427–433. [Google Scholar] [CrossRef]
- Wang, X.; Shi, J.; Niu, Z.; Wang, J.; Zhang, W. MiR-216a-3p regulates the proliferation, apoptosis, migration, and invasion of lung cancer cells via targeting COPB2. Biosci. Biotechnol. Biochem. 2020, 84, 2014–2027. [Google Scholar] [CrossRef]
- Tong, F.; Ying, Y.; Pan, H.; Zhao, W.; Li, H.; Zhan, X. Microrna-466 (Mir-466) functions as a tumor suppressor and prognostic factor in colorectal cancer (CRC). Bosn. J. Basic Med. Sci. 2018, 18, 252–259. [Google Scholar] [CrossRef]
- He, N.; Liu, L.; Ding, J.; Sun, Y.; Xing, H.; Wang, S. MiR-222-3p ameliorates glucocorticoid-induced inhibition of airway epithelial cell repair through down-regulating GILZ expression. J. Recept. Signal Transduct. 2020, 40, 301–312. [Google Scholar] [CrossRef]
- Yang, L.; Liu, Z.M.; Rao, Y.W.; Cui, S.Q.; Wang, H.; Jia, X.J. Downregulation of microRNA-586 Inhibits Proliferation, Invasion and Metastasis and Promotes Apoptosis in Human Osteosarcoma U2-OS Cell Line. Cytogenet. Genome Res. 2015, 146, 268–278. [Google Scholar] [CrossRef]
- Ramanathan, S.; Shenoda, B.B.; Lin, Z.; Alexander, G.M.; Huppert, A.; Sacan, A.; Ajit, S.K. Inflammation potentiates miR-939 expression and packaging into small extracellular vesicles. J. Extracell. Vesicles 2019, 8, 1650595. [Google Scholar] [CrossRef]
- Lin, L.; Wang, Y. MiR-548b-3p regulates proliferation, apoptosis, and mitochondrial function by targeting CIP2A in hepatocellular carcinoma. Biomed Res. Int. 2018, 2018, 7385426. [Google Scholar] [CrossRef]
- Herrera-Van Oostdam, A.S.; Toro-Ortíz, J.C.; López, J.A.; Noyola, D.E.; García-López, D.A.; Durán-Figueroa, N.V.; Martínez-Martínez, E.; Portales-Pérez, D.P.; Salgado-Bustamante, M.; López-Hernández, Y. Placental exosomes isolated from urine of patients with gestational diabetes exhibit a differential profile expression of microRNAs across gestation. Int. J. Mol. Med. 2020, 46, 546–560. [Google Scholar] [CrossRef] [PubMed]
- Mei, L.; Yan, H.; Wang, S.; Guo, C.; Zheng, X.; Yan, B.; Zhao, J.; Yang, A. Upregulation of miR-630 Induced by Oxidative Damage Resists Cell Migration Through Targeting ALCAM in Human Lens Epithelium Cells. Curr. Eye Res. 2020, 45, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Yanaka, Y.; Muramatsu, T.; Uetake, H.; Kozaki, K.I.; Inazawa, J. miR-544a induces epithelial-mesenchymal transition through the activation of WNT signaling pathway in gastric cancer. Carcinogenesis 2015, 36, 1363–1371. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.X.; Wu, X.B.; Zheng, C.W.; Zhang, Q.L.; Zhang, G.Q.; Chen, K.; Zhan, Q.; An, F.M. Mechanistic Investigation on the Regulation of FABP1 by the IL-6/miR-603 Signaling in the Pathogenesis of Hepatocellular Carcinoma. Biomed Res. Int. 2021, 2021, 8579658. [Google Scholar] [CrossRef]
- Cai, W.; Xu, Y.; Yin, J.; Zuo, W.; Su, Z. miR-552-5p facilitates osteosarcoma cell proliferation and metastasis by targeting WIF1. Exp. Ther. Med. 2019, 17, 3781–3788. [Google Scholar] [CrossRef]
- Chen, Y.; Lei, Y.; Lin, J.; Huang, Y.; Zhang, J.; Chen, K.; Sun, S.; Lin, X. The Linc01260 functions as a tumor suppressor via the miR-562/CYLD/NF-κB pathway in non-small cell lung cancer. Onco. Targets. Ther. 2020, 13, 10707–10719. [Google Scholar] [CrossRef] [PubMed]
- Christofides, A.; Papagregoriou, G.; Dweep, H.; Makrides, N.; Gretz, N.; Felekkis, K.; Deltas, C. Evidence for miR-548c-5p regulation of FOXC2 transcription through a distal genomic target site in human podocytes. Cell. Mol. Life Sci. 2020, 77, 2441–2459. [Google Scholar] [CrossRef]
- Kijpaisalratana, N.; Nimsamer, P.; Khamwut, A.; Payungporn, S.; Pisitkun, T.; Chutinet, A.; Utoomprurkporn, N.; Kerr, S.J.; Vongvasinkul, P.; Suwanwela, N.C. Serum miRNA125a-5p, miR-125b-5p, and miR-433-5p as biomarkers to differentiate between posterior circulation stroke and peripheral vertigo. BMC Neurol. 2020, 20, 372. [Google Scholar] [CrossRef]
- Mao, Q.; Liang, X.L.; Zhang, C.L.; Pang, Y.H.; Lu, Y.X. LncRNA KLF3-AS1 in human mesenchymal stem cell-derived exosomes ameliorates pyroptosis of cardiomyocytes and myocardial infarction through miR-138-5p/Sirt1 axis. Stem Cell Res. Ther. 2019, 10, 393. [Google Scholar] [CrossRef]
- Liu, F.; Yu, X.; Huang, H.; Chen, X.; Wang, J.; Zhang, X.; Lin, Q. Upregulation of microRNA-450 inhibits the progression of lung cancer in vitro and in vivo by targeting interferon regulatory factor 2. Int. J. Mol. Med. 2016, 38, 283–290. [Google Scholar] [CrossRef]
- Luo, Y.; Liu, W.; Tang, P.; Jiang, D.; Gu, C.; Huang, Y.; Gong, F.; Rong, Y.; Qian, D.; Chen, J.; et al. MiR-624-5p promoted tumorigenesis and metastasis by suppressing hippo signaling through targeting PTPRB in osteosarcoma cells. J. Exp. Clin. Cancer Res. 2019, 38, 488. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Song, X.; Li, Y.; Ma, T.; Su, P.; Guo, R.; Chen, B.; Zhang, H.; Sang, Y.; Liu, Y.; et al. Targeting the circBMPR2/miR-553/USP4 Axis as a Potent Therapeutic Approach for Breast Cancer. Mol. Ther. Nucleic Acids 2019, 17, 347–361. [Google Scholar] [CrossRef] [PubMed]
- Rongna, A.; Yu, P.; Hao, S.; Li, Y. MiR-876-5p suppresses cell proliferation by targeting Angiopoietin-1 in the psoriasis. Biomed. Pharmacother. 2018, 103, 1163–1169. [Google Scholar] [CrossRef]
- Pei, X.; Li, Y.; Zhu, L.; Zhou, Z. Astrocyte-derived exosomes transfer miR-190b to inhibit oxygen and glucose deprivation-induced autophagy and neuronal apoptosis. Cell Cycle 2020, 19, 906–917. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Li, F.; Shen, K.; Luo, C.; Song, G. LOXL1-AS1/miR-515-5p/STAT3 Positive Feedback Loop Facilitates Cell Proliferation and Migration in Atherosclerosis. J. Cardiovasc. Pharmacol. 2020, 76, 151–158. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Ji, J.; Wang, T.; Wang, M.B.; Chen, X.L. Upregulation of Circulating miR-195-3p in Heart Failure. Cardiology 2017, 138, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Han, H.; Li, X.; Wu, J.; Li, X.; Niu, H.; Li, W. Analysis of lncRNA, miRNA, and mRNA Expression Profiling in Type i IFN and Type II IFN Overexpressed in Porcine Alveolar Macrophages. Int. J. Genomics 2021, 2021, 6666160. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Gu, H.; Wang, L.; Huang, F.; Cai, J. MiR-885-3p is down-regulated in peripheral blood mononuclear cells from T1D patients and regulates the inflammatory response via targeting TLR4/NF-κB signaling. J. Gene Med. 2020, 22, 3145. [Google Scholar] [CrossRef]
- Liao, C.H.; Liu, Y.; Wu, Y.F.; Zhu, S.W.; Cai, R.Y.; Zhou, L.; Yin, X.M. microRNA-329 suppresses epithelial-to-mesenchymal transition and lymph node metastasis in bile duct cancer by inhibiting laminin subunit beta 3. J. Cell. Physiol. 2019, 234, 17786–17799. [Google Scholar] [CrossRef]
- Wang, W.; Zhao, E.; Yu, Y.; Geng, B.; Zhang, W.; Li, X. MiR-216a exerts tumor-suppressing functions in renal cell carcinoma by targeting TLR4. Am. J. Cancer Res. 2018, 8, 476–488. [Google Scholar]
- Goodwin, A.J.; Li, P.; Halushka, P.V.; Cook, J.A.; Sumal, A.S.; Fan, H. Circulating miRNA 887 is differentially expressed in ARDS and modulates endothelial function. Am. J. Physiol. Lung Cell. Mol. Physiol. 2020, 318, L1261–L1269. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.; Li, W.; Yuan, L.; Mei, X.; Hu, W. MicroRNA-106a-3p induces apatinib resistance and activates Janus-Activated Kinase 2 (JAK2)/Signal Transducer and Activator of Transcription 3 (STAT3) by targeting the SOCS system in gastric cancer. Med. Sci. Monit. 2019, 25, 10122–10128. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, D.; Zhao, S.T.; Zhang, Y.; Xu, A.; Hu, Y.Y.; Fang, Z. MiRNA-616 aggravates the progression of bladder cancer by regulating cell proliferation, migration and apoptosis through downregulating SOX7. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 9304–9312. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.H.; Wang, J.; Nie, G.; Chen, Y.J.; Li, X.; Jiang, X.; Cao, W.H. MicroRNA-509-5p functions as an anti-oncogene in breast cancer via targeting SOD2. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3617–3625. [Google Scholar] [CrossRef]
- Siege, S.R.; MacKenzie, J.; Chaplin, G.; Jablonski, N.G.; Griffiths, L. Circulating microRNAs involved in multiple sclerosis. Mol. Biol. Rep. 2012, 39, 6219–6225. [Google Scholar] [CrossRef]
- Magenta, A.; Cencioni, C.; Fasanaro, P.; Zaccagnini, G.; Greco, S.; Sarra-Ferraris, G.; Antonini, A.; Martelli, F.; Capogrossi, M.C. MiR-200c is upregulated by oxidative stress and induces endothelial cell apoptosis and senescence via ZEB1 inhibition. Cell Death Differ. 2011, 18, 1628–1639. [Google Scholar] [CrossRef]
- Chen, Y.; Li, Y.; Gao, H. Long noncoding RNA CASC9 promotes the proliferation and metastasis of papillary thyroid cancer via sponging miR-488-3p. Cancer Med. 2020, 9, 1830–1841. [Google Scholar] [CrossRef]
- Shibayama, Y.; Kubo, Y.; Nakagawa, T.; Iseki, K. MicroRNA-101-5p suppresses the expression of the ras-related protein RAP1A. Biol. Pharm. Bull. 2019, 42, 1332–1336. [Google Scholar] [CrossRef]
- Chhabra, R. Let-7i-5p, miR-181a-2-3p and EGF/PI3K/SOX2 axis coordinate to maintain cancer stem cell population in cervical cancer. Sci. Rep. 2018, 18, 7840. [Google Scholar] [CrossRef]
- Li, Y.; Wu, Y.; Sun, Z.; Wang, R.; Ma, D. MicroRNA-376a inhibits cell proliferation and invasion in glioblastoma multiforme by directly targeting specificity protein 1. Mol. Med. Rep. 2018, 17, 1583–1590. [Google Scholar] [CrossRef]
- Tang, C.; Wang, X.; Ji, C.; Zheng, W.; Yu, Y.; Deng, X.; Zhou, X.; Fang, L. The Role of miR-640: A Potential Suppressor in Breast Cancer via Wnt7b/β-catenin Signaling Pathway. Front. Oncol. 2021, 11, 1142. [Google Scholar] [CrossRef] [PubMed]
- Liang, D.; Wu, X.; Bai, J.; Zhang, L.; Yin, C.; Zhong, W. MiR-300 inhibits invasion and metastasis of osteosarcoma cell MG63 by negatively regulating PTTG1. Nan Fang Yi Ke Da Xue Xue Bao 2021, 41, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Robertson, G.; Pedersen, L.; Lim, E.; Hernandez-Herrera, A.; Rowat, A.C.; Patil, S.L.; Chan, C.K.; Wen, Y.; Zhang, X.; et al. miR-509-3p is clinically significant and strongly attenuates cellular migration and multi-cellular spheroids in ovarian cancer. Oncotarget 2016, 7, 25930. [Google Scholar] [CrossRef] [PubMed]
- Xiao, W.; Yao, E.; Zheng, W.; Tian, F.; Tian, L. miR-337 can be a key negative regulator in melanoma. Cancer Biol. Ther. 2017, 18, 392–399. [Google Scholar] [CrossRef]
- Gao, X.; Xu, D.; Li, S.; Wei, Z.; Li, S.; Cai, W.; Mao, N.; Jin, F.; Li, Y.; Yi, X.; et al. Pulmonary Silicosis Alters MicroRNA Expression in Rat Lung and miR-411-3p Exerts Anti-fibrotic Effects by Inhibiting MRTF-A/SRF Signaling. Mol. Ther. Nucleic Acids 2020, 20, 851–865. [Google Scholar] [CrossRef]
- Lemecha, M.; Morino, K.; Imamura, T.; Iwasaki, H.; Ohashi, N.; Ida, S.; Sato, D.; Sekine, O.; Ugi, S.; Maegawa, H. MiR-494-3p regulates mitochondrial biogenesis and thermogenesis through PGC1-α signalling in beige adipocytes. Sci. Rep. 2018, 8, 15096. [Google Scholar] [CrossRef]
- Yu, D.; Liu, X.; Han, G.; Liu, Y.; Zhao, X.; Wang, D.; Bian, X.; Gu, T.; Wen, L. The let-7 family of microRNAs suppresses immune evasion in head and neck squamous cell carcinoma by promoting PD-L1 degradation. Cell Commun. Signal. 2019, 17, 173. [Google Scholar] [CrossRef]
- Lin, W.; Tang, Y.; Zhao, Y.; Zhao, J.; Zhang, L.; Wei, W.; Chen, J. MiR-144-3p Targets FoxO1 to Reduce Its Regulation of Adiponectin and Promote Adipogenesis. Front. Genet. 2020, 11, 603144. [Google Scholar] [CrossRef]
- Sheedy, P.; Medarova, Z. The fundamental role of miR-10b in metastatic cancer. Am. J. Cancer Res. 2018, 8, 1674–1688. [Google Scholar]
- Gao, C.; Wei, J.; Tang, T.; Huang, Z. Role of microRNA-33a in malignant cells (review). Oncol. Lett. 2020, 20, 2537–2556. [Google Scholar] [CrossRef]
- Lin, Y.; Dan, H.; Lu, J. Overexpression of microRNA-136-3p Alleviates Myocardial Injury in Coronary Artery Disease via the Rho A/ROCK Signaling Pathway. Kidney Blood Press. Res. 2020, 45, 477–496. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Deng, Q.; Lv, Z.; Ling, Y.; Hou, X.; Chen, Z.; Dinglin, X.; Ma, S.; Li, D.; Wu, Y.; et al. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol. Cancer 2019, 18, 181. [Google Scholar] [CrossRef] [PubMed]
- Qiang, Z.; Jin, B.; Peng, Y.; Zhang, Y.; Wang, J.; Chen, C.; Wang, X.; Liu, F. miR-762 modulates thyroxine-induced cardiomyocyte hypertrophy by inhibiting Beclin-1. Endocrine 2019, 66, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Hou, X.; Wu, C.; Han, L.; Li, Q.; Wang, J.; Luo, S. MiR-645 promotes invasiveness, metastasis and tumor growth in colorectal cancer by targeting EFNA5. Biomed. Pharmacother. 2020, 125, 109889. [Google Scholar] [CrossRef]
- Xia, L.H.; Yan, Q.H.; Sun, Q.D.; Gao, Y.P. MiR-411-5p acts as a tumor suppressor in non-small cell lung cancer through targeting PUM1. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 5546–5553. [Google Scholar] [CrossRef]
- Chakrabarti, M.; Ray, S.K. Anti-tumor activities of luteolin and silibinin in glioblastoma cells: Overexpression of miR-7-1-3p augmented luteolin and silibinin to inhibit autophagy and induce apoptosis in glioblastoma in vivo. Apoptosis 2016, 21, 312–328. [Google Scholar] [CrossRef]
- Liu, F.; Zhang, H.; Xie, F.; Tao, D.; Xiao, X.; Huang, C.; Wang, M.; Gu, C.; Zhang, X.; Jiang, G. Hsa_circ_0001361 promotes bladder cancer invasion and metastasis through miR-491-5p/MMP9 axis. Oncogene 2020, 39, 1696–1709. [Google Scholar] [CrossRef]
- Yu, T.; Wang, L.N.; Li, W.; Zuo, Q.F.; Li, M.M.; Zou, Q.M.; Xiao, B. Downregulation of miR-491-5p promotes gastric cancer metastasis by regulating SNAIL and FGFR4. Cancer Sci. 2018, 109, 1393–1403. [Google Scholar] [CrossRef]
- Li, J.; Lin, T.Y.; Chen, L.; Liu, Y.; Dian, M.J.; Hao, W.C.; Lin, X.L.; Li, X.Y.; Li, Y.L.; Lian, M.; et al. Mir-19 regulates the expression of interferon-induced genes and mhc class i genes in human cancer cells. Int. J. Med. Sci. 2020, 17, 953. [Google Scholar] [CrossRef]
- Zhang, J.; Gao, D.; Zhang, H. Upregulation of miR-614 promotes proliferation and inhibits apoptosis in ovarian cancer by suppressing PPP2R2A expression. Mol. Med. Rep. 2018, 17, 6285–6292. [Google Scholar] [CrossRef]
- Gu, Y.; Cheng, Y.; Song, Y.; Zhang, Z.; Deng, M.; Wang, C.; Zheng, G.; He, Z. MicroRNA-493 suppresses tumor growth, invasion and metastasis of lung cancer by regulating E2F1. PLoS ONE 2014, 9, 102602. [Google Scholar] [CrossRef] [PubMed]
- Cimmino, A.; Calin, G.A.; Fabbri, M.; Iorio, M.V.; Ferracin, M.; Shimizu, M.; Wojcik, S.E.; Aqeilan, R.I.; Zupo, S.; Dono, M.; et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 2005, 102, 13944–13949. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, J.; Li, M.; Li, F.; Shao, Y.; Wu, L. microRNA-510-5p promotes thyroid cancer cell proliferation, migration, and invasion through suppressing SNHG15. J. Cell. Biochem. 2019, 120, 11738–11744. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, S.; Yan, B.; Yang, J.; Chen, E. Mir-581/smad7 axis contributes to colorectal cancer metastasis: A bioinformatic and experimental validation-based study. Int. J. Mol. Sci. 2020, 21, 6499. [Google Scholar] [CrossRef]
- Ren, K.; Yu, Y.; Wang, X.; Liu, H.; Zhao, J. MiR-340-3p-HUS1 axis suppresses proliferation and migration in lung adenocarcinoma cells. Life Sci. 2021, 274, 119330. [Google Scholar] [CrossRef]
- Monteleone, N.J.; Lutz, C.S. miR-708-5p: A microRNA with emerging roles in cancer. Oncotarget 2017, 8, 71292–71316. [Google Scholar] [CrossRef]
- Ramírez-Salazar, E.G.; Almeraya, E.V.; López-Perez, T.V.; Patiño, N.; Salmeron, J.; Velázquez-Cruz, R. MicroRNA-548-3p overexpression inhibits proliferation, migration and invasion in osteoblast-like cells by targeting STAT1 and MAFB. J. Biochem. 2020, 168, 203–211. [Google Scholar] [CrossRef]
- Shi, J.; Gong, L.; Chen, L.; Luo, J.; Song, G.; Xu, J.; Lv, Z.; Tao, H.; Xia, Y.; Ye, Z. miR-618 Suppresses Metastasis in Gastric Cancer by Downregulating the Expression of TGF-β2. Anat. Rec. 2019, 302, 931–940. [Google Scholar] [CrossRef]
- Sun, Y.; Zhao, J.; Yin, X.; Yuan, X.; Guo, J.; Bi, J. miR-297 acts as an oncogene by targeting GPC5 in lung adenocarcinoma. Cell Prolif. 2016, 49, 636–643. [Google Scholar] [CrossRef]
- Yao, Y.; Jia, H.; Wang, G.; Ma, Y.; Sun, W.; Li, P. MiR-297 protects human umbilical vein endothelial cells against LPS-induced inflammatory response and apoptosis. Cell. Physiol. Biochem. 2019, 52, 696–707. [Google Scholar] [CrossRef]
- Liu, Z.; Lu, X.; Wen, L.; You, C.; Jin, X.; Liu, J. Hsa_circ_0014879 regulates the radiosensitivity of esophageal squamous cell carcinoma through miR-519-3p/CDC25A axis. Anticancer. Drugs 2021, 33, e349–e361. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.L.; An, B.F.; Zhang, G.C. MiR-27 alleviates myocardial cell damage induced by hypoxia/reoxygenation via targeting TGFBR1 and inhibiting NF-κB pathway. Kaohsiung J. Med. Sci. 2019, 35, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Qian, F.H.; Deng, X.; Zhuang, Q.X.; Wei, B.; Zheng, D.D. MiR-625-5p suppresses inflammatory responses by targeting AKT2 in human bronchial epithelial cells. Mol. Med. Rep. 2019, 19, 1951–1957. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Yokoi, A.; Yamamoto, Y.; Kajiyama, H. ChrXq27.3 miRNA cluster functions in cancer development. J. Exp. Clin. Cancer Res. 2021, 40, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lv, P.; Luo, Y.F.; Zhou, W.Y.; Liu, B.; Zhou, Z.; Shi, Y.Z.; Huang, R.; Peng, C.; He, Z.L.; Wang, J.; et al. miR-373 inhibits autophagy and further promotes apoptosis of cholangiocarcinoma cells by targeting ULK1. Kaohsiung J. Med. Sci. 2020, 36, 429–440. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Qin, S.; Gu, Y.; Pan, H.; Bian, D. Long non-coding RNA NORAD promotes the occurrence and development of non-small cell lung cancer by adsorbing MiR-656-3p. Mol. Genet. Genomic Med. 2019, 7, e757. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Cheng, Y. Inhibition of miR-20 promotes proliferation and autophagy in articular chondrocytes by PI3K/AKT/mTOR signaling pathway. Biomed. Pharmacother. 2018, 97, 607–615. [Google Scholar] [CrossRef]
- Zhang, Y.; Dai, J.; Tang, J.; Zhou, L.; Zhou, M. MicroRNA-649 promotes HSV-1 replication by directly targeting MALT1. J. Med. Virol. 2017, 89, 1069–1079. [Google Scholar] [CrossRef]
- Pepe, F.; Visone, R.; Veronese, A. The glucose-regulated MiR-483-3p influences key signaling pathways in cancer. Cancers 2018, 10, 181. [Google Scholar] [CrossRef]
- Lu, J.; Zhou, L.; Wu, B.; Duan, Y.; Sun, Y.; Gu, L.; Xu, D.; Du, C. MiR-501-3p functions as a tumor suppressor in non-small cell lung cancer by downregulating RAP1A. Exp. Cell Res. 2020, 387, 111752. [Google Scholar] [CrossRef]
- Zhang, D.; Yang, N. MiR-335-5p inhibits cell proliferation, migration and invasion in colorectal cancer through downregulating LDHB. J. Buon 2019, 24, 1128–1136. [Google Scholar] [PubMed]
- Gracia-Cazaña, T.; González, S.; Parrado, C.; Juarranz, Á.; Gilaberte, Y. Influence of the Exposome on Skin Cancer. Actas Dermosifiliogr. 2020, 111, 460–470. [Google Scholar] [CrossRef] [PubMed]
- Pecorelli, A.; Cordone, V.; Messano, N.; Zhang, C.; Falone, S.; Amicarelli, F.; Hayek, J.; Valacchi, G. Altered inflammasome machinery as a key player in the perpetuation of Rett syndrome oxinflammation. Redox Biol. 2020, 28, 101334. [Google Scholar] [CrossRef] [PubMed]
- Valacchi, G.; Sticozzi, C.; Pecorelli, A.; Cervellati, F.; Cervellati, C.; Maioli, E. Cutaneous responses to environmental stressors. Ann. N. Y. Acad. Sci. 2012, 1271, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Valacchi, G.; Virgili, F.; Cervellati, C.; Pecorelli, A. OxInflammation: From subclinical condition to pathological biomarker. Front. Physiol. 2018, 9, 858. [Google Scholar] [CrossRef] [PubMed]
- Vierkötter, A.; Schikowski, T.; Ranft, U.; Sugiri, D.; Matsui, M.; Krämer, U.; Krutmann, J. Airborne particle exposure and extrinsic skin aging. J. Investig. Dermatol. 2010, 130, 2719–2726. [Google Scholar] [CrossRef]
- Magnani, N.D.; Muresan, X.M.; Belmonte, G.; Cervellati, F.; Sticozzi, C.; Pecorelli, A.; Miracco, C.; Marchini, T.; Evelson, P.; Valacchi, G. Skin damage mechanisms related to airborne particulate matter exposure. Toxicol. Sci. 2016, 149, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, J.Y.; Martinez, R.M.; Morocho-Jácome, A.L.; Castillo-Gómez, T.S.; Pereda-Contreras, V.J.; Rosado, C.; Velasco, M.V.R.; Baby, A.R. Skin impacts from exposure to ultraviolet, visible, infrared, and artificial lights—A review. J. Cosmet. Laser Ther. 2021, 23, 1–7. [Google Scholar] [CrossRef]
- Pattison, D.I.; Davies, M.J. Actions of ultraviolet light on cellular structures. EXS 2006, 96, 131–157. [Google Scholar] [CrossRef]
- Svobodova, A.; Walterova, D.; Vostalova, J. Ultraviolet light induced alteration to the skin. Biomed. Pap. Med. Fac. Univ. Palacky. Olomouc. Czech. Repub. 2006, 150, 25–38. [Google Scholar] [CrossRef]
- Fritsche, K. Important differences exist in the dose-response relationship between diet and immune cell fatty acids in humans and rodents. Lipids 2007, 42, 961–979. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Li, C.X.; Li, Y.S.; Lv, J.Y.; Ma, Y.; Shao, T.T.; Xu, L.D.; Wang, Y.Y.; Du, L.; Zhang, Y.P.; et al. MiRNA-miRNA synergistic network: Construction via co-regulating functional modules and disease miRNA topological features. Nucleic Acids Res. 2011, 39, 825–836. [Google Scholar] [CrossRef] [PubMed]
- Fry, R.C.; Rager, J.E.; Bauer, R.; Sebastian, E.; Peden, D.B.; Jaspers, I.; Alexis, N.E. Air toxics and epigenetic effects: Ozone altered microRNAs in the sputum of human subjects. Am. J. Physiol. Lung Cell. Mol. Physiol. 2014, 306, L1129–L1137. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Shafy, H.I.; El-Khateeb, M.A.; Mansour, M.S.M. Treatment of leather industrial wastewater via combined advanced oxidation and membrane filtration. Water Sci. Technol. 2016, 74, 586–594. [Google Scholar] [CrossRef] [PubMed]
- Izzotti, A.; Calin, G.A.; Arrigo, P.; Steele, V.E.; Croce, C.M.; De Flora, S. Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke. FASEB J. 2009, 23, 806–812. [Google Scholar] [CrossRef] [PubMed]
- Izzotti, A.; Pulliero, A. The effects of environmental chemical carcinogens on the microRNA machinery. Int. J. Hyg. Environ. Health 2014, 217, 601–627. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Unger, N.; Harper, K.; Heyes, C. Global Climate and Human Health Effects of the Gasoline and Diesel Vehicle Fleets. GeoHealth 2020, 4, e2019GH000240. [Google Scholar] [CrossRef]
- Izzotti, A.; Pulliero, A. Molecular damage and lung tumors in cigarette smoke-exposed mice. Ann. N. Y. Acad. Sci. 2015, 1340, 75–83. [Google Scholar] [CrossRef]
- Meek, M.D. Ah receptor and estrogen receptor-dependent modulation of gene expression by extracts of diesel exhaust particles. Environ. Res. 1998, 79, 114–121. [Google Scholar] [CrossRef]
- Hoskin, R.; Pambianchi, E.; Pecorelli, A.; Grace, M.; Therrien, J.P.; Valacchi, G.; Lila, M.A. Novel spray dried algae-rosemary particles attenuate pollution-induced skin damage. Molecules 2021, 26, 3781. [Google Scholar] [CrossRef]
- Izzotti, A.; Calin, G.A.; Steele, V.E.; Croce, C.M.; De Flora, S. Relationships of microRNA expression in mouse lung with age and exposure to cigarette smoke and light. FASEB J. 2009, 23, 3243–3250. [Google Scholar] [CrossRef] [PubMed]
| MicroRNA | Fold Change | Function | Reference |
|---|---|---|---|
| hsa-miR-495-5p | >10 | Promotes Th2 differentiation in allergic rhinitis, as a tumor suppressor | [23] |
| hsa-miR-628-5p | >10 | Inhibits osteoblast differentiation via RUNX2 | [24] |
| hsa-miR-361-3p | >10 | Suppresses proliferation, invasion inhibited cells invasive and proliferative abilities, and cell lines invasion and proliferation | [25,26] |
| hsa-miR-875-5p | >10 | Promotes cellular apoptosis and proliferation | [27] |
| hsa-miR-509-3p | >10 | Tumor suppressor | [28,29] |
| hsa-miR-518b | >10 | Suppresses cell proliferation, invasiveness, and migration in colorectal cancer | [30,31] |
| hsa-miR-516b-5p | >10 | Cell proliferation, inducing G1 cell cycle arrest and apoptosis | [32] |
| hsa-miR-381-5p | >10 | Induces apoptosis | [33] |
| hsa-miR-661 | >10 | Promotes proliferation, migration, and metastasis of NSCLC | [34] |
| hsa-miR-216a-3p | >10 | Antitumor functions | [35] |
| hsa-miR-548c-3p | >10 | Inflammatory responses and potential estrogen receptor sensitivity | [36] |
| hsa-miR-106a-3p | >10 | Cell proliferation and autophagy | [37] |
| hsa-miR-616-5p | >10 | Promotes angiogenesis and modulates cell proliferation | [38] |
| hsa-miR-671-3p | >10 | Suppresses proliferation and invasion of breast cancer cells and regulates metabolic processes | [39,40] |
| hsa-miR-544a | >10 | Regulates migration and invasion in colorectal cancer cells | [41] |
| hsa-miR-614 | >10 | Inflammatory process | [42] |
| hsa-miR-525-3p | <10 | Modifies the expression of proinflammatory cytokines; apoptosis | [43] |
| hsa-miR-378c | <10 | Regulates the angiogenic capacity of CD34(+) progenitor cells | [44] |
| hsa-miR-924 | <10 | Suppresses the proliferation, migration, and invasion of NSCLC cells | [45] |
| hsa-miR-522-3p | <10 | Cell proliferation of human glioblastoma cells; modulates the expression of proinflammatory cytokines | [46] |
| hsa-miR-431-5p | <10 | Promotes differentiation and regeneration of cells | [47,48] |
| hsa-miR-770-5p | <10 | Suppresses cell apoptosis and the release of proinflammatory factors | [49] |
| hsa-miR-183-5p | <10 | Tumor suppressor inflammation and alters miRNA expression in the airway epithelium | [50] |
| hsa-miR-598-5p | <10 | Promotes cell proliferation and cell cycle progression in human colorectal carcinoma; elevates apoptosis | [51] |
| hsa-miR-486-5p | <10 | Regulation of heart contraction, muscle contraction, and ion channel activity | [52] |
| hsa-miR-125a-5p | <10 | Regulates stress response, apoptosis, proliferation, angiogenesis, and expression of genes, associated with human lung cancer | [53] |
| hsa-miR-34b-5p | <10 | Regulates stress response, apoptosis, proliferation, angiogenesis, and expression of genes, and is upregulated during cardiac hypertrophy | [54] |
| hsa-miR-301a-3p | <10 | Promotes autophagy and inhibits apoptosis | [55,56] |
| hsa-miR-23c | <10 | Inhibits cell proliferation and induces apoptosis of hepatocellular carcinoma cells; cell growth arrest and apoptosis. | [57] |
| hsa-miR-383-5p | <10 | Oxidative stress and inflammation-related factors | [58] |
| hsa-miR-574-5p | <10 | Promotes the differentiation of human cardiac fibroblasts | [59] |
| hsa-miR-151a-5p | <10 | Regulation of cellular respiration and ATP production through targeting Cytb | [60] |
| hsa-miR-514a-3p | <10 | Attenuates proliferation and increases chemoresistance | [61] |
| hsa-miR-136-3p | <10 | Promotes apoptosis in gastric cancer cells | [62] |
| hsa-miR-1-3p | <10 | Inflammation; regulator of heart adaption after ischemia or ischemic stress | [63] |
| hsa-miR-18a-3p | <10 | Downregulated in aging cells; induces the apoptosis of colon cancer cells | [64] |
| hsa-miR-502-5p | <10 | Enhances early apoptosis and inhibits proliferation of breast cancer cells | [65] |
| hsa-miR-451a | <10 | Apoptosis; inhibits autophagy | [66] |
| hsa-miR-19b-1-5p | <10 | Linked to oxidative stress, inflammation, and atherosclerosis | [67] |
| hsa-miR-873-5p | <0.1 | Tumor suppressor in thyroid cancer by inhibiting the proliferation, migration, and invasion of the cancer cells | [68] |
| hsa-miR-212-5p | <0.1 | Promotes cancer cell apoptosis and suppresses cancer cell proliferation and invasion | [69] |
| hsa-miR-552-5p | <0.1 | Tumorigenesis; progression | [70] |
| hsa-let-7a-3p | <0.1 | Linked to oxidative stress, inflammation, and atherosclerosis | [71] |
| hsa-miR-495-3p | <0.1 | Regulates proliferation, apoptosis, migration, and invasion in metastatic prostate cancer cells. | [68] |
| hsa-miR-767-3p | <0.1 | Promoted cell proliferation in human melanoma cell lines | [72] |
| hsa-miR-27b-5p | <0.1 | Involved in beige and brown adipogenesis after cold exposure | [73] |
| hsa-miR-26a-2-3p | <0.1 | Regulated stress response, apoptosis, proliferation, angiogenesis, and expression of genes | [74] |
| hsa-miR-629-5p | <0.1 | Action on tumor growth and metastasis in hepatocellular carcinoma | [75] |
| hsa-miR-543 | <0.1 | Cell oxidative phosphorylation | [76] |
| hsa-miR-515-5p | <0.1 | Upregulated in placentas from women with preeclampsia | [77] |
| hsa-miR-708-5p | <0.1 | Induces apoptosis and suppresses tumorigenicity in renal cancer cells | [78] |
| hsa-miR-378j | <0.1 | Tumor suppressor | [79] |
| hsa-miR-548az-3p | <0.1 | Alters inflammation | [80] |
| hsa-miR-876-5p | <0.1 | Regulates regulation proliferation, migration, invasion, and glutaminolysis in gastric cancer cells | [37] |
| hsa-miR-656-3p | <0.1 | Suppresses glioma cell proliferation, neurosphere formation, migration, and invasion | [81] |
| hsa-miR-944 | <0.1 | Increases p53 expression in cancer cells | [82] |
| hsa-miR-518e-3p | <0.1 | Tumor suppressor | [83] |
| hsa-miR-373-3p | <0.1 | Promotes the invasion and migration of breast cancer; regulates inflammatory cytokine-mediated chondrocyte proliferation | [32] |
| MicroRNA | Fold Change | Function | Reference |
|---|---|---|---|
| hsa-miR-628-5p | >10 | Inhibits osteoblast differentiation | [84] |
| hsa-miR-15a-3p | >10 | Proliferation; inflammation; apoptosis | [25,26] |
| hsa-miR-548am-3p | >10 | Induces proliferation and migration | [85] |
| hsa-miR-550b-2-5p | >10 | Cancer promotion | [86] |
| hsa-miR-495-5p | >10 | Tumor suppressor; proliferation and differentiation of osteoblasts in mice; inhibits the growth of fibroblasts in hypertrophic scar | [87] |
| hsa-miR-345-3p | >10 | Apoptosis and inflammation | [24,88] |
| hsa-miR-548q | >10 | Induces proliferation and migration | [89] |
| hsa-miR-887-3p | >10 | Pathways in cancer | [86] |
| hsa-miR-877-5p | >10 | Pathways in cancer | [90] |
| hsa-miR-513c-5p | >10 | Pathways in cancer | [91] |
| hsa-miR-422a | >10 | Pathways in cancer | [92] |
| hsa-miR-194-5p | >10 | Pathways in cancer | [93] |
| hsa-miR-378b | >10 | Inflammation and cell cycle | [94] |
| hsa-miR-610 | >10 | Pathways in cancer | [95] |
| hsa-miR-519e-5p | >10 | Atherosclerosis and pathways in cancer | [96] |
| hsa-miR-627-5p | >10 | Cell proliferation and cancer promotion | [97,98] |
| hsa-miR-548au-5p | >10 | Pathways in cancer | [99] |
| hsa-miR-770-5p | >10 | Apoptosis and inflammation | [100] |
| hsa-miR-196b-3p | >10 | Cell proliferation | [101] |
| hsa-miR-330-3p | >10 | Apoptosis and cell proliferation | [102] |
| hsa-miR-617 | >10 | Pathways in cancer | [103] |
| hsa-miR-375 | >10 | Cell proliferation and pathways in cancer | [104] |
| hsa-miR-936 | >10 | Cell proliferation, pathways in cancer, and apoptosis | [105] |
| hsa-miR-657 | >10 | Inflammation | [106] |
| hsa-miR-542-5p | >10 | Mitochondrial dysfunction and inflammation | [107] |
| hsa-miR-136-3p | >10 | Vascularization and pathways in cancer | [108] |
| hsa-miR-409-5p | >10 | Cardiovascular process, proliferation, migration, and cell cycle. | [109,110] |
| hsa-miR-154-3p | >10 | Pathways in cancer | [111,112] |
| hsa-miR-378c | >10 | Proliferation and inhibited apoptosis | [113] |
| hsa-miR-93-3p | >10 | Inflammation and apoptosis | [114] |
| hsa-miR-556-3p | >10 | Cell proliferation and apoptosis | [115] |
| hsa-miR-518c-5p | >10 | Tumor suppressor | [116] |
| hsa-miR-23b-5p | >10 | Cell proliferation and cancer | [32] |
| hsa-miR-504-5p | >10 | Cell proliferation and differentiation | [117] |
| hsa-miR-509-3p | >10 | Cardiovascular process | [118] |
| hsa-miR-514a-3p | >10 | Tumor suppressor | [119] |
| hsa-miR-431-5p | >10 | Cell proliferation and apoptosis | [120] |
| hsa-miR-506-3p | >10 | Cell proliferation and cancer | [121] |
| hsa-miR-645 | >10 | Cell proliferation and cancer | [122] |
| hsa-miR-129-5p | >10 | Inhibits the proliferation and metastasis of gastric cancer cells | [123] |
| hsa-miR-516b-5p | >10 | Migration, cell proliferation, and cancer process | [124] |
| hsa-miR-512-3p | >10 | Apoptosis and cell cycle | [125] |
| hsa-miR-101-5p | >10 | Apoptosis and promotes cell proliferation | [126] |
| hsa-miR-561-5p | >10 | Cell proliferation, G(1)/S transition, and suppresses apoptosis | [127] |
| hsa-miR-194-5p | >10 | Cell proliferation and cancer | [128] |
| hsa-miR-329-3p | >10 | Proliferation, invasion, and suppresses cell apoptosis | [129] |
| hsa-let-7i-3p | >10 | Coronary disease and cancer | [130] |
| hsa-miR-129-2-3p | >10 | Proliferation, invasion, and apoptosis | [131] |
| hsa-miR-548a-5p | >10 | Proliferation and inhibits apoptosis | [132] |
| hsa-miR-887-5p | >10 | Pathways in cancer | [133] |
| hsa-miR-99a-3p | >10 | Cell proliferation and pathways in cancer | [134] |
| hsa-miR-487a-3p | >10 | Cell proliferation and cancer | [135] |
| hsa-miR-378g | >10 | Cancer promotion | [136] |
| hsa-miR-548at-5p | >10 | Neurodegenerative disease | [137] |
| hsa-miR-374c-5p | >10 | Proliferation, apoptosis, and autophagy | [138] |
| hsa-miR-106a-3p | >10 | Proliferation and apoptosis | [139] |
| hsa-miR-92a-2-5p | >10 | Apoptosis and cell proliferation | [140] |
| hsa-miR-616-5p | >10 | Invasion, cell migration, and cancer | [141] |
| hsa-miR-509-5p | >10 | Tumor suppressor | [142] |
| hsa-miR-598-3p | >10 | Cancer process | [21] |
| hsa-miR-873-5p | >10 | Cell migration and cancer | [143] |
| hsa-miR-525-3p | <10 | Cancer cell migration | [21] |
| hsa-miR-500a-5p | <10 | Cell apoptosis and proliferation | [144] |
| hsa-miR-659-3p | <10 | Cell proliferation and cancer; apoptosis | [145] |
| hsa-miR-526b-5p | <10 | Cell proliferation and cancer | [146] |
| hsa-miR-764 | <10 | Cardiac diseases and cancer | [147] |
| hsa-miR-934 | <10 | Cancer progression and inflammation | [148] |
| hsa-miR-516a-5p | <10 | Cell proliferation and cancer | [149] |
| hsa-miR-520f-3p | <10 | DNA repair | [150] |
| hsa-miR-369-5p | <10 | Aerobic glycolysis and pathways in cancer | [151] |
| hsa-miR-613 | <10 | Invasion and cell proliferation | [152] |
| hsa-miR-411-3p | <10 | Proliferation and cancer | [153] |
| hsa-miR-432-3p | <10 | Inflammation | [154] |
| hsa-let-7c-3p | <10 | Apoptosis | [155] |
| hsa-miR-671-5p | <10 | Proliferation and cell cycle | [156] |
| hsa-miR-181d-5p | <10 | Proliferation and angiogenesis | [157] |
| hsa-miR-192-5p | <10 | Hypertension and cancer | [158] |
| hsa-let-7g-3p | <10 | Linked to oxidative stress, inflammation, and atherosclerosis | [159] |
| hsa-miR-1-3p | <10 | Decreases tumor volume in a xenograft model | [68] |
| hsa-miR-515-3p | <10 | Cell proliferation, migration, invasion, and induced apoptosis | [160] |
| hsa-miR-320d | <10 | Apoptosis and cancer | [161] |
| hsa-miR-548aa | <10 | Can alter the inflammatory responses | [162] |
| hsa-miR-502-5p | <10 | Cell proliferation and invasion | [37] |
| hsa-miR-758 | <10 | Proliferation and invasion | [163] |
| hsa-miR-7-1-3p | <10 | Autophagy and cancer process | [164] |
| hsa-miR-324-3p | <10 | Cell proliferation and cancer | [165] |
| hsa-miR-520g-3p | <10 | DNA repair | [166] |
| hsa-miR-576-5p | <10 | Cell invasion and cancer | [151] |
| hsa-miR-520a-3p | <10 | Inhibits tumor progression, indicating its potential role as a tumor suppressor. | [167] |
| hsa-miR-449b-3p | <10 | Proliferation | [168] |
| hsa-miR-211-5p | <10 | Pathways in cancer | [169] |
| hsa-miR-376a-3p | <10 | Coronary artery disease | [170] |
| hsa-miR-939-3p | <10 | Pathways in cancer | [171] |
| hsa-miR-214-3p | <10 | Inhibition of migration and proliferation | [172] |
| hsa-miR-609 | <10 | Cardiovascular process | [173] |
| hsa-miR-29a-5p | <10 | Cardiac myocytes and overall cardiac dysfunction | [174] |
| hsa-miR-449c-3p | <10 | Inhibits NSCLC cell progression | [175] |
| hsa-miR-185-3p | <10 | Proliferation and invasion of cell | [176] |
| hsa-miR-766-3p | <10 | Suppresses apoptosis and facilitates autophagy | [177] |
| hsa-miR-486-5p | <10 | Regulation of heart contraction, muscle contraction, and ion channel activity | [53] |
| hsa-miR-144 | <10 | Tumor inhibitors or tumor suppressors, proliferation, and apoptosis | [53] |
| hsa-miR-664a-5p | <10 | Induces cell differentiation | [178] |
| hsa-miR-32-3p | <10 | Atherosclerosis | [179] |
| hsa-miR-224-3p | <10 | Cell proliferation and promotes apoptosis | [180] |
| hsa-miR-130a-5p | <10 | Myocardial infarction | [181] |
| hsa-miR-378i | <10 | Metabolic pathways, mitochondrial energy homeostasis, and related biological processes | [182] |
| hsa-miR-642b-5p | <10 | Inflammation | [183] |
| hsa-miR-668-3p | <10 | Progression of different types of cancer | [184] |
| hsa-miR-18b-5p | <10 | Progression of different types of cancer; cardiac function | [185] |
| hsa-miR-483-3p | <10 | Apoptosis | [186] |
| hsa-miR-485-3p | <10 | Cell proliferation; pathways in cancer | [187] |
| hsa-miR-200c-5p | <10 | Oxidative stress and cell apoptosis | [188] |
| hsa-miR-126-5p | <10 | Linked to oxidative stress, inflammation, and atherosclerosis | [189] |
| hsa-miR-26b-3p | <10 | Cell proliferation and invasion | [190] |
| hsa-miR-378d | <10 | Proliferation and migration of cancer | [191] |
| hsa-miR-526b-3p | <10 | Regulates the proliferation, invasion, and migration of cancer cells | [192] |
| hsa-miR-575 | <10 | Oncogene | [193] |
| hsa-miR-564 | <10 | Cell proliferation and invasion | [194] |
| hsa-miR-513a-5p | <10 | Induced apoptosis | [195] |
| hsa-miR-548i | <10 | Downregulates the inflammatory cytokines | [196] |
| hsa-miR-188-5p | <10 | Cell proliferation and cancer promotion | [197] |
| hsa-miR-563 | <10 | Cell proliferation and cancer promotion | [198] |
| hsa-miR-139-3p | <10 | Proliferation and invasion | [199] |
| hsa-miR-34a-5p | <0.1 | Inflammation | [200] |
| hsa-miR-34b-5p | <0.1 | P53 effector, cell proliferation, and apoptosis | [201] |
| hsa-miR-371b-5p | <0.1 | Cell proliferation and apoptosis | [202] |
| hsa-let-7f-2-3p | <0.1 | Cell proliferation and apoptosis | [203,204] |
| hsa-miR-557 | <0.1 | Tumor suppressor | [205] |
| hsa-miR-574-5p | <0.1 | Cell cycle and cancer process | [206] |
| hsa-miR-216a-3p | <0.1 | Cell proliferation and apoptosis | [207] |
| hsa-miR-466 | <0.1 | Tumor suppressor | [208] |
| hsa-miR-222-3p | <0.1 | Cell viability, migration, and invasion | [209] |
| hsa-miR-586 | <0.1 | Cell proliferation, invasion, metastasis, and apoptosis | [210] |
| hsa-miR-939-5p | <0.1 | Inflammation | [211] |
| hsa-miR-548b-3p | <0.1 | Proliferation, apoptosis, and mitochondrial function | [212] |
| hsa-miR-517c-3p | <0.1 | Responses to stress; alterations in circulating glucose levels | [213] |
| hsa-miR-630 | <0.1 | Oxidative damage and cell migration | [214] |
| hsa-miR-544a | <0.1 | Pathways in cancer | [215] |
| hsa-miR-603 | <0.1 | Proliferation, migration, invasion, and metastasis | [216] |
| hsa-miR-552-5p | <0.1 | Cell proliferation | [217] |
| hsa-miR-562 | <0.1 | Tumor suppressor | [218] |
| hsa-miR-548 | <0.1 | Cancer cell proliferation, migration, and invasion | [219] |
| hsa-miR-518a- | <0.1 | Tumor suppressor | [220] |
| hsa-miR-433-5p | <0.1 | Cardiovascular process | [32] |
| hsa-miR-138-5p | <0.1 | Cardiac function and pathological damage | [221] |
| hsa-miR-548ad-5p | <0.1 | Cancer cell proliferation, migration, and invasion | [222] |
| hsa-miR-450a-2-3p | <0.1 | Cell proliferation and cancer | [220] |
| hsa-miR-548av-3p | <0.1 | Cancer cell proliferation, migration, and invasion | [223] |
| hsa-miR-624-5p | <0.1 | Cell proliferation and cancer | [220] |
| hsa-miR-553 | <0.1 | Cell proliferation and cancer | [224] |
| hsa-miR-876-5p | <0.1 | Cell proliferation and cancer | [225] |
| hsa-miR-190b | <0.1 | Autophagy and cell cycle | [226] |
| hsa-miR-26a-2-3p | <0.1 | Cell cycle and cancer process | [227] |
| hsa-miR-515-5p | <0.1 | Cardiac function and proliferation cells | [75] |
| hsa-miR-195-3p | <0.1 | Cardiac function and proliferation cells | [228] |
| hsa-miR-365b-5p | <0.1 | Inflammation and cell proliferation | [229] |
| Hsa-miR-885-3p | <0.1 | Inflammation | [230] |
| MicroRNA | Fold Variation | Functions | Reference |
|---|---|---|---|
| hsa-miR-329-3p | >10 | Inhibits cell proliferation in glioma cells | [231] |
| hsa-miR-520g-5p | >10 | DNA repair | [232] |
| hsa-miR-216a- | >10 | Regulates the proliferation, apoptosis, migration, and invasion of lung cancer cells | [151] |
| hsa-miR-548c-3p | >10 | Cancer cell proliferation, migration, and invasion | [233] |
| hsa-miR-129-2-3p | >10 | Inhibits the proliferation and metastasis of gastric cancer cells | [220] |
| hsa-miR-887-5p | <10 | Pathways in cancer | [124] |
| hsa-miR-106a-3p | >10 | Involved in tumorigenesis and highly expressed in gastric cancer | [234] |
| hsa-miR-616-5p | >10 | Progression of bladder cancer by regulating cell proliferation, migration, and apoptosis | [235] |
| hsa-miR-509-5p | >10 | Tumor suppressive effects | [236] |
| hsa-miR-648 | >10 | Post-transcriptional regulators of glioblastoma | [237] |
| hsa-miR-378h | >10 | Metabolic pathways, mitochondrial energy homeostasis, and angiogenic network in tumors | [238] |
| hsa-miR-200c-5p | >10 | Upregulated by oxidative stress and induces endothelial cell apoptosis | [183] |
| hsa-miR-525-3p | <10 | Pathways in cancer | [239] |
| hsa-miR-488-3p | <10 | Pathways in cancer | [44] |
| hsa-miR-101-5p | <10 | Regulates cell proliferation | [240] |
| hsa-let-7g-3p | <10 | Modulates inflammatory responses; pathways in cancer | [241] |
| hsa-miR-376a-2-5p | <10 | Pathways in cancer | [242] |
| hsa-miR-640 | <10 | Pathways in cancer | [243] |
| hsa-miR-300 | <10 | Controls stem cell function and inhibits differentiation | [244] |
| hsa-miR-509-3p | <10 | Pathways in cancer | [245] |
| hsa-miR-548au-5p | <10 | Pathways in cancer | [246] |
| hsa-miR-337-3p | <10 | Pathways in cancer | [100] |
| hsa-miR-411-3p | <10 | Pathways in cancer | [247] |
| hsa-miR-494-3p | <10 | Mitochondrial biogenesis and pathways in cancer | [248] |
| hsa-let-7e-3p | <10 | Pathways in cancer | [249] |
| hsa-miR-144-3p | <10 | Regulates adipogenesis and pathways in cancer | [250] |
| hsa-miR-196b-3p | <10 | Cell proliferation | [251] |
| hsa-miR-10b-5p | <10 | Pathways in cancer | [102] |
| hsa-miR-33a-5p | <10 | Associated with carcinogenesis | [252] |
| hsa-miR-136-3p | <10 | Cardiac function and pathological damage in myocardial tissue, cardiomyocyte apoptosis, oxidative stress, and inflammatory response | [253] |
| hsa-miR-143-3p | <10 | Pathways in cancer | [254] |
| hsa-miR-762 | <10 | Modulates thyroxine-induced cardiomyocyte and pathways in cancer | [255] |
| hsa-miR-582-5p | <10 | Cell proliferation | [256] |
| hsa-miR-645 | <10 | Cell proliferation and apoptosis | [21] |
| hsa-miR-411-5p | <10 | Cell proliferation and pathways in cancer | [257] |
| hsa-miR-7-1-3p | <10 | Inhibits autophagy and induces apoptosis in glioblastoma | [258] |
| hsa-miR-18a-3p | <10 | Progression of different types of cancer; cardiac function. | [259] |
| hsa-miR-99a-3p | <10 | Cell proliferation and pathways in cancer | [186] |
| hsa-miR-491-5p | <10 | Cell proliferation and pathways in cancer | [135] |
| hsa-miR-19b-1-5p | <10 | Pathways in cancer | [260,261] |
| hsa-miR-614 | <10 | Cell proliferation and pathways in cancer | [262] |
| hsa-miR-493-5p | <0.1 | Cell proliferation | [263] |
| hsa-miR-15b-3p | <0.1 | Proliferation, inflammation, and apoptosis | [264] |
| hsa-miR-510-5p | <0.1 | Cell proliferation | [265] |
| hsa-miR-485-5p | <0.1 | Cell proliferation and pathways in cancer | [266] |
| hsa-miR-581 | <0.1 | Induces proliferation and migration | [188] |
| hsa-miR-340-3p | <0.1 | Cell proliferation and pathways in cancer | [267] |
| hsa-miR-708-5p | <0.1 | Cell proliferation and pathways in cancer | [268] |
| hsa-miR-548j-3p | <0.1 | Induces proliferation and migration | [269] |
| hsa-miR-618 | <0.1 | Cell proliferation and pathways in cancer | [270] |
| hsa-miR-885-3p | <0.1 | Inflammatory response and pathways in cancer | [271] |
| hsa-miR-297 | <0.1 | Oncogene, inflammatory response, and apoptosis | [231] |
| hsa-miR-518a-5p | <0.1 | Tumor suppressor | [272,273] |
| hsa-let-7f-2-3p | <0.1 | Proliferation and apoptosis | [32] |
| hsa-miR-519b-3p | <0.1 | Radiosensitivity of radio-resistant cells and pathways in cancer | [205] |
| hsa-miR-27b-5p | <0.1 | Prevents atherosclerosis by inhibiting inflammatory responses | [274] |
| hsa-miR-625-5p | <0.1 | Inflammation and inhibits cardiac hypertrophy | [275] |
| hsa-miR-548ah-5p | <0.1 | Induces proliferation and migration | [276] |
| hsa-miR-892c-3p | <0.1 | Pathways in cancer | [86] |
| hsa-miR-373-3p | <0.1 | Inhibits autophagy | [277] |
| hsa-miR-656-3p | <0.1 | Induces proliferation and migration | [278] |
| hsa-miR-20a-3p | <0.1 | Proliferation and autophagy | [279] |
| hsa-miR-518a-3p | <0.1 | Tumor suppressor | [280] |
| hsa-miR-649 | <0.1 | Pathways in cancer | [32] |
| hsa-miR-483-3p | <0.1 | Pathways in cancer and cardiac response | [281] |
| hsa-miR-501-3p | <0.1 | Cell proliferation, clonogenicity, migration, and invasion | [282] |
| hsa-miR-335-5p | <0.1 | Cell proliferation, migration, and invasion. | [283] |
| hsa-miR-129-5p | <0.1 | Proliferation, invasion, and apoptosis | [284] |
| hsa-miR-34c-5p | <0.1 | Proliferation and apoptosis | [132] |
| hsa-miR-548ao-5p | <0.1 | Promotes proliferation and inhibits apoptosis | [202] |
| hsa-miR-624-5p | <0.1 | Cell proliferation and pathways in cancer | [133] |
| Diesel | Ozone | UV | |
|---|---|---|---|
| Apoptosis | 16 | 36 | 12 |
| Cell cycle | 7 | 24 | 11 |
| Inflammation | 11 | 21 | 7 |
| DNA repair | 1 | 4 | 2 |
| Pathways in cancer | 23 | 72 | 39 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Valacchi, G.; Pambianchi, E.; Coco, S.; Pulliero, A.; Izzotti, A. MicroRNA Alterations Induced in Human Skin by Diesel Fumes, Ozone, and UV Radiation. J. Pers. Med. 2022, 12, 176. https://doi.org/10.3390/jpm12020176
Valacchi G, Pambianchi E, Coco S, Pulliero A, Izzotti A. MicroRNA Alterations Induced in Human Skin by Diesel Fumes, Ozone, and UV Radiation. Journal of Personalized Medicine. 2022; 12(2):176. https://doi.org/10.3390/jpm12020176
Chicago/Turabian StyleValacchi, Giuseppe, Erika Pambianchi, Simona Coco, Alessandra Pulliero, and Alberto Izzotti. 2022. "MicroRNA Alterations Induced in Human Skin by Diesel Fumes, Ozone, and UV Radiation" Journal of Personalized Medicine 12, no. 2: 176. https://doi.org/10.3390/jpm12020176
APA StyleValacchi, G., Pambianchi, E., Coco, S., Pulliero, A., & Izzotti, A. (2022). MicroRNA Alterations Induced in Human Skin by Diesel Fumes, Ozone, and UV Radiation. Journal of Personalized Medicine, 12(2), 176. https://doi.org/10.3390/jpm12020176









