Folate Pathway Gene Single Nucleotide Polymorphisms and Neural Tube Defects: A Systematic Review and Meta-Analysis
Abstract
1. Introduction
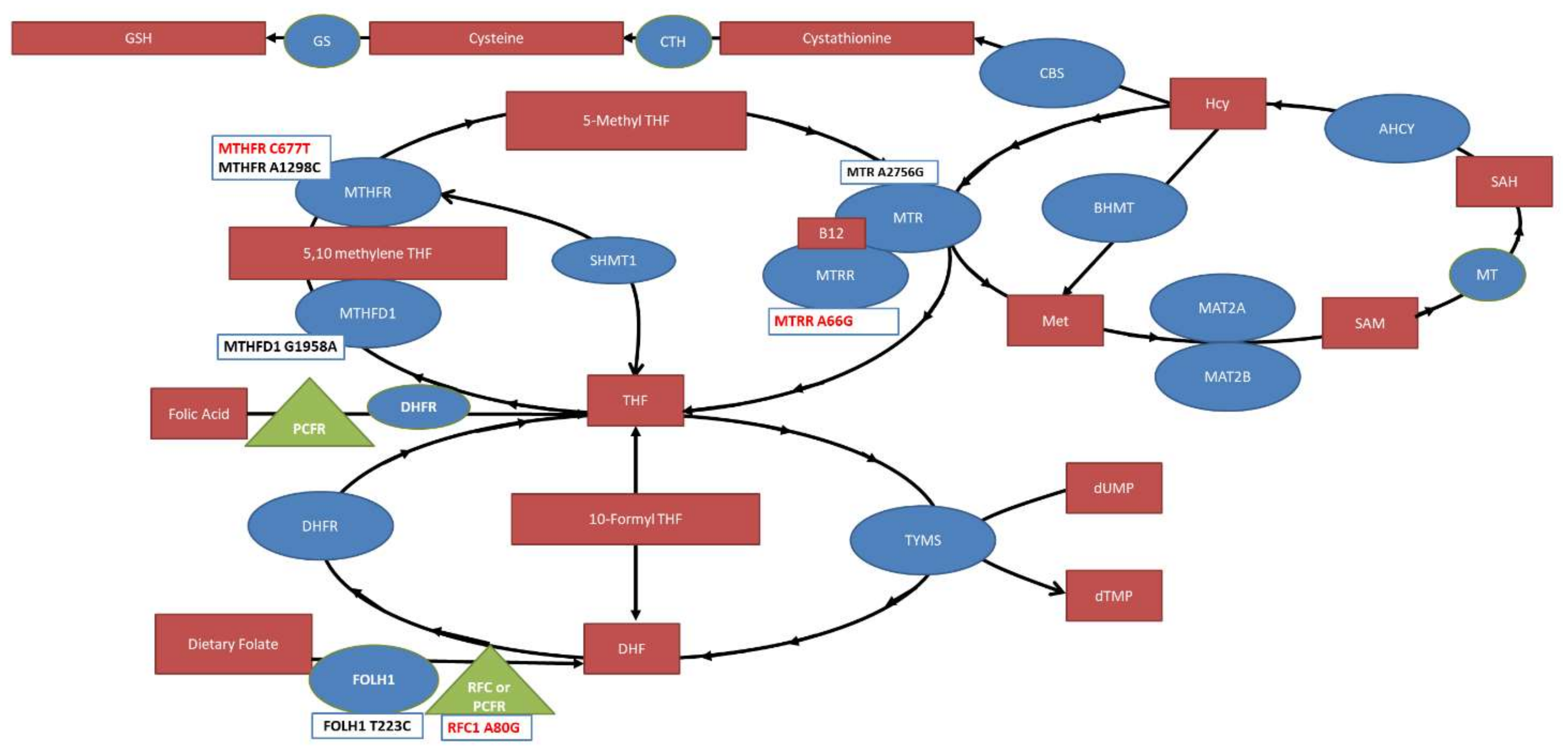
1.1. Folate Absorption and Transport
1.2. Folate Interconversion: Reduction of Folate and the Folate Cycle
1.3. Methylation Cycle
1.4. Neurodevelopmental Disorders and Neural Tube Defects: Links to Folate Metabolism
2. Materials and Methods
2.1. Search Strategy and Identification of Relevant Studies
2.2. Data Extraction
2.3. Statistical Analysis
2.4. Quality Assessment
3. Results
3.1. Characteristics of Eligible Studies
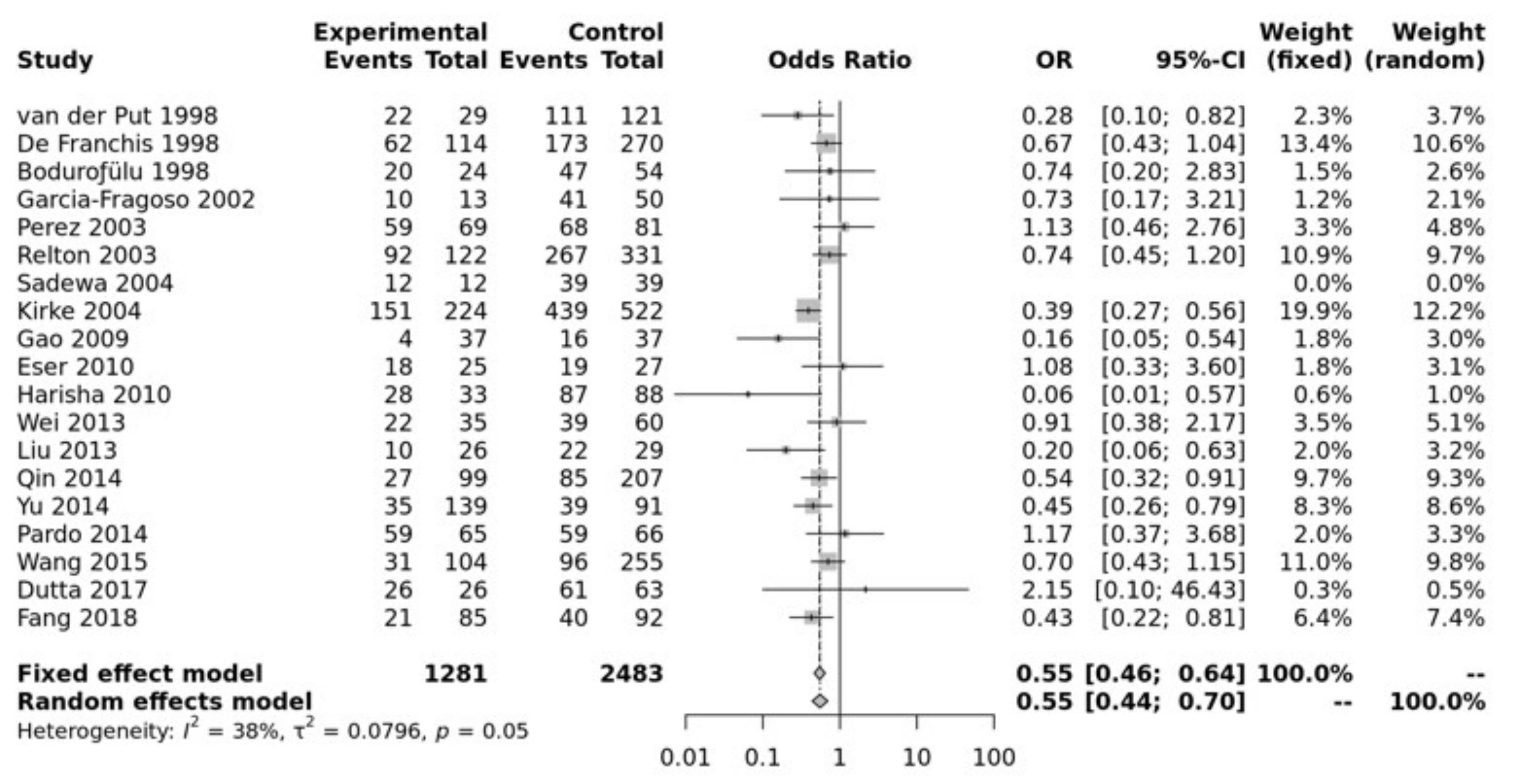
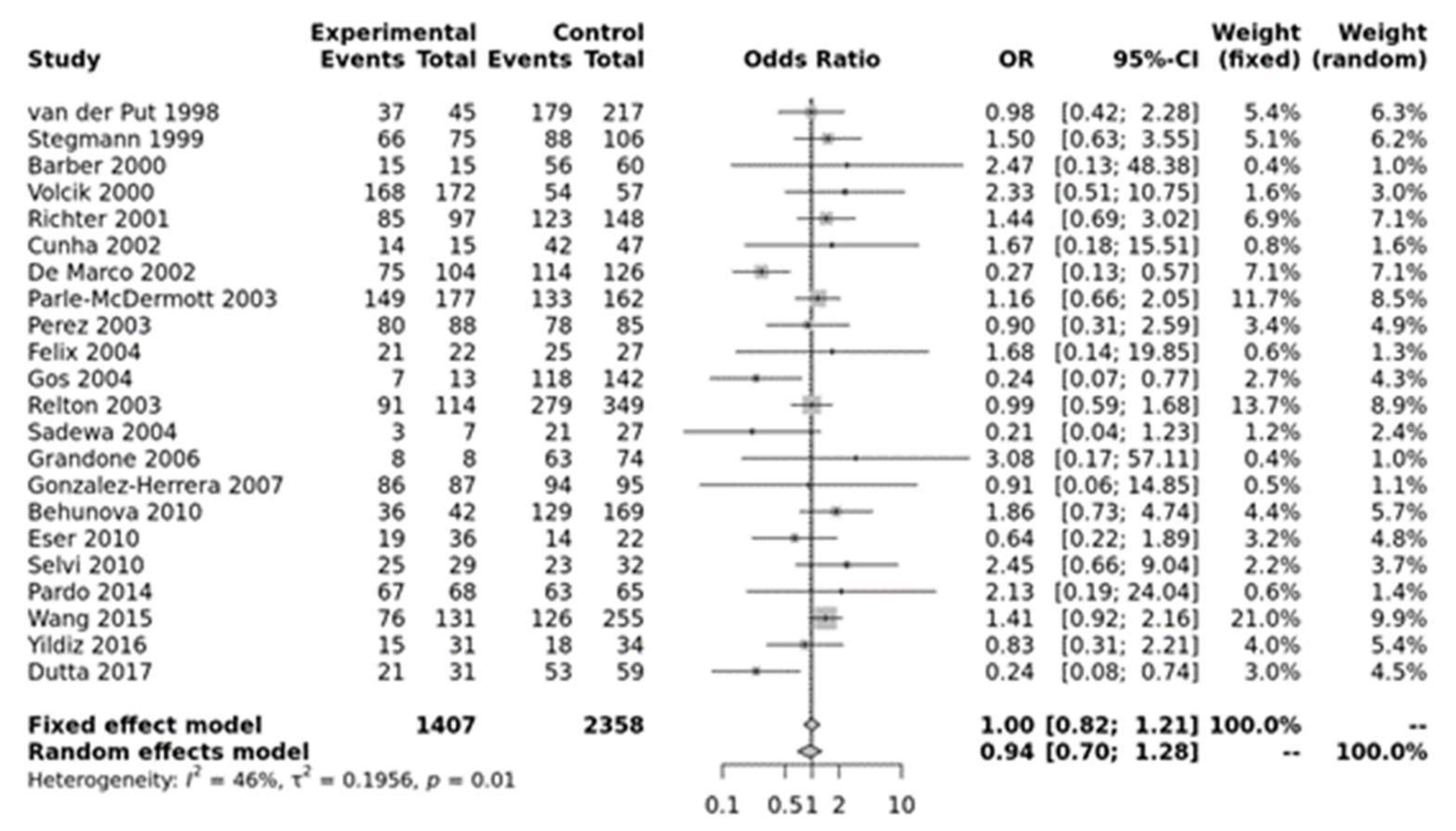
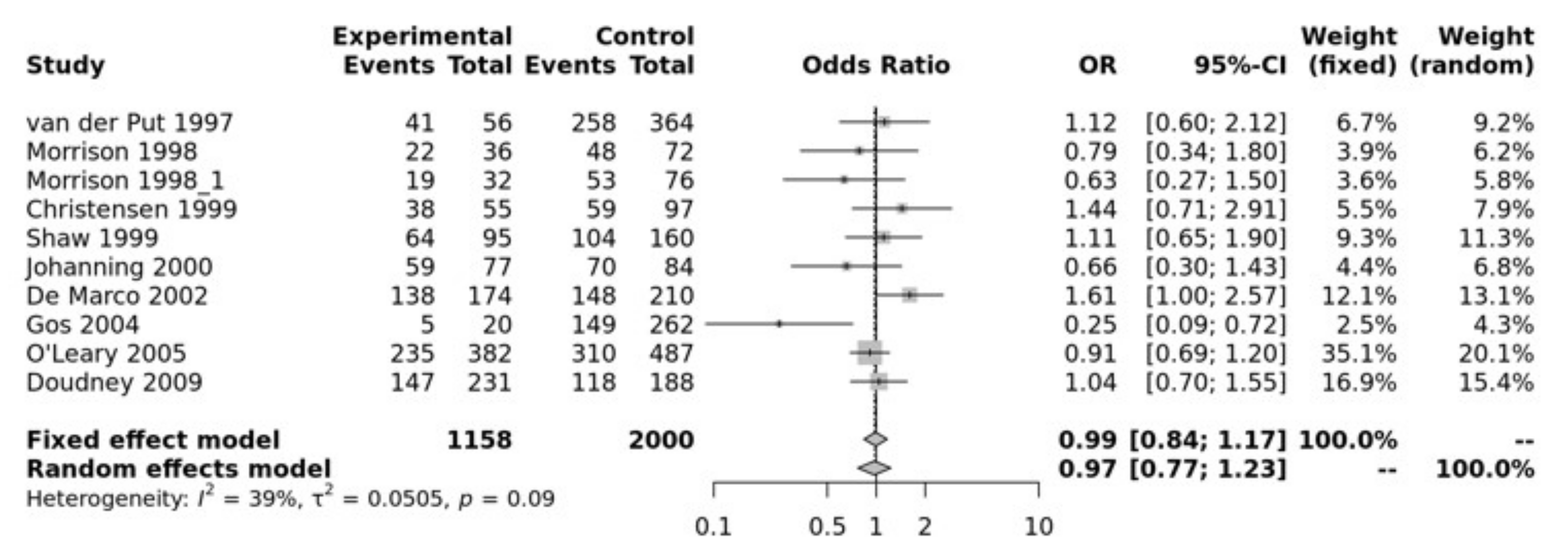
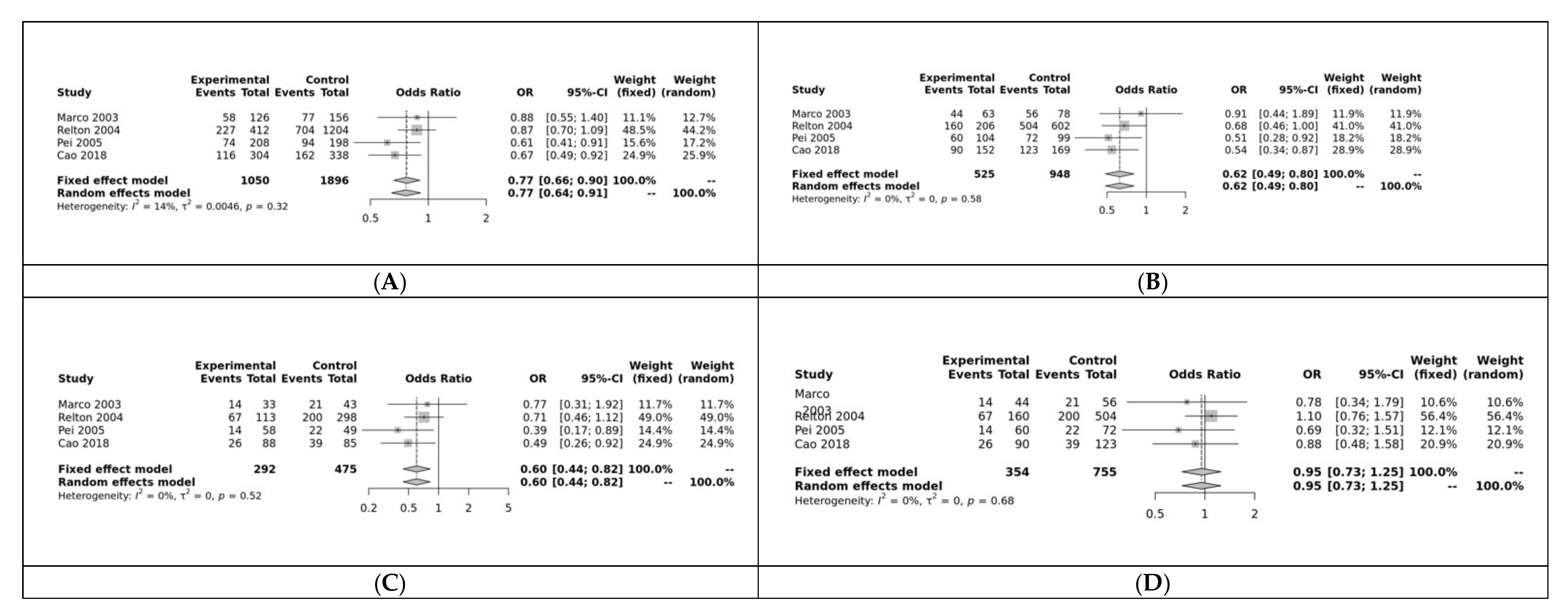
3.2. Meta-Analysis Results
3.3. Quality Assessment and Inter-Rater Agreement
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Frye, R.E.; Slattery, J.C.; Quadros, E.V. Folate metabolism abnormalities in autism: Potential biomarkers. Biomark. Med. 2017, 11, 687–699. [Google Scholar] [CrossRef] [PubMed]
- Rossignol, D.A.; Frye, R.E. Cerebral Folate Deficiency, Folate Receptor Alpha Autoantibodies and Leucovorin (Folinic Acid) Treatment in Autism Spectrum Disorders: A Systematic Review and Meta-Analysis. J. Pers. Med. 2021, 11, 1141. [Google Scholar] [CrossRef] [PubMed]
- Gil Martínez, V.; Avedillo Salas, A.; Santander Ballestín, S. Vitamin Supplementation and Dementia: A Systematic Review. Nutrients 2022, 14, 1033. [Google Scholar] [CrossRef] [PubMed]
- Visentin, M.; Diop-Bove, N.; Zhao, R.; Goldman, I.D. The intestinal absorption of folates. Annu. Rev. Physiol. 2014, 76, 251–274. [Google Scholar] [CrossRef] [PubMed]
- Boyer, J.L.; Soroka, C.J. Bile formation and secretion: An update. J. Hepatol. 2021, 75, 190–201. [Google Scholar] [CrossRef]
- Christianson, A.; Howson, C.P.; Modell, B. March of Dimes: Global Report on Birth Defect, the Hidden Toll of Dying and Disabled Children; March of Dimes Birth Defects Foundation: New York, NY, USA, 2006; pp. 10–16. [Google Scholar]
- Pitkin, R.M. Folate and neural tube defects. Am. J. Clin. Nutr. 2007, 85, 285S–288S. [Google Scholar] [CrossRef]
- Copp, A.J.; Greene, N.D.; Murdoch, J.N. The genetic basis of mammalian neurulation. Nat. Rev. Genet. 2003, 4, 784–793. [Google Scholar] [CrossRef]
- Smithells, R.W.; Sheppard, S.; Schorah, C.J. Vitamin dificiencies and neural tube defects. Arch. Dis. Child. 1976, 51, 944–950. [Google Scholar] [CrossRef]
- Wolff, T.; Witkop, C.T.; Miller, T.; Syed, S.B. Folic acid supplementation for the prevention of neural tube defects: An update of the evidence for the US Preventive Services Task Force. Ann. Intern. Med. 2009, 150, 632–639. [Google Scholar] [CrossRef]
- Laurence, K. Prevention of neural tube defects by improvement in maternal diet and preconceptional folic acid supplementation. Prog. Clin. Biol. Res. 1985, 163, 383–388. [Google Scholar]
- Christiansen, M.; Garne, E. Prevention of neural tube defects with periconceptional folic acid supplementation in Europe. Ugeskr. Laeger 2005, 167, 2875–2876. [Google Scholar]
- Berry, R.J.; Li, Z.; Erickson, J.D.; Li, S.; Moore, C.A.; Wang, H.; Mulinare, J.; Zhao, P.; Wong, L.-Y.C.; Gindler, J. Prevention of neural-tube defects with folic acid in China. N. Engl. J. Med. 1999, 341, 1485–1490. [Google Scholar] [CrossRef]
- Kibar, Z.; Capra, V.; Gros, P. Toward understanding the genetic basis of neural tube defects. Clin. Genet. 2007, 71, 295–310. [Google Scholar] [CrossRef] [PubMed]
- De-Regil, L.M.; Peña-Rosas, J.P.; Fernández-Gaxiola, A.C.; Rayco-Solon, P. Effects and safety of periconceptional oral folate supplementation for preventing birth defects. Cochrane Database Syst Rev. 2015, 12. [Google Scholar] [CrossRef]
- Bassuk, A.G.; Kibar, Z. Genetic basis of neural tube defects. Semin. Pediatric Neurol. 2009, 16, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, C.M.; Robien, K.; McLeod, H.L. Cancer pharmacogenetics: Polymorphisms, pathways and beyond. Nat. Rev. Cancer 2003, 3, 912–920. [Google Scholar] [CrossRef]
- Molloy, A.M.; Brody, L.C.; Mills, J.L.; Scott, J.M.; Kirke, P.N. The search for genetic polymorphisms in the homocysteine/folate pathway that contribute to the etiology of human neural tube defects. Birth Defects Res. Part A Clin. Mol. Teratol. 2009, 85, 285–294. [Google Scholar] [CrossRef]
- Richter, B.; Stegmann, K.; Röper, B.; Böddeker, I.; Ngo, E.T.; Koch, M.C. Interaction of folate and homocysteine pathway genotypes evaluated in susceptibility to neural tube defects (NTD) in a German population. J. Hum. Genet. 2001, 46, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Collin, S.M.; Metcalfe, C.; Zuccolo, L.; Lewis, S.J.; Chen, L.; Cox, A.; Davis, M.; Lane, J.; Donovan, J.; Smith, G.D. Association of Folate-Pathway Gene Polymorphisms with the Risk of Prostate Cancer: A Population-Based Nested Case-Control Study, Systematic Review, and Meta-analysisFolate-Pathway Gene Polymorphisms and Prostate Cancer. Cancer Epidemiol. Biomark. Prev. 2009, 18, 2528–2539. [Google Scholar] [CrossRef]
- Van der Linden, I.J.; Afman, L.A.; Heil, S.G.; Blom, H.J. Genetic variation in genes of folate metabolism and neural-tube defect risk. Proc. Nutr. Soc. 2006, 65, 204–215. [Google Scholar] [CrossRef]
- Greene, N.D.; Copp, A.J. Neural tube defects. Annu. Rev. Neurosci. 2014, 37, 221. [Google Scholar] [CrossRef] [PubMed]
- Boduroglu, K.; Alanay, Y.; Alikasifoglu, M.; Aktas, D.; Tunçbilek, E. Analysis of MTHFR 1298A> C in addition to MTHFR 677C> T polymorphism as a risk factor for neural tube defects in the Turkish population. Turk. J. Pediatrics 2005, 47, 327. [Google Scholar]
- Kirke, P.N.; Mills, J.L.; Molloy, A.M.; Brody, L.C.; O’Leary, V.B.; Daly, L.; Murray, S.; Conley, M.; Mayne, P.D.; Smith, O. Impact of the MTHFR C677T polymorphism on risk of neural tube defects: Case-control study. BMJ 2004, 328, 1535–1536. [Google Scholar] [CrossRef]
- De Marco, P.; Calevo, M.G.; Moroni, A.; Arata, L.; Merello, E.; Finnell, R.H.; Zhu, H.; Andreussi, L.; Cama, A.; Capra, V. Study of MTHFR and MS polymorphisms as risk factors for NTD in the Italian population. J. Hum. Genet. 2002, 47, 319–324. [Google Scholar] [CrossRef]
- Parle-McDermott, A.; Mills, J.L.; Kirke, P.N.; O’Leary, V.B.; Swanson, D.A.; Pangilinan, F.; Conley, M.; Molloy, A.M.; Cox, C.; Scott, J.M. Analysis of the MTHFR 1298A→ C and 677C→ T polymorphisms as risk factors for neural tube defects. J. Hum. Genet. 2003, 48, 190–193. [Google Scholar] [CrossRef] [PubMed]
- Shaw, G.M.; Rozen, R.; Finnell, R.H.; Wasserman, C.R.; Lammer, E.J. Maternal vitamin use, genetic variation of infant methylenetetrahydrofolate reducatase, and risk for spina bifida. Am. J. Epidemiol. 1998, 148, 30–37. [Google Scholar] [CrossRef]
- Relton, C.; Wilding, C.; Pearce, M.; Laffling, A.; Jonas, P.; Lynch, S.; Tawn, E.; Burn, J. Gene–gene interaction in folate-related genes and risk of neural tube defects in a UK population. J. Med. Genet. 2004, 41, 256–260. [Google Scholar] [CrossRef]
- Godbole, K.; Gayathri, P.; Ghule, S.; Sasirekha, B.V.; Kanitkar-Damle, A.; Memane, N.; Suresh, S.; Sheth, J.; Chandak, G.R.; Yajnik, C.S. Maternal one-carbon metabolism, MTHFR and TCN2 genotypes and neural tube defects in India. Birth Defects Res. Part A: Clin. Mol. Teratol. 2011, 91, 848–856. [Google Scholar] [CrossRef]
- De Franchis, R.; Buoninconti, A.; Mandato, C.; Pepe, A.; Sperandeo, M.P.; Del Gado, R.; Capra, V.; Salvaggio, E.; Andria, G.; Mastroiacovo, P. The C677T mutation of the 5, 10-methylenetetrahydrofolate reductase gene is a moderate risk factor for spina bifida in Italy. J. Med. Genet. 1998, 35, 1009–1013. [Google Scholar] [CrossRef] [PubMed]
- Mornet, E.; Muller, F.; Lenvoisé-Furet, A.; Delezoide, A.-L.; Col, J.-Y.; Simon-Bouy, B.; Serre, J.-L. Screening of the C677T mutation on the methylenetetrahydrofolate reductase gene in French patients with neural tube defects. Hum. Genet. 1997, 100, 512–514. [Google Scholar] [CrossRef] [PubMed]
- Moher, D.; Altman, D.G.; Liberati, A.; Tetzlaff, J. PRISMA statement. Epidemiology 2011, 22, 128. [Google Scholar] [CrossRef] [PubMed]
- Stang, A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur. J. Epidemiol. 2010, 25, 603–605. [Google Scholar] [CrossRef] [PubMed]
- Van der Put, N.; Thomas, C.; Eskes, T.; Trijbels, F.; Steegers-Theunissen, R.; Mariman, E.; De Graaf-Hess, A.; Smeitink, J.; Blom, H. Altered folate and vitamin B12 metabolism in families with spina bifida offspring. QJM Mon. J. Assoc. Physicians 1997, 90, 505–510. [Google Scholar] [CrossRef]
- Garcia-Fragoso, L.; García-García, I.; de la Vega, A.; Renta, J.; Cadilla, C.L. Presence of the 5, 10-methylenetetrahydrofolate reductase C677T mutation in Puerto Rican patients with neural tube defects. J. Child Neurol. 2002, 17, 30–32. [Google Scholar] [CrossRef] [PubMed]
- Perez, A.B.A.; D’Almeida, V.; Vergani, N.; de Oliveira, A.C.; de Lima, F.T.; Brunoni, D. Methylenetetrahydrofolate reductase (MTHFR): Incidence of mutations C677T and A1298C in Brazilian population and its correlation with plasma homocysteine levels in spina bifida. Am. J. Med. Genet. Part A 2003, 119, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Sadewa, A.H.; Sutomo, R.; Istiadjid, M.; Nishiyama, K.; Shirakawa, T.; Matsuo, M.; Nishio, H. C677T mutation in the MTHFR gene was not found in patients with frontoethmoidal encephalocele in East Java, Indonesia. Pediatrics Int. 2004, 46, 409–414. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.M.; Tang, J.H.; Cao, H.X.; Ding, J.H.; Wu, J.Z.; Wang, J.; Liu, Y.T.; Li, S.P.; Su, P.; Matsuo, K.; et al. MTHFR polymorphisms, dietary folate intake and breast cancer risk in Chinese women. J. Hum. Genet. 2009, 54, 414–418. [Google Scholar] [CrossRef]
- Eser, B.; Cosar, M.; Eser, O.; Erdogan, M.O.; Aslan, A.; Yildiz, H.; Boyaci, G.; Buyukbas, S.; Solak, M. 677C>T and 1298A>C polymorphisms of methylenetetrahydropholate reductase gene and biochemical parameters in Turkish population with spina bifida occulta. Turk. Neurosurg. 2010, 20, 9–15. [Google Scholar] [PubMed]
- Harisha, P.N.; Devi, B.I.; Christopher, R.; Kruthika-Vinod, T.P. Impact of 5, 10-methylenetetrahydrofolate reductase gene polymorphism on neural tube defects. J. Neurosurg. Pediatrics 2010, 6, 364–367. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Qi, J.; Yu, X.; Zhu, J.; Zhang, L.; Ning, Q.; Luo, X. Investigations of single nucleotide polymorphisms in folate pathway genes in Chinese families with neural tube defects. J. Neurol. Sci. 2014, 337, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Nasri, K.; Midani, F.; Kallel, A.; Ben Jemaa, N.; Aloui, M.; Boulares, M.; Lassoued, M.; Ben Halima, M.; Ben Wafi, S.; Soussi, M.; et al. Association of MTHFR C677T, MTHFR A1298C, and MTRR A66G Polymorphisms with Neural Tube Defects in Tunisian Parents. Pathobiology 2019, 86, 190–200. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Bai, B.-L.; Liu, X.-Z.; Miao, C.-Y.; Li, H. Association of folate metabolism genes MTRR and MTHFR with complex congenital abnormalities among Chinese population in Shanxi Province, China. Zhongguo Dang Dai Er Ke Za Zhi. 2014, 16, 840–845. [Google Scholar] [PubMed]
- Yu, J.; Chen, B.; Zhang, G.; Fu, S.; Li, P. The 677 C→T Mutation in the Methylenetetrahydrofolate Reductase (MTHFR) Gene in Five Chinese Ethnic Groups. Hum. Hered. 2000, 50, 268–270. [Google Scholar] [CrossRef] [PubMed]
- Pardo, R.; Suazo, J.; Castillo, S.; Vargas, M.; Zalavari, A.; Santos, J.L.; Blanco, R.; Rotter, K.; Solar, M.; Tapia, E. Methylenetetrahydrofolate reductase polymorphisms as risk factors for myelomeningocele. Rev. Med. Chil. 2014, 142, 587–592. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, Y.; Ji, W.; Qin, H.; Wu, H.; Xu, D.; Turtuohut, T.; Wang, Z. Variants in MTHFR gene and neural tube defects susceptibility in China. Metab. Brain Dis. 2015, 30, 1017–1026. [Google Scholar] [CrossRef] [PubMed]
- Dutta, H.K.; Borbora, D.; Baruah, M.; Narain, K. Evidence of gene-gene interactions between MTHFD1 and MTHFR in relation to anterior encephalocele susceptibility in Northeast India. Birth Defects Res. 2017, 109, 432–444. [Google Scholar] [CrossRef]
- Fang, Y.; Zhang, R.; Zhi, X.; Zhao, L.; Cao, L.; Wang, Y.; Cai, C. Association of main folate metabolic pathway gene polymorphisms with neural tube defects in Han population of Northern China. Child’s Nerv. Syst. 2018, 34, 725–729. [Google Scholar] [CrossRef]
- Stegmann, K.; Zietgler, A.; Ngo, E.T.; Kohlschmidt, N.; Schröter, B.; Ermert, A.; Koch, M.C. Linkage disequilibrium of MTHFR genotypes 677C/T-1298A/C in the German population and association studies in probands with neural tube defects(NTD). Am. J. Med Genet. 1999, 87, 23–29. [Google Scholar] [CrossRef]
- Barber, R.; Shalat, S.; Hendricks, K.; Joggerst, B.; Larsen, R.; Suarez, L.; Finnell, R. Investigation of Folate Pathway Gene Polymorphisms and the Incidence of Neural Tube Defects in a Texas Hispanic Population. Mol. Genet. Metab. 2000, 70, 45–52. [Google Scholar] [CrossRef]
- A Volcik, K.; Blanton, S.H.; Tyerman, G.H.; Jong, S.T.; Rott, E.J.; Palge, T.Z.; Romaine, N.K.; Northrup, H. Methylenetetrahydrofolate reductase and spina bifida: Evaluation of level of defect and maternal genotypic risk in Hispanics. Am. J. Med Genet. 2000, 95, 21–27. [Google Scholar] [CrossRef]
- Cunha, A.L.; Hirata, M.H.; Kim, C.; Guerra-Shinohara, E.M.; Nonoyama, K.; Hirata, R.D.C. Metabolic effects of C677T and A1298C mutations at the MTHFR gene in Brazilian children with neural tube defects. Clin. Chim. Acta. 2002, 318, 139–143. [Google Scholar] [CrossRef]
- Félix, T.M.; Leistner, S.; Giugliani, R. Metabolic effects and the methylenetetrahydrofolate reductase (MTHFR) polymorphism associated with neural tube defects in southern Brazil. Birth Defects Res. Part A: Clin. Mol. Teratol. 2004, 70, 459–463. [Google Scholar] [CrossRef] [PubMed]
- Gos, M.; Sliwerska, E.; Szpecht-Potocka, A. Mutation incidence in folate metabolism genes and regulatory genes in Polish families with neural tube defects. J. Appl. Genet. 2004, 45, 363–368. [Google Scholar] [PubMed]
- Grandone, E.; Corrao, A.M.; Colaizzo, D.; Vecchione, G.; Di Girgenti, C.; Paladini, D.; Sardella, L.; Pellegrino, M.; Zelante, L.; Martinelli, P.; et al. Homocysteine metabolism in families from southern Italy with neural tube defects: Role of genetic and nutritional determinants. Prenat. Diagn. 2006, 26, 1–5. [Google Scholar] [CrossRef]
- Gonzalez-Herrera, L.; Castillo-Zapata, I.; Garcia-Escalante, G.; Pinto-Escalante, D. A1298C polymorphism of the MTHFR gene and neural tube defects in the state of Yucatan, Mexico. Birth Defects Res A Clin Mol Teratol. 2007, 79, 622–626. [Google Scholar] [CrossRef]
- Behunova, J.; Klimcakova, L.; Zavadilikova, E.; Potocekova, D.; Sykora, P.; Podracka, L. Methylenetetrahydrofolate reductase gene polymorphisms and neural tube defects epidemiology in the Slovak population. Birth Defects Res. Part A: Clin. Mol. Teratol. 2010, 88, 695–700. [Google Scholar] [CrossRef]
- Selvi, N.; Ergun, R.; Gube, O.; Yİldİrİ, U.; Tetİk, A.; Gunduz, C.; Ulman, I.; Eroglu, Z. Genetic Polymorphism of Methylenetetrahydrofolate Reductase. Middle-East Journal of Scientific Research. 2010, 6, 93–98. [Google Scholar]
- Yildiz, S.H.; Erdogan, M.O.; Solak, M.; Eser, O.; Terzi, E.A.; Eser, B.; Kocabaş, V.; Aslan, A. Lack of association between the methylenetetrahydropholate reductase gene A1298C polymorphism and neural tube defects in a Turkish study group. Genet Mol Res. 2016, 15. [Google Scholar] [CrossRef]
- Morrison, K.; Papapetrou, C.; Hol, F.A.; Mariman, E.C.M.; Lynch, S.A.; Burn, J.; Edwards, Y.H. Susceptibility to spina bifida; an association study of five candidate genes. Ann. Hum. Genet. 1998, 62, 379–396. [Google Scholar] [CrossRef]
- Christensen, B.; Arbour, L.; Tran, P.; Leclerc, D.; Sabbaghian, N.; Platt, R.; Gilfix, B.M.; Rosenblatt, D.S.; Gravel, R.A.; Forbes, P.; et al. Genetic polymorphisms in methylenetetrahydrofolate reductase and methionine synthase, folate levels in red blood cells, and risk of neural tube defects. Am. J. Med. Genet. 1999, 84, 151–157. [Google Scholar] [CrossRef]
- Johanning, G.L.; Tamura, T.; Johnston, K.E.; Wenstrom, K.D. Comorbidity of 5,10-methylenetetrahydrofolate reductase and methionine synthase gene polymorphisms and risk for neural tube defects. J. Med Genet. 2000, 37, 949–951. [Google Scholar] [CrossRef]
- O’Leary, V.B.; Mills, J.L.; Parle-McDermott, A.; Pangilinan, F.; Molloy, A.M.; Cox, C.; Weiler, A.; Conley, M.; Kirke, P.N.; Scott, J.M.; et al. Screening for new MTHFR polymorphisms and NTD risk. Am. J. Med. Genet. Part A. 2005, 138, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Doudney, K.; Grinham, J.; Whittaker, J.; Lynch, S.; Thompson, D.; Moore, G.; Copp, A.L.; Greene, N.; Stanier, P. Evaluation of folate metabolism gene polymorphisms as risk factors for open and closed neural tube defects. Am. J. Med Genet. Part A. 2009, 149A, 1585–1589. [Google Scholar] [CrossRef] [PubMed]
- Wilson, A.; Platt, R.; Wu, Q.; Leclerc, D.; Christensen, B.; Yang, H.; Gravel, R.A.; Rozen, R. A Common Variant in Methionine Synthase Reductase Combined with Low Cobalamin (Vitamin B12) Increases Risk for Spina Bifida. Mol. Genet. Metab. 1999, 67, 317–323. [Google Scholar] [CrossRef]
- Pietrzyk, J.J.; Bik-Multanowski, M.; Salnak, M.; Twardowska, M. Polymorphisms of the 5,10-methylenetetrahydrofolate and the methionine synthase reductase genes as independent risk factors for spina bifida. J. Appl. Genet. 2003, 44, 111–113. [Google Scholar] [PubMed]
- Pei, L.; Zhu, H.; Retn, A.; Li, Z.; Hao, L.; Finnell, R.; Li, Z. Reduced folate carrier gene is a risk factor for neural tube defects in a Chinese population. Birth Defects Res. Part A: Clin. Mol. Teratol. 2005, 73, 430–433. [Google Scholar] [CrossRef] [PubMed]
- Cao, L.; Wang, Y.; Zhang, R.; Dong, L.; Cui, H.; Fang, Y.; Zhao, L.; Shi, O.; Cai, C. Association of neural tube defects with gene polymorphisms in one-carbon metabolic pathway. Child’s Nerv. Syst. 2018, 34, 277–284. [Google Scholar] [CrossRef]
- Kotova, N.; Maichuk, V.; Fedorenko, O. The differentiated approach to prevention of neural tube defects in children. Georgian Med. News. 2018, 274, 52–59. [Google Scholar]
- Crider, K.S.; Yang, T.P.; Berry, R.J.; Bailey, L.B. Folate and DNA methylation: A review of molecular mechanisms and the evidence for folate’s role. Adv. Nutr. 2012, 3, 21–38. [Google Scholar] [CrossRef]
- Zheng, J.; Lu, X.; Liu, H.; Zhao, P.; Li, K.; Li, L. MTHFD1 polymorphism as maternal risk for neural tube defects: A meta-analysis. Neurol. Sci. 2015, 36, 607–616. [Google Scholar] [CrossRef]
- Friso, S.; Choi, S.-W.; Girelli, D.; Mason, J.B.; Dolnikowski, G.G.; Bagley, P.J.; Olivieri, O.; Jacques, P.F.; Rosenberg, I.H.; Corrocher, R. A common mutation in the 5, 10-methylenetetrahydrofolate reductase gene affects genomic DNA methylation through an interaction with folate status. Proc. Natl. Acad. Sci. USA 2002, 99, 5606–5611. [Google Scholar] [CrossRef] [PubMed]
- Kondo, A.; Kamihira, O.; Ozawa, H. Neural tube defects: Prevalence, etiology and prevention. Int. J. Urol. 2009, 16, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Frosst, P.; Blom, H.; Milos, R.; Goyette, P.; Sheppard, C.A.; Matthews, R.; Boers, G.; Den Heijer, M.; Kluijtmans, L.; Van Den Heuve, L. A candidate genetic risk factor for vascular disease: A common mutation in methylenetetrahydrofolate reductase. Nat. Genet. 1995, 10, 111–113. [Google Scholar] [CrossRef]
- Weisberg, I.; Tran, P.; Christensen, B.; Sibani, S.; Rozen, R. A second genetic polymorphism in methylenetetrahydrofolate reductase (MTHFR) associated with decreased enzyme activity. Mol. Genet. Metab. 1998, 64, 169–172. [Google Scholar] [CrossRef]
- Li, Y.N.; Gulati, S.; Baker, P.J.; Brody, L.C.; Banerjee, R.; Kruger, W.D. Cloning, mapping and RNA analysis of the human methionine synthase gene. Hum. Mol. Genet. 1996, 5, 1851–1858. [Google Scholar] [CrossRef]
- Selzer, R.R.; Rosenblatt, D.S.; Laxova, R.; Hogan, K. Adverse effect of nitrous oxide in a child with 5, 10-methylenetetrahydrofolate reductase deficiency. N. Engl. J. Med. 2003, 349, 45–50. [Google Scholar] [CrossRef]
- Sutherland, H.G.; Hermile, H.; Sanche, R.; Menon, S.; Lea, R.A.; Haupt, L.M.; Griffiths, L.R. Association study of mthfd 1 coding polymorphisms r 134 k and r 653 q with migraine susceptibility. Headache J. Head Face Pain. 2014, 54, 1506–1514. [Google Scholar] [CrossRef] [PubMed]
- Christensen, K.E.; Dahhou, M.; Kramer, M.S.; Rozen, R. The MTHFD1 1958G>A variant is associated with elevated C-reactive protein and body mass index in Canadian women from a premature birth cohort. Mol. Genet. Metab. 2014, 111, 390–392. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Jing, Y.; Sun, Z. Lack of association between MTHFD1 G401A polymorphism and ovarian cancer susceptibility. Tumor Biol. 2014, 35, 3385–3389. [Google Scholar] [CrossRef]
- Da Silva, L.M.; Galbiatti, A.L.; Ruiz, M.T.; Raposo, L.S.; Maniglia, J.V.; Pavarino, E.C.; Goloni-Bertollo, E.M. MTHFD1 G1958A, BHMT G742A, TC2 C776G and TC2 A67G polymorphisms and head and neck squamous cell carcinoma risk. Mol. Biol. Rep. 2012, 39, 887–893. [Google Scholar] [CrossRef]
- García-González, I.; Flores-Ocampo, J.; Alejos-Mex, R.; Herrera-Sánchez, L.; González-Herrera, L. G894T (NOS3) and G1958A (MTHFD1) gene polymorphisms and risk of ischemic heart disease in Yucatan, Mexico. Clin. Investig. Arterioscler. Publ. Of. Soc. Esp. Arterioscler. 2014, 27, 64–73. [Google Scholar]
- Jiang, J.; Zhang, Y.; Wei, L.; Sun, Z.; Liu, Z. Association between MTHFD1 G1958A polymorphism and neural tube defects susceptibility: A meta-analysis. PLoS ONE 2014, 9, e101169. [Google Scholar] [CrossRef]
- Keller, M.D.; Ganesh, J.; Heltzer, M.; Paessler, M.; Bergqvist, A.C.; Baluarte, H.J.; Watkins, D.; Rosenblatt, D.S.; Orange, J.S. Severe combined immunodeficiency resulting from mutations in MTHFD1. Pediatrics 2013, 131, e629–e634. [Google Scholar] [CrossRef] [PubMed]
- Lorenc, A.; Seremak-Mrozikiewicz, A.; Barlik, M.; Wolski, H.; Drews, K. The role of 401a>G polymorphism of methylenetetrahydrofolate dehydrogenase gene (MTHFD1) in fetal hypotrophy. Ginekol. Pol. 2014, 85, 494–499. [Google Scholar] [CrossRef]
- Murthy, J.; Gurramkonda, V.B.; Lakkakula, B.V. Significant association of MTHFD1 1958G>A single nucleotide polymorphism with nonsyndromic cleft lip and palate in Indian population. Med. Oral Patol. Oral Y Cir. Bucal 2014, 19, e616–e621. [Google Scholar] [CrossRef]
- Carroll, N.; Pangilinan, F.; Molloy, A.M.; Troendle, J.; Mills, J.L.; Kirke, P.N.; Brody, L.C.; Scott, J.M.; Parle-McDermott, A. Analysis of the MTHFD1 promoter and risk of neural tube defects. Hum. Genet. 2009, 125, 247–256. [Google Scholar] [CrossRef]
- Parle-McDermott, A.; Pangilinan, F.; O’Brien, K.K.; Mills, J.L.; Magee, A.M.; Troendle, J.; Sutton, M.; Scott, J.M.; Kirke, P.N.; Molloy, A.M.; et al. A common variant in MTHFD1L is associated with neural tube defects and mRNA splicing efficiency. Hum. Mutat. 2009, 30, 1650–1656. [Google Scholar] [CrossRef]
- Green, R.F.; Byrne, J.; Crider, K.S.; Gallagher, M.; Koontz, D.; Berry, R.J. Folate-related gene variants in Irish families affected by neural tube defects. Front. Genet. 2013, 4, 223. [Google Scholar] [CrossRef]

| MTHFR C677T | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Model | Ethnicity | Number of Studies | Test of Association | Test of Heterogeneity | Bias | ||||
| OR | 95% CI | p-Value | Model | p-Value | I^2 | p | |||
| Allele contrast (A vs. a) | Overall | 19 | 0.722 | [0.6337; 0.8227] | 0.000001 | Random | 0.0022 | 0.55 | 0.75 |
| Asian | 10 | 0.629 | [0.5021; 0.7878] | 0.00005 | Random | 0.0040 | 0.63 | 0.33 | |
| Caucasian | 7 | 0.7676 | [0.6462; 0.9118] | 0.003 | Random | 0.0825 | 0.46 | 0.47 | |
| Mixed | 2 | 0.9273 | [0.6956; 1.2363] | 0.61 | Fixed | 0.5556 | |||
| Recessive model (AA vs. Aa + aa) | Overall | 19 | 0.682 | [0.5759; 0.8077] | 0.000009 | Random | 0.0302 | 0.42 | 0.51 |
| Asian | 10 | 0.5676 | [0.4623; 0.6970] | 0.00000006 | Fixed | 0.2346 | 0.23 | 0.74 | |
| Caucasian | 7 | 0.7817 | [0.6053; 1.0094] | 0.06 | Random | 0.0517 | 0.52 | 0.72 | |
| Mixed | 2 | 0.8206 | [0.5697; 1.1819] | 0.29 | Fixed | 0.3387 | |||
| Dominant model (AA + Aa vs. aa) | Overall | 18 | 0.6441 | [0.5219; 0.7948] | 0.00004 | Random | 0.0177 | 0.46 | 0.87 |
| Asian | 9 | 0.5954 | [0.4205; 0.8430] | 0.003 | Random | 0.0069 | 0.62 | 0.23 | |
| Caucasian | 7 | 0.5727 | [0.4648; 0.7057] | 0.0000001 | Fixed | 0.5767 | 0.00 | 0.28 | |
| Mixed | 2 | 1.3088 | [0.6597; 2.5965] | 0.44 | Fixed | 0.8181 | |||
| Homozygote (AA vs. aa) | Overall | 18 | 0.5519 | [0.4380; 0.6955] | 0.0000005 | Random | 0.0539 | 0.38 | 0.89 |
| Asian | 9 | 0.4627 | [0.3204; 0.6681] | 0.00004 | Random | 0.0774 | 0.44 | 0.25 | |
| Caucasian | 7 | 0.5443 | [0.4334; 0.6837] | 0.0000002 | Fixed | 0.2066 | 0.29 | 0.52 | |
| Mixed | 2 | 1.1424 | [0.5639; 2.3145] | 0.71 | Fixed | 0.9638 | |||
| Heterozygous (AA vs. Aa) | Overall | 19 | 0.7328 | [0.6141; 0.8745] | 0.0006 | Random | 0.0442 | 0.39 | 0.46 |
| Asian | 10 | 0.6034 | [0.4824; 0.7546] | 0.00001 | Fixed | 0.3125 | 0.14 | 0.80 | |
| Caucasian | 7 | 0.8635 | [0.6633; 1.1240] | 0.28 | Random | 0.0675 | 0.49 | 0.93 | |
| Mixed | 2 | 0.7676 | [0.5231; 1.1265] | 0.18 | Fixed | 0.2662 | 0.19 | ||
| RFC A80G | |||||||||
| Allele contrast (A vs. a) | Overall | 4 | 0.7729 | [0.6607; 0.9043] | 0.001 | Fixed | 0.3230 | 0.14 | 0.53 |
| Asian | 2 | 0.6468 | [0.5053; 0.8280] | 0.0005 | Fixed | 0.7203 | 0.00 | ||
| Caucasian | 2 | 0.8721 | [0.7117; 1.0688] | 0.19 | Fixed | 0.9875 | 0.00 | ||
| Recessive model (AA vs. Aa + aa) | Overall | 4 | 0.8223 | [0.6382; 1.0596] | 0.13 | Fixed | 0.4688 | 0.00 | 0.15 |
| Asian | 2 | 0.6321 | [0.4062; 0.9838] | 0.04 | Fixed | 0.6188 | 0.00 | ||
| Caucasian | 2 | 0.9352 | [0.6864; 1.2743] | 0.67 | Fixed | 0.6064 | 0.00 | ||
| Dominant model (AA + Aa vs. aa) | Overall | 4 | 0.6248 | [0.4858; 0.8036] | 0.0002 | Fixed | 0.5797 | 0.00 | 0.76 |
| Asian | 2 | 0.5304 | [0.3676; 0.7653] | 0.0007 | Fixed | 0.8762 | 0.00 | ||
| Caucasian | 2 | 0.7229 | [0.5115; 1.0217] | 0.07 | Fixed | 0.4832 | 0.00 | ||
| Homozygote (AA vs. aa) | Overall | 4 | 0.6028 | [0.4410; 0.8238] | 0.002 | Fixed | 0.5159 | 0.00 | 0.55 |
| Asian | 2 | 0.4536 | [0.2756; 0.7467] | 0.002 | Fixed | 0.6543 | 0.00 | ||
| Caucasian | 2 | 0.7246 | [0.4852; 1.0821] | 0.12 | Fixed | 0.8798 | 0.00 | ||
| Heterozygous (AA vs. Aa) | Overall | 4 | 0.9533 | [0.7268; 1.2504] | 0.73 | Fixed | 0.6769 | 0.00 | 0.03 |
| Asian | 2 | 0.8028 | [0.5006; 1.2876] | 0.36 | Fixed | 0.6384 | 0.00 | ||
| Caucasian | 2 | 1.0374 | [0.7447; 1.4451] | 0.83 | Fixed | 0.4605 | 0.00 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Almekkawi, A.K.; AlJardali, M.W.; Daadaa, H.M.; Lane, A.L.; Worner, A.R.; Karim, M.A.; Scheck, A.C.; Frye, R.E. Folate Pathway Gene Single Nucleotide Polymorphisms and Neural Tube Defects: A Systematic Review and Meta-Analysis. J. Pers. Med. 2022, 12, 1609. https://doi.org/10.3390/jpm12101609
Almekkawi AK, AlJardali MW, Daadaa HM, Lane AL, Worner AR, Karim MA, Scheck AC, Frye RE. Folate Pathway Gene Single Nucleotide Polymorphisms and Neural Tube Defects: A Systematic Review and Meta-Analysis. Journal of Personalized Medicine. 2022; 12(10):1609. https://doi.org/10.3390/jpm12101609
Chicago/Turabian StyleAlmekkawi, Ahmad K., Marwa W. AlJardali, Hicham M. Daadaa, Alison L. Lane, Ashley R. Worner, Mohammad A. Karim, Adrienne C. Scheck, and Richard E. Frye. 2022. "Folate Pathway Gene Single Nucleotide Polymorphisms and Neural Tube Defects: A Systematic Review and Meta-Analysis" Journal of Personalized Medicine 12, no. 10: 1609. https://doi.org/10.3390/jpm12101609
APA StyleAlmekkawi, A. K., AlJardali, M. W., Daadaa, H. M., Lane, A. L., Worner, A. R., Karim, M. A., Scheck, A. C., & Frye, R. E. (2022). Folate Pathway Gene Single Nucleotide Polymorphisms and Neural Tube Defects: A Systematic Review and Meta-Analysis. Journal of Personalized Medicine, 12(10), 1609. https://doi.org/10.3390/jpm12101609








