Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe
Abstract
1. Introduction
2. Core Challenges
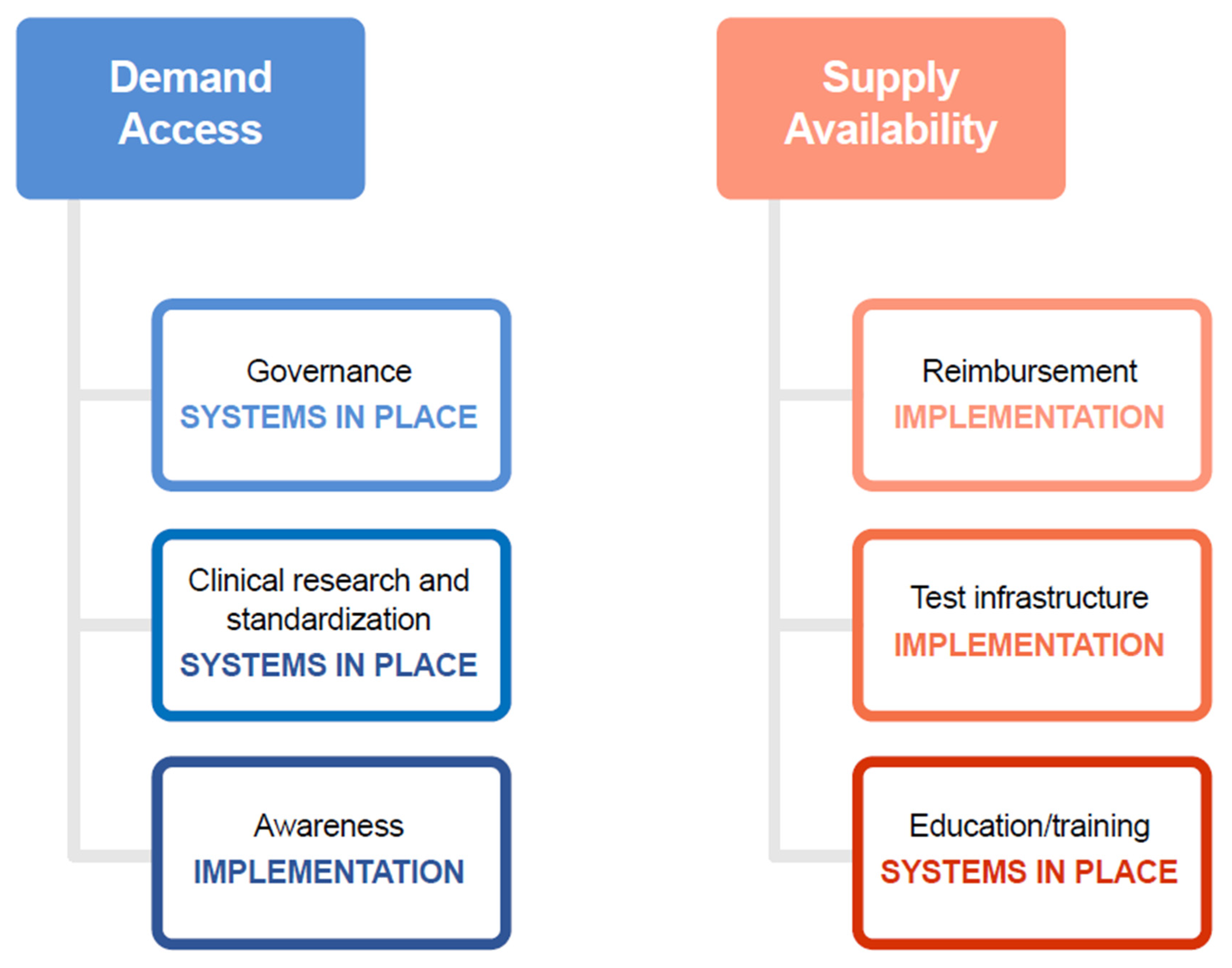
2.1. Key Challenges to the Demand for Tests
2.1.1. Governance
- Insufficient harmonization of clinical infrastructures (e.g., data capture through electronic health records and limited laboratory and analytic services) due to the absence of national strategies for the implementation of personalized medicine approaches, confounded further by the absence of a pan-stakeholder group (i.e., physicians, patients, payers) well placed to advise on the implementation of NGS.
- Lack of oversight in the implementation of policies and guidelines, as well as a lack of a regulatory framework to aid implementation. For instance, in France, implementation of ESMO recommendations currently varies between different institutions and laboratories, due to varying awareness of such guidelines.
- A dynamic environment, with increasing numbers of innovative MGTOs and clinical trial designs, and greater volumes of genomic data from high-throughput technologies. Healthcare systems are often not ready to respond, and there are typically logistical challenges related to data management and sharing due to General Data Protection Regulation requirements and ever-increasing demands for data storage, and difficulties with data collection and harmonization.
- A wide range of country-specific governance challenges that require resolution (summarized in Table S1).
2.1.2. Clinical Standardization
- Among clinicians, there are variations in the patterns of ordering large gene panels (>50 genes), due to lack of validation and standardization of in-house biomarkers and testing procedures, as well as varying application of International Organization for Standardization (ISO)-based accreditation (ISO 15189) [29] between hospitals. Standardization between hospitals may be difficult to achieve due to the complexity of NGS, but can be aided by the use of commercial NGS test providers that have achieved clinical validation for particular biomarkers [7].
- Insufficient guidance on evidence requirements to demonstrate clinical utility of NGS.
- Although some MGTOs have demonstrated clinical activity across multiple tumor types sharing the same molecular alteration (e.g., NTRK fusion, microsatellite instability, DNA mismatch repair-deficiencies [30,31]), leading to broad or even tumor-agnostic approvals, there remains insufficient understanding of the varying efficacy in different tumor types. For example, the poly(ADP-ribose) polymerase inhibitor olaparib leads to an improvement in progression-free survival of ~3 years in patients with BRCA1/2-altered ovarian tumors [32], compared with up to ~7 months for BRCA1/2-altered breast, pancreatic, or prostate cancer [33,34,35]. Furthermore, dabrafenib plus trametinib has demonstrated varying objective response rates of 61–68% between patients with previously untreated melanoma and non-small cell lung cancer [36,37].
- Factors affecting choice of test type, such as cancer type, age, gender, high and low scoring of tumor mutational burden, and challenges related to quality assurance and reliability of assay results.
2.1.3. Awareness and Education
2.2. Key Challenges to the Supply of Tests
2.2.1. Reimbursement
- Lack of value assessment processes for diagnostics, including NGS, in many countries and a generalized lack of formal pathways for reimbursement. There is also a lack of pathways for reimbursement that are time-bound and flexible and that accommodate the unique characteristics of the advanced diagnostic [43].
- Timing to access for advanced diagnostics is currently a major barrier to adoption.
- Absence of a widely accepted health technology assessment (HTA) framework for demonstrating the value of diagnostic methods, including NGS, as well as other health technologies, to improve access to targeted therapies.
- Inconsistency across countries in whether NGS should be evaluated and reimbursed at the generic level or for the class of test (e.g., based on the number of base pairs sequenced/size of the gene panel). Inconsistent evaluations are also a result of a lack of evidence frameworks for NGS, including broadly agreed-upon standards to demonstrate clinical utility, mitigate uncertainty, and secure public/private reimbursement [44].
- In some countries, lack of reimbursement codes for NGS or insufficient coverage for laboratories to conduct NGS.
- Insufficient laboratory/hospital budgets to cover growing volumes of NGS testing, and siloed budgets between diagnostics and hospitals, leading to infrastructural challenges (e.g., obtaining an appropriately trained workforce, data sharing and data storage infrastructure capabilities, and digital health recording within healthcare systems) [43]. Such budget insufficiency is also seen more broadly for non-academic stakeholders co-investing in a public–private partnership, e.g., via an innovation fund.
2.2.2. Infrastructure
- Differences in centralized/decentralized testing infrastructure requirements.
- Lack of alignment of availability of bioinformatics technology (e.g., data management, analysis, and storage) across Europe.
- Limited knowledge and capacity for interpreting NGS reports.
- Variable and inconsistent methods of ordering biomarker testing.
- Insufficient funding for NGS.
- Insufficient efforts to resource and train a specialized workforce.
- Lack of harmonization across datasets, limiting application of artificial intelligence.
- Varying levels of centralization and infrastructure capacity across Europe, leading to several country-specific recommendations (Table S5).
2.2.3. Access to Tests Linked to Evidence Generation and Supporting Infrastructure
- Lack of national registries.
- Data standardization and harmonization.
- Data protection legislation.
- Lack of European-level regulatory guidance for institutions conducting RWD-based studies that sufficiently address concerns from payers or regulatory bodies, including RWD quality and comprehensiveness, data standardization, methodological challenges and lack of transparency, failure to collect or prioritize data relevant to coverage decision-making (health resource utilization), and small sample sizes [46,54,55].
- Biased RWE due to connections with groups assessing reimbursement.
- Incorporation of RWE into the value assessment process, which is critical for securing reimbursement and enabling an RWE-based learning healthcare system to continually study, learn, and improve precision oncology.
- Various country-specific challenges (Table S6).
3. Relevant Recent and Upcoming NGS-Related Initiatives
4. Detailed Recommendations
4.1. Governance
- Evidence generation: Demonstration of the clinical utility of NGS to patients, both at baseline and at tumor progression, is critical in supporting national initiatives and harmonized implementation of policies and guidelines. As NGS typically succeeds on a case-by-case basis, this would allow for selection of the most appropriate patients to receive NGS. Traditional randomized clinical trials may not easily generate such evidence but can be aided through innovative clinical trial designs and RWD-based studies [45].
- Up-to-date strategies: Alignment of national precision medicine strategies across Europe with the ever-changing environment of NGS and innovative treatment options. This can avert the risk of local resistance that inhibits the implementation of European NGS policies, such as Europe’s BCP or ESMO recommendations. Divergent interpretations in France of the ESMO recommendations highlight the need for more formal recommendations from the country’s National Cancer Institute.
- Maximal European cooperation: European-level cooperation is required to discuss challenges of patient access to NGS and to ensure equal implementation across the Member States of the European Commission’s 1+ Million Genomes initiative, as well as other initiatives, including EHDEN and the European Health Data Space [26,74].
- NGS standardization: National strategies must aim to: regulate NGS standardization and quality; clinical interpretation of genomic variants (based on, e.g., the ESMO Scale for Clinical Actionability of molecular Targets (ESCAT)) [78,79] and access to evolving treatment options; methods of measuring tumor mutational burden; efficient collection and use of tumor samples; privacy and ethics in relation to data sharing and participation in international clinical databases; data interoperability for use; and the tumor types that incorporate RNA (for gene fusions and, in the near future, gene expressions), circulating tumor DNA, or epigenetic analysis as part of testing. This is necessary for consistency between laboratories performing genomic testing, including consistency regarding quality, the test/platform used, and the interpretation of results, particularly in those countries without laboratory accreditation. Patients and patient advocates should be involved in the discussion of privacy and ethics to advance the application of robust patient consent procedures in all Member States, with clear guidance on who should and can obtain consent, and educational initiatives to equip clinicians accordingly [80].
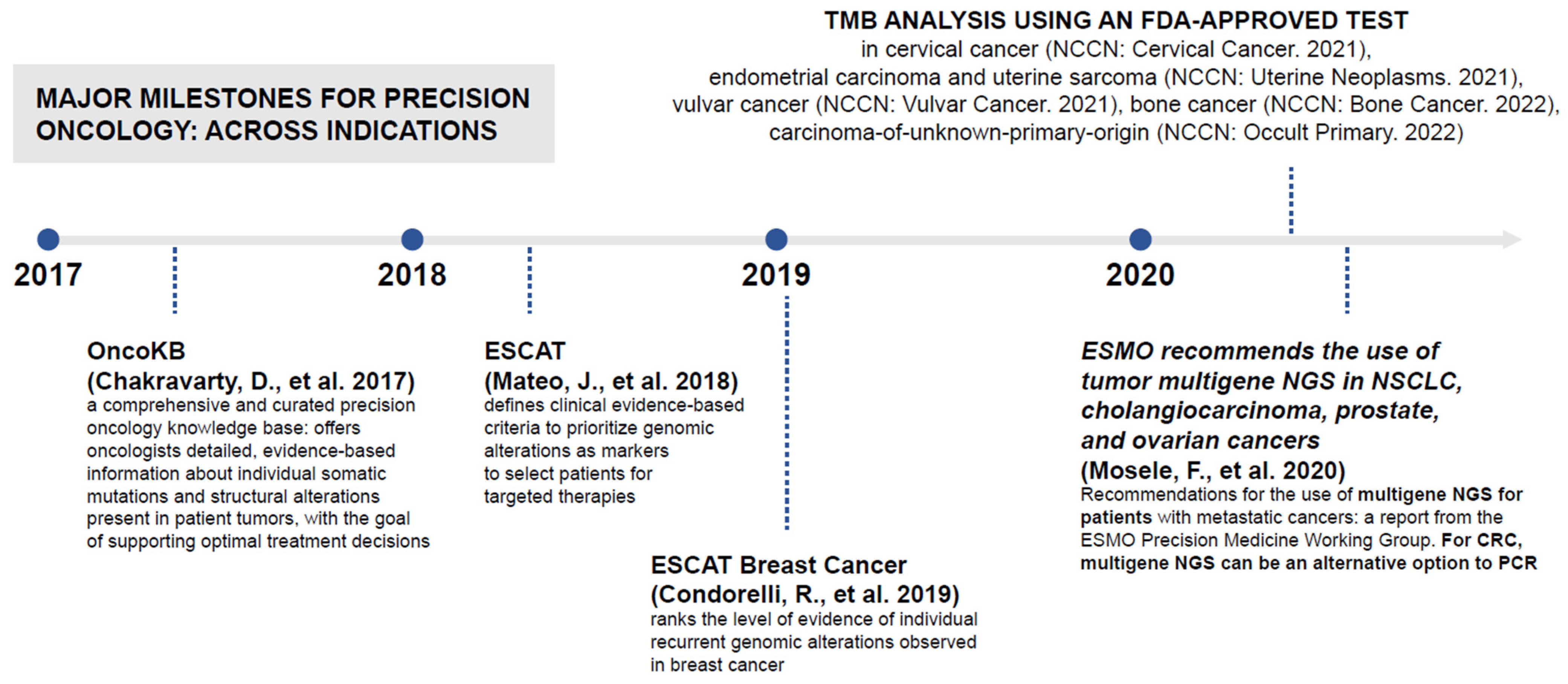
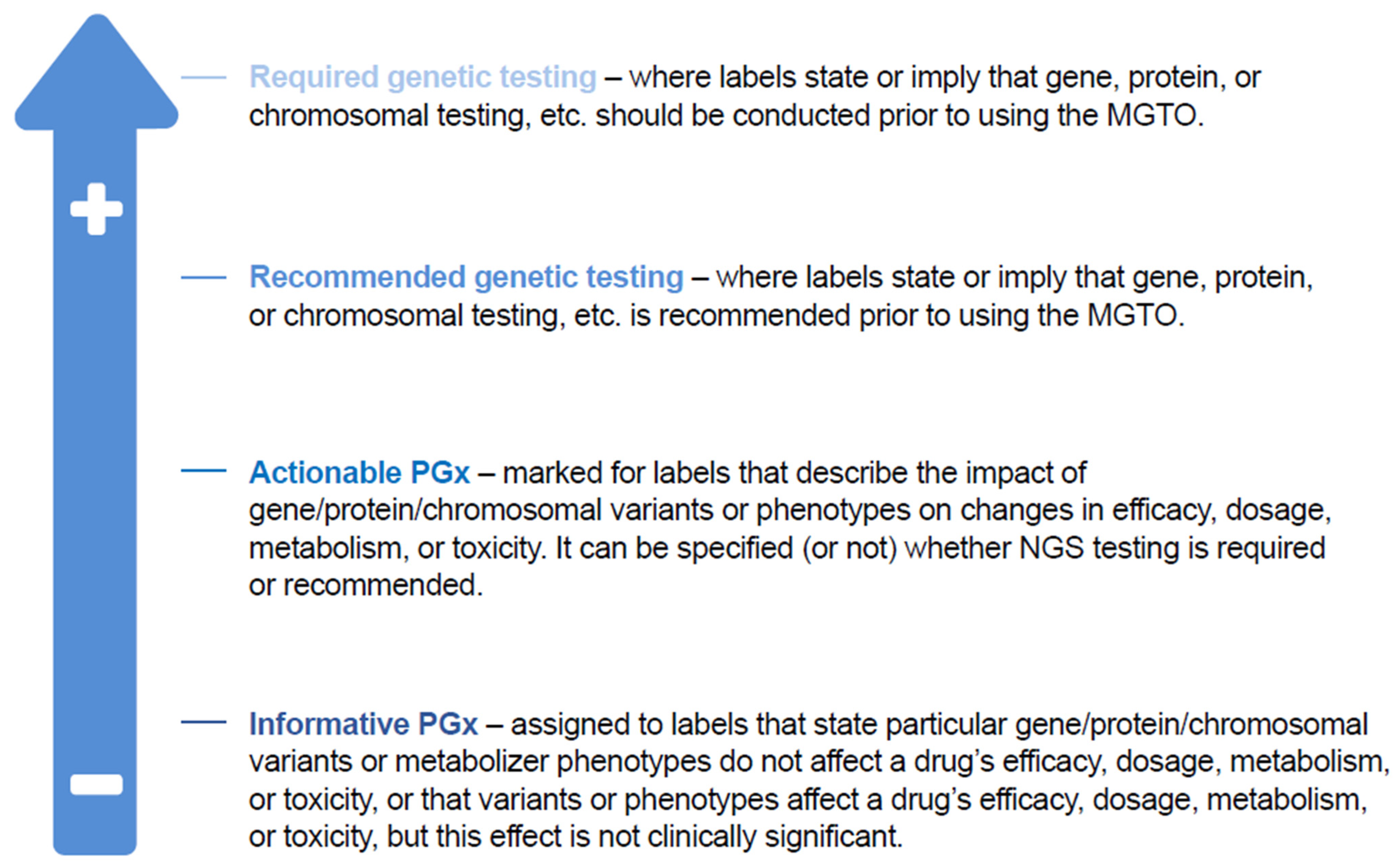
- Clear classification: National strategies should classify the use of NGS testing in the context of clinical decision-making and delivery of care. A US method exists to classify pharmacogenomic information from a particular gene–drug combination as clinically actionable (PharmGKB, Stanford, CA, USA; Figure 4) [81]; however, translating this classification system to routine clinical practice would require a clear definition of the aim of NGS testing (i.e., diagnosis, assessing immediate treatments decisions, identifying actionable alterations), clear delineation of the difference between genomic insights based on clinical validation (i.e., companion diagnostics as required by the new IVDR or based on Software as a Medical Device) and insights based on research-use-only techniques, and attention to the importance of the pharmacogenomic information that can be vital for disease prevention and for understanding potential drug toxicities. Consistent interpretation of NGS results depends on standardized MTB evidence scales, which are reliant on validated bioinformatics analysis pathways, variant calling, and pipeline analysis. OncoKB is a knowledge base offering evidence-based information about individual somatic mutations and structural alterations present in patient tumors [82]; it has an important role in improving reproducibility and accuracy of MTB findings. ESCAT also helps to rank targets for precision oncology based on clinical evidence of their utility and to prioritize targets for clinical use [78,79].
- Clinically useful MTBs: MTBs are critical to accommodate the ever-evolving environment of NGS and precision oncology, to harmonize interpretation of NGS results and thus aid translation into clinical action and benefit [25]. To maximize its clinical utility, it is necessary to clearly define the purpose of an MTB, to establish guidelines on minimal membership, operational requirements, and measures of quality, and to secure global harmonization in the sharing of clinical data experiences between institutions. It is also critical to accommodate virtual MTBs in a time- and location-independent manner between cancer care teams comprising experts across several disciplines. MTBs must be reactive and adaptive to recent clinical trial results and capture patient outcomes to validate clinical decision-making as well as support a learning healthcare system. Several further country-specific recommendations are summarized in Table 1.
4.2. Clinical Standardization
- Clear, up-to-date, and dynamic guidance: Guidance describing when and where NGS should be performed, including the genes and patient populations to be analyzed, minimum standards for testing to ensure high-quality NGS, and limiting NGS to laboratories with ISO-based accreditation (according to the ISO 15189 standards) [29], as a basis for establishing an external quality assessment system.
- NGS workflow standardization and optimization: Standardization and tracking of the entire NGS workflow, including the pre-analytical (i.e., tissue quality, tumor content quantification, DNA fragmentation, and library preparation), DNA sequencing, coverage, and data-analysis steps. Standardization is also essential to avoid outliers when using NGS for precision-oncology-based research. In addition, optimization of the NGS workflow, particularly in relation to turnaround time, is key. Europe’s BCP offers a model aimed at ensuring high-quality innovative treatments and diagnostics, with an EU Network aiming to link recognized National Comprehensive Cancer Centers in every member state by 2025 and guarantee access to such centers for 90% of eligible patients by 2030 [26]. Such centers should involve external stakeholders to promote multicenter collaboration.
- European collaboration to address the IVDR: In-house laboratory in vitro diagnostic tests are essential to ensure broad coverage of healthcare in many diagnostic fields and will be impacted by IVDR, which limits their use to situations in which appropriate certified in vitro diagnostic tests are not available [73]. Diagnostic processes, for instance to validate individual genomic alterations, may be complicated by the IVDR. Moreover, many current NGS tests are not performed according to IVDR requirements, which will lead to a steep increase in the number of tests requiring certification [73]. Compliance with guidelines may require test revalidation and significant replacement of technology, but, given limited funding in some countries, the result may be an increase in laboratory reliance on commercial providers leading to concentration among laboratories. In addition, in the dynamic and fast-moving fields of precision oncology and molecular pathology, laboratory-developed tests play a central role, as they often drive diagnostic innovation in collaboration with industry when no approved testing options exist. Nonetheless, the IVDR, established in 2017, is key to ensuring high-quality, standardized, and high-throughput testing, and it aims to guarantee safe and effective use of the corresponding medicinal product [83]. This may address many of the challenges detailed above, and laboratories must prepare for this legislation. In that regard, laboratories should appoint a small team dedicated to ensuring compliance with IVDR regulations and prepare an inventory of all currently implemented commercially available and laboratory-developed tests. Furthermore, laboratories must engage in European collaboration as part of multicenter studies to generate data related to laboratory-developed tests and obtain appropriate certification, and consult published IVDR guidance documents from the European Commission (although some of these documents are still pending) [73].
4.3. Awareness and Education
- Physician, molecular pathologist, and bioinformatician education: Education should equip physicians, molecular pathologists, and bioinformaticians to order, interpret, and use genomic evidence to guide treatment, from students and trainees (e.g., updated undergraduate genomics curriculum) to consultants and associate specialists, providing advice at clinical genetics services and genomic laboratory hubs [80]. MTBs and clinical-decision-support systems should be used to assist this process. It is important that regular, objective meetings designed for postgraduate training be organized and that they be able to indicate clear arrangements and plans for infrastructure or reimbursement.
- Involvement of patient advisory groups: With the assistance of patient advisory groups, patients’ preferences should be considered throughout the regulatory (e.g., value assessments) and scientific processes, and it is key to manage patients’ expectations of NGS. For example, patient organizations in the Netherlands are requesting the release of NGS reports to both patient and clinician, which will help to raise awareness of NGS and precision oncology across all tumor types.
- Broader national initiatives: There should be wider use of national initiatives, such as the Health Education England’s Genomics Education Programme and the NHS’s Genomics Clinical Reference Group (Box 2), which serve as examples of connecting genomic data sources and combining data with clinical RWD. There are no equivalents in Portugal and Italy, and such national initiatives could also benefit Germany’s multiple ongoing initiatives (e.g., German Cancer Aid). As part of integrity and harmonization, the UK initiatives should be used as an approach to connect genomic data sources and combine data with clinical RWD in a centralized, aggregated manner across Europe.
- Framework from Europe’s BCP: Europe’s BCP could provide a framework for these recommendations in its recruitment of social workers, teachers, and nurses to improve public health literacy on cancer risks and determinants, as well as to educate patients on healthy behavior and on how they can live well following cancer treatment. Europe’s BCP also aims to use training and continuous education of healthcare professionals, including on digital skills, artificial intelligence, genomics, and personalized medicine, to build a stronger multidisciplinary cancer workforce. This will help Member States to address skills gaps and equip their health workforce with personnel trained in cancer prevention, early detection, diagnosis, treatment, rehabilitation, and survivorship. Furthermore, as part of the “Partnership on Personalised Medicine” (due to be set up in 2023), Europe’s BCP will establish and support priorities for personalized-medicine-related research and education and provide guidance on the implementation of personalized medicine approaches into routine clinical care. Moreover, the “European Initiative to Understand Cancer” aims to increase understanding of cancer development in order to identify individuals at high risk of developing cancer and therefore better facilitate personalized approaches to cancer prevention and care. Finally, Europe’s BCP’s Comprehensive Cancer Centers, planned to be established by 2025, will also play a key role in the above recommendations; for instance, they will facilitate diagnostic- and treatment-related training, research, and clinical trials across the EU, helping to improve patients’ access to these aspects of personalized medicine [26].
- To advise on implementation of NHS long-term commitments and development of the NHS Genome Medicine Service.
- To facilitate an annual review of the National Genomic Test Directory.
- To enhance awareness and implementation of genomics across all clinical specialties.
- To drive improvements in personalized medicine.
- To advise on, review, and develop genomics guidance and testing specifications.
4.4. Reimbursement
- Standardized, comprehensive value assessment frameworks: Value assessment frameworks dedicated to advanced diagnostics must be standardized across Europe to incorporate the perspective of multiple stakeholders on a wider range of factors than is currently the case. These include: short- and long-term clinical utility as NGS moves away from the one-biomarker-one-therapy approach, patient preferences and utility, uncertainties in data, cost-effectiveness and cost-utility analyses that consider the full economic impact of all components of the intervention, ethical and legal implications, and equity-of-access and uptake requirements [88,89,90]. The frameworks should also enable use of innovative clinical trial designs and RWD-based studies, and take into account other patient perspectives, including quality of life, work productivity, caregiver and family burden, unmet need, burden of illness, and patient-reported outcome and experience measures. To more closely link reimbursement of the therapy and the diagnostic test, value assessment frameworks should incorporate the value of both [88]. Finally, value assessment frameworks should recognize the value of NGS in providing hereditary, diagnostic, prognostic, and treatment-response-related genomic information [91], or of using NGS for clinical trial allocation as a part of a patient’s current and future care plan.
- Value assessment framework-guided evidence generation: Standardization of fast and efficient reimbursement of advanced diagnostics across Europe, and consequent harmonization of access, depends on value assessment framework-guided evidence generation demonstrating the clinical utility of NGS through innovative clinical trial designs and RWD-based studies assessing the safety and efficacy of NGS-based MGTOs. Evidence generation also requires guidance from regulatory bodies to ensure that requirements are met. In this way, improved evidence generation should convince payers of the value of these technologies in informing clinical decision-making. For example, NGS can offer better turnaround times than simultaneous hotspot panel testing, and thus time-to-therapy initiation, so that initial high costs may be offset by overall longer-term cost-effectiveness and the improved survival benefits associated with precision oncology [75,92]. There is scope for strengthening the case through increased access to MGTOs and improved incremental benefit compared with current treatments [76].
4.5. Infrastructure
- Aligned methodologies across Europe: Harmonization of methodologies and analysis between laboratories at a national and European level will make it more feasible to tackle the huge volumes of data generated by NGS, which require significant infrastructure for secure data storage, analysis, and interpretation. European guidelines, analogous to those already in force in the US [93], are needed to promote alignment. Existing infrastructure should be leveraged with national genomics policy initiatives.
- Use of artificial intelligence: Infrastructure for the bioinformatics pipeline should incorporate artificial intelligence, which has already been demonstrated as improving NGS-based diagnostics in accuracy of variant identification [94], distinguishing between benign and pathogenic variants [95], user-friendliness of electronic health record systems [96], and linking multiple types of “omics” (e.g., genomics, transcriptomics, proteomics) and clinical data to better inform treatment decisions [97]. Artificial intelligence may also aid secure, harmonized data storage in line with privacy laws [98], and datasets to which artificial intelligence algorithms could be applied must be standardized and appropriately prepared.
- National infrastructural initiatives: Projects critical for patient diagnosis and management must be developed, for example with the goal of the UK’s Genomic Medicine Service (within the NHS) of sequencing 500,000 whole genomes by 2023/2024 [62]. Such initiatives depend on a national network of genomic laboratory hubs, with the relevant NGS provision, data, and informatics infrastructure. This is easier in countries with a formalized healthcare service (e.g., the UK, the Netherlands [99], and Estonia [100]) than in countries with insufficient funding/capacity (e.g., Italy and France) or a federal structure (e.g., Germany). All national initiatives should incorporate a common language between institutions/laboratories to facilitate data integration and sharing, as well as elements of data protection.
- Centralized, standardized centers for NGS excellence: Solutions must be found to fulfil the size, complexity, and cost of the information technology infrastructure required for NGS platforms, and the bioinformatic and molecular pathology expertise required to interpret NGS data. A promising option is to establish centralized, standardized centers of excellence in each country for MTBs and molecular oncology, capable of issuing standardized, clear recommendations that are backed by professional laboratories performing high-quality, end-to-end NGS services. Such a centralized approach may be particularly effective in small countries, such as Slovenia [101]. As the cost of NGS platforms decreases in the future and the technology becomes simpler to use [30,102,103], a decentralized model may become more appropriate, with several cancer centers carrying out sequencing and sending the results to a central bioinformatics core; such a model requires consistent guidelines to ensure standardization of preanalytical sequencing factors [104], and may increase turnaround time [105]. It is important to note, however, that the relative degree of (de)centralization appropriate will vary between countries, depending on specific factors (e.g., technology/personnel available).
- In-house and commercial NGS platforms: In-house platforms for NGS often offer reduced costs [106], increased availability of raw data and DNA/RNA left for extraction, and opportunities for training of medical oncologists and pathologists. Although in-house platforms may also demonstrate improved turnaround times compared to commercial tests [107,108], this is highly dependent on the number of samples for in-house platforms (turnaround time for commercial platforms, which may offer increased sample numbers, is consistent) [107]. Nonetheless, commercial tests may enable incorporation of recently approved biomarkers more easily [109], and in-house platforms, although feasible for the wet/bench portion of NGS, are more challenging for smaller institutions when considering bioinformatics, clinical informatics, and data analytics, versus commercial platforms [110]. Careful selection of the most appropriate NGS platform and expertise is critical, and depends on prior assessment of laboratory workflow, structure, and NGS requirements and aims [106,111]. Careful review of current laboratory personnel, with additional training if appropriate, is necessary to ensure availability of sufficient expertise [111]. Regulatory requirements are also key to ensure that patients are not at risk of misdiagnosis or of missing out on MGTOs. Whichever platform is chosen, it must comply with the incoming IVDR.
- Public–private partnerships: The involvement of local laboratories, public–private partnerships, and connectivity between local hospitals can assist broad knowledge generation and the translation of advanced diagnostics and innovative therapies in precision oncology to the community setting. Public–private partnerships lead to better communication and data sharing between the industry, genomic centers, and clinics [43].
4.6. Access and RWE
- Standardized, global, and centralized registries: As lack of data standardization can limit access to and interpretation of RWE across Europe, networks of standardized, global (or at least pan-European), and centralized multi-stakeholder registries collecting data related to diagnosis, treatments, and outcome, must be created. To permit data interoperability, pooling from disparate resources, and meaningfully comparable and reproducible results, all data variables collected should comply with the FAIR (Findable, Accessible, Interoperable and Reusable) Principles on data management [112] and be harmonized according to global data standards, terminologies, and models. Patient-reported outcomes will also be needed to enable adjustment for potential RWE bias. The federated network of the EHDEN [74] and the Cancer Inequalities Registry planned for 2021 as part of Europe’s BCP [26] offer potential platforms that could be usefully exploited. Such platforms may aid the governance-related infrastructural challenges detailed earlier.
- Consensus guidelines for RWE: Pan-European regulation could give payers, regulators, and industry clearer guidance on the feasibility and implementation of RWE in clinical practice. European consensus guidelines could help to standardize RWD-based studies by providing guidance on methodology, analysis, and reporting of NGS results, establishing best practices for data sharing in line with General Data Protection Regulation and for advising on assessment of clinical utility of NGS-based testing [54]. RWD-based studies respecting these guidelines should address payer and regulatory-body concerns regarding RWE, and thus aid their incorporation into value assessment frameworks. Current frameworks are lacking in useful examples of RWE to demonstrate value, and there is a clear need for expert panel discussions to provide these. Guidelines on how best to report RWD-based studies in medical journals are also needed, with few journals currently providing such recommendations [113].
5. Conclusions
- Evidence generation demonstrating the clinical value of NGS through traditional and innovative clinical trials and standardized, guideline-driven RWD-based studies.
- Converging approval of diagnostics and medicine to secure availability.
- A dynamic, collaborative, and multi-stakeholder national precision-medicine strategy that is aligned across Europe, with European collaboration.
- Up-to-date guidance specifying when and where NGS should be performed, along with minimum standards for testing. Updating the European Commission’s 2003 recommendations on cancer screening [114] to reflect the challenges detailed in this review is critical for providing proper guidance and establishing equitable early diagnosis through NGS.
- Integration and ongoing update of national guidelines covering NGS testing, in collaboration with scientific medical societies.
- Use of MTBs and clinical decision-supporting systems to align treatment strategies, based on profound expertise in cancer genomics.
- Increased awareness and education of stakeholders, including physicians, patients, and payers.
- Standardized value assessment frameworks that incorporate values from the perspective of multiple stakeholders.
- Sufficient and consistently adjusted budgeting for molecular testing, with significant investment in bioinformatics and artificial intelligence-based infrastructure for NGS data storage, analysis, and interpretation.
- Depending on the country-specific conditions, centralized, standardized centers with laboratories performing high-quality, in-house, and commercial NGS, or professional laboratories offering end-to-end services.
- Standardized, pan-European, and centralized multi-stakeholder registries collecting RWD related to diagnosis, treatments, and outcomes, with consensus guidelines at the European level to standardize RWD-based studies.
- Utilization of horizon-scanning techniques to identify future challenges related to innovative testing techniques, such as NGS, thus preparing stakeholders for their implementation into healthcare [115].
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Malone, E.R.; Oliva, M.; Sabatini, P.J.B.; Stockley, T.L.; Siu, L.L. Molecular profiling for precision cancer therapies. Genome Med. 2020, 12, 8. [Google Scholar] [CrossRef]
- Phillips, K.A.; Douglas, M.P.; Wordsworth, S.; Buchanan, J.; Marshall, D.A. Availability and funding of clinical genomic sequencing globally. BMJ Glob. Health 2021, 6, e004415. [Google Scholar] [CrossRef] [PubMed]
- Trédan, O.; Wang, Q.; Pissaloux, D.; Cassier, P.; de la Fouchardière, A.; Fayette, J.; Desseigne, F.; Ray-Coquard, I.; de la Fouchardière, C.; Frappaz, D.; et al. Molecular screening program to select molecular-based recommended therapies for metastatic cancer patients: Analysis from the ProfiLER trial. Ann. Oncol. 2019, 30, 757–765. [Google Scholar] [CrossRef] [PubMed]
- Massard, C.; Michiels, S.; Ferté, C.; Le Deley, M.-C.; Lacroix, L.; Hollebecque, A.; Verlingue, L.; Ileana, E.; Rosellini, S.; Ammari, S.; et al. High-throughput genomics and clinical outcome in hard-to-treat advanced cancers: Results of the MOSCATO 01 trial. Cancer Discov. 2017, 7, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Kamps, R.; Brandão, R.D.; van den Bosch, B.J.; Paulussen, A.D.C.; Xanthoulea, S.; Blok, M.J.; Romano, A. Next-generation sequencing in oncology: Genetic diagnosis, risk prediction and cancer classification. Int. J. Mol. Sci. 2017, 18, 308. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.P.; Shum, E.; Rajdev, L.; Cheng, H.; Goel, S.; Perez-Soler, R.; Halmos, B. Impact and diagnostic gaps of comprehensive genomic profiling in real-world clinical practice. Cancers 2020, 12, 1156. [Google Scholar] [CrossRef]
- US Food and Drug Administration (FDA). List of Cleared or Approved Companion Diagnostic Devices (In Vitro and Imaging Tools). Available online: https://www.fda.gov/medical-devices/in-vitro-diagnostics/list-cleared-or-approved-companion-diagnostic-devices-in-vitro-and-imaging-tools (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Non-Small Cell Lung Cancer. Version: 6.2021. Available online: https://www.nccn.org/professionals/physician_gls/PDF/nscl.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Prostate Cancer. Version: 1.2022. Available online: https://www.nccn.org/professionals/physician_gls/pdf/prostate.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Colon Cancer. Version: 3.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/colon.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Ovarian Cancer/Fallopian Tube Cancer/Primary Peritoneal Cancer. Version 3.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/ovarian.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Breast Cancer. Version: 8.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/breast.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Bone Cancer. Version: 1.2022. Available online: https://www.nccn.org/professionals/physician_gls/pdf/bone.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Occult Primary. Version: 1.2022. Available online: https://www.nccn.org/professionals/physician_gls/pdf/occult.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Vulvar Cancer. Version: 3.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/vulvar.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Uterine Neoplasms. Version: 4.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/uterine.pdf (accessed on 29 September 2021).
- National Comprehensive Cancer Network (NCCN). NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®): Cervical Cancer. Version: 1.2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/cervical.pdf (accessed on 29 September 2021).
- Mosele, F.; Remon, J.; Mateo, J.; Westphalen, C.; Barlesi, F.; Lolkema, M.; Normanno, N.; Scarpa, A.; Robson, M.; Meric-Bernstam, F.; et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: A report from the ESMO Precision Medicine Working Group. Ann. Oncol. 2020, 31, 1491–1505. [Google Scholar] [CrossRef]
- Sicklick, J.K.; Kato, S.; Okamura, R.; Schwaederle, M.; Hahn, M.E.; Williams, C.B.; De, P.; Krie, A.; Piccioni, D.E.; Miller, V.A.; et al. Molecular profiling of cancer patients enables personalized combination therapy: The I-PREDICT study. Nat. Med. 2019, 25, 744–750. [Google Scholar] [CrossRef]
- van der Velden, D.L.; Hoes, L.R.; van der Wijngaart, H.; van Berge Henegouwen, J.M.; van Werkhoven, E.; Roepman, P.; Schilsky, R.L.; de Leng, W.W.J.; Huitema, A.D.R.; Nuijen, B.; et al. The Drug Rediscovery protocol facilitates the expanded use of existing anticancer drugs. Nature 2019, 574, 127–131. [Google Scholar] [CrossRef]
- International Quality Network for Pathology (IQNPath), European Cancer Patient Coalition; European Federation of Pharmaceutical Industries and Associations (EFPIA). Unlocking the Potential of Precision Medicine in Europe: Improving Cancer Care through Broader Access to Quality Biomarker Testing. Policy Recommendations. February 2021. Available online: http://www.iqnpath.org/wp-content/uploads/2021/02/unlocking-the-potential-of-precision-medicine-in-europe.pdf (accessed on 29 September 2021).
- Leitsalu, L.; Haller, T.; Esko, T.; Tammesoo, M.-L.; Alavere, H.; Snieder, H.; Perola, M.; Ng, P.C.; Mägi, R.; Milani, L.; et al. Cohort profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int. J. Epidemiol. 2015, 44, 1137–1147. [Google Scholar] [CrossRef]
- Narayanasamy, S.; Markina, V.; Thorogood, A.; Blazkova, A.; Shabani, M.; Knoppers, B.M.; Prainsack, B.; Koesters, R. Genomic sequencing capacity, data retention, and personal access to raw data in Europe. Front. Genet. 2020, 11, 303. [Google Scholar] [CrossRef]
- Ayuso, C.; Millán, J.M.; Mancheño, M.; Dal-Ré, R. Informed consent for whole-genome sequencing studies in the clinical setting. Proposed recommendations on essential content and process. Eur. J. Hum. Genet. 2013, 21, 1054–1059. [Google Scholar] [CrossRef][Green Version]
- van de Haar, J.; Hoes, L.; Voest, E. Advancing molecular tumour boards: Highly needed to maximise the impact of precision medicine. ESMO Open 2019, 4, e000516. [Google Scholar] [CrossRef]
- European Commission. Communication from the Commission to the European Parliament and the Council: Europe’s Beating Cancer Plan. Available online: https://ec.europa.eu/health/sites/default/files/non_communicable_diseases/docs/eu_cancer-plan_en.pdf (accessed on 29 September 2021).
- Priestley, P.; Baber, J.; Lolkema, M.P.; Steeghs, N.; de Bruijn, E.; Shale, C.; Duyvesteyn, K.; Haidari, S.; van Hoeck, A.; Onstenk, W.; et al. Pan-cancer whole-genome analyses of metastatic solid tumours. Nature 2019, 575, 210–216. [Google Scholar] [CrossRef]
- Trein, P.; Wagner, J. Governing personalized health: A scoping review. Front. Genet. 2021, 12, 650504. [Google Scholar] [CrossRef]
- Schneider, F.; Maurer, C.; Friedberg, R.C. International Organization for Standardization (ISO) 15189. Ann. Lab. Med. 2017, 37, 365–370. [Google Scholar] [CrossRef]
- Adashek, J.J.; Subbiah, V.; Kurzrock, R. From tissue-agnostic to n-of-one therapies: (R)evolution of the precision paradigm. Trends Cancer 2021, 7, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Lavacchi, D.; Roviello, G.; D’Angelo, A. Tumor-agnostic treatment for cancer: When how is better than where. Clin. Drug Investig. 2020, 40, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Moore, K.; Colombo, N.; Scambia, G.; Kim, B.G.; Oaknin, A.; Friedlander, M.; Lisyanskaya, A.; Floquet, A.; Leary, A.; Sonke, G.S.; et al. Maintenance olaparib in patients with newly diagnosed advanced ovarian cancer. N. Engl. J. Med. 2018, 379, 2495–2505. [Google Scholar] [CrossRef] [PubMed]
- Robson, M.; Im, S.-A.; Senkus, E.; Xu, B.; Domchek, S.M.; Masuda, N.; Delaloge, S.; Li, W.; Tung, N.; Armstrong, A.; et al. Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N. Engl. J. Med. 2017, 377, 523–533. [Google Scholar] [CrossRef]
- Golan, T.; Hammel, P.; Reni, M.; Van Cutsem, E.; Macarulla, T.; Hall, M.J.; Park, J.-O.; Hochhauser, D.; Arnold, D.; Oh, D.-Y.; et al. Maintenance olaparib for germline BRCA-mutated metastatic pancreatic cancer. N. Engl. J. Med. 2019, 381, 317–327. [Google Scholar] [CrossRef] [PubMed]
- de Bono, J.; Mateo, J.; Fizazi, K.; Saad, F.; Shore, N.; Sandhu, S.; Chi, K.N.; Sartor, O.; Agarwal, N.; Olmos, D.; et al. Olaparib for metastatic castration-resistant prostate cancer. N. Engl. J. Med. 2020, 382, 2091–2102. [Google Scholar] [CrossRef] [PubMed]
- Robert, C.; Grob, J.J.; Stroyakovskiy, D.; Karaszewska, B.; Hauschild, A.; Levchenko, E.; Chiarion Sileni, V.; Schachter, J.; Garbe, C.; Bondarenko, I.; et al. Five-year outcomes with dabrafenib plus trametinib in metastatic melanoma. N. Engl. J. Med. 2019, 381, 626–636. [Google Scholar] [CrossRef] [PubMed]
- Odogwu, L.; Mathieu, L.; Blumenthal, G.; Larkins, E.; Goldberg, K.B.; Griffin, N.; Bijwaard, K.; Lee, E.Y.; Philip, R.; Jiang, X.; et al. FDA approval summary: Dabrafenib and trametinib for the treatment of metastatic non-small cell lung cancers harboring BRAF V600E mutations. Oncologist 2018, 23, 740–745. [Google Scholar] [CrossRef] [PubMed]
- McDonald, C.J.; Huff, S.M.; Suico, J.G.; Hill, G.; Leavelle, D.; Aller, R.; Forrey, A.; Mercer, K.; DeMoor, G.; Hook, J.; et al. LOINC, a universal standard for identifying laboratory observations: A 5-year update. Clin. Chem. 2003, 49, 624–633. [Google Scholar] [CrossRef] [PubMed]
- Spanish Society of Pathological Anatomy—International Academy of Pathology. Who We Are. Available online: https://www.seap.es/quienes-somos (accessed on 29 September 2021).
- Horgan, D.; Ciliberto, G.; Conte, P.; Baldwin, D.; Seijo, L.; Montuenga, L.M.; Paz-Ares, L.; Garassino, M.; Penault-Llorca, F.; Galli, F.; et al. Bringing greater accuracy to europe’s healthcare systems: The unexploited potential of biomarker testing in oncology. Biomed. Hub. 2020, 5, 182–223. [Google Scholar] [CrossRef]
- European Cancer Patient Coalition (ECPC); European Alliance for Personalised Medicine (EAPM). Patient Awareness on Biomarkers across Europe. Available online: https://ecpc.org/wp-content/uploads/2019/08/ECPC-patient-awareness-on-biomarkers-across-europe-2.pdf (accessed on 29 September 2021).
- Monkhorst, K.; Samsom, K.; Schipper, L.; Roepman, P.; Bosch, L.; de Bruijn, E.; Hoes, L.R.; Riethorst, I.; Schoenmaker, L.; van der Kolk, L.; et al. Validation of whole genome sequencing in routine clinical practice. Ann. Oncol. 2020, 31, S784. [Google Scholar] [CrossRef]
- European Confederation of Pharmaceutical Entrepreneurs AISBL (EUCOPE). Developing an Advanced Diagnostics Ecosystem in Europe: A Proposal for Change. Available online: https://www.eucope.org/wp-content/uploads/2021/03/eucope-advanced-diagnostics-white-paper.pdf (accessed on 29 September 2021).
- Parkinson, D.R.; Johnson, B.E.; Sledge, G.W. Making personalized cancer medicine a reality: Challenges and opportunities in the development of biomarkers and companion diagnostics. Clin. Cancer Res. 2012, 18, 619–624. [Google Scholar] [CrossRef]
- Office of Health Economics (OHE); European Personalised Medicine Association (EPEMED). The Value of Knowing and Knowing the Value: Improving the Health Technology Assessment of Complementary Diagnostics. Available online: https://www.ohe.org/sites/default/files/WP_EpemedOHE_final.pdf (accessed on 29 September 2021).
- The London School of Economics and Political Science; Gill, J.; Fontrier, A.-M.; Miracolo, A.; Kanavos, P. Access to Personalised Oncology in Europe. Available online: https://www.efpia.eu/media/580518/access-to-personalised-oncology-in-europe.pdf (accessed on 29 September 2021).
- Govaerts, L.; Simoens, S.; Van Dyck, W.; Huys, I. Shedding light on reimbursement policies of companion diagnostics in European countries. Value Health 2020, 23, 606–615. [Google Scholar] [CrossRef]
- Janiaud, P.; Serghiou, S.; Ioannidis, J.P.A. New clinical trial designs in the era of precision medicine: An overview of definitions, strengths, weaknesses, and current use in oncology. Cancer Treat. Rev. 2019, 73, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Tsimberidou, A.M.; Müller, P.; Ji, Y. Innovative trial design in precision oncology. Semin. Cancer Biol. 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Dienstmann, R.; Garralda, E.; Aguilar, S.; Sala, G.; Viaplana, C.; Ruiz-Pace, F.; González-Zorelle, J.; LoGiacco, D.G.; Ogbah, Z.; Masdeu, L.R.; et al. Evolving landscape of molecular prescreening strategies for oncology early clinical trials. JCO Precis. Oncol. 2020, 4, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Kennedy-Martin, T.; Curtis, S.; Faries, D.; Robinson, S.; Johnston, J. A literature review on the representativeness of randomized controlled trial samples and implications for the external validity of trial results. Trials 2015, 16, 495. [Google Scholar] [CrossRef] [PubMed]
- Dickson, D.; Johnson, J.; Bergan, R.; Owens, R.; Subbiah, V.; Kurzrock, R. The master observational trial: A new class of master protocol to advance precision medicine. Cell 2020, 180, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Cave, A.; Kurz, X.; Arlett, P. Real-world data for regulatory decision making: Challenges and possible solutions for Europe. Clin. Pharmacol. Ther. 2019, 106, 36–39. [Google Scholar] [CrossRef]
- Deverka, P.A.; Douglas, M.P.; Phillips, K.A. Use of real-world evidence in us payer coverage decision-making for next-generation sequencing-based tests: Challenges, opportunities, and potential solutions. Value Health 2020, 23, 540–550. [Google Scholar] [CrossRef]
- Feinberg, B.A.; Gajra, A.; Zettler, M.E.; Phillips, T.D.; Phillips, E.G., Jr.; Kish, J.K. Use of real-world evidence to support FDA approval of oncology drugs. Value Health 2020, 23, 1358–1365. [Google Scholar] [CrossRef]
- Tarricone, R.; Rognoni, C. What can health systems learn from COVID-19? Eur. Heart J. Suppl. 2020, 22, P4–P7. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Pruneri, G.; De Braud, F.; Sapino, A.; Aglietta, M.; Vecchione, A.; Giusti, R.; Marchiò, C.; Scarpino, S.; Baggi, A.; Bonetti, G.; et al. Next-generation sequencing in clinical practice: Is it a cost-saving alternative to a single-gene testing approach? Pharmacoecon. Open 2021, 5, 285–298. [Google Scholar] [CrossRef]
- Yu, T.M.; Morrison, C.; Gold, E.J.; Tradonsky, A.; Arnold, R.J.G. Budget impact of next-generation sequencing for molecular assessment of advanced non-small cell lung cancer. Value Health 2018, 21, 1278–1285. [Google Scholar] [CrossRef]
- Signorovitch, J.; Zhou, Z.; Ryan, J.; Anhorn, R.; Chawla, A. Budget impact analysis of comprehensive genomic profiling in patients with advanced non-small cell lung cancer. J. Med. Econ. 2019, 22, 140–150. [Google Scholar] [CrossRef] [PubMed]
- Anhorn, R.; Roberts, G.; Skovhus, M.; Khorshid, M. Impact of comprehensive genomic profiling of patients with first line non-small cell lung cancer in the UK. Value Health 2017, 20, PA575. [Google Scholar] [CrossRef]
- Stark, Z.; Dolman, L.; Manolio, T.A.; Ozenberger, B.; Hill, S.L.; Caulfied, M.J.; Levy, Y.; Glazer, D.; Wilson, J.; Lawler, M.; et al. Integrating genomics into healthcare: A global responsibility. Am. J. Hum. Genet. 2019, 104, 13–20. [Google Scholar] [PubMed]
- Pinto, C.; Biffoni, M.; Popoli, P.; Marchetti, A.; Marchetti, P.; Martini, N.; Normanno, N. Molecular tests and target therapies in oncology: Recommendations from the Italian workshop. Future Oncol. 2021, 17, 3529–3539. [Google Scholar] [CrossRef]
- Edsjö, A.; Friedman, M.; Rosenquist, R. Genomic Medicine Sweden—A national initiative for the broad introduction of precision medicine in Swedish healthcare [article in Swedish]. Lakartidningen 2021, 118, 21023. [Google Scholar]
- NHS England. Genomic Laboratory Hubs. Available online: https://www.england.nhs.uk/genomics/genomic-laboratory-hubs/ (accessed on 29 September 2021).
- Helsebiblioteket. National Action Program with Guidelines for Diagnosis, Treatment and Follow-up of Lung Cancer, Mesothelioma and Thymoma. 4 January 2021. Available online: https://www.helsebiblioteket.no/retningslinjer/lungekreft/forord (accessed on 29 September 2021).
- Garrido, P.; Conde, E.; de Castro, J.; Gómez-Román, J.J.; Felip, E.; Pijuan, L.; Isla, D.; Sanz, J.; Paz-Ares, L.; López-Ríos, F. Updated guidelines for predictive biomarker testing in advanced non-small-cell lung cancer: A National Consensus of the Spanish Society of Pathology and the Spanish Society of Medical Oncology. Clin. Transl. Oncol. 2020, 22, 989–1003. [Google Scholar] [CrossRef]
- Colomer, R.; Aranda-López, I.; Albanell, J.; García-Caballero, T.; Ciruelos, E.; López-García, M.; Cortés, J.; Rojo, F.; Martín, M.; Palacios-Calvo, J. Biomarkers in breast cancer: A consensus statement by the Spanish Society of Medical Oncology and the Spanish Society of Pathology. Clin. Transl. Oncol. 2018, 20, 815–826. [Google Scholar] [CrossRef]
- Arbeitsgemeinschaft Gynäkologische Onkologie (AGO). Breast Cancer Guidelines 2021: Diagnosis and Treatment of Patients with Early and Advanced Breast Cancer. Available online: https://www.ago-online.de/fileadmin/ago-online/downloads/_leitlinien/kommission_mamma/2021/englisch/_ohne_Literartur/2021E_04_Pathology_MASTER_final_20210301.pdf (accessed on 29 September 2021).
- Kunskapsbanken. Quality Appendix for Breast Pathology (KVAST Appendix). Available online: https://kunskapsbanken.cancercentrum.se/diagnoser/brostcancer/vardprogram/kvalitetsdokument-for--patologi/#chapter--Forord (accessed on 29 September 2021).
- European Commission. European Health Union: Protecting the Health of Europeans and Collectively Responding to Cross-Border Health Crises. Available online: https://ec.europa.eu/info/strategy/priorities-2019-2024/promoting-our-european-way-life/european-health-union_en (accessed on 29 September 2021).
- Berns, A.; Ringborg, U.; Celis, J.E.; Heitor, M.; Aaronson, N.K.; Abou-Zeid, N.; Adami, H.O.; Apostolidis, K.; Baumann, M.; Bardelli, A.; et al. Towards a cancer mission in Horizon Europe: Recommendations. Mol. Oncol. 2020, 14, 1589–1615. [Google Scholar] [CrossRef]
- Lubbers, B.R.; Schilhabel, A.; Cobbaert, C.M.; Gonzalez, D.; Dombrink, I.; Brüggemann, M.; Bitter, W.M.; van Dongen, J.J.M. The new EU regulation on in vitro diagnostic medical devices: Implications and preparatory actions for diagnostic laboratories. Hemasphere 2021, 5, e568. [Google Scholar] [CrossRef]
- European Health Data & Evidence Network (EHDEN). Vision, Mission & Objectives. Available online: https://www.ehden.eu/vision-and-mission/ (accessed on 29 September 2021).
- Chawla, A.; Janku, F.; Wheler, J.J.; Miller, V.A.; Ryan, J.; Anhorn, R.; Zhou, Z.; Signorovitch, J. Estimated cost of anticancer therapy directed by comprehensive genomic profiling in a single-center study. JCO Precis. Oncol. 2018, 2, 1–11. [Google Scholar] [CrossRef]
- Steuten, L.; Goulart, B.; Meropol, N.J.; Pritchard, D.; Ramsey, S.D. Cost effectiveness of multigene panel sequencing for patients with advanced non–small-cell lung cancer. JCO Clin. Cancer Inform. 2019, 3, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Tan, O.; Shrestha, R.; Cunich, M.; Schofield, D.J. Application of next-generation sequencing to improve cancer management: A review of the clinical effectiveness and cost-effectiveness. Clin. Genet. 2018, 93, 533–544. [Google Scholar] [CrossRef] [PubMed]
- Mateo, J.; Chakravarty, D.; Dienstmann, R.; Jezdic, S.; Gonzalez-Perez, A.; Lopez-Bigas, N.; Ng, C.K.Y.; Bedard, P.L.; Tortora, G.; Douillard, J.Y.; et al. A framework to rank genomic alterations as targets for cancer precision medicine: The ESMO Scale for Clinical Actionability of molecular Targets (ESCAT). Ann. Oncol. 2018, 29, 1895–1902. [Google Scholar] [CrossRef] [PubMed]
- Condorelli, R.; Mosele, F.; Verret, B.; Bachelot, T.; Bedard, P.L.; Cortes, J.; Hyman, D.M.; Juric, D.; Krop, I.; Bieche, I.; et al. Genomic alterations in breast cancer: Level of evidence for actionability according to ESMO Scale for Clinical Actionability of molecular Targets (ESCAT). Ann. Oncol. 2019, 30, 365–373. [Google Scholar] [CrossRef] [PubMed]
- Barwell, J.; Snape, K.; Wedderburn, S. The new genomic medicine service and implications for patients. Clin. Med. 2019, 19, 273–277. [Google Scholar]
- Kim, J.A.; Ceccarelli, R.; Lu, C.Y. Pharmacogenomic biomarkers in US FDA-approved drug labels (2000–2020). J. Pers. Med. 2021, 11, 179. [Google Scholar] [CrossRef] [PubMed]
- Chakravarty, D.; Gao, J.; Phillips, S.M.; Kundra, R.; Zhang, H.; Wang, J.; Rudolph, J.E.; Yaeger, R.; Soumerai, T.; Nissan, M.H.; et al. OncoKB: A precision oncology knowledge base. JCO Precis. Oncol. 2017, 2017, PO.17.00011. [Google Scholar] [CrossRef] [PubMed]
- Ritzhaupt, A.; Hayes, I.; Ehmann, F. Implementing the EU in vitro diagnostic regulation—A European regulatory perspective on companion diagnostics. Expert Rev. Mol. Diagn. 2020, 20, 565–567. [Google Scholar] [CrossRef]
- Genomics Education Programme, Health Education England (HEE). Genomics Education Programme: Preparing the Healthcare Workforce to Harness the Power of Genomic Medicine. Available online: https://www.genomicseducation.hee.nhs.uk/about-us/ (accessed on 29 September 2021).
- NHS England. Genomics. Available online: https://www.england.nhs.uk/commissioning/spec-services/npc-crg/e01-genomics/ (accessed on 29 September 2021).
- NHS England. National Cancer Drugs Fund List (Including List of NICE Approved and Baseline Funded Drugs/Indications from 1st April 2016 with Criteria for Use). ver1.172. 25-Nov-20. Available online: https://www.england.nhs.uk/wp-content/uploads/2017/04/national-cdf-list-v1.172.pdf (accessed on 29 September 2021).
- Assessment in Medicine (AiM). Reimbursement of Medical Devices in Germany. Available online: https://www.iges.com/sites/igesgroup/iges.de/myzms/content/e6/e1621/e10211/e22175/e22650/e22652/e22723/AiM_Reimbursement_of_Medical_Devices_in_Germany_2020_21_ger.pdf (accessed on 29 September 2021).
- Faulkner, E.; Holtorf, A.-P.; Walton, S.; Liu, C.Y.; Lin, H.; Biltaj, E.; Brixner, D.; Barr, C.; Oberg, J.; Shandhu, G.; et al. Being precise about precision medicine: What should value frameworks incorporate to address precision medicine? A report of the personalized precision medicine special interest group. Value Health 2020, 23, 529–539. [Google Scholar] [CrossRef]
- Mandelblatt, J.S.; Ramsey, S.D.; Lieu, T.A.; Phelps, C.E. Evaluating frameworks that provide value measures for health care interventions. Value Health 2017, 20, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Wurcel, V.; Cicchetti, A.; Garrison, L.; Kip, M.M.A.; Koffijberg, H.; Kolbe, A.; Leeflang, M.M.G.; Merlin, T.; Mestre-Ferrandiz, J.; Oortwijn, W.; et al. The value of diagnostic information in personalised healthcare: A comprehensive concept to facilitate bringing this technology into healthcare systems. Public Health Genom. 2019, 22, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Regier, D.A.; Weymann, D.; Buchanan, J.; Marshall, D.A.; Wordsworth, S. Valuation of health and nonhealth outcomes from next-generation sequencing: Approaches, challenges, and solutions. Value Health 2018, 21, 1043–1047. [Google Scholar] [CrossRef] [PubMed]
- Pennell, N.A.; Mutebi, A.; Zhou, Z.-Y.; Ricculli, M.L.; Tang, W.; Wang, H.; Guerin, A.; Arnhart, T.; Dalal, A.; Sasane, M.; et al. Economic impact of next-generation sequencing versus single-gene testing to detect genomic alterations in metastatic non–small-cell lung cancer using a decision analytic model. JCO Precis. Oncol. 2019, 3, 1–9. [Google Scholar] [CrossRef]
- Roy, S.; Coldren, C.; Karunamurthy, A.; Kip, N.S.; Klee, E.W.; Lincoln, S.E.; Leon, A.; Pullambhatla, M.; Temple-Smolkin, R.L.; Voelkerding, K.V.; et al. Standards and guidelines for validating next-generation sequencing bioinformatics pipelines: A joint recommendation of the Association for Molecular Pathology and the College of American Pathologists. J. Mol. Diagn. 2018, 20, 4–27. [Google Scholar] [CrossRef]
- Poplin, R.; Chang, P.-C.; Alexander, D.; Schwartz, S.; Colthurst, T.; Ku, A.; Newburger, D.; Dijamco, J.; Nguyen, N.; Afshar, P.T.; et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 2018, 36, 983–987. [Google Scholar] [CrossRef]
- Sundaram, L.; Gao, H.; Padigepati, S.R.; McRae, J.F.; Li, Y.; Kosmicki, J.A.; Fritzilas, N.; Hakenberg, J.; Dutta, A.; Shon, J.; et al. Predicting the clinical impact of human mutation with deep neural networks. Nat. Genet. 2018, 50, 1161–1170. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhu, L.; Roberts, R.; Tong, W. Toward clinical implementation of next-generation sequencing-based genetic testing in rare diseases: Where are we? Trends Genet. 2019, 35, 852–867. [Google Scholar] [CrossRef]
- Zhang, L.; Lv, C.; Jin, Y.; Cheng, G.; Fu, Y.; Yuan, D.; Tao, Y.; Guo, Y.; Ni, X.; Shi, T. Deep learning-based multi-omics data integration reveals two prognostic subtypes in high-risk neuroblastoma. Front. Genet. 2018, 9, 477. [Google Scholar] [CrossRef]
- Langmead, B.; Nellore, A. Cloud computing for genomic data analysis and collaboration. Nat. Rev. Genet. 2018, 19, 208–219. [Google Scholar] [CrossRef] [PubMed]
- Boomsma, D.I.; Wijmenga, C.; Slagboom, E.P.; Swertz, M.A.; Karssen, L.C.; Abdellaoui, A.; Ye, K.; Guryev, V.; Vermaat, M.; van Dijk, F.; et al. The Genome of the Netherlands: Design, and project goals. Eur. J. Hum. Genet. 2014, 22, 221–227. [Google Scholar] [CrossRef]
- Metspalu, A. Estonian Genome Project—before the take-off and take-off. Bioinformatics 2002, 18, S152. [Google Scholar] [CrossRef] [PubMed]
- Zwitter, M.; Čufer, T.; Vrankar, M.; Kern, I.; Štupnik, T.; Rozman, A.; Koprivnikar, H.; Zadnik, V. Lung cancer in Slovenia. J. Thorac. Oncol. 2019, 14, 1327–1331. [Google Scholar] [CrossRef] [PubMed]
- Colomer, R.; Mondejar, R.; Romero-Laorden, N.; Alfranca, A.; Sanchez-Madrid, F.; Quintela-Fandino, M. When should we order a next generation sequencing test in a patient with cancer? EClinicalMedicine 2020, 25, 100487. [Google Scholar] [CrossRef] [PubMed]
- Denny, J.C.; Collins, F.S. Precision medicine in 2030-seven ways to transform healthcare. Cell 2021, 184, 1415–1419. [Google Scholar] [CrossRef]
- Hynes, S.O.; Pang, B.; James, J.A.; Maxwell, P.; Salto-Tellez, M. Tissue-based next generation sequencing: Application in a universal healthcare system. Br. J. Cancer 2017, 116, 553–560. [Google Scholar] [CrossRef]
- Beukers, A.G.; Jenkins, F.; van Hal, S.J. Centralised or localised pathogen whole genome sequencing: Lessons learnt from implementation in a clinical diagnostic laboratory. Front. Cell. Infect. Microbiol. 2021, 11, 636290. [Google Scholar] [CrossRef]
- Johnston, K.M.; Sheffield, B.S.; Yip, S.; Lakzadeh, P.; Qian, C.; Nam, J. Costs of in-house genomic profiling and implications for economic evaluation: A case example of non-small cell lung cancer (NSCLC). J. Med. Econ. 2020, 23, 1123–1129. [Google Scholar] [CrossRef]
- Heeke, S.; Benzaquen, J.; Long-Mira, E.; Audelan, B.; Lespinet, V.; Bordone, O.; Lalvée, S.; Zahaf, K.; Poudenx, M.; Humbert, O.; et al. In-house implementation of tumor mutational burden testing to predict durable clinical benefit in non-small cell lung cancer and melanoma patients. Cancers 2019, 11, 1271. [Google Scholar] [CrossRef]
- Sadri, N.; Miller, T.E.; Yang, M.; Dowlati, A.; Friedman, J.D.; Bajor, D.L.; Chang, R.Y.; Willis, J.E. Clinical utility of reflex testing using focused next generation sequencing for management of patients with advanced lung adenocarcinoma. J. Clin. Oncol. 2018, 36, e24199. [Google Scholar] [CrossRef]
- Gondos, A.; Paz-Ares, L.G.; Saldana, D.; Thomas, M.; Mascaux, C.; Bubendorf, L.; Barlesi, F. Genomic testing among patients (pts) with newly diagnosed advanced non-small cell lung cancer (aNSCLC) in the United States: A contemporary clinical practice patterns study. J. Clin. Oncol. 2020, 38, 9592. [Google Scholar] [CrossRef]
- Gullapalli, R.R. Evaluation of commercial next-generation sequencing bioinformatics software solutions. J. Mol. Diagn. 2020, 22, 147–158. [Google Scholar] [CrossRef]
- Akkari, Y.; Smith, T.; Westfall, J.; Lupo, S. Implementation of cancer next-generation sequencing testing in a community hospital. Cold Spring Harb. Mol. Case Stud. 2019, 5, a003707. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, M.D.; Dumontier, M.; Aalbersberg, I.J.; Appleton, G.; Axton, M.; Baak, A.; Blomberg, N.; Boiten, J.W.; da Silva Santos, L.B.; Bourne, P.E.; et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci. Data 2016, 3, 160018. [Google Scholar] [CrossRef] [PubMed]
- Oehrlein, E.M.; Graff, J.S.; Perfetto, E.M.; Mullins, C.D.; Dubois, R.W.; Anyanwu, C.; Onukwugha, E. Peer-reviewed journal editors’ views on real-world evidence. Int. J. Technol. Assess. Health Care 2018, 34, 111–119. [Google Scholar] [CrossRef]
- Official Journal of the European Union. Council Recommendation of 2 December 2003 on Cancer Screening. Available online: https://ec.europa.eu/jrc/sites/default/files/2_December_2003%20cancer%20screening.pdf (accessed on 29 September 2021).
- Hines, P.; Hiu Yu, L.; Guy, R.H.; Brand, A.; Papaluca-Amati, M. Scanning the horizon: A systematic literature review of methodologies. BMJ Open 2019, 9, e026764. [Google Scholar] [CrossRef] [PubMed]
| Country | MTB Usage | Recommendations |
|---|---|---|
| Belgium | Most large centers have MTBs, and precision projects to initiate national MTBs have been started, which will enable the exchange of expertise. In general, MTBs tend to be convened on an ad hoc basis and attendance is irregular, e.g., oncologists may not always attend |
|
| Bulgaria | The decision to conduct biomarker screening is made by interdisciplinary MTBs | |
| France | MTBs are required and mandatory for decision-making concerning all oncology patients (public and private hospitals) and take place in a health facility, a group of health establishments, an oncology network, or as part of coordination centers in oncology (3C). The organizational procedures for MTBs are defined by article D. 6124-131 of the Public Health Code. MTBs must be carried out in the presence of at least three physicians from different specialties working with patients, providing a relevant opinion on all the procedures envisaged. The doctor then informs the patient and gives a personalized program of care. For complicated cases, local, regional, or national molecular MTBs are available and can connect to early-phase clinical trials. Fifteen “national reference networks for rare adult cancers” have been labeled by the French National Cancer Institute and provide national MTBs |
|
| Germany | Decision for large-scale NGS is dependent upon MTBs, which are integrated at most, if not all, university hospitals. Hospitals in Germany run weekly MTBs on a national and international scale (600–1000 patients per year, per site). MTBs typically have a structure for reporting the patient-decision process and requirements for data storage. The German government has encouraged health insurance companies to provide significant funding for translational projects at universities, including the roll out of MTBs, which means there is a network of working groups as well as a standardization of reporting and quality control. At least 100,000 patients in Germany qualify for CGP every year and the current system is not scalable, as due to sectoral limitations in patient care, most tumor samples and cases are not transferred into centralized genome centers for in-depth molecular analysis and MTB-based therapy evaluation |
|
| Israel | MTBs are not overburdened and are conducted in an efficient manner, with a broad spectrum of expertise. MTBs are weekly/bi-weekly 2 h sessions that cover every tumor type; there has been an MTB in Israel (Hadassah) running for more than 8 years, and it has become partially virtual to allow sharing of expertise between local hospitals. There are no strict rules on the required composition of an MTB, although they must include pathologists, oncologists, geneticists, and radiotherapists. Referral to MTBs is increased when discussions are virtual; recommendations are provided to physicians to refer the patient to a local MTB (the process is not regulated). Some cases may then be sent to Foundation Medicine, Inc., for further discussion |
|
| Italy | Italy is currently working to create a national MTB in collaboration with regional MTBs. A national agency, in charge of evaluating the efficiency of regional healthcare systems, is attempting to provide national guidelines for regional MTBs. Some regions (such as Regione Veneto) have also provided individual guidance for MTBs (e.g., structure, criteria to access molecular profiling, access to off-label therapies, reimbursement procedures). However, not all hospitals are able to incorporate the required expertise, and these hospitals should defer to centralized MTBs to discuss complex patient cases. MTBs in Italy use the ESCAT and OncoKB scales (for less frequently reported genetic variants) to guide treatment decisions; only targeted therapies with a specific level of evidence (Level 1 or 1A) can be recommended. MTBs have a strict procedure for obtaining access in order to limit patient numbers, with access only recommended by multidisciplinary boards for particular tumor types (and not by physicians or patients). Some precision medicine trials are ongoing in the country |
|
| The Netherlands | Determining eligibility for MTBs is performed on a case-by-case basis | |
| Poland | Typically, three teams are involved in MTBs, with representatives from radiology, pathology, and clinical teams (the clinical team is often the most important team, as they provide an overview of all patient information). Clinically, there are low numbers of molecular diagnostic and genetic experts in Poland, challenging the implementation of MTBs | |
| Portugal | The Portuguese Oncology Institute of Porto was the first to implement an MTB in Portugal; the MTB typically includes medical oncologists, pathologists, and molecular geneticists/pathologists, in addition to the treating physician | |
| Republic of Ireland | Although use of MTBs is variable (Republic of Ireland does not currently have a high number of patients included in MTBs), they enable international collaboration and are useful in rare cancer types. Some clinical colleagues use international (e.g., co-operative group) MTBs, most commonly as part of research or in clinical trials, and there are initiatives to facilitate organization of MTBs |
|
| Slovenia | MTBs are established in large cancer centers. Molecular testing is performed based on ESCAT guidelines. All additional indications for NGS testing are made by MTBs | |
| Spain | No national MTBs, only regional, and these mainly occur within large hospitals that use NGS | |
| Sweden | Diagnostics and the treatment of cancer are driven by firm clinical guidelines, which involve MTBs and the treating clinicians | |
| United Kingdom | MTBs are uncommon but will evolve as further NGS data become available, especially with the roll-out of genomic hubs by NHS England |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Horgan, D.; Curigliano, G.; Rieß, O.; Hofman, P.; Büttner, R.; Conte, P.; Cufer, T.; Gallagher, W.M.; Georges, N.; Kerr, K.; et al. Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe. J. Pers. Med. 2022, 12, 72. https://doi.org/10.3390/jpm12010072
Horgan D, Curigliano G, Rieß O, Hofman P, Büttner R, Conte P, Cufer T, Gallagher WM, Georges N, Kerr K, et al. Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe. Journal of Personalized Medicine. 2022; 12(1):72. https://doi.org/10.3390/jpm12010072
Chicago/Turabian StyleHorgan, Denis, Giuseppe Curigliano, Olaf Rieß, Paul Hofman, Reinhard Büttner, Pierfranco Conte, Tanja Cufer, William M. Gallagher, Nadia Georges, Keith Kerr, and et al. 2022. "Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe" Journal of Personalized Medicine 12, no. 1: 72. https://doi.org/10.3390/jpm12010072
APA StyleHorgan, D., Curigliano, G., Rieß, O., Hofman, P., Büttner, R., Conte, P., Cufer, T., Gallagher, W. M., Georges, N., Kerr, K., Penault-Llorca, F., Mastris, K., Pinto, C., Van Meerbeeck, J., Munzone, E., Thomas, M., Ujupan, S., Vainer, G. W., Velthaus, J.-L., & André, F. (2022). Identifying the Steps Required to Effectively Implement Next-Generation Sequencing in Oncology at a National Level in Europe. Journal of Personalized Medicine, 12(1), 72. https://doi.org/10.3390/jpm12010072









