Validation of an NGS Panel Designed for Detection of Actionable Mutations in Tumors Common in Latin America
Abstract
:1. Introduction
2. Materials and Methods
2.1. Panel Design
2.2. Sample Information
2.3. Control Samples
2.4. DNA Extraction, Quantification, and Sample Quality Control
2.5. Library Preparation
2.6. Target Enrichment
2.7. Sequencing Run Set-Up
2.8. Bioinformatic Analyses
2.9. Variant Filtering and Sequence Quality Reporting
2.10. Germline Variant Calling
3. Results
3.1. Panel Design and Sequencing Metrics
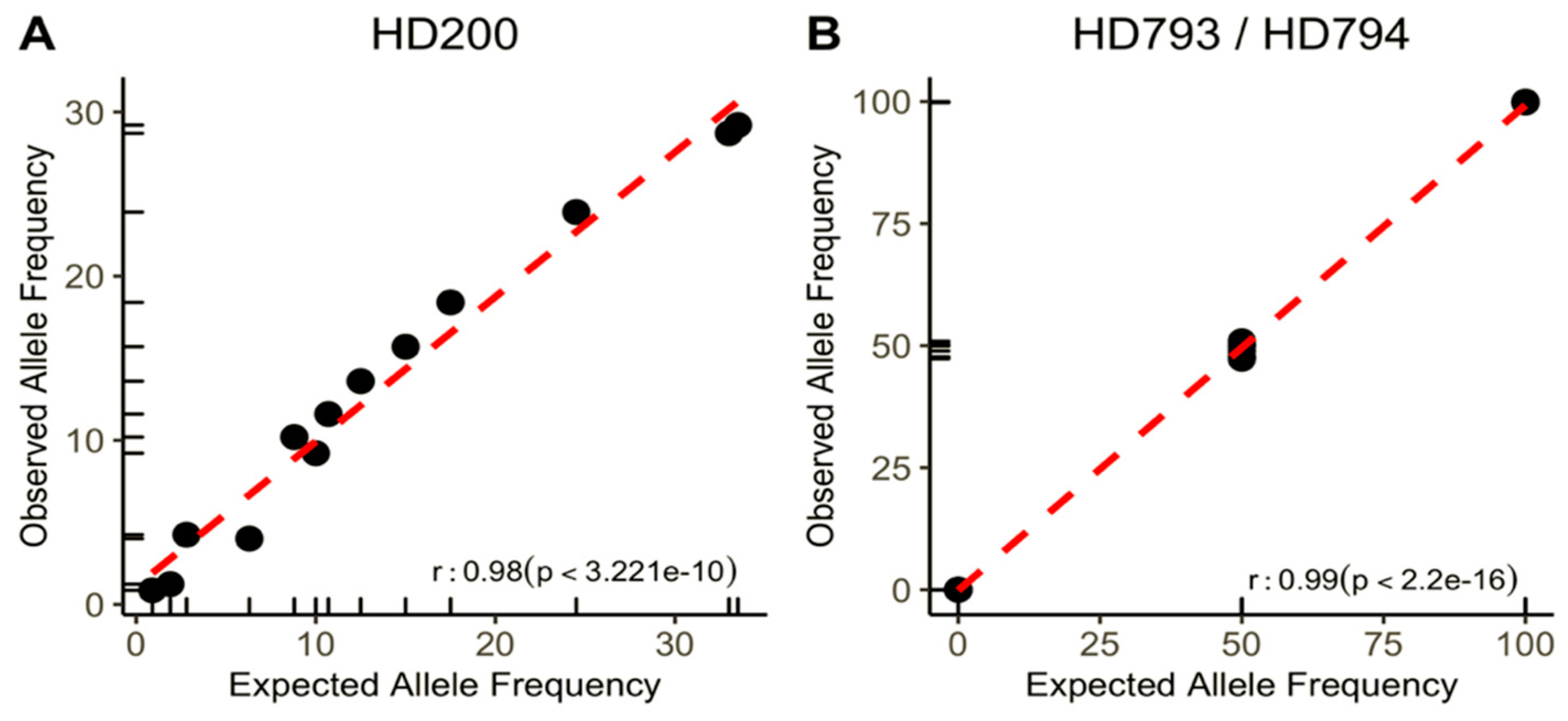
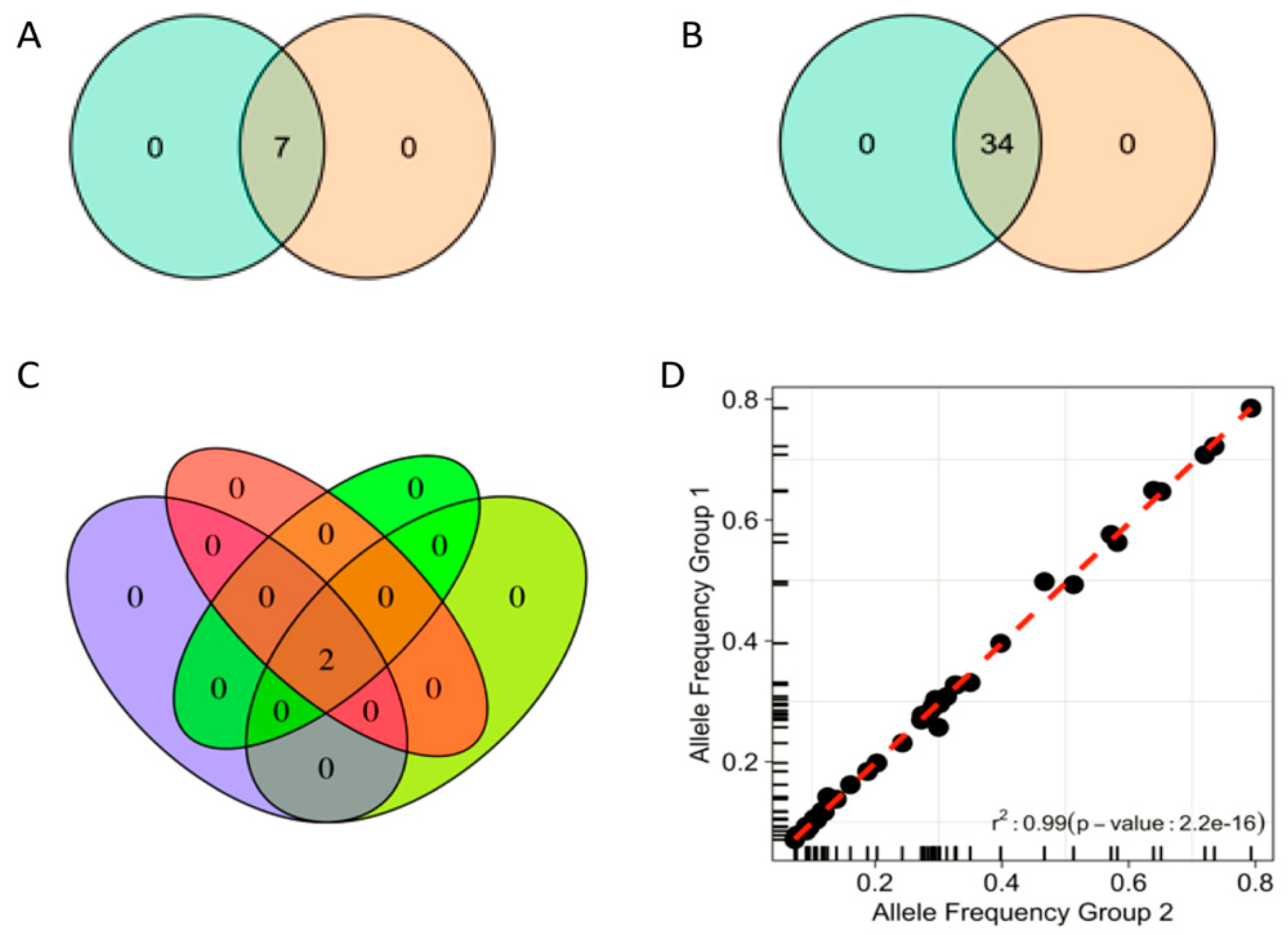
3.2. Panel Performance
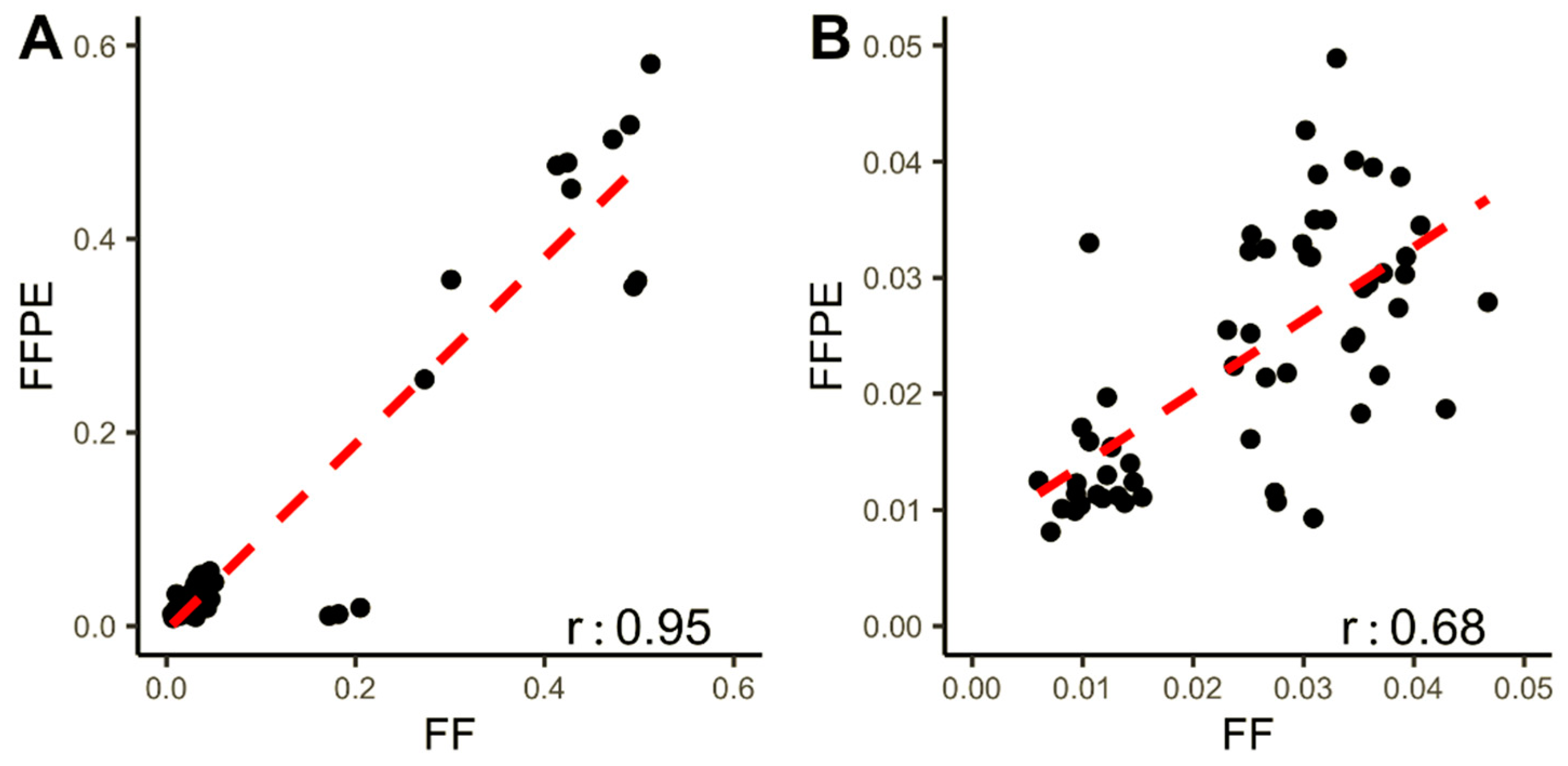
3.3. Comparison between FFPE, Fresh Frozen, and Blood gDNA
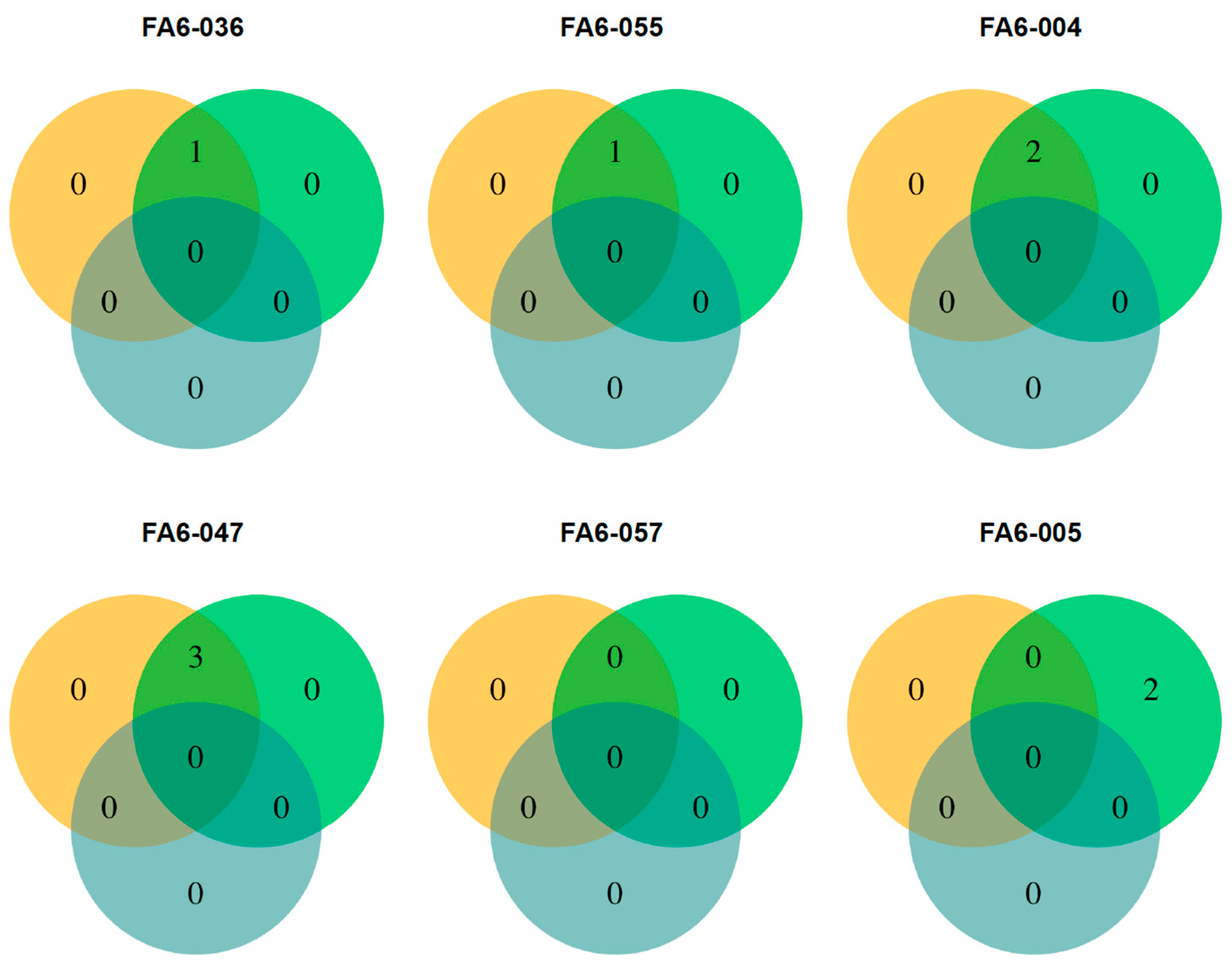
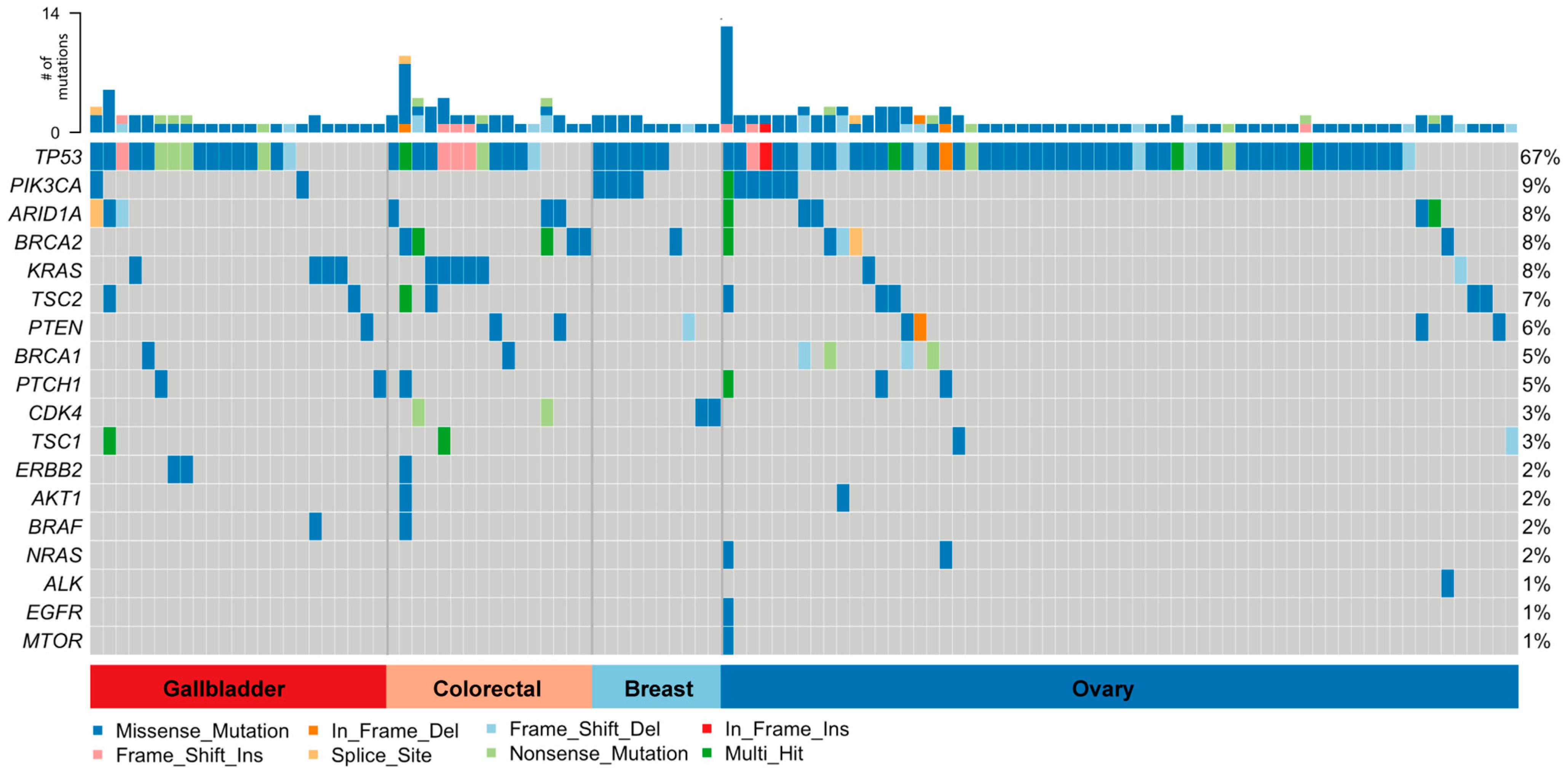
3.4. Validation of the Assay in Clinical Samples
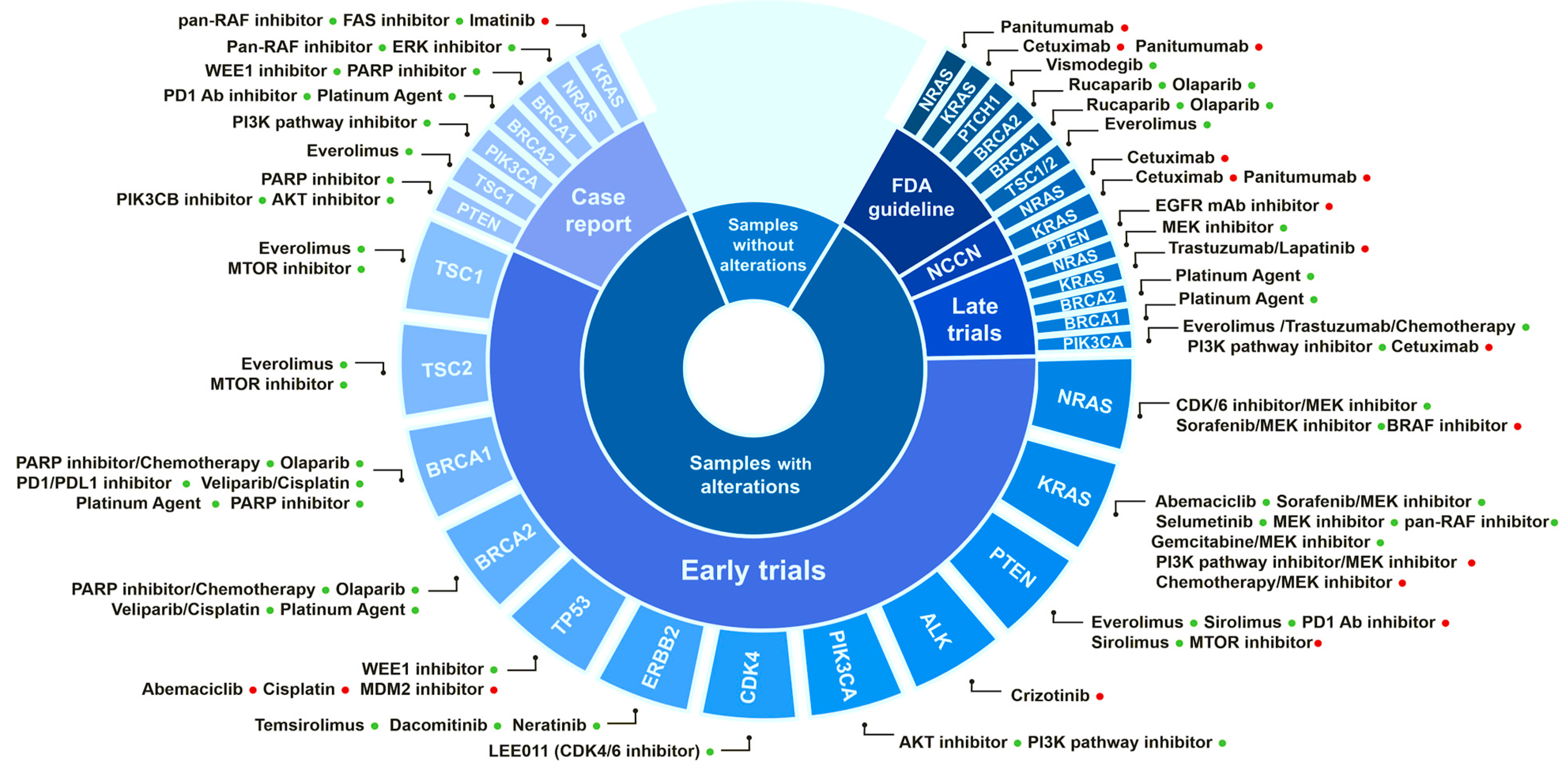
3.5. Identification of Biomarkers for Targeted Therapies
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shyr, D.; Liu, Q. Next Generation Sequencing in Cancer Research and Clinical Application. Biol. Proced. Online 2013, 15, 4. [Google Scholar] [CrossRef] [Green Version]
- Wakai, T.; Prasoon, P.; Hirose, Y.; Shimada, Y.; Ichikawa, H.; Nagahashi, M. Next-Generation Sequencing-Based Clinical Sequencing: Toward Precision Medicine in Solid Tumors. Int. J. Clin. Oncol. 2019, 24, 115–122. [Google Scholar] [CrossRef]
- Martin-Liberal, J.; Hierro, C.; Ochoa de Olza, M.; Rodon, J. Immuno-Oncology: The Third Paradigm in Early Drug Development. Target Oncol. 2017, 12, 125–138. [Google Scholar] [CrossRef]
- Garralda, E.; Dienstmann, R.; Piris-Giménez, A.; Braña, I.; Rodon, J.; Tabernero, J. New Clinical Trial Designs in the Era of Precision Medicine. Mol. Oncol. 2019, 13, 549–557. [Google Scholar] [CrossRef] [PubMed]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carioli, G.; Bertuccio, P.; Malvezzi, M.; Rodriguez, T.; Levi, F.; Boffetta, P.; La Vecchia, C.; Negri, E. Cancer Mortality Predictions for 2019 in Latin America. Int. J. Cancer 2020, 147, 619–632. [Google Scholar] [CrossRef]
- Sierra, M.S.; Soerjomataram, I.; Antoni, S.; Laversanne, M.; Piñeros, M.; de Vries, E.; Forman, D. Cancer Patterns and Trends in Central and South America. Cancer Epidemiol. 2016, 44 (Suppl. 1), S23–S42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Torres, Á.; Oliver, J.; Frecha, C.; Montealegre, A.L.; Quezada-Urbán, R.; Díaz-Velásquez, C.E.; Vaca-Paniagua, F.; Perdomo, S. Cancer Genomic Resources and Present Needs in the Latin American Region. Public Health Genom. 2017, 20, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Harismendy, O.; Kim, J.; Xu, X.; Ohno-Machado, L. Evaluating and Sharing Global Genetic Ancestry in Biomedical Datasets. J. Am. Med. Inform. Assoc. 2019, 26, 457–461. [Google Scholar] [CrossRef] [PubMed]
- Spratt, D.E.; Chan, T.; Waldron, L.; Speers, C.; Feng, F.Y.; Ogunwobi, O.O.; Osborne, J.R. Racial/Ethnic Disparities in Genomic Sequencing. JAMA Oncol. 2016, 2, 1070–1074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, J.; Hu, Z.; Mahal, B.A.; Zhao, S.D.; Kensler, K.H.; Pi, J.; Hu, X.; Zhang, Y.; Wang, Y.; Jiang, J.; et al. Integrated Analysis of Genetic Ancestry and Genomic Alterations across Cancers. Cancer Cell 2018, 34, 549–560.e9. [Google Scholar] [CrossRef] [Green Version]
- Popejoy, A.B.; Fullerton, S.M. Genomics Is Failing on Diversity. Nature 2016, 538, 161–164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bustamante, C.D.; Burchard, E.G.; De la Vega, F.M. Genomics for the World. Nature 2011, 475, 163–165. [Google Scholar] [CrossRef] [PubMed]
- Sirugo, G.; Williams, S.M.; Tishkoff, S.A. The Missing Diversity in Human Genetic Studies. Cell 2019, 177, 26–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chakravarty, D.; Gao, J.; Phillips, S.M.; Kundra, R.; Zhang, H.; Wang, J.; Rudolph, J.E.; Yaeger, R.; Soumerai, T.; Nissan, M.H.; et al. OncoKB: A Precision Oncology Knowledge Base. JCO Precis. Oncol. 2017, 2017, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Griffith, M.; Spies, N.C.; Krysiak, K.; McMichael, J.F.; Coffman, A.C.; Danos, A.M.; Ainscough, B.J.; Ramirez, C.A.; Rieke, D.T.; Kujan, L.; et al. CIViC Is a Community Knowledgebase for Expert Crowdsourcing the Clinical Interpretation of Variants in Cancer. Nat. Genet. 2017, 49, 170–174. [Google Scholar] [CrossRef] [Green Version]
- Wagner, A.H.; Walsh, B.; Mayfield, G.; Tamborero, D.; Sonkin, D.; Krysiak, K.; Deu-Pons, J.; Duren, R.P.; Gao, J.; McMurry, J.; et al. A Harmonized Meta-Knowledgebase of Clinical Interpretations of Somatic Genomic Variants in Cancer. Nat. Genet. 2020, 52, 448–457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.M.; Datto, M.; Duncavage, E.J.; Kulkarni, S.; Lindeman, N.I.; Roy, S.; Tsimberidou, A.M.; Vnencak-Jones, C.L.; Wolff, D.J.; Younes, A.; et al. Standards and Guidelines for the Interpretation and Reporting of Sequence Variants in Cancer: A Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J. Mol. Diagn. 2017, 19, 4–23. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce Framework for Analyzing Next-Generation DNA Sequencing Data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fang, L.T.; Afshar, P.T.; Chhibber, A.; Mohiyuddin, M.; Fan, Y.; Mu, J.C.; Gibeling, G.; Barr, S.; Asadi, N.B.; Gerstein, M.B.; et al. An Ensemble Approach to Accurately Detect Somatic Mutations Using SomaticSeq. Genome Biol. 2015, 16, 197. [Google Scholar] [CrossRef] [Green Version]
- Benjamin, D.; Sato, T.; Cibulskis, K.; Getz, G.; Stewart, C.; Lichtenstein, L. Calling Somatic SNVs and Indels with Mutect2. BioRxiv Bioinform. 2019. [Google Scholar] [CrossRef] [Green Version]
- Koboldt, D.C.; Zhang, Q.; Larson, D.E.; Shen, D.; McLellan, M.D.; Lin, L.; Miller, C.A.; Mardis, E.R.; Ding, L.; Wilson, R.K. VarScan 2: Somatic Mutation and Copy Number Alteration Discovery in Cancer by Exome Sequencing. Genome Res. 2012, 22, 568–576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lai, Z.; Markovets, A.; Ahdesmaki, M.; Chapman, B.; Hofmann, O.; McEwen, R.; Johnson, J.; Dougherty, B.; Barrett, J.C.; Dry, J.R. VarDict: A Novel and Versatile Variant Caller for Next-Generation Sequencing in Cancer Research. Nucleic Acids Res. 2016, 44, e108. [Google Scholar] [CrossRef] [PubMed]
- Wilm, A.; Aw, P.P.K.; Bertrand, D.; Yeo, G.H.T.; Ong, S.H.; Wong, C.H.; Khor, C.C.; Petric, R.; Hibberd, M.L.; Nagarajan, N. LoFreq: A Sequence-Quality Aware, Ultra-Sensitive Variant Caller for Uncovering Cell-Population Heterogeneity from High-Throughput Sequencing Datasets. Nucleic Acids Res. 2012, 40, 11189–11201. [Google Scholar] [CrossRef] [Green Version]
- Saunders, C.T.; Wong, W.S.W.; Swamy, S.; Becq, J.; Murray, L.J.; Cheetham, R.K. Strelka: Accurate Somatic Small-Variant Calling from Sequenced Tumor-Normal Sample Pairs. Bioinformatics 2012, 28, 1811–1817. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tamborero, D.; Rubio-Perez, C.; Deu-Pons, J.; Schroeder, M.P.; Vivancos, A.; Rovira, A.; Tusquets, I.; Albanell, J.; Rodon, J.; Tabernero, J.; et al. Cancer Genome Interpreter Annotates the Biological and Clinical Relevance of Tumor Alterations. Genome Med. 2018, 10, 25. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional Annotation of Genetic Variants from High-Throughput Sequencing Data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Jennings, L.J.; Arcila, M.E.; Corless, C.; Kamel-Reid, S.; Lubin, I.M.; Pfeifer, J.; Temple-Smolkin, R.L.; Voelkerding, K.V.; Nikiforova, M.N. Guidelines for Validation of Next-Generation Sequencing-Based Oncology Panels: A Joint Consensus Recommendation of the Association for Molecular Pathology and College of American Pathologists. J. Mol. Diagn. 2017, 19, 341–365. [Google Scholar] [CrossRef] [Green Version]
- GATK. (How to) Filter Variants Either with VQSR or by Hard-Filtering. Available online: https://gatk.broadinstitute.org/hc/en-us/articles/360035531112--How-to-Filter-variants-either-with-VQSR-or-by-hard-filtering (accessed on 6 July 2021).
- Sukhai, M.A.; Misyura, M.; Thomas, M.; Garg, S.; Zhang, T.; Stickle, N.; Virtanen, C.; Bedard, P.L.; Siu, L.L.; Smets, T.; et al. Somatic Tumor Variant Filtration Strategies to Optimize Tumor-Only Molecular Profiling Using Targeted Next-Generation Sequencing Panels. J. Mol. Diagn. 2019, 21, 261–273. [Google Scholar] [CrossRef] [Green Version]
- Tourneau, C.L.; Delord, J.-P.; Gonçalves, A.; Gavoille, C.; Dubot, C.; Isambert, N.; Campone, M.; Trédan, O.; Massiani, M.-A.; Mauborgne, C.; et al. Molecularly Targeted Therapy Based on Tumour Molecular Profiling versus Conventional Therapy for Advanced Cancer (SHIVA): A Multicentre, Open-Label, Proof-of-Concept, Randomised, Controlled Phase 2 Trial. Lancet Oncol. 2015, 16, 1324–1334. [Google Scholar] [CrossRef]
- Prentice, L.M.; Miller, R.R.; Knaggs, J.; Mazloomian, A.; Hernandez, R.A.; Franchini, P.; Parsa, K.; Tessier-Cloutier, B.; Lapuk, A.; Huntsman, D.; et al. Formalin Fixation Increases Deamination Mutation Signature but Should Not Lead to False Positive Mutations in Clinical Practice. PLoS ONE 2018, 13, e0196434. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guyard, A.; Boyez, A.; Pujals, A.; Robe, C.; Tran Van Nhieu, J.; Allory, Y.; Moroch, J.; Georges, O.; Fournet, J.-C.; Zafrani, E.-S.; et al. DNA Degrades during Storage in Formalin-Fixed and Paraffin-Embedded Tissue Blocks. Virchows Arch. 2017, 471, 491–500. [Google Scholar] [CrossRef]
- Nagahashi, M.; Shimada, Y.; Ichikawa, H.; Nakagawa, S.; Sato, N.; Kaneko, K.; Homma, K.; Kawasaki, T.; Kodama, K.; Lyle, S.; et al. Formalin-Fixed Paraffin-Embedded Sample Conditions for Deep Next Generation Sequencing. J. Surg. Res. 2017, 220, 125–132. [Google Scholar] [CrossRef]
- Ebbert, M.T.W.; Wadsworth, M.E.; Staley, L.A.; Hoyt, K.L.; Pickett, B.; Miller, J.; Duce, J.; Alzheimer’s Disease Neuroimaging Initiative; Kauwe, J.S.K.; Ridge, P.G. Evaluating the Necessity of PCR Duplicate Removal from Next-Generation Sequencing Data and a Comparison of Approaches. BMC Bioinform. 2016, 17 (Suppl. 7), 239. [Google Scholar] [CrossRef] [Green Version]
- Barnell, E.K.; Ronning, P.; Campbell, K.M.; Krysiak, K.; Ainscough, B.J.; Sheta, L.M.; Pema, S.P.; Schmidt, A.D.; Richters, M.; Cotto, K.C.; et al. Standard Operating Procedure for Somatic Variant Refinement of Sequencing Data with Paired Tumor and Normal Samples. Genet. Med. 2019, 21, 972–981. [Google Scholar] [CrossRef] [Green Version]
- Parikh, K.; Huether, R.; White, K.; Hoskinson, D.; Beaubier, N.; Dong, H.; Adjei, A.A.; Mansfield, A.S. Tumor Mutational Burden From Tumor-Only Sequencing Compared With Germline Subtraction From Paired Tumor and Normal Specimens. JAMA Netw. Open 2020, 3, e200202. [Google Scholar] [CrossRef] [Green Version]
| Gene | Drugs | Tumor Type | Evidence 1 |
|---|---|---|---|
| AKT1 * | AZD-5363 | Breast cancer Ovarian cancer Endometrial cancer | B |
| ALK | Ceritinib Crizotinib Alectinib Brigatinib Lorlatinib | Non-small cell lung cancer | A |
| ARID1A * | Trastuzumab ENMD-2076 Bevacizumab Everolimus | Breast cancer Ovarian clear cell cancer Renal cell carcinoma | C |
| BRAF * | Encorafenib + Cetuximab Vemurafenib Dabrafenib Trametinib + Dabrafenib Cobimetinib + Vemurafenib Trametinib Encorafenib + Binimetinib Vemurafenib + Cobimetinib, Trametinib + Dabrafenib Vemurafenib + Cobimetinib Encorafenib + Panitumumab | Colorectal cancer Melanoma Non-small cell lung cancer Anaplastic thyroid cancer Hairy cell leukemia Pilocytic astrocytoma Ganglioglioma Pleomorphic xanthoastrocytoma | A |
| BRCA1 * | Olaparib Niraparib Rucaparib Talazoparib | Ovarian cancer Peritoneal serous carcinoma Breast cancer Prostate cancer Ovary/fallopian tube Pancreatic cancer | A |
| BRCA2 * | Olaparib Rucaparib Talazoparib | Ovarian cancer Peritoneal serous carcinoma Breast cancer Prostate cancer Ovary/fallopian tube | A |
| CDK4 * | Palbociclib Abemaciclib | Liposarcoma | B |
| EGFR | Erlotinib Afatinib Osimertinib Gefitinib Dacomitinib | Non-small cell lung cancer | A |
| ERBB2 | Trastuzumab Fam-Trastuzumab deruxtecan-nxki Trastuzumab + Pembrolizumab Afatinib | Breast cancer Gastric adenocarcinoma Gastroesophageal junction adenocarcinoma Non-small cell lung cancer | A |
| ESR1 | Anastrozole Fulvestrant Palbociclib | Breast cancer | B |
| IDH2 | Enasidenib | Acute myeloid leukemia | A |
| KIT | Sunitinib Imatinib Regorafenib Sorafenib Ripretinib | Gastrointestinal stromal tumor Melanoma | A |
| KRAS * | Cetuximab Panitumumab Erlotinib Lapatinib Regorafenib Selumetinib Gefitinib Afatinib Icotinib Irinotecan | Colorectal cancer Non-small cell lung cancer | A |
| MET | Crizotinib Capmatinib Tepotinib | Non-small cell lung cancer | A |
| MTOR * | Everolimus Temsirolimus | Renal cell carcinoma Bladder Cancer | B |
| NRAS * | Cetuximab Panitumumab | Colorectal cancer | A |
| PDGFRA | Imatinib Sunitinib Regorafenib | Gastrointestinal stromal tumor | A |
| PI3KCA * | Buparlisib Serabelisib Alpelisib Copanlisib | Breast cancer | A |
| PTCH1 * | Vismodegib | Skin basal cell carcinoma Squamous cell carcinoma Medulloblastoma | A |
| PTEN * | Everolimus Pembrolizumab Cetuximab Sorafenib | Renal cell carcinoma Glioma Head and neck squamous cell carcinoma Colorectal cancer Hepatocellular carcinoma | B |
| ROS1 | Crizotinib Alectinib Ceritinib | Non-small cell lung cancer | C |
| SMO | Vismodegib | Skin basal cell carcinoma | B |
| TP53 * | Prognosis | Various | A |
| TSC1 * | Everolimus | Giant cell astrocytoma Renal cell carcinoma Renal angiomyolipoma | A |
| TSC2 * | MTOR inhibitors | Giant cell astrocytoma Renal cell carcinoma Renal angiomyolipoma | A |
| Classification of Variants | Total Variants | Unique Variants |
|---|---|---|
| Germline | 55 | 26 |
| Putative Novel Germline | 4 | 3 |
| Somatic | 125 | 86 |
| Putative Somatic | 13 | 13 |
| Putative Novel Somatic | 59 | 45 |
| Total | 256 | 173 |
| Gene | Mutation | Drug | Effect |
|---|---|---|---|
| BRCA1 | E1609 *, L702Wfs * 5, N1745Tfs * 20, Q1273 *, V370I | Rucaparib (PARP inhibitor) Olaparib (PARP inhibitor) | Responsive |
| BRCA2 | A2603S, D1796Mfs * 9, K3327Nfs * 13, L1114V, splice_acceptor_variant, T2783Afs * 13, T2790I, I1364M, L398P, D635G, R2034C | ||
| KRAS | A146V, Q61H G12A, G12D, G12V, L19F, Q25 * fs * 1 | Panitumumab (EGFR mAb inhibitor) Cetuximab (EGFR mAb inhibitor) | Resistant |
| NRAS | G12C, Q61R | Panitumumab (EGFR mAb inhibitor) Cetuximab (EGFR mAb inhibitor) | Resistant |
| PIK3CA | H1047R, E545A, E545K, E542K, R88Q, N345S, E579K | Alpesilib + Fulvestrant | Responsive |
| PTCH1 | R441H, D717N, H1240R, P725S, V580A, T677A, N871D | Vismodegib (SHH inhibitor) | Responsive |
| TSC1 | K375Sfs * 30, L826Q, L827Q, T582S | Everolimus (MTOR inhibitor) | Responsive |
| TSC2 | R1729C, S1530L, K533delK, A460T, A950T, D1084G, P1771L, S1096C, T154I |
| # of Samples | Gene | Mutation | Drugs | Evidence | Tumor Tested |
|---|---|---|---|---|---|
| 1 | BRCA1 | V370I | Rucaparib (PARP inhibitor) Olaparib (PARP inhibitor) WEE1 inhibitor Platinum agent (chemotherapy) Veliparib; Cisplatin (PARP inhibitor; chemotherapy) | FDA guidelines Case report Early trials | OV BRCA BRCA OV OV |
| 2 | ERBB2 | L755S | Dacomitinib (Pan ERBB inhibitor) Neratinib (ERBB2 inhibitor) Temsirolimus (MTOR inhibitor) | Early trials | NSCLC CANCER, LUAD |
| 4 | KRAS | G12A G12V Q61H G12D | Panitumumab (EGFR mAb inhibitor) Cetuximab (EGFR mAb inhibitor) Trastuzumab; Lapatinib (ERBB2 mAb inhibitor; ERBB2 inhibitor) Gemcitabine; MEK inhibitor (chemotherapy; MEK inhibitor) MEK inhibitor Selumetinib (MEK inhibitor) PI3K pathway inhibitor; MEK inhibitor Abemaciclib (CDK4/6 inhibitor) Imatinib (BCR-ABL inhibitor and KIT inhibitor) | FDA guidelines FDA guidelines Late trials Early trials Early trials Early trials Early trials Early trials Case report | COREAD LUAD PA NSCLC, HC, BT, L L PA L L GIST |
| 1 | PIK3CA | E545K | PI3K pathway inhibitor Everolimus; Trastuzumab; chemotherapy (MTOR inhibitor; ERBB2 mAb inhibitor; chemotherapy) Cetuximab (EGFR mAb inhibitor) AKT inhibitor PI3K pathway inhibitor PI3K pathway inhibitor | FDA guidelines Late trials Late trials Early trials Early trials Case report | BRCA BRCA COREAD BRCA ED, OV, CESC BLCA, HNSC, L |
| 1 | PTCH1 | P725S | Vismodegib (SHH inhibitor) | FDA guidelines | BCC, MB |
| 11 | TP53 | E171 * G244S G266V K321Ifs * 10 L257P R280T V173Gfs * 10 R213 * R248W R273C W53 * R273H R248Q Q192 * C238F | MDM2 inhibitor Abemaciclib (CDK4/6 inhibitor) Cisplatin (chemotherapy) WEE1 inhibitor | Early trials Early trials Early trials Early trials | LIP BRCA FGCT, MGCT OV |
| 1 | TSC1 | L826Q | Everolimus (MTOR inhibitor) | FDA guidelines Early trials Case report | GCA, RA BLCA ST, S, R |
| 2 | TSC2 | D1084G S1096C | Everolimus (MTOR inhibitor) | FDA guidelines | GCA, RA |
| 1 | ARID1A # | Splice acceptor variant | (EZH2 inhibitor) (PD1 inhibitor) (PARP inhibitor) (ATR inhibitor) | Pre-clinical Pre-clinical Pre-clinical Pre-clinical | OV OV CANCER CANCER |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salvo, M.; González-Feliú, E.; Toro, J.; Gallegos, I.; Maureira, I.; Miranda-González, N.; Barajas, O.; Bustamante, E.; Ahumada, M.; Colombo, A.; et al. Validation of an NGS Panel Designed for Detection of Actionable Mutations in Tumors Common in Latin America. J. Pers. Med. 2021, 11, 899. https://doi.org/10.3390/jpm11090899
Salvo M, González-Feliú E, Toro J, Gallegos I, Maureira I, Miranda-González N, Barajas O, Bustamante E, Ahumada M, Colombo A, et al. Validation of an NGS Panel Designed for Detection of Actionable Mutations in Tumors Common in Latin America. Journal of Personalized Medicine. 2021; 11(9):899. https://doi.org/10.3390/jpm11090899
Chicago/Turabian StyleSalvo, Mauricio, Evelin González-Feliú, Jessica Toro, Iván Gallegos, Ignacio Maureira, Nicolás Miranda-González, Olga Barajas, Eva Bustamante, Mónica Ahumada, Alicia Colombo, and et al. 2021. "Validation of an NGS Panel Designed for Detection of Actionable Mutations in Tumors Common in Latin America" Journal of Personalized Medicine 11, no. 9: 899. https://doi.org/10.3390/jpm11090899
APA StyleSalvo, M., González-Feliú, E., Toro, J., Gallegos, I., Maureira, I., Miranda-González, N., Barajas, O., Bustamante, E., Ahumada, M., Colombo, A., Armisén, R., Villamán, C., Ibañez, C., Bravo, M. L., Sanhueza, V., Spencer, M. L., de Toro, G., Morales, E., Bizama, C., ... Marcelain, K. (2021). Validation of an NGS Panel Designed for Detection of Actionable Mutations in Tumors Common in Latin America. Journal of Personalized Medicine, 11(9), 899. https://doi.org/10.3390/jpm11090899








