Translational Research in the Era of Precision Medicine: Where We Are and Where We Will Go
Abstract
1. Introduction
2. The Evolution of Translational Precision Medicine Research
3. Real-World Data for Translational Research
- -
- Classification: with ontological inconsistencies at registry, procedural, and research levels.
- -
- Quality: with syntactic (e.g., uterine cancer in a man), semantic (e.g., erroneous meaning assignments), or research (e.g., inconsistent correlations) relevance.
- -
- Privacy and intellectual property.
- -
- Technical: relative to informatics or computational limits.
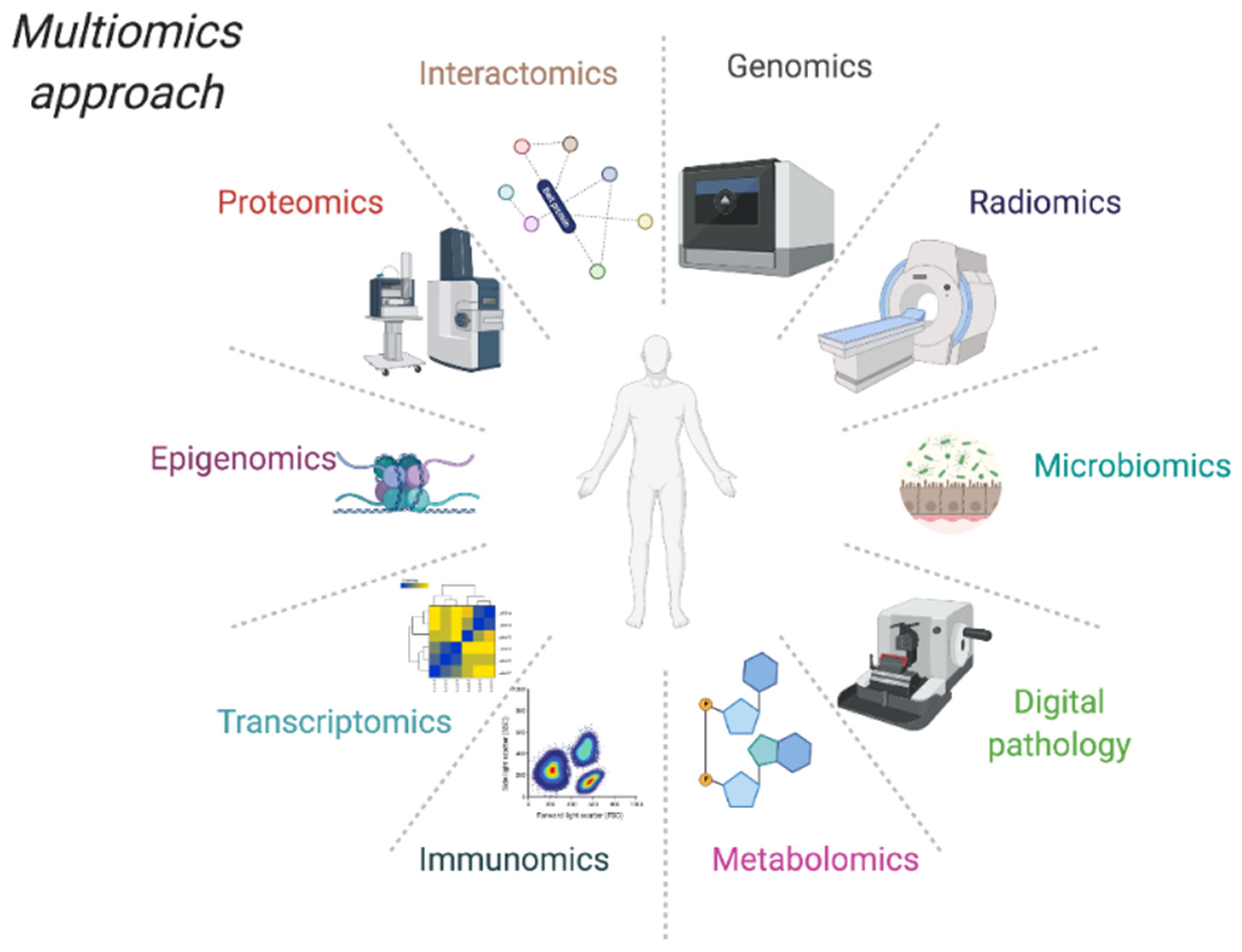
4. Omics Data for Translational Research
5. GerSom and GENERAtOR Projects: Italian Initiatives
- A.
- Mini-bots: software realized for task automation and standardization, such as data recognition and collection, process selection and projection, preliminary data analysis, validation and reporting, or rapid learning solutions, in which the AI tool automatically learns and optimizes its performances during its own activity. These mini-bots are characterized by explainable AI applications, in which explicit algorithms process data whose integrity is guaranteed from the semantic and ontological point of view by the attending researcher. Being explicit algorithms, the human intervention is always possible, and the given output is directly comprehensible for the average scientist-user, granting process transparency, repeatability, and traceability in every phase of the translational analysis.Different mini-bots can be realized: one of the most popular examples are: the guardian bot, thought to automatically warn the researchers in case specific events occur (e.g., collection of out of range values); process bot, that identifies deviations from selected guidelines or from the expected behavior of a specific phenomenon; advanced data manager bot that collect and make actionable data of different sources and type (e.g., elastic search and text mining tools that integrate into e-platform lab reports, clinical charts and records, surgical reports, or visits).
- B.
- Avatar: these tools are represented by advanced algorithms, specifically trained to create decisional support systems able to predict clinical outcomes, such as prognosis, treatment related toxicities or complications, therapy results, or diagnostic performances of a specific approach. These Avatars may represent a digital twin of the single patient.Avatars may successfully be used in the setup of virtual trials that will for sure boost the potentialities of these approaches.
- C.
- Synthetic data packages: these totally anonymized, General Data Protection Regulation (GDPR) compliant by design, data packages could be used to generate and develop translational and clinical studies in certified and protected virtual environments in which innovative data analysis techniques, coming from knowledge domains other than the traditional biomedical ones, can be successfully applied in the framework of the most fruitful open innovation paradigms.
- D.
- Advanced radiomics and quantitative bio-imaging analysis tools. These image analysis platforms will enrich the value of standard clinical imaging with new decisional variables and translation meaning, thanks to the extraction of certified radiomics features. In this way also the institutional imaging data-lake can be successfully made actionable, flanking the image scientist in both his clinical and research activities [59,60].
- E.
- Informatics solutions aiming to integrate data extracted from portable devices (i.e., fitness bracelets and other types of wearables) in the innovative framework of patient generated RWD, e-health 2.0 clinical trials.
6. The CERVGEN Project: A Next Step towards Precision Medicine in Cervical Cancer
7. Data Privacy/Security
8. Discussion
9. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ledford, H. Translational Research: The Full Cycle. Nature 2008, 453, 843–845. [Google Scholar] [CrossRef] [PubMed]
- Koga, T.; Chaim, I.A.; Benitez, J.A.; Markmiller, S.; Parisian, A.D.; Hevner, R.F.; Turner, K.M.; Hessenauer, F.M.; D’Antonio, M.; Nguyen, N.D.; et al. Longitudinal Assessment of Tumor Development Using Cancer Avatars Derived from Genetically Engineered Pluripotent Stem Cells. Nat. Commun. 2020, 11, 550. [Google Scholar] [CrossRef] [PubMed]
- Gargiulo, G. Next-Generation in Vivo Modeling of Human Cancers. Front. Oncol. 2018, 8, 429. [Google Scholar] [CrossRef]
- Kijewska, M.; Viski, C.; Turrell, F.; Fitzpatrick, A.; van Weverwijk, A.; Gao, Q.; Iravani, M.; Isacke, C.M. Using an In-Vivo Syngeneic Spontaneous Metastasis Model Identifies ID2 as a Promoter of Breast Cancer Colonisation in the Brain. Breast Cancer Res. 2019, 21, 4. [Google Scholar] [CrossRef]
- Filippini, D.; Agosto, S.D.; Delfino, P.; Simbolo, M.; Piro, G.; Rusev, B.; Veghini, L.; Cantù, C.; Lupo, F.; Ugel, S.; et al. Immunoevolution of Mouse Pancreatic Organoid Isografts from Preinvasive to Metastatic Disease. Sci. Rep. 2019, 9, 12286. [Google Scholar] [CrossRef]
- Lupo, F.; Piro, G.; Torroni, L.; Delfino, P.; Trovato, R.; Rusev, B.; Fiore, A.; Filippini, D.; De Sanctis, F.; Manfredi, M.; et al. Organoid-Transplant Model Systems to Study the Effects of Obesity on the Pancreatic Carcinogenesis in Vivo. Front. Cell Dev. Biol. 2020, 8, 308. [Google Scholar] [CrossRef]
- Kent Lloyd, K.C.; Robinson, P.N.; MacRae, C.A. Animal-Based Studies Will Be Essential for Precision Medicine. Sci. Transl. Med. 2016, 8, 352ed12. [Google Scholar] [CrossRef]
- Dance, A. Medical Histories. Nature 2016, 537, S52–S53. [Google Scholar] [CrossRef]
- Druker, B.J.; Sawyers, C.L.; Kantarjian, H.; Resta, D.J.; Reese, S.F.; Ford, J.M.; Capdeville, R.; Talpaz, M. Activity of a Specific Inhibitor of the BCR-ABL Tyrosine Kinase in the Blast Crisis of Chronic Myeloid Leukemia and Acute Lymphoblastic Leukemia with the Philadelphia Chromosome. N. Engl. J. Med. 2001, 344, 1038–1042. [Google Scholar] [CrossRef]
- de Leon, J.; Susce, M.T.; Murray-Carmichael, E. The AmpliChip CYP450 Genotyping Test: Integrating a New Clinical Tool. Mol. Diagn. Ther. 2006, 10, 135–151. [Google Scholar] [CrossRef]
- Ginsburg, G.S.; Phillips, K.A. Precision Medicine: From Science To Value. Health Aff. 2018, 37, 694–701. [Google Scholar] [CrossRef]
- The ICGC/TCGA. Pan-Cancer Analysis of Whole Genomes Consortium Pan-Cancer Analysis of Whole Genomes. Nature 2020, 578, 82–93. [Google Scholar] [CrossRef]
- Rheinbay, E.; Nielsen, M.M.; Abascal, F.; Wala, J.A.; Shapira, O.; Tiao, G.; Hornshøj, H.; PCAWG Drivers and Functional Interpretation Working Group; PCAWG Structural Variation Working Group; PCAWG Consortium; et al. Analyses of Non-Coding Somatic Drivers in 2,658 Cancer Whole Genomes. Nature 2020, 578, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, L.B.; Kim, J.; Haradhvala, N.J.; Huang, M.N.; Ng, A.W.; Wu, Y.; Boot, A.; Covington, K.R.; Gordenin, D.A.; Bergstrom, E.N.; et al. The Repertoire of Mutational Signatures in Human Cancer. Nature 2020, 578, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Roberts, N.D.; Wala, J.A.; Shapira, O.; Schumacher, S.E.; Kumar, K.; Khurana, E.; Waszak, S.; PCAWG Structural Variation Working Group; PCAWG Consortium; et al. Patterns of Somatic Structural Variation in Human Cancer Genomes. Nature 2020, 578, 112–121. [Google Scholar] [CrossRef] [PubMed]
- Gerstung, M.; Jolly, C.; Leshchiner, I.; Dentro, S.C.; Gonzalez, S.; Rosebrock, D.; Mitchell, T.J.; Rubanova, Y.; PCAWG Evolution & Heterogeneity Working Group; PCAWG Consortium; et al. The Evolutionary History of 2658 Cancers. Nature 2020, 578, 122–128. [Google Scholar] [CrossRef]
- Calabrese, C.; Davidson, N.R.; Demircioğlu, D.; Fonseca, N.A.; He, Y.; Kahles, A.; Lehmann, K.-V.; PCAWG Transcriptome Core Group; PCAWG Transcriptome Working Group; PCAWG Consortium; et al. Genomic Basis for RNA Alterations in Cancer. Nature 2020, 578, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Dreyer, S.B.; Jamieson, N.B.; Cooke, S.L.; Valle, J.W.; McKay, C.J.; Biankin, A.V.; Chang, D.K. PRECISION-Panc: The Next Generation Therapeutic Development Platform for Pancreatic Cancer. Clin. Oncol. 2020, 32, 1–4. [Google Scholar] [CrossRef]
- Froeling, F.E.M.; Casolino, R.; Pea, A.; Biankin, A.V.; Chang, D.K. Molecular Subtyping and Precision Medicine for Pancreatic Cancer. J. Clin. Med. 2021, 10, 149. [Google Scholar] [CrossRef]
- Abernethy, A.P.; Etheredge, L.M.; Ganz, P.A.; Wallace, P.; German, R.R.; Neti, C.; Bach, P.B.; Murphy, S.B. Rapid-Learning System for Cancer Care. JCO 2010, 28, 4268–4274. [Google Scholar] [CrossRef] [PubMed]
- Sinsky, C.; Colligan, L.; Li, L.; Prgomet, M.; Reynolds, S.; Goeders, L.; Westbrook, J.; Tutty, M.; Blike, G. Allocation of Physician Time in Ambulatory Practice: A Time and Motion Study in 4 Specialties. Ann. Intern. Med. 2016, 165, 753. [Google Scholar] [CrossRef]
- Makady, A.; de Boer, A.; Hillege, H.; Klungel, O.; Goettsch, W. What Is Real-World Data? A Review of Definitions Based on Literature and Stakeholder Interviews. Value Health 2017, 20, 858–865. [Google Scholar] [CrossRef]
- Lewis, J.R.R.; Kerridge, I.; Lipworth, W. Use of Real-World Data for the Research, Development, and Evaluation of Oncology Precision Medicines. JCO Precis. Oncol. 2017, 1–11. [Google Scholar] [CrossRef]
- Meldolesi, E.; van Soest, J.; Dinapoli, N.; Dekker, A.; Damiani, A.; Gambacorta, M.A.; Valentini, V. Medicine Is a Science of Uncertainty and an Art of Probability (Sir W. Osler). Radiother. Oncol. 2015, 114, 132–134. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Ashrafian, H.; Ghafur, S.; Fontana, G.; Gardner, C.; Prime, M. Challenges for the Evaluation of Digital Health Solutions—A Call for Innovative Evidence Generation Approaches. NPJ Digit. Med. 2020, 3, 110. [Google Scholar] [CrossRef] [PubMed]
- Househ, M.; Aldosari, B. The Hazards of Data Mining in Healthcare. Stud. Health Technol. Inform. 2017, 238, 80–83. [Google Scholar]
- Schneeweiss, S.; Glynn, R.J. Real-World Data Analytics Fit for Regulatory Decision-Making. Am. J. Law. Med. 2018, 44, 197–217. [Google Scholar] [CrossRef]
- Wright, A.; Sittig, D.F. A Four-Phase Model of the Evolution of Clinical Decision Support Architectures. Int. J. Med. Inform. 2008, 77, 641–649. [Google Scholar] [CrossRef][Green Version]
- Sievers, H.; Joos, A.; Hiligsmann, M. Real-World Evidence: Perspectives on Challenges, Value, and Alignment of Regulatory and National Health Technology Assessment Data Collection Requirements. Int. J. Technol. Assess Health Care 2021, 37, e40. [Google Scholar] [CrossRef]
- Marazzi, F.; Tagliaferri, L.; Masiello, V.; Moschella, F.; Colloca, G.F.; Corvari, B.; Sanchez, A.M.; Capocchiano, N.D.; Pastorino, R.; Iacomini, C.; et al. GENERATOR Breast DataMart—The Novel Breast Cancer Data Discovery System for Research and Monitoring: Preliminary Results and Future Perspectives. JPM 2021, 11, 65. [Google Scholar] [CrossRef] [PubMed]
- Wise, J.; Möller, A.; Christie, D.; Kalra, D.; Brodsky, E.; Georgieva, E.; Jones, G.; Smith, I.; Greiffenberg, L.; McCarthy, M.; et al. The Positive Impacts of Real-World Data on the Challenges Facing the Evolution of Biopharma. Drug Discov. Today 2018, 23, 788–801. [Google Scholar] [CrossRef] [PubMed]
- Collins, R. What Makes UK Biobank Special? Lancet 2012, 379, 1173–1174. [Google Scholar] [CrossRef]
- Chen, Z.; Chen, J.; Collins, R.; Guo, Y.; Peto, R.; Wu, F.; Li, L.; on behalf of the China Kadoorie Biobank (CKB) Collaborative Group. China Kadoorie Biobank of 0.5 Million People: Survey Methods, Baseline Characteristics and Long-Term Follow-Up. Int. J. Epidemiol. 2011, 40, 1652–1666. [Google Scholar] [CrossRef] [PubMed]
- Nagai, A.; Hirata, M.; Kamatani, Y.; Muto, K.; Matsuda, K.; Kiyohara, Y.; Ninomiya, T.; Tamakoshi, A.; Yamagata, Z.; Mushiroda, T.; et al. Overview of the BioBank Japan Project: Study Design and Profile. J. Epidemiol. 2017, 27, S2–S8. [Google Scholar] [CrossRef]
- Stark, Z.; Dolman, L.; Manolio, T.A.; Ozenberger, B.; Hill, S.L.; Caulfied, M.J.; Levy, Y.; Glazer, D.; Wilson, J.; Lawler, M.; et al. Integrating Genomics into Healthcare: A Global Responsibility. Am. J. Hum. Genet. 2019, 104, 13–20. [Google Scholar] [CrossRef]
- Caron, M.; Choquet-Kastylevsky, G.; Joubert-Caron, R. Cancer Immunomics Using Autoantibody Signatures for Biomarker Discovery. Mol. Cell Proteom. 2007, 6, 1115–1122. [Google Scholar] [CrossRef] [PubMed]
- Di Sante, G.; Tredicine, M.; Rolla, S.; Di Pino, A.; Ria, F. Past and Future of the Molecular Characterization of the T Cell Repertoire: Some Highlights of Eli Sercarz’s Contributions. Crit. Rev. Immunol. 2020, 40, 249–253. [Google Scholar] [CrossRef]
- Pandolfi, F.; Cianci, R.; Casciano, F.; Pagliari, D.; De Pasquale, T.; Landolfi, R.; Di Sante, G.; Kurnick, J.T.; Ria, F. Skewed T-Cell Receptor Repertoire: More than a Marker of Malignancy, a Tool to Dissect the Immunopathology of Inflammatory Diseases. J. Biol. Regul. Homeost. Agents 2011, 25, 153–161. [Google Scholar]
- Finn, O.J. Immune Response as a Biomarker for Cancer Detection and a Lot More. N. Engl. J. Med. 2005, 353, 1288–1290. [Google Scholar] [CrossRef]
- He, Y.; Mohamedali, A.; Huang, C.; Baker, M.S.; Nice, E.C. Oncoproteomics: Current Status and Future Opportunities. Clin. Chim. Acta 2019, 495, 611–624. [Google Scholar] [CrossRef]
- Price, N.D.; Magis, A.T.; Earls, J.C.; Glusman, G.; Levy, R.; Lausted, C.; McDonald, D.T.; Kusebauch, U.; Moss, C.L.; Zhou, Y.; et al. A Wellness Study of 108 Individuals Using Personal, Dense, Dynamic Data Clouds. Nat. Biotechnol. 2017, 35, 747–756. [Google Scholar] [CrossRef]
- Ogilvie, L.A.; Wierling, C.; Kessler, T.; Lehrach, H.; Lange, B.M.H. Predictive Modeling of Drug Treatment in the Area of Personalized Medicine. Cancer Inform. 2015, 14, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Prosperi, M.; Min, J.S.; Bian, J.; Modave, F. Big Data Hurdles in Precision Medicine and Precision Public Health. BMC Med. Inform. Decis. Mak. 2018, 18, 139. [Google Scholar] [CrossRef]
- Hulsen, T.; Jamuar, S.; Moody, A.; Karnes, J.; Varga, O.; Hedensted, S.; Spreafico, R.; Hafler, D.; McKinney, E. From Big Data to Precision Medicine. Front. Med. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Heindl, A.; Nawaz, S.; Yuan, Y. Mapping Spatial Heterogeneity in the Tumor Microenvironment: A New Era for Digital Pathology. Lab. Investig. 2015, 95, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Barsoum, I.; Tawedrous, E.; Faragalla, H.; Yousef, G.M. Histo-Genomics: Digital Pathology at the Forefront of Precision Medicine. Diagnosis 2019, 6, 203–212. [Google Scholar] [CrossRef]
- Kent, W.; Sugnet, C.; Furey, T.; Roskin, K.; Pringle, T.; Zahler, A.; Haussler, D. The Human Genome Browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef]
- Zhou, X.; Maricque, B.; Xie, M.; Li, D.; Sundaram, V.; Martin, E.A.; Koebbe, B.C.; Nielsen, C.; Hirst, M.; Farnham, P.; et al. The Human Epigenome Browser at Washington University. Nat. Methods 2011, 8, 989–990. [Google Scholar] [CrossRef]
- Martin, T.C.; Yet, I.; Tsai, P.-C.; Bell, J.T. CoMET: Visualisation of Regional Epigenome-Wide Association Scan Results and DNA Co-Methylation Patterns. BMC Bioinform. 2015, 16. [Google Scholar] [CrossRef] [PubMed]
- Hoadley, K.A.; Yau, C.; Wolf, D.M.; Cherniack, A.D.; Tamborero, D.; Ng, S.; Leiserson, M.D.M.; Niu, B.; McLellan, M.D.; Uzunangelov, V.; et al. Multiplatform Analysis of 12 Cancer Types Reveals Molecular Classification within and across Tissues of Origin. Cell 2014, 158, 929–944. [Google Scholar] [CrossRef]
- Issa, I.A.; Noureddine, M. Colorectal Cancer Screening: An Updated Review of the Available Options. World J. Gastroenterol. 2017, 23, 5086–5096. [Google Scholar] [CrossRef]
- Zolotovskaia, M.; Sorokin, M.; Garazha, A.; Borisov, N.; Buzdin, A. Molecular Pathway Analysis of Mutation Data for Biomarkers Discovery and Scoring of Target Cancer Drugs. In Nucleic Acid Detection and Structural Investigations: Methods and Protocols; Astakhova, K., Bukhari, S.A., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2020; pp. 207–234. ISBN 978-1-07-160138-9. [Google Scholar]
- Cheung, P.K.; Ma, M.H.; Tse, H.F.; Yeung, K.F.; Tsang, H.F.; Chu, M.K.M.; Kan, C.M.; Cho, W.C.S.; Ng, L.B.W.; Chan, L.W.C.; et al. The Applications of Metabolomics in the Molecular Diagnostics of Cancer. Expert Rev. Mol. Diagn. 2019, 19, 785–793. [Google Scholar] [CrossRef]
- Cammarota, G.; Ianiro, G.; Ahern, A.; Carbone, C.; Temko, A.; Claesson, M.J.; Gasbarrini, A.; Tortora, G. Gut Microbiome, Big Data and Machine Learning to Promote Precision Medicine for Cancer. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 635–648. [Google Scholar] [CrossRef]
- Armour, C.R.; Nayfach, S.; Pollard, K.S.; Sharpton, T.J. A Metagenomic Meta-Analysis Reveals Functional Signatures of Health and Disease in the Human Gut Microbiome. mSystems 2019, 4. [Google Scholar] [CrossRef]
- Elinav, E.; Garrett, W.S.; Trinchieri, G.; Wargo, J. The Cancer Microbiome. Nat. Rev. Cancer 2019, 19, 371–376. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A.; Manghi, P.; Asnicar, F.; Pasolli, E.; Armanini, F.; Zolfo, M.; Beghini, F.; Manara, S.; Karcher, N.; Pozzi, C.; et al. Metagenomic Analysis of Colorectal Cancer Datasets Identifies Cross-Cohort Microbial Diagnostic Signatures and a Link with Choline Degradation. Nat. Med. 2019. [Google Scholar] [CrossRef]
- Laghi, L.; Ricciardiello, L. The Changing Approach for Identifying Hereditary Colorectal Cancer Syndromes. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 593–594. [Google Scholar] [CrossRef] [PubMed]
- Dinapoli, N.; Alitto, A.R.; Vallati, M.; Gatta, R.; Autorino, R.; Boldrini, L.; Damiani, A.; Valentini, V. Moddicom: A Complete and Easily Accessible Library for Prognostic Evaluations Relying on Image Features. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; pp. 771–774. [Google Scholar]
- Zwanenburg, A.; Vallières, M.; Abdalah, M.A.; Aerts, H.J.W.L.; Andrearczyk, V.; Apte, A.; Ashrafinia, S.; Bakas, S.; Beukinga, R.J.; Boellaard, R.; et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-Based Phenotyping. Radiology 2020, 295, 328–338. [Google Scholar] [CrossRef]
- Buttarelli, M.; Babini, G.; Raspaglio, G.; Filippetti, F.; Battaglia, A.; Ciucci, A.; Ferrandina, G.; Petrillo, M.; Marino, C.; Mancuso, M.; et al. A Combined ANXA2-NDRG1-STAT1 Gene Signature Predicts Response to Chemoradiotherapy in Cervical Cancer. J. Exp. Clin. Cancer Res. 2019, 38, 279. [Google Scholar] [CrossRef]
- Vayena, E.; Dzenowagis, J.; Brownstein, J.S.; Sheikh, A. Policy Implications of Big Data in the Health Sector. Bull. World Health Organ. 2018, 96, 66–68. [Google Scholar] [CrossRef] [PubMed]
- Pastorino, R.; De Vito, C.; Migliara, G.; Glocker, K.; Binenbaum, I.; Ricciardi, W.; Boccia, S. Benefits and Challenges of Big Data in Healthcare: An Overview of the European Initiatives. Eur. J. Public Health 2019, 29, 23–27. [Google Scholar] [CrossRef] [PubMed]
- Riba, M.; Sala, C.; Toniolo, D.; Tonon, G. Big Data in Medicine, the Present and Hopefully the Future. Front. Med. 2019, 6. [Google Scholar] [CrossRef] [PubMed]
- Abul-Husn, N.S.; Kenny, E.E. Personalized Medicine and the Power of Electronic Health Records. Cell 2019, 177, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Prasser, F.; Kohlbacher, O.; Mansmann, U.; Bauer, B.; Kuhn, K.A. Data Integration for Future Medicine (DIFUTURE). Methods Inf. Med. 2018, 57, e57–e65. [Google Scholar] [CrossRef] [PubMed]
- Palombo, F.; De Paoli, P.; De Maria, R. Alleanza Contro Il Cancro: The Accreditation System of the Excellence Network of Italian Cancer Centers in the Precision Medicine Era. Tumori 2015, 101 (Suppl. S1), S64–S66. [Google Scholar] [CrossRef]
- Jones, D.T. Setting the Standards for Machine Learning in Biology. Nat. Rev. Mol. Cell Biol. 2019, 20, 659–660. [Google Scholar] [CrossRef]
- Leonelli, S. Data—From Objects to Assets. Nature 2019, 574, 317–320. [Google Scholar] [CrossRef]
- Finlayson, S.G.; Bowers, J.D.; Ito, J.; Zittrain, J.L.; Beam, A.L.; Kohane, I.S. Adversarial Attacks on Medical Machine Learning. Science 2019, 363, 1287–1289. [Google Scholar] [CrossRef]
- Taddeo, M.; Floridi, L. How AI Can Be a Force for Good. Science 2018, 361, 751–752. [Google Scholar] [CrossRef] [PubMed]
- Aronson, S.J.; Rehm, H.L. Building the Foundation for Genomics in Precision Medicine. Nature 2015, 526, 336–342. [Google Scholar] [CrossRef]
- Hood, L.; Friend, S.H. Predictive, Personalized, Preventive, Participatory (P4) Cancer Medicine. Nat. Rev. Clin. Oncol. 2011, 8, 184–187. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Yang, L.; Hu, H.; Chen, J.; Shen, B. How to Become a Smart Patient in the Era of Precision Medicine? In Healthcare and Big Data Management; Shen, B., Ed.; Advances in Experimental Medicine and Biology; Springer: Singapore, 2017; pp. 1–16. ISBN 978-981-10-6041-0. [Google Scholar]
| EU-Code | Acronym | Title | Start Date | End Date |
|---|---|---|---|---|
| 951724 | B1MG | Beyond 1M Genomes | 1 June 2020 | 31 May 2023 |
| 715772 | BabyVir | The role of the virome in shaping the gut ecosystem during the first year of life | 1 April 2017 | 30 September 2022 |
| 777090 | Back-UP | Personalized Prognostic Models to Improve Well-being and Return to Work After Neck and Low Back Pain | 1 January 2018 | 30 April 2021 |
| 115974 | BEAt-DKD | Biomarker Enterprise to Attack DKD | 1 September 2016 | 31 August 2021 |
| 821511 | BIOMAP | Biomarkers in Atopic Dermatitis and Psoriasis | 1 April 2019 | 31 March 2024 |
| 679586 | BUMP BETTER | Understanding the metaphysics of pregnancy | 1 April 2016 | 31 March 2021 |
| 876362 | CHARM | Challenging environments tolerant Smart systems for IoT and AI | 1 June 2020 | 31 May 2023 |
| 825775 | CINECA | Common Infrastructure for National Cohorts in Europe, Canada, and Africa | 1 January 2019 | 31 December 2022 |
| 821520 | ConcePTION | Building an ecosystem for better monitoring and communicating of medication safety in pregnancy and breastfeeding: validated and regulatory endorsed workflows for fast, optimized evidence generation | 1 April 2019 | 31 March 2024 |
| 765158 | COsMIC | COmbatting disorders of adaptive immunity with Systems MedICine | 1 January 2018 | 31 December 2021 |
| 949850 | DCUBATION | Redefining the term ‘Incubation period’ using large-scale digital data | 1 November 2020 | 31 October 2025 |
| 806968 | EHDEN | European Health Data and Evidence Network | 1 November 2018 | 30 April 2024 |
| 724115 | ENABLE | European Academy for Biomedical Science | 1 July 2016 | 30 June 2021 |
| 824160 | EnTimement | ENtrainment and synchronization at multiple TIME scales in the MENTal foundations of expressive gesture | 1 January 2019 | 31 December 2022 |
| 779282 | ERA PerMed | ERA-Net Cofund in Personalized Medicine | 1 December 2017 | 30 November 2022 |
| 806948 | ESCulab | European Screening Centre; Unique Library for Attractive Biology | 1 December 2018 | 30 November 2023 |
| 964333 | EU-Africa PerMed | Building links between Europe and Africa in personalized medicine | 1 February 2021 | 31 January 2025 |
| 825843 | EU-STANDS4PM | A European standardization framework for data integration and data-driven in silico models for personalized medicine | 1 January 2019 | 31 December 2021 |
| 952103 | EuCanImage | A European Cancer Image Platform Linked to Biological and Health Data for Next-Generation Artificial Intelligence and Precision Medicine in Oncology | 1 October 2020 | 30 September 2024 |
| 825903 | euCanSHare | An EU-Canada joint infrastructure for next-generation multi-Study Heart research | 1 December 2018 | 30 November 2022 |
| 824753 | FETFX | Stimulating effects of Future and Emerging Technologies through communication and outreach | 1 January 2019 | 30 June 2021 |
| 101017549 | GenoMed4ALL | Genomics and Personalized Medicine for all though Artificial Intelligence in Hematological Diseases | 1 January 2021 | 31 December 2024 |
| 945334 | Gravitate-Health | Empowering and Equipping Europeans with health information for Active Personal Health Management and Adherence to Treatment | 1 November 2020 | 31 October 2025 |
| 823939 | GreenX4Drug | Green Enantioselective Halogenation for Drug Discovery and Manufacture | 1 April 2019 | 31 March 2023 |
| 116026 | HARMONY | Healthcare Alliance for Resourceful Medicines Offensive against Neoplasms in HematologY | 1 January 2017 | 31 December 2021 |
| 957532 | HEART.FM | Maximizing the Therapeutic Potential of Music through Tailored Therapy with Physiological Feedback in Cardiovascular Disease | 1 November 2020 | 30 April 2022 |
| 874694 | IC2PerMed | Integrating China in the International Consortium for Personalized Medicine | 1 January 2020 | 31 December 2023 |
| 731366 | ICPerMed | Secretariat for the International Consortium for Personalized Medicine (IC PerMed) | 1 November 2016 | 30 April 2021 |
| 964197 | ICPerMed | Secretariat for the International Consortium for Personalized Medicine (ICPerMed) | 1 March 2021 | 29 February 2024 |
| 853981 | IDEA-FAST | Identifying Digital Endpoints to Assess FAtigue, Sleep and acTivities in daily living in Neurodegenerative disorders and Immune-mediated inflammatory diseases | 1 November 2019 | 30 April 2025 |
| 831514 | Immune-Image | Specific Imaging of Immune Cell Dynamics Using Novel Tracer Strategies | 1 October 2019 | 30 September 2024 |
| 101016775 | INTERVENE | International consortium for integrative genomics prediction | 1 January 2021 | 31 December 2025 |
| 826121 | iPC | Individualized Pediatric Cure: Cloud-based virtual-patient models for precision pediatric oncology | 1 January 2019 | 31 May 2023 |
| 825821 | iReceptor Plus | Architecture and tools for the query of antibody and t-cell receptor sequencing data repositories for enabling improved personalized medicine and immunotherapy | 1 January 2019 | 31 December 2022 |
| 681043 | JPsustaiND | Coordination Action in support of the sustainability and globalization of the Joint Programming Initiative on Neurodegenerative Diseases | 1 November 2015 | 31 October 2021 |
| 101017453 | KATY | Knowledge At the Tip of Your fingers: Clinical Knowledge for Humanity | 1 January 2021 | 31 December 2024 |
| 678304 | LEASP | Learning spatiotemporal patterns in longitudinal image data sets of the aging brain | 1 September 2016 | 31 August 2021 |
| 732592 | Lifebrain | Healthy minds from 0-100 years. Optimizing the use of European brain imaging cohorts | 1 January 2017 | 30 June 2022 |
| 777377 | LITMUS | Liver Investigation: Testing Marker Utility in Steatohepatitis | 1 November 2017 | 31 October 2022 |
| 965286 | MAESTRIA | Machine Learning Artificial Intelligence Early Detection Stroke Atrial Fibrillation | 1 March 2021 | 28 February 2026 |
| 873262 | MAGELIA | A disruptive Magnetically Enhanced Library preparation platform for Next Generation Sequencing | 1 August 2019 | 31 July 2021 |
| 820820 | MOBILISE-D | Connecting digital mobility assessment to clinical outcomes for regulatory and clinical endorsement | 1 April 2019 | 31 March 2024 |
| 893699 | MODIRen | Integrative metabolomics and genomics analysis for the development of markers of inherited kidney diseases: a personalized medicine approach | 4 January 2019 | 3 January 2023 |
| 806975 | NECESsITY | NEw Clinical Endpoints in primary Sjögren’s Syndrome: an Interventional Trial based on stratifYing patients | 1 January 2019 | 31 December 2024 |
| 724334 | NOSUDEP | A Wearable Electronics Approach To Reduce Mortality in Epilepsy | 1 September 2017 | 28 February 2023 |
| 825410 | ONCOBIOME | Gut OncoMicrobiome Signatures (GOMS) associated with cancer incidence, prognosis and prediction of treatment response. | 1 January 2019 | 30 June 2024 |
| 693124 | ONOFF | Perception of voices that do not exist: Tracking the temporal signatures of auditory hallucinations | 1 September 2016 | 31 December 2021 |
| 946050 | PACE | Platform for Rapid Development of Personalized Nanomedicine Drug Delivery Systems | 1 September 2020 | 28 February 2022 |
| 101016851 | PANCAIM | Pancreatic cancer AI for genomics and personalized Medicine | 1 January 2021 | 31 December 2024 |
| 951773 | PerMedCoE | HPC/Exascale Centre of Excellence in Personalized Medicine | 1 October 2020 | 30 September 2023 |
| 115976 | PHAGO | Inflammation and AD: modulating microglia function focusing on TREM2 and CD33 | 1 November 2016 | 31 October 2021 |
| 716079 | PREDICT | PREcision medicine Drug combination testing In neuroblastoma organoids to guide Clinical Trials | 1 March 2017 | 28 February 2022 |
| 754425 | PROMINENT | Personalized Medicine in Diabetic Chronic Disease Management | 1 September 2017 | 31 August 2022 |
| 754907 | R-LiNK | Optimizing response to Li treatment through personalized evaluation of individuals with bipolar I disorder: the R-LiNK initiative | 1 January 2018 | 31 December 2022 |
| 115902 | RADAR-CNS | Remote Assessment of Disease and Relapse in Central Nervous System Disorders | 1 April 2016 | 31 March 2022 |
| 825746 | RECODID | Integrated human data repositories for infectious disease-related international cohorts to foster personalized medicine approaches to infectious disease research | 1 January 2019 | 31 December 2022 |
| 825812 | REGIONS4PERMED | interregional coordination for a fast and deep uptake of personalized health | 1 November 2018 | 30 April 2023 |
| 857491 | REMODEL | Research models in infection, cancer and regeneration: replacement and translation | 1 November 2019 | 31 October 2022 |
| 801540 | RESCUE | Local Training Network on REgenerative medicine and Stem Cell technology in UtrEcht | 1 June 2018 | 31 May 2023 |
| 847912 | RESCUER | RESistance under Combinatorial treatment in ER+ and ER- breast cancer. | 1 January 2020 | 31 December 2024 |
| 825046 | SAPHIRE | Securing Adoption of Personalized Health in REgions | 1 December 2018 | 31 May 2022 |
| 874556 | SINO-EU-PerMed | Widening Sino-EU policy and research cooperation in Personalized Medicine | 1 January 2020 | December 2023 |
| 826117 | Smart4Health | Citizen-centered EU-EHR exchange for personalized health | 1 January 2019 | 28 February 2023 |
| 875534 | SOPHIA | Stratification of Obese Phenotypes to Optimize Future Obesity Therapy | 1 June 2020 | 31 May 2025 |
| 733112 | SPIDIA4P SPIDIA | Standardization of generic Pre-analytical procedures for In-vitro DIAgnostics for Personalized Medicine | 1 January 2017 | 30 June 2021 |
| 825884 | SYNCHROS | SYNergies for Cohorts in Health: integrating the ROle of all Stakeholders | 1 January 2019 | 31 December 2021 |
| 733100 | SYSCID | A Systems medicine approach to chronic inflammatory disease | 1 January 2017 | 31 March 2022 |
| 730994 | TERRINet | The European Robotics Research Infrastructure Network | 1 December 2017 | 30 November 2021 |
| 821283 | TransBioLine | Translational Safety Biomarker Pipeline: Enabling development and implementation of novel safety biomarkers in clinical trials and diagnosis of disease | 1 February 2019 | 31 January 2024 |
| 831458 | Trials@Home | Center of Excellence—Remote Decentralized Clinical Trials | 1 September 2019 | 31 August 2024 |
| 668353 | U-PGx | Ubiquitous Pharmacogenomics (U-PGx): Making actionable pharmacogenomic data and effective treatment optimization accessible to every European citizen | 1 January 2016 | 30 June 2021 |
| 820755 | VALUE-Dx | The value of diagnostics to combat antimicrobial resistance by optimizing antibiotic use | 1 April 2019 | 31 March 2023 |
| 826421 | VirtualBrainCloud | Personalized Recommendations for Neurodegenerative Disease | 1 December 2018 | 30 November 2022 |
| 824128 | VIRTUALTIMES | Exploring and Modifying the Sense of Time in Virtual Environments | 1 January 2019 | 31 December 2022 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Maria Marchiano, R.; Di Sante, G.; Piro, G.; Carbone, C.; Tortora, G.; Boldrini, L.; Pietragalla, A.; Daniele, G.; Tredicine, M.; Cesario, A.; et al. Translational Research in the Era of Precision Medicine: Where We Are and Where We Will Go. J. Pers. Med. 2021, 11, 216. https://doi.org/10.3390/jpm11030216
De Maria Marchiano R, Di Sante G, Piro G, Carbone C, Tortora G, Boldrini L, Pietragalla A, Daniele G, Tredicine M, Cesario A, et al. Translational Research in the Era of Precision Medicine: Where We Are and Where We Will Go. Journal of Personalized Medicine. 2021; 11(3):216. https://doi.org/10.3390/jpm11030216
Chicago/Turabian StyleDe Maria Marchiano, Ruggero, Gabriele Di Sante, Geny Piro, Carmine Carbone, Giampaolo Tortora, Luca Boldrini, Antonella Pietragalla, Gennaro Daniele, Maria Tredicine, Alfredo Cesario, and et al. 2021. "Translational Research in the Era of Precision Medicine: Where We Are and Where We Will Go" Journal of Personalized Medicine 11, no. 3: 216. https://doi.org/10.3390/jpm11030216
APA StyleDe Maria Marchiano, R., Di Sante, G., Piro, G., Carbone, C., Tortora, G., Boldrini, L., Pietragalla, A., Daniele, G., Tredicine, M., Cesario, A., Valentini, V., Gallo, D., Babini, G., D’Oria, M., & Scambia, G. (2021). Translational Research in the Era of Precision Medicine: Where We Are and Where We Will Go. Journal of Personalized Medicine, 11(3), 216. https://doi.org/10.3390/jpm11030216











