Estrogen Sulfotransferase (SULT1E1): Its Molecular Regulation, Polymorphisms, and Clinical Perspectives
Abstract
1. Introduction
2. Expression of SULT1E1
3. Sulfation of Estrogens and Thyroid Hormones by SULT1E1
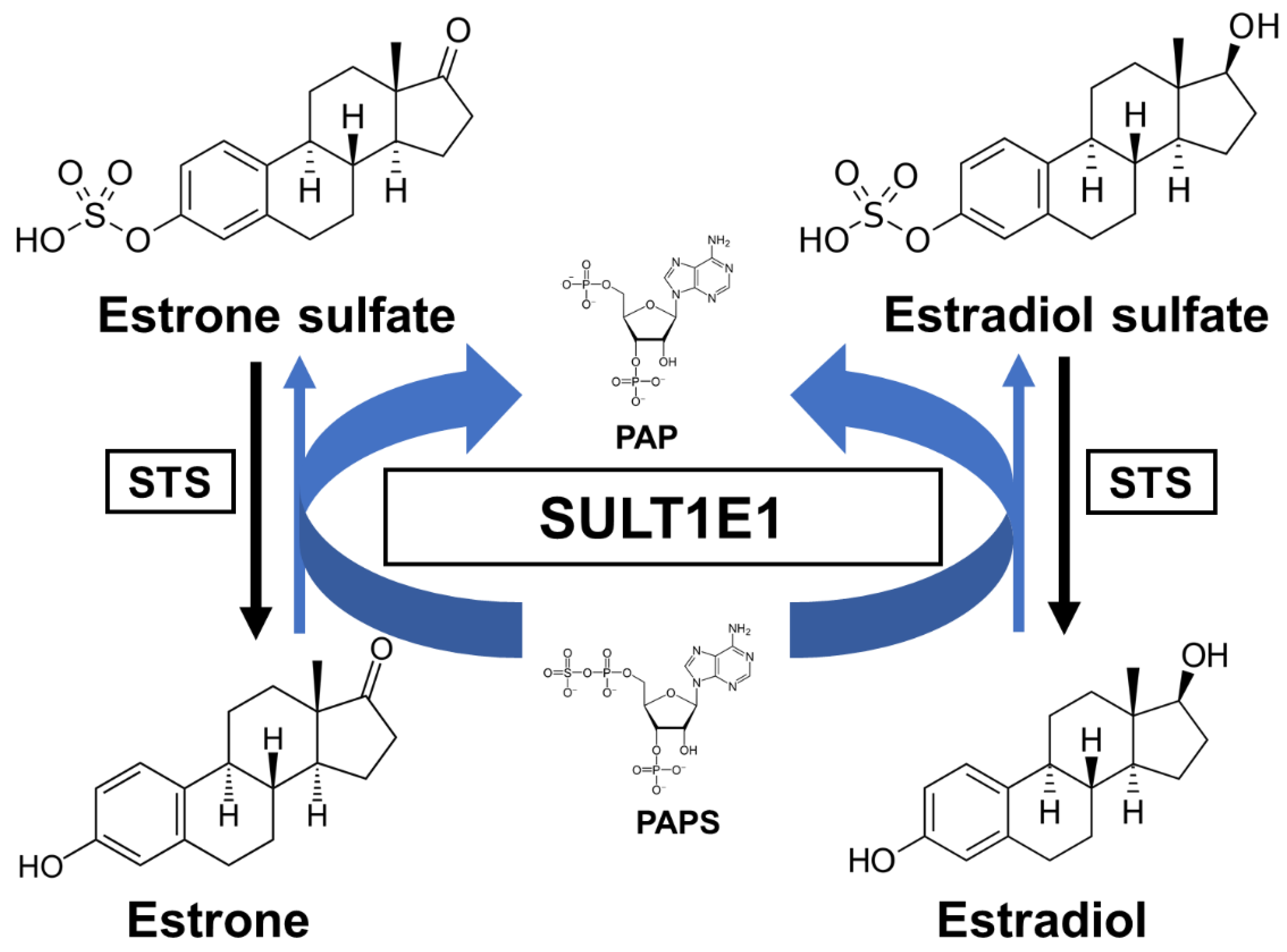
3.1. Sulfation of Estrogens
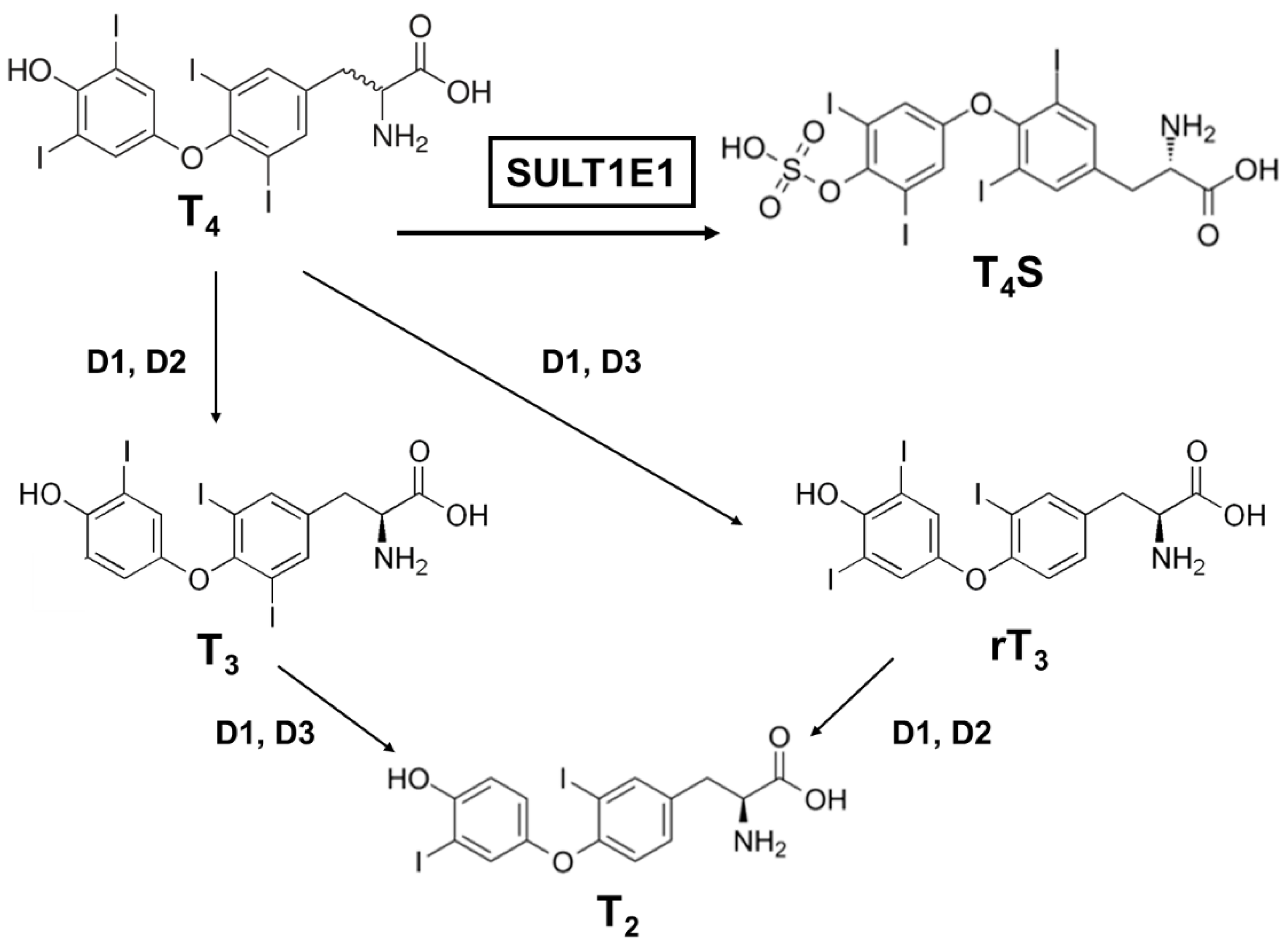
3.2. Sulfation of Thyroid Hormones
4. Sulfation of Other Substrates by SULT1E1
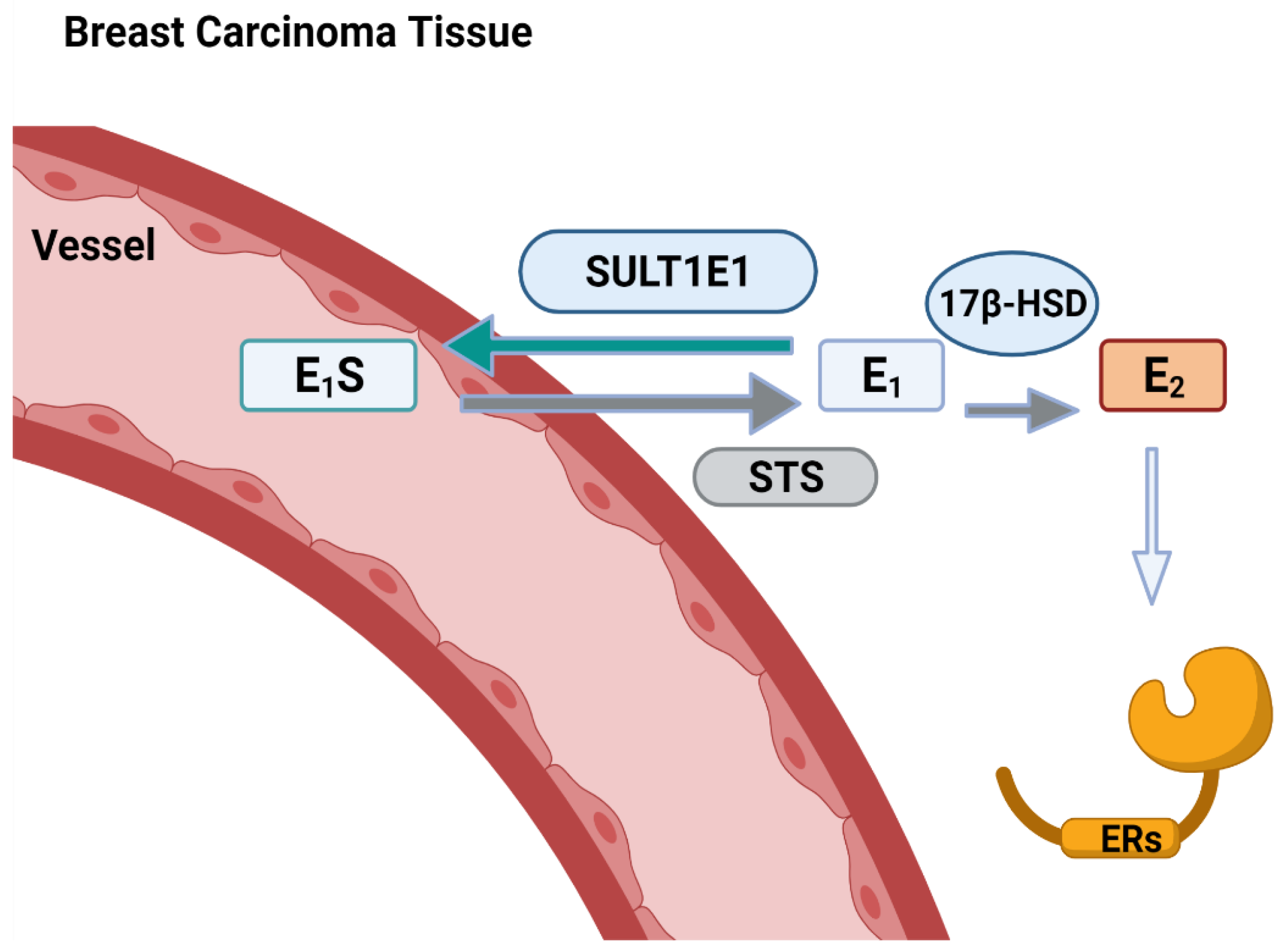
5. SULT1E1 and Diseases
6. Functional Variants of SULT1E1 and Current Research Status
| Type | Position 1 | SNP ID 2 | Effect | Reference |
|---|---|---|---|---|
| Intron | c.772+369T>C | rs3775777 | Treatment failure on abiraterone acetate with mCRPC | [83] |
| c.369+1930A>C | rs4149534 | |||
| c.369+402T>C | rs10019305 | |||
| c.-9-899G>A | rs3775770 | |||
| c.-10+771C>A | rs4149527 | |||
| c.-10+655G>A | rs3775768 | |||
| c.-9-469G>A | rs3822172 | Lower survival rate in colorectal cancer | [89] | |
| c.772+856G>T,C,A | rs1238574 | |||
| c.369+1653T>C | rs3775775 | Decreased survival rate from breast cancer | [84] | |
| 5′UTR | c.-64G>A | rs3736599 | Lower DHEA sulfate levels in the menopausal transition of European-American population | [85] |
| May strongly contribute to risk for endometrial carcinogenesis in Caucasians | [86] | |||
| Higher bone mineral density of distal radius and calcaneus in Korean women | [87] | |||
| Missense | 95C>T (Ala32Val) | rs34547148 | Increased Km value for the sulfation of E2 | [88] |
| 64G>A (Asp22Tyr) | rs11569705 |
7. Future Directions for Clinical Integrations
| Impacted Amino Acids | Substrate 1 | Alteration | SNP ID 2 |
|---|---|---|---|
| Arg256 | PAPS | Not reported | - |
| Phe254 | E2, 4-OH TCB | Phe254Cys | rs746067466 |
| Met247 | 4-OH TCB | Met247Ile | rs1188553969 |
| Ile246 | 4-OH TCB, TBBPA | Ile246Leu | rs1413235220 |
| Tyr239 | E2, 4-OH TCB | Not reported | - |
| Phe228 | PAPS | Not reported | - |
| Thr226 | PAPS | Thr226Ser | rs756363002 |
| Asn168 | 4-OH TCB, TBBPA | Asn168Ser | rs1265277815 |
| Val145 | 4-OH TCB | Val145Leu | rs200443686 |
| Phe141 | E2, 4-OH TCB, TBBPA, 3-OH BDE47 | Phe141Leu | rs1220949195 |
| Phe138 | TBBPA | Not reported | - |
| Ser137 | PAPS, E2 | Ser137Pro | rs1208507410 |
| Arg129 | PAPS | Arg129Gln | rs774700339 |
| His107 | PAPS, E2, 4-OH TCB, TBBPA | His107Arg | rs1316115370 |
| Lys105 | PAPS, E2, 4-OH TCB, TBBPA, 3-OH BDE47 | Not reported | - |
| Cys83 | 3-OH BDE47 | Cys83Phe | rs1431397129 |
| Phe80 | E2, 4-OH TCB, TBBPA, 3-OH BDE47 | Not reported | - |
| Trp52 | PAPS | Not reported | - |
| Thr51 | PAPS | Thr51Ile | rs1170826222 |
| Thr51Ala | rs761632873 | ||
| Thr50 | PAPS | Not reported | - |
| Gly49 | PAP | Gly49Val | rs1460190031 |
| Gly49Ser | rs1210226778 | ||
| Ser48 | PAP | Ser48Cys | rs1336407598 |
| Ser48Pro | rs1052854963 | ||
| Lys47 | PAPS, E2 | Lys47Glu | rs1361781887 |
| Pro46 | 4-OH TCB, TBBPA | Pro46Leu | rs771011878 |
| Phe23 | 4-OH TCB | Phe23Cys | rs1400776691 |
| Asp22 | 4-OH TCB | Asp22Asn | rs11569705 |
| Asp22Tyr | |||
| Tyr20 | PAP-E2, 4-OH TCB, TBBPA | Tyr20Cys | rs778407495 |
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gamage, N.; Barnett, A.; Hempel, N.; Duggleby, R.G.; Windmill, K.F.; Martin, J.L.; McManus, M.E. Human sulfotransferases and their role in chemical metabolism. Toxicol. Sci. 2006, 90, 5–22. [Google Scholar] [CrossRef]
- Liu, L.; Klaassen, C.D. Regulation of hepatic sulfotransferases by steroidal chemicals in rats. Drug Metab. Dispos. 1996, 24, 854–858. [Google Scholar] [PubMed]
- Sonoda, J.; Xie, W.; Rosenfeld, J.M.; Barwick, J.L.; Guzelian, P.S.; Evans, R.M. Regulation of a xenobiotic sulfonation cascade by nuclear pregnane X receptor (PXR). Proc. Natl. Acad. Sci. USA 2002, 99, 13801–13806. [Google Scholar] [CrossRef]
- Song, C.S.; Echchgadda, I.; Baek, B.S.; Ahn, S.C.; Oh, T.; Roy, A.K.; Chatterjee, B. Dehydroepiandrosterone sulfotransferase gene induction by bile acid activated farnesoid X receptor. J. Biol. Chem. 2001, 276, 42549–42556. [Google Scholar] [CrossRef]
- Qatanani, M.; Zhang, J.; Moore, D.D. Role of the constitutive androstane receptor in xenobiotic-induced thyroid hormone metabolism. Endocrinology 2005, 146, 995–1002. [Google Scholar] [CrossRef]
- Tien, E.S.; Matsui, K.; Moore, R.; Negishi, M. The nuclear receptor constitutively active/androstane receptor regulates type 1 deiodinase and thyroid hormone activity in the regenerating mouse liver. J. Pharmacol. Exp. Ther. 2007, 320, 307–313. [Google Scholar] [CrossRef]
- Falany, C.N. Enzymology of human cytosolic sulfotransferases. FASEB J. 1997, 11, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Klaassen, C.D.; Boles, J.W. Sulfation and sulfotransferases 5: The importance of 3′-phosphoadenosine 5′-phosphosulfate (PAPS) in the regulation of sulfation. FASEB J. 1997, 11, 404–418. [Google Scholar] [CrossRef]
- Kakuta, Y.; Pedersen, L.G.; Carter, C.W.; Negishi, M.; Pedersen, L.C. Crystal structure of estrogen sulphotransferase. Nat. Struct. Biol. 1997, 4, 904–908. [Google Scholar] [CrossRef] [PubMed]
- Kakuta, Y.; Petrotchenko, E.V.; Pedersen, L.C.; Negishi, M. The sulfuryl transfer mechanism. Crystal structure of a vanadate complex of estrogen sulfotransferase and mutational analysis. J. Biol. Chem. 1998, 273, 27325–27330. [Google Scholar] [CrossRef]
- Pedersen, L.C.; Petrotchenko, E.; Shevtsov, S.; Negishi, M. Crystal structure of the human estrogen sulfotransferase-PAPS complex: Evidence for catalytic role of Ser137 in the sulfuryl transfer reaction. J. Biol. Chem. 2002, 277, 17928–17932. [Google Scholar] [CrossRef] [PubMed]
- Shevtsov, S.; Petrotchenko, E.V.; Pedersen, L.C.; Negishi, M. Crystallographic analysis of a hydroxylated polychlorinated biphenyl (OH-PCB) bound to the catalytic estrogen binding site of human estrogen sulfotransferase. Environ. Health Perspect. 2003, 111, 884–888. [Google Scholar] [CrossRef] [PubMed]
- Allali-Hassani, A.; Pan, P.W.; Dombrovski, L.; Najmanovich, R.; Tempel, W.; Dong, A.; Loppnau, P.; Martin, F.; Thornton, J.; Edwards, A.M.; et al. Structural and chemical profiling of the human cytosolic sulfotransferases. PLoS Biol. 2007, 5, e97. [Google Scholar]
- Kodama, S.; Negishi, M. Sulfotransferase genes: Regulation by nuclear receptors in response to xeno/endo-biotics. Drug Metab. Rev. 2013, 45, 441–449. [Google Scholar] [CrossRef]
- Falany, C.N.; Krasnykh, V.; Falany, J.L. Bacterial expression and characterization of a cDNA for human liver estrogen sulfotransferase. J. Steroid Biochem. Mol. Biol. 1995, 52, 529–539. [Google Scholar] [CrossRef]
- Zhang, H.; Varlamova, O.; Vargas, F.M. Sulfuryl transfer: The catalytic mechanism of human estrogen sulfotransferase. J. Biol. Chem. 1998, 273, 10888–10892. [Google Scholar] [CrossRef]
- Faucher, F.; Lacoste, L.; Dufort, I.; Luu-The, V. High metabolization of catecholestrogens by type 1 estrogen sulfotransferase (hEST1). J. Steroid Biochem. Mol. Biol. 2001, 77, 83–86. [Google Scholar] [CrossRef]
- Faucher, F.; Lacoste, L.; Luu-The, V. Human type 1 estrogen sulfotransferase: Catecholestrogen metabolism and potential involvement in cancer promotion. Ann. N. Y. Acad. Sci. 2002, 963, 221–228. [Google Scholar] [CrossRef]
- Adjei, A.A.; Weinshilboum, R.M. Catecholestrogen sulfation: Possible role in carcinogenesis. Biochem. Biophys. Res. Commun. 2002, 292, 402–408. [Google Scholar] [CrossRef]
- Her, C.; Aksoy, I.A.; Kimura, S.; Brandriff, B.F.; Wasmuth, J.J.; Weinshilboum, R.M. Human estrogen sulfotransferase gene (STE): Cloning, structure, and chromosomal localization. Genomics 1995, 29, 16–23. [Google Scholar] [CrossRef]
- Stanley, E.L.; Hume, R.; Visser, T.J.; Coughtrie, M.W. Differential expression of sulfotransferase enzymes involved in thyroid hormone metabolism during human placental development. J. Clin. Endocrinol. Metab. 2001, 86, 5944–5955. [Google Scholar] [CrossRef]
- Rubin, G.L.; Harrold, A.J.; Mills, J.A.; Falany, C.N.; Coughtrie, M.W. Regulation of sulphotransferase expression in the endometrium during the menstrual cycle, by oral contraceptives and during early pregnancy. Mol. Hum. Reprod. 1999, 5, 995–1002. [Google Scholar] [CrossRef]
- Song, W.C.; Qian, Y.; Li, A.P. Estrogen sulfotransferase expression in the human liver: Marked interindividual variation and lack of gender specificity. J. Pharmacol. Exp. Ther. 1998, 284, 1197–1202. [Google Scholar]
- Li, Y.; Xu, Y.; Li, X.; Qin, Y.; Hu, R. Effects of PPAR-α agonist and IGF-1 on estrogen sulfotransferase in human vascular endothelial and smooth muscle cells. Mol. Med. Rep. 2013, 8, 133–139. [Google Scholar] [CrossRef]
- Xu, Y.; Yang, X.; Wang, Z.; Li, M.; Ning, Y.; Chen, S.; Yin, L.; Li, X. Estrogen sulfotransferase (SULT1E1) regulates inflammatory response and lipid metabolism of human endothelial cells via PPARγ. Mol. Cell Endocrinol. 2013, 369, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Yuan, X.; Lu, D.; Guo, L.; Wu, B. Farnesoid X receptor regulates SULT1E1 expression through inhibition of PGC1α binding to HNF4α. Biochem. Pharmacol. 2017, 145, 202–209. [Google Scholar] [CrossRef]
- Liu, X.; Xue, R.; Yang, C.; Gu, J.; Chen, S.; Zhang, S. Cholestasis-induced bile acid elevates estrogen level via farnesoid X receptor-mediated suppression of the estrogen sulfotransferase SULT1E1. J. Biol. Chem. 2018, 293, 12759–12769. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Negishi, M. RORα phosphorylation by casein kinase 1α as glucose signal to regulate estrogen sulfation in human liver cells. Biochem. J. 2020, 477, 3583–3598. [Google Scholar] [CrossRef]
- Gong, H.; Guo, P.; Zhai, Y.; Zhou, J.; Uppal, H.; Jarzynka, M.J.; Song, W.C.; Cheng, S.Y.; Xie, W. Estrogen deprivation and inhibition of breast cancer growth in vivo through activation of the orphan nuclear receptor liver X receptor. Mol. Endocrinol. 2007, 21, 1781–1790. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Jarzynka, M.J.; Cole, T.J.; Lee, J.H.; Wada, T.; Zhang, B.; Gao, J.; Song, W.C.; DeFranco, D.B.; Cheng, S.Y.; et al. Glucocorticoids antagonize estrogens by glucocorticoid receptor-mediated activation of estrogen sulfotransferase. Cancer Res. 2008, 68, 7386–7393. [Google Scholar] [CrossRef]
- Sueyoshi, T.; Green, W.D.; Vinal, K.; Woodrum, T.S.; Moore, R.; Negishi, M. Garlic extract diallyl sulfide (DAS) activates nuclear receptor CAR to induce the Sult1e1 gene in mouse liver. PLoS ONE 2011, 6, e21229. [Google Scholar] [CrossRef]
- Yi, M.; Fashe, M.; Arakawa, S.; Moore, R.; Sueyoshi, T.; Negishi, M. Nuclear receptor CAR-ERalpha signaling regulates the estrogen sulfotransferase gene in the liver. Sci. Rep. 2020, 10, 5001. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Yokobori, K.; Negishi, M. PXR phosphorylated at Ser350 transduces a glucose signal to repress the estrogen sulfotransferase gene in human liver cells and fasting signal in mouse livers. Biochem. Pharmacol. 2020, 180, 114197. [Google Scholar] [CrossRef]
- Fashe, M.; Hashiguchi, T.; Yi, M.; Moore, R.; Negishi, M. Phenobarbital-induced phosphorylation converts nuclear receptor RORalpha from a repressor to an activator of the estrogen sulfotransferase gene Sult1e1 in mouse livers. FEBS Lett. 2018, 592, 2760–2768. [Google Scholar] [CrossRef] [PubMed]
- Poisson Paré, D.; Song, D.; Luu-The, V.; Han, B.; Li, S.; Liu, G.; Labrie, F.; Pelletier, G. Expression of Estrogen Sulfotransferase 1E1 and Steroid Sulfatase in Breast Cancer: A Immunohistochemical Study. Breast Cancer 2009, 3, 9–21. [Google Scholar]
- Song, W.C.; Melner, M.H. Steroid transformation enzymes as critical regulators of steroid action in vivo. Endocrinology 2000, 141, 1587–1589. [Google Scholar] [CrossRef][Green Version]
- Falany, C.N.; Wheeler, J.; Oh, T.S.; Falany, J.L. Steroid sulfation by expressed human cytosolic sulfotransferases. J. Steroid Biochem. Mol. Biol. 1994, 48, 369–375. [Google Scholar] [CrossRef]
- Falany, C.N.; Comer, K.A.; Dooley, T.P.; Glatt, H. Human dehydroepiandrosterone sulfotransferase. Purification, molecular cloning, and characterization. Ann. N. Y. Acad. Sci. 1995, 774, 59–72. [Google Scholar] [CrossRef]
- Aksoy, I.A.; Wood, T.C.; Weinshilboum, R. Human liver estrogen sulfotransferase: Identification by cDNA cloning and expression. Biochem. Biophys. Res. Commun. 1994, 200, 1621–1629. [Google Scholar] [CrossRef]
- Schrag, M.L.; Cui, D.; Rushmore, T.H.; Shou, M.; Ma, B.; Rodrigues, A.D. Sulfotransferase 1E1 is a low km isoform mediating the 3-O-sulfation of ethinyl estradiol. Drug Metab. Dispos. 2004, 32, 1299–1303. [Google Scholar] [CrossRef]
- Kester, M.H.; van Dijk, C.H.; Tibboel, D.; Hood, A.M.; Rose, N.J.; Meinl, W.; Pabel, U.; Glatt, H.; Falany, C.N.; Coughtrie, M.W.; et al. Sulfation of thyroid hormone by estrogen sulfotransferase. J. Clin. Endocrinol. Metab. 1999, 84, 2577–2580. [Google Scholar] [CrossRef] [PubMed]
- Ung, D.; Nagar, S. Variable sulfation of dietary polyphenols by recombinant human sulfotransferase (SULT) 1A1 genetic variants and SULT1E1. Drug Metab. Dispos. 2007, 35, 740–746. [Google Scholar] [CrossRef] [PubMed]
- Edavana, V.K.; Yu, X.; Dhakal, I.B.; Williams, S.; Ning, B.; Cook, I.T.; Caldwell, D.; Falany, C.N.; Kadlubar, S. Sulfation of fulvestrant by human liver cytosols and recombinant SULT1A1 and SULT1E1. Pharmgenom. Pers. Med. 2011, 4, 137–145. [Google Scholar]
- Edavana, V.K.; Dhakal, I.B.; Yu, X.; Williams, S.; Kadlubar, S. Sulfation of 4-hydroxy toremifene: Individual variability, isoform specificity, and contribution to toremifene pharmacogenomics. Drug Metab. Dispos. 2012, 40, 1210–1215. [Google Scholar] [CrossRef] [PubMed]
- Honma, W.; Shimada, M.; Sasano, H.; Ozawa, S.; Miyata, M.; Nagata, K.; Ikeda, T.; Yamazoe, Y. Phenol sulfotransferase, ST1A3, as the main enzyme catalyzing sulfation of troglitazone in human liver. Drug Metab. Dispos. 2002, 30, 944–949. [Google Scholar] [CrossRef]
- Squirewell, E.J.; Duffel, M.W. The effects of endoxifen and other major metabolites of tamoxifen on the sulfation of estradiol catalyzed by human cytosolic sulfotransferases hSULT1E1 and hSULT1A1*1. Drug Metab. Dispos. 2015, 43, 843–850. [Google Scholar] [CrossRef]
- Falany, J.L.; Macrina, N.; Falany, C.N. Sulfation of tibolone and tibolone metabolites by expressed human cytosolic sulfotransferases. J. Steroid Biochem. Mol. Biol. 2004, 88, 383–391. [Google Scholar] [CrossRef]
- Buirchell, B.J.; Hahnel, R. Metabolism of estradiol-17beta in human endometrium during the menstrual cycle. J. Steroid Biochem. 1975, 6, 1489–1494. [Google Scholar] [CrossRef]
- Falany, J.L.; Azziz, R.; Falany, C.N. Identification and characterization of cytosolic sulfotransferases in normal human endometrium. Chem. Biol. Interact. 1998, 109, 329–339. [Google Scholar] [CrossRef]
- Kotov, A.; Falany, J.L.; Wang, J.; Falany, C.N. Regulation of estrogen activity by sulfation in human Ishikawa endometrial adenocarcinoma cells. J. Steroid Biochem. Mol. Biol. 1999, 68, 137–144. [Google Scholar] [CrossRef]
- Falany, J.L.; Falany, C.N. Regulation of estrogen sulfotransferase in human endometrial adenocarcinoma cells by progesterone. Endocrinology 1996, 137, 1395–1401. [Google Scholar] [CrossRef][Green Version]
- Moreau, X.; Lejeune, P.J.; Jeanningros, R. Kinetics of red blood cell T3 uptake in hypothyroidism with or without hormonal replacement, in the rat. J. Endocrinol. Investig. 1999, 22, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.Y.; Seo, S.; Choi, K.H.; Yun, J. Evaluation of phototoxicity of tattoo pigments using the 3 T3 neutral red uptake phototoxicity test and a 3D human reconstructed skin model. Toxicol. In Vitro 2020, 65, 104813. [Google Scholar] [CrossRef] [PubMed]
- Leiner, K.A.; Mackenzie, D.S. Central regulation of thyroidal status in a teleost fish: Nutrient stimulation of T4 secretion and negative feedback of T3. J. Exp. Zool. Comp. Exp. Biol. 2003, 298, 32–43. [Google Scholar] [CrossRef] [PubMed]
- Mihasan, M.; Brandsch, R. A predicted T4 secretion system and conserved DNA-repeats identified in a subset of related Arthrobacter plasmids. Microbiol. Res. 2016, 191, 32–37. [Google Scholar] [CrossRef]
- St Germain, D.L.; Galton, V.A.; Hernandez, A. Minireview: Defining the roles of the iodothyronine deiodinases: Current concepts and challenges. Endocrinology 2009, 150, 1097–1107. [Google Scholar] [CrossRef]
- Wu, S.Y.; Huang, W.S.; Polk, D.; Florsheim, W.H.; Green, W.L.; Fisher, D.A. Identification of thyroxine-sulfate (T4S) in human serum and amniotic fluid by a novel T4S radioimmunoassay. Thyroid 1992, 2, 101–105. [Google Scholar] [CrossRef]
- Chatterjee, B.; Song, C.S.; Kim, J.M.; Roy, A.K. Androgen and estrogen sulfotransferases of the rat liver: Physiological function, molecular cloning, and in vitro expression. Chem. Biol. Interact. 1994, 92, 273–279. [Google Scholar] [CrossRef]
- Kester, M.H.; Kaptein, E.; Roest, T.J.; van Dijk, C.H.; Tibboel, D.; Meinl, W.; Glatt, H.; Coughtrie, M.W.; Visser, T.J. Characterization of human iodothyronine sulfotransferases. J. Clin. Endocrinol. Metab. 1999, 84, 1357–1364. [Google Scholar] [CrossRef]
- Wang, J.; Falany, J.L.; Falany, C.N. Expression and characterization of a novel thyroid hormone-sulfating form of cytosolic sulfotransferase from human liver. Mol. Pharmacol. 1998, 53, 274–282. [Google Scholar] [CrossRef]
- Li, X.; Clemens, D.L.; Anderson, R.J. Sulfation of iodothyronines by human sulfotransferase 1C1 (SULT1C1). Biochem. Pharmacol. 2000, 60, 1713–1716. [Google Scholar] [CrossRef]
- Kester, M.; Coughtrie, M.W.; Visser, T.J. Sulfation of Thyroid Hormones. In Human Cytosolic Sulfotransferases; Pacifici, G.M., Coughtrie, M.W., Eds.; CRC Press/Taylor & Francis Group: Boca Raton, FL, USA, 2005; pp. 121–134. [Google Scholar]
- Glatt, H.; Boeing, H.; Engelke, C.E.; Ma, L.; Kuhlow, A.; Pabel, U.; Pomplun, D.; Teubner, W.; Meinl, W. Human cytosolic sulphotransferases: Genetics, characteristics, toxicological aspects. Mutat. Res. 2001, 482, 27–40. [Google Scholar] [CrossRef]
- Weinshilboum, R.M.; Otterness, D.M.; Aksoy, I.A.; Wood, T.C.; Her, C.; Raftogianis, R.B. Sulfation and sulfotransferases 1: Sulfotransferase molecular biology: cDNAs and genes. FASEB J. 1997, 11, 3–14. [Google Scholar] [CrossRef]
- Kiehlbauch, C.C.; Lam, Y.F.; Ringer, D.P. Homodimeric and heterodimeric aryl sulfotransferases catalyze the sulfuric acid esterification of N-hydroxy-2-acetylaminofluorene. J. Biol. Chem. 1995, 270, 18941–18947. [Google Scholar] [CrossRef] [PubMed]
- Salehi, B.; Venditti, A.; Sharifi-Rad, M.; Kregiel, D.; Sharifi-Rad, J.; Durazzo, A.; Lucarini, M.; Santini, A.; Souto, E.B.; Novellino, E.; et al. The Therapeutic Potential of Apigenin. Int. J. Mol. Sci. 2019, 20, 1305. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.; de Bruijn, W.J.C.; van Zadelhoff, A.; Lin, Z.; Vincken, J.P. Browning of Epicatechin (EC) and Epigallocatechin (EGC) by Auto-Oxidation. J. Agric. Food Chem. 2020, 68, 13879–13887. [Google Scholar] [CrossRef] [PubMed]
- Salehi, B.; Mishra, A.P.; Nigam, M.; Sener, B.; Kilic, M.; Sharifi-Rad, M.; Fokou, P.V.T.; Martins, N.; Sharifi-Rad, J. Resveratrol: A Double-Edged Sword in Health Benefits. Biomedicines 2018, 6, 91. [Google Scholar] [CrossRef]
- Jana, K.; Yin, X.; Schiffer, R.B.; Chen, J.J.; Pandey, A.K.; Stocco, D.M.; Grammas, P.; Wang, X. Chrysin, a natural flavonoid enhances steroidogenesis and steroidogenic acute regulatory protein gene expression in mouse Leydig cells. J. Endocrinol. 2008, 197, 315–323. [Google Scholar] [CrossRef]
- Li, Y.; Yao, J.; Han, C.; Yang, J.; Chaudhry, M.T.; Wang, S.; Liu, H.; Yin, Y. Quercetin, Inflammation and Immunity. Nutrients 2016, 8, 167. [Google Scholar] [CrossRef]
- Pasqualini, J.R.; Chetrite, G.; Blacker, C.; Feinstein, M.C.; Delalonde, L.; Talbi, M.; Maloche, C. Concentrations of estrone, estradiol, and estrone sulfate and evaluation of sulfatase and aromatase activities in pre- and postmenopausal breast cancer patients. J. Clin. Endocrinol. Metab. 1996, 81, 1460–1464. [Google Scholar]
- Mady, E.A.; Ramadan, E.E.; Ossman, A.A. Sex steroid hormones in serum and tissue of benign and malignant breast tumor patients. Dis. Markers 2000, 16, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Bonorden, M.J.; Greany, K.A.; Wangen, K.E.; Phipps, W.R.; Feirtag, J.; Adlercreutz, H.; Kurzer, M.S. Consumption of Lactobacillus acidophilus and Bifidobacterium longum do not alter urinary equol excretion and plasma reproductive hormones in premenopausal women. Eur. J. Clin. Nutr. 2004, 58, 1635–1642. [Google Scholar] [CrossRef]
- Thomas, H.V.; Key, T.J.; Allen, D.S.; Moore, J.W.; Dowsett, M.; Fentiman, I.S.; Wang, D.Y. A prospective study of endogenous serum hormone concentrations and breast cancer risk in post-menopausal women on the island of Guernsey. Br. J. Cancer 1997, 76, 401–405. [Google Scholar] [CrossRef]
- Pasqualini, J.R.; Cortes-Prieto, J.; Chetrite, G.; Talbi, M.; Ruiz, A. Concentrations of estrone, estradiol and their sulfates, and evaluation of sulfatase and aromatase activities in patients with breast fibroadenoma. Int. J. Cancer 1997, 70, 639–643. [Google Scholar] [CrossRef]
- Falany, J.L.; Falany, C.N. Expression of cytosolic sulfotransferases in normal mammary epithelial cells and breast cancer cell lines. Cancer Res. 1996, 56, 1551–1555. [Google Scholar] [PubMed]
- Chetrite, G.S.; Paris, J.; Shields-Botella, J.; Philippe, J.C.; Pasqualini, J.R. Effect of nomegestrol acetate on human estrogen sulfotransferase activity in the hormone-dependent MCF-7 and T-47D breast cancer cell lines. Anticancer Res. 2003, 23, 4651–4655. [Google Scholar]
- Qian, Y.; Deng, C.; Song, W.C. Expression of estrogen sulfotransferase in MCF-7 cells by cDNA transfection suppresses the estrogen response: Potential role of the enzyme in regulating estrogen-dependent growth of breast epithelial cells. J. Pharmacol. Exp. Ther. 1998, 286, 555–560. [Google Scholar]
- Qian, Y.M.; Sun, X.J.; Tong, M.H.; Li, X.P.; Richa, J.; Song, W.C. Targeted disruption of the mouse estrogen sulfotransferase gene reveals a role of estrogen metabolism in intracrine and paracrine estrogen regulation. Endocrinology 2001, 142, 5342–5350. [Google Scholar] [CrossRef] [PubMed]
- Chai, X.; Guo, Y.; Jiang, M.; Hu, B.; Li, Z.; Fan, J.; Deng, M.; Billiar, T.R.; Kucera, H.R.; Gaikwad, N.W.; et al. Oestrogen sulfotransferase ablation sensitizes mice to sepsis. Nat. Commun. 2015, 6, 7979. [Google Scholar] [CrossRef]
- Leiter, E.H.; Chapman, H.D. Obesity-induced diabetes (diabesity) in C57BL/KsJ mice produces aberrant trans-regulation of sex steroid sulfotransferase genes. J. Clin. Investig. 1994, 93, 2007–2013. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; He, J.; Shi, X.; Stefanovic-Racic, M.; Xu, M.; O’Doherty, R.M.; Garcia-Ocana, A.; Xie, W. Sex-specific effect of estrogen sulfotransferase on mouse models of type 2 diabetes. Diabetes 2012, 61, 1543–1551. [Google Scholar] [CrossRef]
- Agarwal, N.; Alex, A.B.; Farnham, J.M.; Patel, S.; Gill, D.; Buckley, T.H.; Stephenson, R.A.; Cannon-Albright, L. Inherited variants in SULT1E1 and response to abiraterone acetate by men with metastatic castration refractory prostate cancer. J. Urol. 2016, 196, 1112–1116. [Google Scholar] [CrossRef]
- Choi, J.Y.; Lee, K.M.; Park, S.K.; Noh, D.Y.; Ahn, S.H.; Chung, H.W.; Han, W.; Kim, J.S.; Shin, S.G.; Jang, I.J.; et al. Genetic polymorphisms of SULT1A1 and SULT1E1 and the risk and survival of breast cancer. Cancer Epidemiol. Biomark. Prev. 2005, 14, 1090–1095. [Google Scholar] [CrossRef] [PubMed]
- Rebbeck, T.R.; Su, H.I.; Sammel, M.D.; Lin, H.; Tran, T.V.; Gracia, C.R.; Freeman, E.W. Effect of hormone metabolism genotypes on steroid hormone levels and menopausal symptoms in a prospective population-based cohort of women experiencing the menopausal transition. Menopause 2010, 17, 1026–1034. [Google Scholar] [CrossRef][Green Version]
- Hirata, H.; Hinoda, Y.; Okayama, N.; Suehiro, Y.; Kawamoto, K.; Kikuno, N.; Rabban, J.T.; Chen, L.M.; Dahiya, R. CYP1A1, SULT1A1, and SULT1E1 polymorphisms are risk factors for endometrial cancer susceptibility. Cancer 2008, 112, 1964–1973. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.A.; Choi, J.Y.; Shin, C.S.; Hong, Y.C.; Chung, H.; Kang, D. SULT1E1 genetic polymorphisms modified the association between phytoestrogen consumption and bone mineral density in healthy Korean women. Calcif. Tissue Int. 2006, 79, 152–159. [Google Scholar] [CrossRef] [PubMed]
- Adjei, A.A.; Thomae, B.A.; Prondzinski, J.L.; Eckloff, B.W.; Wieben, E.D.; Weinshilboum, R.M. Human estrogen sulfotransferase (SULT1E1) pharmacogenomics: Gene resequencing and functional genomics. Br. J. Pharmacol. 2003, 139, 1373–1382. [Google Scholar] [CrossRef]
- Li, S.; Xie, L.; Du, M.; Xu, K.; Zhu, L.; Chu, H.; Chen, J.; Wang, M.; Zhang, Z.; Gu, D. Association study of genetic variants in estrogen metabolic pathway genes and colorectal cancer risk and survival. Arch. Toxicol. 2018, 92, 1991–1999. [Google Scholar] [CrossRef]
- Collin, L.J.; Cronin-Fenton, D.P.; Ahern, T.P.; Christiansen, P.M.; Damkier, P.; Ejlertsen, B.; Hamilton-Dutoit, S.; Kjærsgaard, A.; Silliman, R.A.; Sørensen, H.T.; et al. Cohort Profile: The Predictors of Breast Cancer Recurrence (ProBe CaRE) Premenopausal Breast Cancer Cohort Study in Denmark. BMJ Open 2018, 8, e021805. [Google Scholar] [CrossRef]
- Wei, Z.; Li, M.; Rebbeck, T.; Li, H. U-statistics-based tests for multiple genes in genetic association studies. Ann. Hum. Genet. 2008, 72, 821–833. [Google Scholar] [CrossRef]
- Hoh, B.P.; Deng, L.; Julia-Ashazila, M.J.; Zuraihan, Z.; Nur-Hasnah, M.; Nur-Shafawati, A.R.; Hatin, W.I.; Endom, I.; Zilfalil, B.A.; Khalid, Y.; et al. Fine-scale population structure of Malays in Peninsular Malaysia and Singapore and implications for association studies. Hum. Genom. 2015, 9, 16. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gosavi, R.A.; Knudsen, G.A.; Birnbaum, L.S.; Pedersen, L.C. Mimicking of estradiol binding by flame retardants and their metabolites: A crystallographic analysis. Environ. Health Perspect. 2013, 121, 1194–1199. [Google Scholar] [CrossRef] [PubMed]
| Gene ID 1 | Locus 2 | Alias 1 | Number of Amino Acids 3 | Number of Exons 1 |
|---|---|---|---|---|
| SULT1A1 | Chr 16p11.2 | HAST1/HAST2, P-PST, PST, ST1A1, ST1A3, STP, STP1, TSPST1 | 295 (isoform a) | 13 |
| 217 (isoform b) | ||||
| SULT1A2 | Chr 16p11.2 | HAST4, P-PST, P-PST 2, ST1A2, STP2, TSPST2 | 295 (isoform 1) | 8 |
| 262 (isoform 2) | ||||
| SULT1A3 | Chr 16p11.2 | HAST, HAST3, M-PST, ST1A3, ST1A3/ST1A4, ST1A4, ST1A5, STM, TL-PST | 295 | 8 |
| SULT1A4 | Chr 16p11.2 | HAST3, M-PST, ST1A3, ST1A3/ST1A4, ST1A4, STM, TL-PST | 295 | 8 |
| SULT1B1 | Chr 4q13.3 | ST1B1, ST1B2, SULT1B2 | 296 | 10 |
| SULT1C2 | Chr 2q12.3 | ST1C1, ST1C2, SULT1C1, humSULTC2 | 296 (isoform a) | 9 |
| 307 (isoform b) | ||||
| SULT1C3 | Chr 2q12.3 | ST1C3 | 304 (isoform 1) | 10 |
| 304 (isoform 2) | ||||
| SULT1C4 | Chr 2q12.3 | SULT1C, SULT1C2 | 302 (isoform 1) | 7 |
| 227 (isoform 2) | ||||
| SULT1E1 | Chr 4q13.3 | EST, EST-1, ST1E1, STE | 294 | 9 |
| SULT2A1 | Chr 19q13.33 | DHEA-ST, DHEA-ST8, DHEAS, HST, ST2, ST2A1, ST2A3, STD, SULT2A3, hSTa | 285 | 6 |
| SULT2B1 | Chr 19q13.33 | ARCI14, HSST2 | 350 (isoform a) | 7 |
| 365 (isoform b) | ||||
| SULT4A1 | Chr 22q13.31 | BR-STL-1, BRSTL1, DJ388M5.3, NST, SULTX3, hBR-STL-1 | 284 | 11 |
| SULT6B1 | Chr 2p22.2 | ST6B1 | 304 (isoform 1) | 9 |
| 265 (isoform 2) | ||||
| SUPl1C2P1 | Chr 2q12.3 | SULT1C1P | pseudogene | 4 |
| SULT1C2P2 | Chr 2q12.3 | pseudogene | ||
| SULT1D1P | Chr 4q13.3 | SULT1D1 | pseudogene | |
| SULT6B2P | Chr 12p12.1 | pseudogene |
| Species | RefSeq 1 | RefSeq mRNA 2 | RefSeq Protein 3 | Number of Exons 1 |
|---|---|---|---|---|
| Homo sapiens (human) | NC_000004.12 | NM_005420.3 | NP_005411.1 | 9 |
| Mus musculus (mouse) | NC_000071.7 | NM_023135.2 | NP_075624.2 | 8 |
| Rattus norvegicus (rat) | NC_005113.4 | NM_012883.2 | NP_037015.2 | 10 |
| Bos taurus (cow) | NC_037333.1 | NM_177488.3 | NP_803454.2 | 9 |
| Oryctolagus cuniculus (rabbit) | NC_013683.1 | XM_002717123.2 | XP_002717169.1 | 8 |
| Sus scrofa (pig) | NC_010450.4 | NM_213992.1 | NP_999157.1 | 9 |
| Equus caballus (horse) | NC_009146.3 | NM_001081918.1 | NP_001075387.1 | 8 |
| Gene ID | Nuclear Receptor | Species | Tissue | Reference |
|---|---|---|---|---|
| NR3A1 | ERα | Mouse | Liver tissue | [32] |
| NR3C1 | GR | Mouse | Liver tissue | [30] |
| NR1C1 | PPARα | Human | Vascular endothelial cell | [24] |
| Smooth muscle cell | ||||
| NR1C3 | PPARγ | Human | Endothelial cell | [25] |
| NR1H2, H3 | LXR | Mouse | Uterine | [29] |
| NR1H4 | FXR | Human | Liver cell line | [26] |
| Human | Liver tissue | [27] | ||
| NR1I2 | PXR | Human | Liver cell line | [33] |
| Mouse | Liver tissue | |||
| NR1I3 | CAR | Mouse | Liver tissue | [31,32] |
| NR2A1 | HNF4α | Human | Liver tissue | [27] |
| NR1F1 | RORα | Human | Liver cell line | [28] |
| Mouse | Liver tissue | [34] |
| Substrate | Compound Characteristics | Km | Reference |
|---|---|---|---|
| E1 | Agonist of ER | ~0.17 µM | [39] |
| E2 | Most active agonist of ER | 5 ± 0.8 nM | [16] |
| 29 nM | [38] | ||
| EE2 | Agonist of GPER and ER | 6.7 ± 0.1 nM | [40] |
| DHEA | Partial agonist of AR and ER | ~0.85 µM | [37] |
| 4.57 ± 0.07 µM | [40] | ||
| T4 | Thyroid prohormone | 22.6 ± 1.0 µM | [41] |
| T3 | Receptor active iodothyronine | 25.7 ± 10.4 µM | |
| rT3 | Receptor inactive iodothyronine | 2.15 ± 1.45 µM | |
| T2 | Breakdown metabolite of triiodothyronine | 4.75 ± 1.25 µM | |
| Apigenin | Common dietary flavonoid | 5.3 ± 0.65 µM | [42] |
| Epicatechin | Antioxidative flavonoid | 0.96 ± 0.17 mM | |
| Resveratrol | Antioxidative flavonoid | 6.88 ± 1.12 µM | |
| Chrysin | Flavonoid in bee pollen or propolis | 4.5 ± 0.65 µM | |
| Quercetin | Flavonoid in plants or fruits | 2.0 ± 0.34 µM | |
| Fulvestrant | Steroidal ER antagonist | 0.2 ± 0.02 µM | [43] |
| 4-OH-TOR | Hydroxy metabolite of TOR (nonsteroidal agonist-antagonist of ER) | 6.4 ± 0.09 µM | [44] |
| Troglitazone | PPAR agonist | 8.5 ± 0.44 µM | [45] |
| Endoxifen | Active metabolite of Tamoxifen (nonsteroidal antagonist of ER) | 24 ± 5 µM | [46] |
| 4-OH TAM | Hydroxy metabolite of Tamoxifen | 24 ± 5 µM | |
| N-des TAM | N-demethyl metabolite of Tamoxifen | 96 ± 52 µM | |
| Tibolone | Selective tissue estrogenic activity regulator | 19.5 ± 2.8 µM | [47] |
| 3α-OH-TIB | Hydroxy metabolite of TIB | 6.6 ± 2.2 µM | |
| 3β-OH-TIB | Hydroxy metabolite of TIB | 2.1 ± 0.5 µM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yi, M.; Negishi, M.; Lee, S.-J. Estrogen Sulfotransferase (SULT1E1): Its Molecular Regulation, Polymorphisms, and Clinical Perspectives. J. Pers. Med. 2021, 11, 194. https://doi.org/10.3390/jpm11030194
Yi M, Negishi M, Lee S-J. Estrogen Sulfotransferase (SULT1E1): Its Molecular Regulation, Polymorphisms, and Clinical Perspectives. Journal of Personalized Medicine. 2021; 11(3):194. https://doi.org/10.3390/jpm11030194
Chicago/Turabian StyleYi, MyeongJin, Masahiko Negishi, and Su-Jun Lee. 2021. "Estrogen Sulfotransferase (SULT1E1): Its Molecular Regulation, Polymorphisms, and Clinical Perspectives" Journal of Personalized Medicine 11, no. 3: 194. https://doi.org/10.3390/jpm11030194
APA StyleYi, M., Negishi, M., & Lee, S.-J. (2021). Estrogen Sulfotransferase (SULT1E1): Its Molecular Regulation, Polymorphisms, and Clinical Perspectives. Journal of Personalized Medicine, 11(3), 194. https://doi.org/10.3390/jpm11030194







