Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Setting
2.3. Inclusion and Exclusion Criteria
2.4. Recruitment
2.4.1. Living Patients
2.4.2. Personal Representatives
2.4.3. Informed Consent
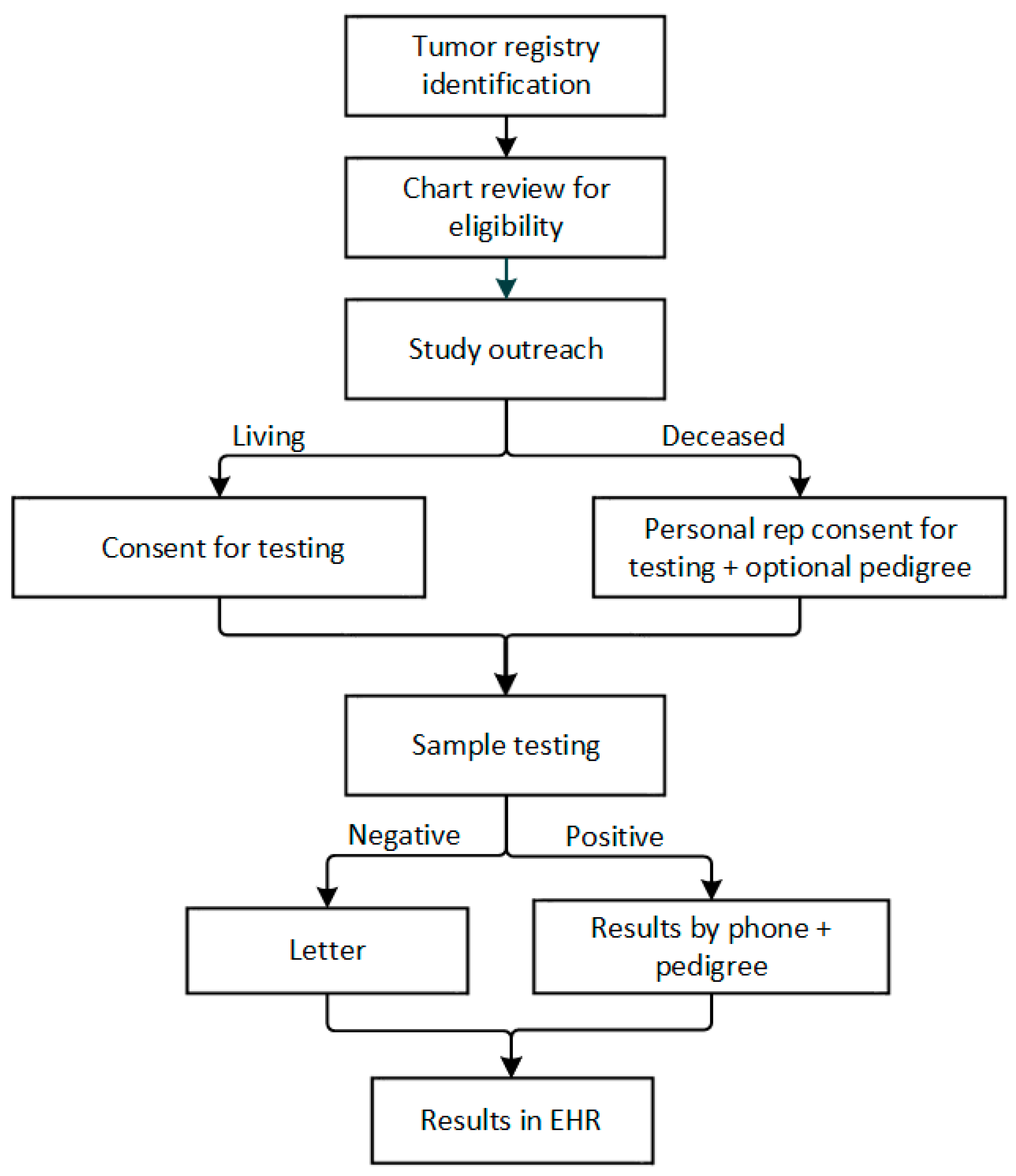
2.5. Study Procedures
2.5.1. Genetic testing
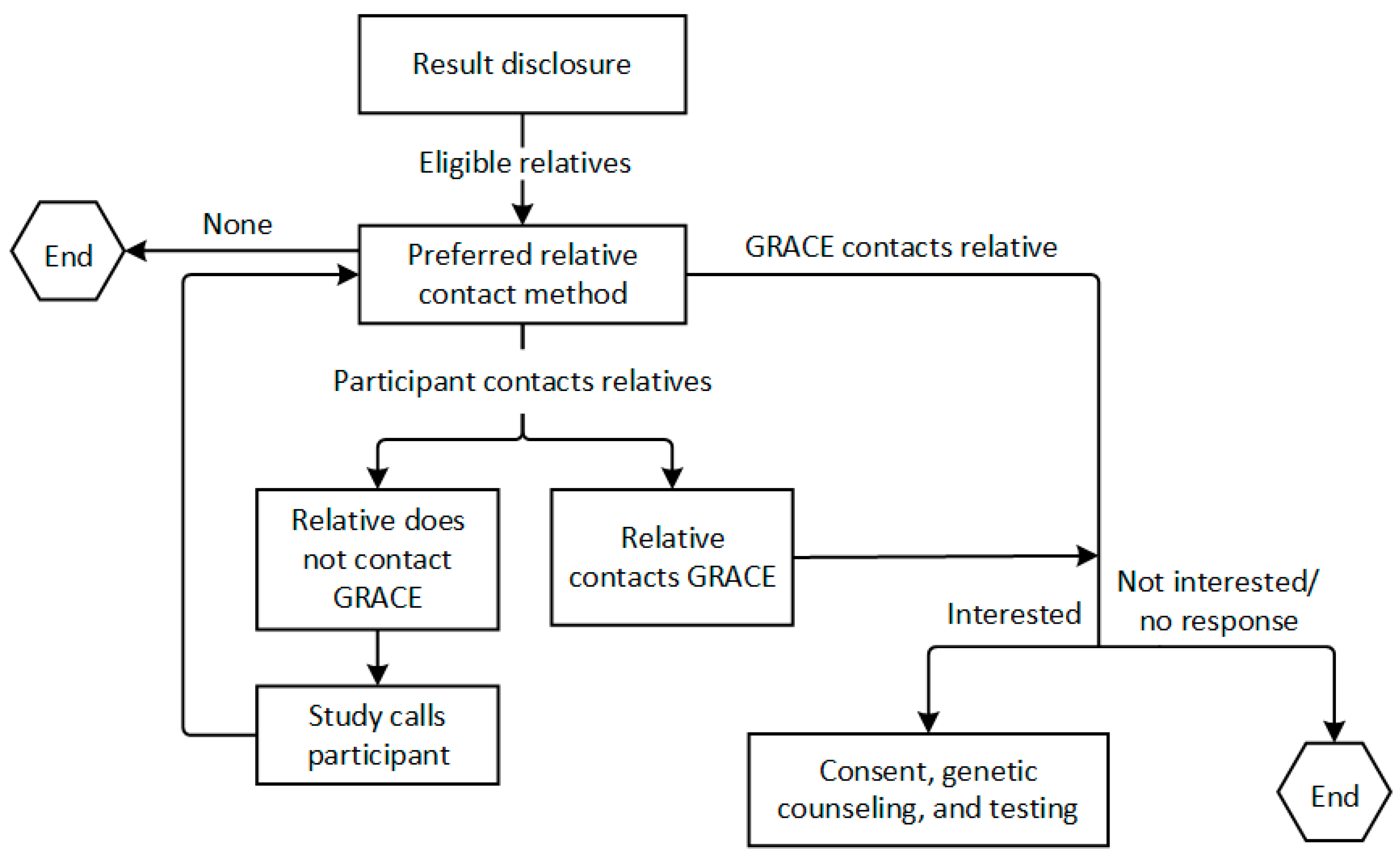
2.5.2. Result Disclosure
2.5.3. Cascade Testing
2.5.4. Data Collection
2.5.5. Interviews
2.6. Data Analysis
2.6.1. Evaluate the Feasibility of Ovarian Cancer Traceback Testing
2.6.2. Stakeholder Perspectives
3. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- American Cancer Society. Breast Cancer Facts & Figures 2015–2016. Available online: https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-statistics/breast-cancer-facts-and-figures/breast-cancer-facts-and-figures-2015-2016 (accessed on 16 January 2019).
- Norquist, B.M.; Harrell, M.I.; Brady, M.F.; Walsh, T.; Lee, M.K.; Gulsuner, S.; Bernards, S.S.; Casadei, S.; Yi, Q.; Burger, R.A.; et al. Inherited Mutations in Women with Ovarian Carcinoma. JAMA Oncol. 2016, 2, 482–490. [Google Scholar] [CrossRef] [PubMed]
- Alsop, K.; Fereday, S.; Meldrum, C.; deFazio, A.; Emmanuel, C.; George, J.; Dobrovic, A.; Birrer, M.J.; Webb, P.M.; Stewart, C.; et al. BRCA Mutation Frequency and Patterns of Treatment Response in BRCA Mutation-Positive Women with Ovarian Cancer: A Report from the Australian Ovarian Cancer Study Group. J. Clin. Oncol. 2012, 30, 2654–2663. [Google Scholar] [CrossRef]
- Zhang, S.; Royer, R.; Li, S.; McLaughlin, J.R.; Rosen, B.; Risch, H.A.; Fan, I.; Bradley, L.; Shaw, P.A.; Narod, S.A. Frequencies of BRCA1 and BRCA2 Mutations among 1342 Unselected Patients with Invasive Ovarian Cancer. Gynecol. Oncol. 2011, 121, 353–357. [Google Scholar] [CrossRef]
- Nelson, H.D.; Fu, R.; Goddard, K.; Mitchell, J.P.; Okinaka-Hu, L.; Pappas, M.; Zakher, B.U.S. Preventive Services Task Force Evidence Syntheses, Formerly Systematic Evidence Reviews. In Risk Assessment, Genetic Counseling, and Genetic Testing for BRCA-Related Cancer: Systematic Review to Update the U.S. Preventive Services Task Force Recommendation; Agency for Healthcare Research and Quality (US): Rockville, MD, USA, 2013. [Google Scholar]
- Manickam, K.; Buchanan, A.H.; Schwartz, M.L.B.; Hallquist, M.L.G.; Williams, J.L.; Rahm, A.K.; Rocha, H.; Savatt, J.M.; Evans, A.E.; Butry, L.M.; et al. Exome Sequencing-Based Screening for BRCA1/2 Expected Pathogenic Variants among Adult Biobank Participants. JAMA Netw. Open 2018, 1, e182140. [Google Scholar] [CrossRef] [PubMed]
- Bellcross, C.; Hermstad, A.; Tallo, C.; Stanislaw, C. Validation of Version 3.0 of the Breast Cancer Genetics Referral Screening Tool (B-Rst™). Genet. Med. 2019, 21, 181–184. [Google Scholar] [CrossRef]
- Bellcross, C.A.; Lemke, A.A.; Pape, L.S.; Tess, A.L.; Meisner, L.T. Evaluation of a Breast/Ovarian Cancer Genetics Referral Screening Tool in a Mammography Population. Genet. Med. 2009, 11, 783–789. [Google Scholar] [CrossRef]
- Couch, F.J.; Shimelis, H.; Hu, C.; Hart, S.N.; Polley, E.C.; Na, J.; Hallberg, E.; Moore, R.; Thomas, A.; Lilyquist, J.; et al. Associations between Cancer Predisposition Testing Panel Genes and Breast Cancer. JAMA Oncol. 2017, 3, 1190–1196. [Google Scholar] [CrossRef]
- Drohan, B.; Roche, C.A.; Cusack, J.C., Jr.; Hughes, K.S. Hereditary Breast and Ovarian Cancer and Other Hereditary Syndromes: Using Technology to Identify Carriers. Ann. Surg. Oncol. 2012, 19, 1732–1737. [Google Scholar] [CrossRef] [PubMed]
- Robinson, L.S.; Hendrix, A.; Xie, X.J.; Yan, J.; Pirzadeh-Miller, S.; Pritzlaff, M.; Read, P.; Pass, S.; Euhus, D.; Ross, T.S. Prediction of Cancer Prevention: From Mammogram Screening to Identification of BRCA1/2 Mutation Carriers in Underserved Populations. EBioMedicine 2015, 2, 1827–1833. [Google Scholar] [CrossRef][Green Version]
- Wood, M.E.; Kadlubek, P.; Pham, T.H.; Wollins, D.S.; Lu, K.H.; Weitzel, J.N.; Neuss, M.N.; Hughes, K.S. Quality of Cancer Family History and Referral for Genetic Counseling and Testing among Oncology Practices: A Pilot Test of Quality Measures as Part of the American Society of Clinical Oncology Quality Oncology Practice Initiative. J. Clin. Oncol. 2014, 32, 824–829. [Google Scholar] [CrossRef] [PubMed]
- National Collaborating Centre for Cancer (UK). National Collaborating Centre for Cancer (UK). National Institute for Health and Clinical Excellence: Guidance. In Familial Breast Cancer: Classification and Care of People at Risk of Familial Breast Cancer and Management of Breast Cancer and Related Risks in People with a Family History of Breast Cancer; National Collaborating Centre for Cancer (UK): Cardiff, UK, 2013. [Google Scholar]
- Moyer, V.A. Risk Assessment, Genetic Counseling, and Genetic Testing for BRCA-Related Cancer in Women: U.S. Preventive Services Task Force Recommendation Statement. Ann. Intern. Med. 2014, 160, 271–281. [Google Scholar] [CrossRef] [PubMed]
- Daly, M.B.; Pilarski, R.; Axilbund, J.E.; Berry, M.; Buys, S.S.; Crawford, B.; Farmer, M.; Friedman, S.; Garber, J.E.; Khan, S.; et al. Genetic/Familial High-Risk Assessment: Breast and Ovarian, Version 2.2015. J. Natl. Compr. Canc. Netw. 2016, 14, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Warner, E.; Hill, K.; Causer, P.; Plewes, D.; Jong, R.; Yaffe, M.; Foulkes, W.D.; Ghadirian, P.; Lynch, H.; Couch, F.; et al. Prospective Study of Breast Cancer Incidence in Women with a BRCA1 or BRCA2 Mutation under Surveillance with and without Magnetic Resonance Imaging. J. Clin. Oncol. 2011, 29, 1664–1669. [Google Scholar] [CrossRef] [PubMed]
- Kuhl, C.; Weigel, S.; Schrading, S.; Arand, B.; Bieling, H.; König, R.; Tombach, B.; Leutner, C.; Rieber-Brambs, A.; Nordhoff, D.; et al. Prospective Multicenter Cohort Study to Refine Management Recommendations for Women at Elevated Familial Risk of Breast Cancer: The Eva Trial. J. Clin. Oncol. 2010, 28, 1450–1457. [Google Scholar] [CrossRef]
- Long, E.F.; Ganz, P.A. Cost-Effectiveness of Universal BRCA1/2 Screening: Evidence-Based Decision Making. JAMA Oncol. 2015, 1, 1217–1218. [Google Scholar] [CrossRef]
- Manchanda, R.; Legood, R.; Burnell, M.; McGuire, A.; Raikou, M.; Loggenberg, K.; Wardle, J.; Sanderson, S.; Gessler, S.; Side, L.; et al. Cost-Effectiveness of Population Screening for BRCA Mutations in Ashkenazi Jewish Women Compared with Family History-Based Testing. J. Natl. Cancer Inst. 2015, 107, 380. [Google Scholar] [CrossRef]
- Anderson, K.; Jacobson, J.S.; Heitjan, D.F.; Zivin, J.G.; Hershman, D.; Neugut, A.I.; Grann, V.R. Cost-Effectiveness of Preventive Strategies for Women with a BRCA1 or a BRCA2 Mutation. Ann. Intern. Med. 2006, 144, 397–406. [Google Scholar] [CrossRef]
- Khoury, M.J.; Coates, R.J.; Evans, J.P. Evidence-Based Classification of Recommendations on Use of Genomic Tests in Clinical Practice: Dealing with Insufficient Evidence. Genet. Med. 2010, 12, 680–683. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Office of Public Health Genomics—Genomic Tests by Levels of Evidence. Available online: https://www.cdc.gov/genomics/gtesting/index.htm (accessed on 18 December 2018).
- U.S. Department of Health and Human Services. Healthy People 2030 Objectives and Data. Available online: https://health.gov/healthypeople/objectives-and-data (accessed on 16 July 2021).
- Samimi, G.; Bernardini, M.Q.; Brody, L.C.; Caga-Anan, C.F.; Campbell, I.G.; Chenevix-Trench, G.; Couch, F.J.; Dean, M.; de Hullu, J.A.; Domchek, S.M.; et al. Traceback: A Proposed Framework to Increase Identification and Genetic Counseling of BRCA1 and BRCA2 Mutation Carriers through Family-Based Outreach. J. Clin. Oncol. 2017, 35, 2329–2337. [Google Scholar] [CrossRef]
- Kurian, A.W.; Ward, K.C.; Howlader, N.; Deapen, D.; Hamilton, A.S.; Mariotto, A.; Miller, D.; Penberthy, L.S.; Katz, S.J. Genetic Testing and Results in a Population-Based Cohort of Breast Cancer Patients and Ovarian Cancer Patients. J. Clin. Oncol. 2019, 37, 1305–1315. [Google Scholar] [CrossRef]
- University of San Francisco Helen Diller Family Comprehensive Cancer Center. New Nccn Guidelines for Hereditary Breast and Ovarian Cancer. Available online: https://brca.ucsf.edu/news/new-nccn-guidelines-hereditary-breast-and-ovarian-cancer (accessed on 16 July 2021).
- Torre, L.A.; Trabert, B.; DeSantis, C.E.; Miller, K.D.; Samimi, G.; Runowicz, C.D.; Gaudet, M.M.; Jemal, A.; Siegel, R.L. Ovarian Cancer Statistics, 2018. CA Cancer J. Clin. 2018, 68, 284–296. [Google Scholar] [CrossRef] [PubMed]
- Mafficini, A.; Simbolo, M.; Parisi, A.; Rusev, B.; Luchini, C.; Cataldo, I.; Piazzola, E.; Sperandio, N.; Turri, G.; Franchi, M.; et al. BRCA Somatic and Germline Mutation Detection in Paraffin Embedded Ovarian Cancers by Next-Generation Sequencing. Oncotarget 2016, 7, 1076–1083. [Google Scholar] [CrossRef]
- Petersen, A.H.; Aagaard, M.M.; Nielsen, H.R.; Steffensen, K.D.; Waldstrøm, M.; Bojesen, A. Post-Mortem Testing; Germline BRCA1/2 Variant Detection Using Archival Ffpe Non-Tumor Tissue. A New Paradigm in Genetic Counseling. Eur. J. Hum. Genet. 2016, 24, 1104–1111. [Google Scholar] [CrossRef]
- Enyedi, M.Z.; Jaksa, G.; Pintér, L.; Sükösd, F.; Gyuris, Z.; Hajdu, A.; Határvölgyi, E.; Priskin, K.; Haracska, L. Simultaneous Detection of BRCA Mutations and Large Genomic Rearrangements in Germline DNA and Ffpe Tumor Samples. Oncotarget 2016, 7, 61845–61859. [Google Scholar] [CrossRef]
- National Institutes of Health; National Cancer Institute. Par-18-616: Traceback Testing: Increasing Identification and Genetic Counseling of Mutation Carriers through Family-Based Outreach (U01 Clinical Trial Optional); National Institutes of Health: Bethesda, MA, USA; National Cancer Institute: Rockville, MA, USA, 2021. [Google Scholar]
- Harris, P.A.; Taylor, R.; Thielke, R.; Payne, J.; Gonzalez, N.; Conde, J.G. Research Electronic Data Capture (Redcap)—A Metadata-Driven Methodology and Workflow Process for Providing Translational Research Informatics Support. J. Biomed. Inform. 2009, 42, 377–381. [Google Scholar] [CrossRef]
- Harris, P.A.; Taylor, R.; Minor, B.L.; Elliott, V.; Fernandez, M.; O’Neal, L.; McLeod, L.; Delacqua, G.; Delacqua, F.; Kirby, J.; et al. The Redcap Consortium: Building an International Community of Software Platform Partners. J. Biomed. Inform. 2019, 95, 103208. [Google Scholar] [CrossRef]
- Landrum, M.J.; Lee, J.M.; Benson, M.; Brown, G.R.; Chao, C.; Chitipiralla, S.; Gu, B.; Hart, J.; Hoffman, D.; Jang, W.; et al. Clinvar: Improving Access to Variant Interpretations and Supporting Evidence. Nucleic Acids Res. 2018, 46, D1062–D1067. [Google Scholar] [CrossRef]
- National Center for Biotechnology Information; U.S. National Library of Medicine. Dbgap/Database of Genotypes and Phenotypes. 2021. Available online: https://www.ncbi.nlm.nih.gov/gap/ (accessed on 1 October 2021).
- Roberts, M.C.; Dotson, W.D.; DeVore, C.S.; Bednar, E.M.; Bowen, D.J.; Ganiats, T.G.; Green, R.F.; Hurst, G.M.; Philp, A.R.; Ricker, C.N.; et al. Delivery of Cascade Screening for Hereditary Conditions: A Scoping Review of the Literature. Health Aff. 2018, 37, 801–808. [Google Scholar] [CrossRef] [PubMed]
- Henrikson, N.B.; Wagner, J.K.; Hampel, H.; DeVore, C.; Shridhar, N.; Williams, J.L.; Donohue, K.E.; Kullo, I.; Prince, A.E.R. What Guidance Does Hipaa Offer to Providers Considering Familial Risk Notification and Cascade Genetic Testing? J. Law Biosci. 2020, 7, lsaa071. [Google Scholar] [CrossRef] [PubMed]
- Palinkas, L.A.; Horwitz, S.M.; Green, C.A.; Wisdom, J.P.; Duan, N.; Hoagwood, K. Purposeful Sampling for Qualitative Data Collection and Analysis in Mixed Method Implementation Research. Adm. Policy Ment. Health 2015, 42, 533–544. [Google Scholar] [CrossRef]
- Lin, J.; Sharaf, R.N.; Saganty, R.; Ahsan, D.; Feit, J.; Khoury, A.; Bergeron, H.; Chapman-Davis, E.; Cantillo, E.; Holcomb, K.; et al. Achieving Universal Genetic Assessment for Women with Ovarian Cancer: Are We There Yet? A Systematic Review and Meta-Analysis. Gynecol. Oncol. 2021, 162, 506–516. [Google Scholar] [CrossRef]
- National Cancer Institute. Ovarian, Fallopian Tube, and Primary Peritoneal Cancer—Patient Version. Available online: https://www.cancer.gov/types/ovarian (accessed on 16 July 2021).
- National Comprehensive Cancer Network. Genetic/Familial High-Risk Assessment: Breast, Ovarian and Pancreatic, Version 1.2022—11 August 2021. Available online: https://www.nccn.org/professionals/physician_gls/pdf/genetics_bop.pdf (accessed on 16 July 2021).
- Moss, H.A.; Samimi, G.; Havrilesky, L.J.; Sherman, M.E.; Myers, E.R. Estimating the Number of Potential Family Members Eligible for BRCA1 and BRCA2 Mutation Testing in a “Traceback” Approach. Genet. Epidemiol. 2018, 42, 117–122. [Google Scholar] [CrossRef]
- DiNucci, A.; Henrikson, N.B.; Jonas, M.C.; Basra, S.; Blasi, P.; Brown, J.; Esplin, E.D.; Hassen, D.; Hao, J.; Hu, Y.; et al. Feasibility and Assessment of a Cascade Traceback Screening Program (Facts): Protocol for a Multisite Study to Implement and Assess an Ovarian Cancer Traceback Cascade Testing Program. J. Pers. Med. 2021, 11, 543. [Google Scholar] [CrossRef]
- Wang, Y.; Golesworthy, B.; Cuggia, A.; Domecq, C.; Chaudhury, P.; Barkun, J.; Metrakos, P.; Asselah, J.; Bouganim, N.; Gao, Z.H.; et al. Oncology Clinic-Based Germline Genetic Testing for Exocrine Pancreatic Cancer Enables Timely Return of Results and Unveils Low Uptake of Cascade Testing. J. Med. Genet. 2021. [Google Scholar] [CrossRef] [PubMed]
- Dugan, R.B.; Wiesner, G.L.; Juengst, E.T.; O’Riordan, M.; Matthews, A.L.; Robin, N.H. Duty to Warn at-Risk Relatives for Genetic Disease: Genetic Counselors’ Clinical Experience. Am. J. Med. Genet. C Semin. Med. Genet. 2003, 119c, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Clayton, E.W.; Evans, B.J.; Hazel, J.W.; Rothstein, M.A. The Law of Genetic Privacy: Applications, Implications, and Limitations. J. Law Biosci. 2019, 6, 1–36. [Google Scholar] [CrossRef] [PubMed]
- Menko, F.H.; Jeanson, K.N.; Bleiker, E.M.A.; van Tiggelen, C.W.M.; Hogervorst, F.B.L.; Ter Stege, J.A.; Ait Moha, D.; van der Kolk, L.E. The Uptake of Predictive DNA Testing in 40 Families with a Pathogenic BRCA1/BRCA2 Variant. An Evaluation of the Proband-Mediated Procedure. Eur. J. Hum. Genet. 2020, 28, 1020–1027. [Google Scholar] [CrossRef]
- Marshall, K.E.; Hexemer, A.; Seelman, S.L.; Fatica, M.K.; Blessington, T.; Hajmeer, M.; Kisselburgh, H.; Atkinson, R.; Hill, K.; Sharma, D.; et al. Lessons Learned from a Decade of Investigations of Shiga Toxin-Producing Escherichia Coli Outbreaks Linked to Leafy Greens, United States and Canada. Emerg. Infect. Dis. 2020, 26, 2319–2328. [Google Scholar] [CrossRef]
- Gill, M.J.; Towns, D.; Allaire, S.; Meyers, G. Transmission of Human Immunodeficiency Virus through Blood Transfusion: The Use of Lookback and Traceback Approaches to Optimize Recipient Identification in a Regional Population. Transfusion 1997, 37, 513–516. [Google Scholar] [CrossRef] [PubMed]
- Ghinai, I.; Pray, I.W.; Navon, L.; O’Laughlin, K.; Saathoff-Huber, L.; Hoots, B.; Kimball, A.; Tenforde, M.W.; Chevinsky, J.R.; Layer, M.; et al. E-Cigarette Product Use, or Vaping, among Persons with Associated Lung Injury—Illinois and Wisconsin, April–September 2019. MMWR Morb. Mortal. Wkly. Rep. 2019, 68, 865–869. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. CLIA Law & Regulations. Available online: https://www.cdc.gov/clia/law-regulations.html (accessed on 6 June 2021).
| Study Participants | |
|---|---|
| Inclusion Criteria | Exclusion Criteria |
|
|
| Cascade testing | |
| Inclusion criteria | Exclusion criteria |
|
|
| APC | FH | NF2 | SDHB |
| ATM | FLCN | NTHL1 | SDHC |
| AXIN2 | GREM1 | PALB2 | SDHD |
| BAP1 | HOXB13 | PDGFRA | SMAD4 |
| BARD1 | KIT | PMS2 | SMARCA4 |
| BMPR1A | MAX | POLD1 | SMARCB1 |
| BRCA1 | MEN1 | POLE | STK11 |
| BRCA2 | MET | PRKAR1A | TEME127 |
| BRIP1 | MITF | PTCH1 | TP53 |
| CDC73 | MLH1 | PTEN | TSC1 |
| CDH1 | MSH2 | RAD51C | TSC2 |
| CDK4 | MSH3 | RAD51D | VHL |
| CDKN2A | MSH6 | RB1 | |
| CHEK2 | MUTYH | RET | |
| DICER1 | NBN | SDHA | |
| EPCAM | NF1 | SDHAF2 |
| Topics | Domains |
|---|---|
| Background knowledge and baseline opinions |
|
| Participant’s experience in the study |
|
| Accuracy of tumor registries and pathology reports to identify patients with a correct diagnosis |
| Number of deceased patients with contact information for next of kin in EHR |
| Success rate to locate patients or contact next of kin for deceased patients |
| Uptake of genetic testing among contacted patients or next of kin |
| Availability of archived pathology specimens for germline genetic testing |
| Uptake of cascade testing among at-risk relatives |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kauffman, T.L.; Prado, Y.K.; Reyes, A.A.; Zepp, J.M.; Sawyer, J.; White, L.L.; Martucci, J.; Salas, S.B.; Vertrees, S.; Rope, A.F.; et al. Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol. J. Pers. Med. 2021, 11, 1194. https://doi.org/10.3390/jpm11111194
Kauffman TL, Prado YK, Reyes AA, Zepp JM, Sawyer J, White LL, Martucci J, Salas SB, Vertrees S, Rope AF, et al. Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol. Journal of Personalized Medicine. 2021; 11(11):1194. https://doi.org/10.3390/jpm11111194
Chicago/Turabian StyleKauffman, Tia L., Yolanda K. Prado, Ana A. Reyes, Jamilyn M. Zepp, Jennifer Sawyer, Larissa Lee White, Jessica Martucci, Suzanne Bianca Salas, Sarah Vertrees, Alan F. Rope, and et al. 2021. "Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol" Journal of Personalized Medicine 11, no. 11: 1194. https://doi.org/10.3390/jpm11111194
APA StyleKauffman, T. L., Prado, Y. K., Reyes, A. A., Zepp, J. M., Sawyer, J., White, L. L., Martucci, J., Salas, S. B., Vertrees, S., Rope, A. F., Weinmann, S., Henrikson, N. B., Lee, S. S.-J., Feigelson, H. S., & Hunter, J. E. (2021). Feasibility of a Traceback Approach for Using Pathology Specimens to Facilitate Genetic Testing in the Genetic Risk Analysis in Ovarian Cancer (GRACE) Study Protocol. Journal of Personalized Medicine, 11(11), 1194. https://doi.org/10.3390/jpm11111194






