1. Introduction
The coronavirus disease, designated as COVID-19 and declared as pandemic by the World Health Organization is caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) [
1,
2,
3,
4]. COVID-19 has infected around 224 million people across the globe and has resulted in more than 4.86 million deaths till date (
https://www.worldometers.info/coronavirus, accessed on 12 October 2021). While as the individuals more than 50 years of age were prone to higher severity and mortality of the disease, the common comorbidities included hypertension, obesity, and diabetes [
5,
6]. COVID-19 has exhibited geographical variations in its spread and mortality and it has caused more morbidity and mortality in Europe and USA [
7,
8,
9].
SARS-CoV-2 is an enveloped RNA virus with three structural protein components which are described as M (membrane), E (envelop) and S (spike) proteins [
10,
11]. The SARS-COV-2 makes its entry into the host cells through the S proteins. The two subunits of S protein, S1 and S2 are used by the virus to attach to the host cell membrane. The C-terminal domain (CTD) of S1 subunits is vital for the interaction and attachment of S proteins to the protein receptors of the host cell [
4,
12,
13].
Just like the SARS-CoV, the human angiotensin converting enzyme-2 (ACE2) has emerged as the target for the SARS-CoV-2 [
14,
15]. ACE2 is expressed ubiquitously in different organs of the human body that include heart, lungs, kidneys, intestines and endothelium [
16,
17,
18,
19]. ACE2, a type I transmembrane zinc finger glycoprotein, is a mono-carboxypeptidase (with 805 amino acid residues and MW of 100 KD) and converts angiotensin 1 to angiotensin 1-9 [
20]. The gene for ACE2, present on chromosome X (Xp22) comprises of 39.98 kb of genomic DNA and has 18 exons [
20]. Many ACE2 variants are associated with several common diseases, whose incidence depends on the balance in the renin-angiotensin-aldosterone (RAAS) pathway. Hypertension is associated with rs1514283, rs2074192, rs233575, rs4646155, rs4646176, rs2285666, rs879922, rs2106809, rs4646188, rs4240157, rs4830542, rs2158083, and rs879922 [
21,
22]. The role of ACE2 is very critical since the inhibition of RAAS pathway leads to upregulation of ACE2 which alleviates ARDS and myocarditis symptoms in COVID-19 patients but at the same time increased ACE2 expression may increase the entry of virus into host cells, complication the overall clinical picture [
23].
Geographic distribution of ACE I (insertion in intron 16) allele was summarized by Saab et al. and its frequency increases eastwards and westwards from the Middle East [
24]. The genetic polymorphism of ACE2 gene is well known world over with racial and ethnic variations [
25,
26] having varying influences on the altered functions of RAAS pathway [
27]. The polymorphism of ACE2 gene has been associated with hypertension in Chinese [
28,
29], Canadian [
30], Indian [
31] and Brazilian populations [
32]. The ACE2 polymorphism has been associated with varying degree of disease severity and clinical outcomes of COVID-19, with absence of ACE D/D genotype conferring protection against severe lung injury [
33]. The African Americans with high frequency of D allele have depicted higher mortality rates in USA [
34,
35]. Similarly, higher fatalities have been reported from many parts of Europe (
https://ec.europa.eu/eurostat/statistics-explained/index.php/Main_Page, accessed on 12 October 2021). In contrast lower mortality rates have been observed in Asian patients who harbor high frequency of II genotype as compared to DD genotype [
36] and (
https://www.worldometers.info/coronavirus/coronavirus-death-rate, accessed on 12 October 2021). The D allele results in an increased angiotensin-2 level (as a consequence of increased ACE-1 and decreased ACE-2 levels) that leads to increased microvascular permeability resulting in pulmonary edema and other complications [
37,
38].
In the current scenario, the knowledge of ACE2 gene polymorphism in different populations assumes a huge significance. However, although there have been many published reports on the ACE2 gene polymorphism from different parts of the globe, we came across only one report from Saudi Arabia suggesting a link between the ACE2-rs4646994 I/D polymorphism and the obesity [
39]. To the best of our knowledge, the current study on the genetic polymorphism ACE2-rs4646994 I/D and ACE2 rs4240157 C > T in the ACE2 gene is the first one from Saudi Arabia and was carried out in the COVID-19 patients with different disease severity/outcomes and healthy controls.
2. Methodology
2.1. Study Population
This population-based case-control study (prospective cohort study) was conducted on COVID-19 patients and 150 age matched healthy controls. COVID-19 laboratory confirmation was defined as a positive result on RT-qPCR (SARS-CoV-2 RT-qPCR detection kit) of nasal and oropharyngeal swab specimens. We selected COVID-19 positive cases confirmed by a real-time reverse transcription polymerase chain reaction (rRT-PCR) test.
This study was a collaborative study. Specimens from positive cases of COVID-19 were collected from different hospitals in Saudi Arabia (Bisha, Abha and Tabuk). The sample collection was conducted between September 2020 and April 2021.
2.2. Ethics Approvals
The ethical approvals were obtained from local RELOC committees of College of Medicine, University of Bisha (Ref. No. UBCOM/H-06-BH-087 (05/25)), University of Tabuk (Ref. No: KAEK2020/4/4) and College of Medicine, King Khalid University, Abha (Ref. No. KKU-H-06-B-091) in accordance with human subjects and complied with the principles of the Helsinki Declaration. Informed consent was obtained before collecting samples from all patients and control subjects.
2.3. Data Collection
All the subjects were interviewed using a structured questionnaire regarding epidemiological/demographic data, past history of CAD, T2D, and history of addiction particularly smoking, family history of any significant diseases.
2.3.1. Sample Collection from COVID-19 Patients
About 3 mL of peripheral blood sample was collected in an EDTA or Lavender top tube for all COVID-19 patients. The Blood specimens were immediately stored at −20 °C to −30 °C. The COVID-19 patients who’s who oxygen saturation was less than 60 and needed mechanical ventilatory support in ICU were classified as having severe COVID-19 disease.
2.3.2. Sample Collection from Healthy Controls
All healthy age matched controls specimens have been timed around routine blood draws that are part of routine workout, and hence did not require additional phlebotomy and all participants were provided written informed consent form. About 3 mL peripheral blood was collected in EDTA tubes. The Blood specimens were immediately stored at −20 °C to −30 °C.
2.4. Genomic DNA Extraction
Genomic DNA was extracted using DNeasy Blood K (Qiagen, Hilden, Germany) as per the manufacturer’s instructions. The extracted DNA was dissolved in nuclease-free water and stored at 4 °C until use. Quality and integrity of DNA were checked by NanoDrop™ (Thermo Scientific, Waltham, MA, USA). All DNA samples from COVID-19 and controls were screened for purity by measuring optical density (OD) at 260 nm (OD260) and 280 nm (OD280). The λ260/λ280 ratios ranged from 1.83–1.99 indicating good quality DNA.
2.5. Angiotensin-Converting Enzyme Genotyping
The Angiotensin-converting enzyme-2 genotyping (ACE2-rs4646994 I/D) was done by mutation specific PCR and amplification refractory mutation system (ARMS-PCR) PCR was used to detect ACE2 rs4240157 T > C genotyping. The ARMS primers were designed by using the Primer3 software as depicted in
Table 1.
The PCR was done in a reaction volume of 25 µL containing template DNA (50 ng), Fo—0.25 µL, R0—0.25 µL, RI—0.25 µL, RI—0.25 µL of 25 pmol of each primer and 10 µL from GoTaq® Green Master Mix (cat no M7122) (Promega, Madison, WI, USA). The final volume of 25 µL was adjusted by adding nuclease free double distilled water (ddH2O). Finally, 2 µL of DNA was added from each patient. The thermocycling conditions used were: 95 °C for 10 min followed by 40 cycles of 95 °C for 35 s, annealing temperature ACE2-rs4646994 I/D (58 °C) and ACE2 rs4240157 C > T (60 °C) gene polymorphism for 40 s, 72 °C for 43 s followed by the final extension at 72 °C for 10 min.
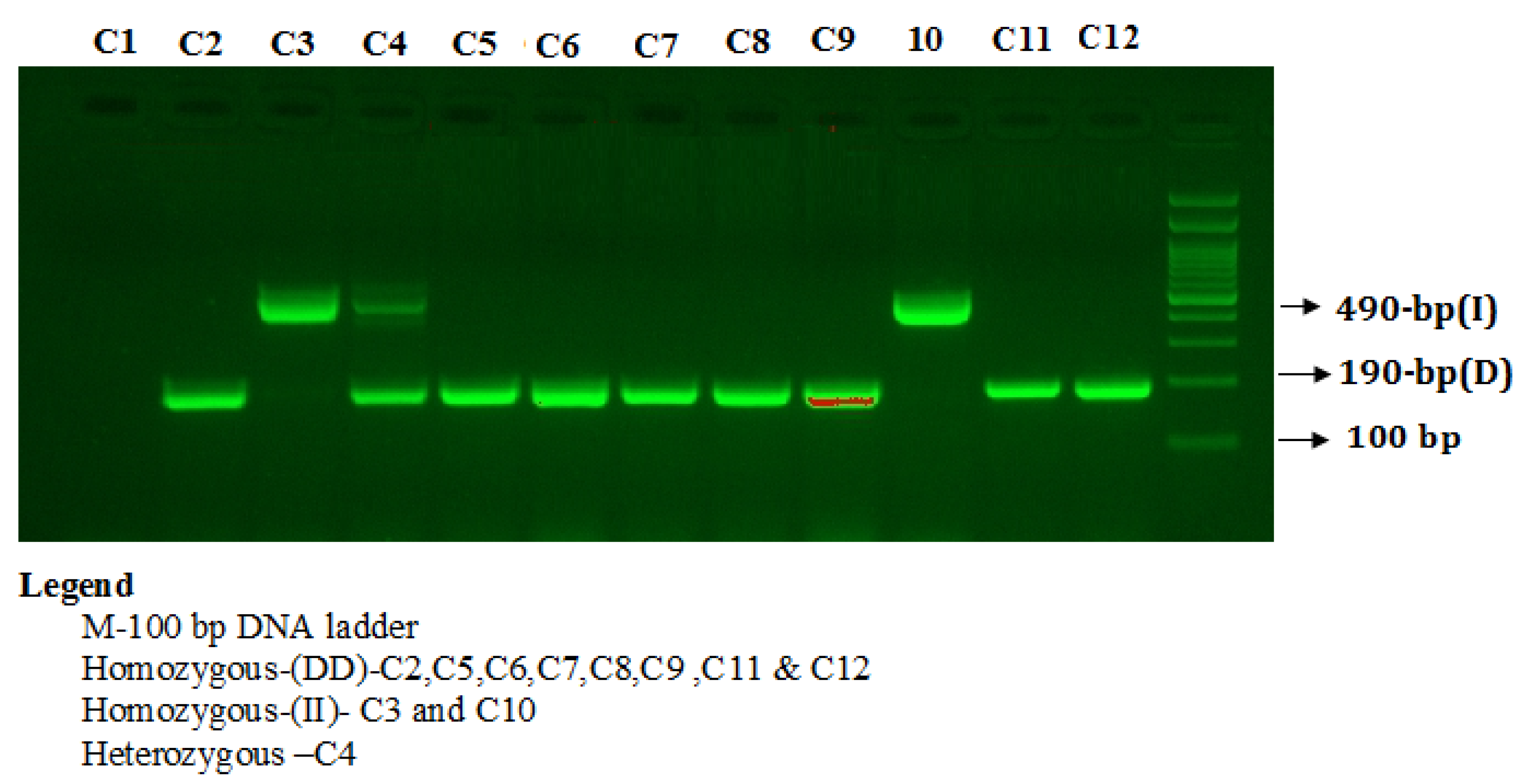
PCR products were separated on 2% agarose gel stained with 2 µL of sybre safe stain (Thermo Scientific, Waltham, MA, USA) and visualized on a UV trans illuminator from Bio-Rad (Hercules, CA, USA). The PCR products were electrophoresed on a 2% agarose gel stained with ethidium bromide to visualize three patterns: I/I (490-bp fragment), D/D (190-bp fragment), and I/D (both 490- and 190-bp fragments) as depicted in
Figure 1.
Primers FO and RO flank the exon of the ACE2 rs4240157 T > C gene, resulting in a band of 386 bp to act as a control for DNA quality and quantity. Primers FI and RO amplify a wild-type allele (T allele), generating a band of 244 bp, and primers FO and RI generate a band of 194 bp from the mutant allele (C allele) as depicted in
Figure 2.
2.6. Statistical Analysis
Deviations from Hardy–Weinberg disequilibrium (HWD) were calculated by chi-square (χ2) goodness-of-fit test. Group differences were compared using Student’s two-sample t-test or one-way analysis of variance (ANOVA) for continuous variables and chi-squared for categorical variables. Differences in the ACE-rs4646994 I/D, and ACE2 rs4240157 C > T allele and genotype frequencies between groups were evaluated using Chi-square test. The associations between ACE2-rs4646994 I/D, ACE2 rs4240157 C > T genotypes and risk of COVID-19 patients were estimated by computing the odds ratios (ORs), risk ratios (RRs) and risk differences (RDs) with 95% confidence intervals (CIs). Allele frequencies among cases as well as controls were evaluated by using the chi-square Hardy–Weinberg equilibrium (HWE) test. A p-value < 0.05 was considered significant. All statistical analyses were performed SPSS 16.0.
4. Discussion
Coronaviruses are divided into α, β, γ, and δ genera on the basis of the target host [
1,
2,
3]. Out of these, mammals are infected by α and β-CoV whereas γ and δ-CoV genera tend to infect birds. The current COVID-19 pandemic is found to be caused by SARS-CoV-2 [
3,
4]. It is a β-CoV that is enveloped, non-segmented and positive-sense RNA virus. Genome sequencing results showed that this newly discovered virus shares 96.2% identity with bat CoV RaTG13 and 79.5% identity with SARS-CoV [
10].
4.1. Role of RAAS in SARS-CoV-2 Infection
Different ACE genotypes are believed to be associated with development of acute respiratory distress syndrome and several studies have reported that the distribution frequency of ACE insertion/insertion (II) genotype might have a significant influence on COVID-19 mortality [
36,
40]. It has been reported that ACE2 levels correlate with susceptibility to SARS-CoV-2 infection and men have a higher ACE2 expression in lung than women and Asian people express ACE2 higher than Caucasian and African American populations [
41,
42].
4.2. Association of ACE2-D/I with Gender
Angiotensin converting enzyme is a metallopeptidase that converts angiotensin I (AT-I) to angiotensin II (AT-II) that acts as a vasoconstrictor in addition to its other functions. ACE also degrades bradykinin, a vasodilator substance. ACE2 is also a known SARS virus receptor [
14]. Both ACE1 and ACE2 are highly polymorphic. Polymorphisms present in ACE1 are just as important as those in ACE2. As it was reported that the conversion of Ang II to Ang (1–7) by ACE2 is higher in males than females [
43] because ACE2 gene is located on the X chromosome and men express more ACE2 than women that depends on the allelic expressions by women, therefore women may be considered of lower sensitivity against the SARS-CoV-2 infection and its lethal effects [
43,
44].
Several research reports in mice have shown the protective effect of ACE2, whereby severe lung failure is associated with ACE2 downregulation. Immediately after SARS-CoV-2 infection, there is a downregulation of ACE2 [
37,
38,
45]. ACE gene on chromosome 17 consists of 26 exons and 25 introns. A functional insertion-deletion (I/D) polymorphism of 287 bp Alu repeat sequence has been reported in the intron 16 of the ACE gene [
44,
46]. Jeong et al. have reported that the deletion (D) and insertion allele (I) are associated with an increased and decreased ACE level and enzyme activity, respectively [
47]. The ACE2 gene exhibits a high degree of genetic polymorphism [
42]. Itoyama et al. [
48] reported the correlation of ACE-D allele with the incidence of pneumonia in SARS patients and the death of subjects with acute respiratory distress syndrome. These results tempted us to investigate the role of ACE2 I/D polymorphism in susceptibility to SARS-CoV-2 infection and related mortality. Our results indicated higher overall frequencies of deletion polymorphism in COVID-19 patients (67%) than the insertion polymorphism (33%). The distribution of homozygote insertion (II) genotype is seen to be higher in the healthy population (26.66%) compared with the COVID-19 patients (13.67%). The frequency of heterozygote ID genotype is comparable between COVID-19 patients (37.60%) and control (33.34%) groups. We observed a significant difference in the distribution of homozygote deletion (DD) genotype between COVID-19 cases and controls (
p < 0.03) (
Table 3).
4.3. Distribution of ACE2-II, DI and DD Genotypes in the World Populations
The distribution of ACE2-II, DI and DD genotypes in the SARS-CoV-2 patients in the world populations has been summarized in the
Table 13 [
49]. The frequency of DD genotype in our patient population was high (48.71%) corresponding to the frequency of Brazil (49%), Finland (42.61%), Turkey (34.44%), Iran (37.50%), Spain (37.84%), Russia (39.38%), Italy (37.61%).
Lee et al., reported that populations in France, Italy and Spain have shown high
D allele frequency between 82% and 87% [
50]. Similarly, African Americans in the United States have been reported to have the highest
D allele frequency (89%) compared with white Americans (69%) [
33]. Conversely, populations in East Asian countries such as China, Japan, Taiwan and Korea have a high frequency of the II genotype [
51]. Pati et al. [
52] reported that the low DD genotype frequency and high II genotype frequency in the ACE gene is strongly correlated with the relatively low mortality rate of COVID-19 among these populations [
52]. A higher mortality rate has been shown among the European populations as in the case of black ethnicity in the United States [
34]. In the present study, we observed a positive correlation of allele D with infection and mortality rate of COVID-19.
Recent studies have also shown that the ethnic variations of the ACE I/D genotype tend to correlate with the variations in outcomes where populations with a high D genotype frequency tend to experience higher mortality rates [
23]. Our results indicated that in the codominant model, the ACE2-DD genotype was strongly associated with increased COVID-19 mortality OR 8.25 (95%) CI = (2.40 to 28.34),
p < 0.008 (
Table 7). Mortality of SARS-CoV-2 subjects is dependent on a wide range of complex phenomena and genetic mutations are one among them [
53,
54,
55,
56]. The prevalence of the DD genotype is higher in patients with severe lung infections and is significantly correlated with a high death rate. Mortality related to SARS-CoV-2 infections has been associated to various comorbid conditions such as hypertension, diabetes, hyperlipidemia, coronary artery disease and renal disease [
23,
57,
58,
59,
60,
61,
62,
63,
64]. High producer of ACE, the DD genotype has been associated with susceptibility to hypertension. [
34,
35,
55,
62], Type 2 diabetes [
59,
60], coronary artery disease [
57,
59] indicating a possible role of ACE I/D polymorphism with SARS-CoV-2 related mortality. Our results reported a significant correlation of ACE2-DD genotypes with advanced age of the COVID-19 patients (
p < 0.035), with T2D (
p < 0.0013), hypertension (
p < 0.0031) and coronary artery disease (
p < 0.0001) [
Table 5]. Our study revealed that age is a significant risk factor for severe disease. Results showed that the there was a significant correlation between the ACE2-rs4646994 I and D genotypes with respect to the Age of the COVID-19 patients (
p < 0.035). Advanced age is a significant risk factor for mortality from COVID-19, the important reason being the associated increased frequency of comorbid conditions with advancing age [
61,
63].
4.4. Role of ACE2 rs4240157 T > C Gene Polymorphism in COVID-19 Severity
Our results indicated a significant correlation of ACE2 rs4240157 T > C genotypes with the COVID-19 severity among the COVID-19 patients with T2D (
p < 0.007), with coronary artery disease (
p < 0.049) and hypertension (
p < 0.006) (
Table 10). Recent studies identified ACE2 polymorphisms that might influence disease severity and indicated that out of 10 studied SNPs, 5 polymorphisms (rs6632680, rs4830965, rs1476524, rs4240157 and rs2048683) indicated an association with higher tissue specific expression of ACE2 resulting in hospitalization whereas rs1548474 polymorphism showed correlation with low tissue expression and lesser severity [
23,
37,
38,
39,
40]. Variation in circulating ACE2 levels was speculated to be controlled by genetic factors including rs2106809 polymorphism [
39,
63]. Xiao et al., [
64] have earlier reported that a point mutation in the ACE2 gene (Leu584Ala) facilitates entry of SARS-CoV-1 into host cells. Recent studies have proven that several amino acid variants can potentially affect the interaction between the viral S1 protein and ACE2 receptors and thus the level of infection [
65]. Different amino acid residues expressed within the ACE2 receptor were observed to be very relevant either by promoting or preventing viral infection. A total of 13 ACE2 polymorphisms enhanced ACE2/S1 recognition, thereby facilitating SARS-CoV-2 infection while as in contrast, 18 SNPs hindered interactions between ACE2 and S1, thereby reduces the infection rate [
62]. Patel et al., [
59] reported that ACE 2 rs2074192, rs4240157 and rs4646188 variations in T2D exhibited higher risk with hypertension among persons of Australian descent. ACE2 s4240157 (
p < 0.001) and rs4646156 (
p < 0.037) are correlated with increased blood pressure [
64]. Our results reported higher frequency of heterozygosity among SARS-CoV-2 cases of ACE2-rs4240157 TC genotypes (40.17%) followed by TT genotypes (35%), and CC genotypes (24.78%) respectively [
Table 8].
In our study, the ACE2 rs4240157 T > C gene polymorphism observed between COVID-19 patients and controls was statistically significant (
p < 0.0007). Wooster et al., [
55] reported that ACE2 rs4240157 polymorphisms is associated with COVID-19 disease severity as it might be inducing higher tissue specific expression of ACE2 resulting in the hospitalization of COVID-19 patients. Pouladi et al., [
62] reported the association ACE2 rs4240157 T > C gene polymorphism with hypertension and other related heart diseases. Our results indicated ACE2-CT genotype association with COVID-19 severity with an OR 2.18 (95% CI) (1.92–3.99),
p < 0.010 and also ACE2-CC genotype was linked with COVID-19 severity with an OR 2.66 (95% CI) (1.53–4.62),
p < 0.005 (
Table 4). We observed that ACE2-CC genotype was associated with increased COVID-19 mortality with OR 3.66 (95%) CI = (1.34 to 9.97),
p < 0.011 (
Table 10). Similarly, in recessive inheritance model, ACE2-(TT + CT) genotype was associated with increased COVID-19 mortality with OR 5.0 (95%) CI = (2.0640–12.437),
p < 0.004.
Despite the relatively small sample size, our results indicate that there is a close association between the ACE I/D gene polymorphism with clinical severity of COVID-19 disease. In addition, advanced age, coronary artery disease and diabetes were independent risk factors for the development of severe COVID-19 disease and mortality. The ACE-II genotype was a protective factor against the development of severe COVID-19 disease. Since this study is a first of its kind from Saudi Arabia, we believe that the relation of ACE2 DD genotype with the disease severity and clinical outcome in COVID-19 patients should be further investigated with more extensive studies. However, none of the studied SNPs are common in the general population. This finding might help in the recognition of people less and more prone to COVID-19.