Metabolomics Biomarkers of Prostate Cancer: A Systematic Review
Abstract
1. Introduction
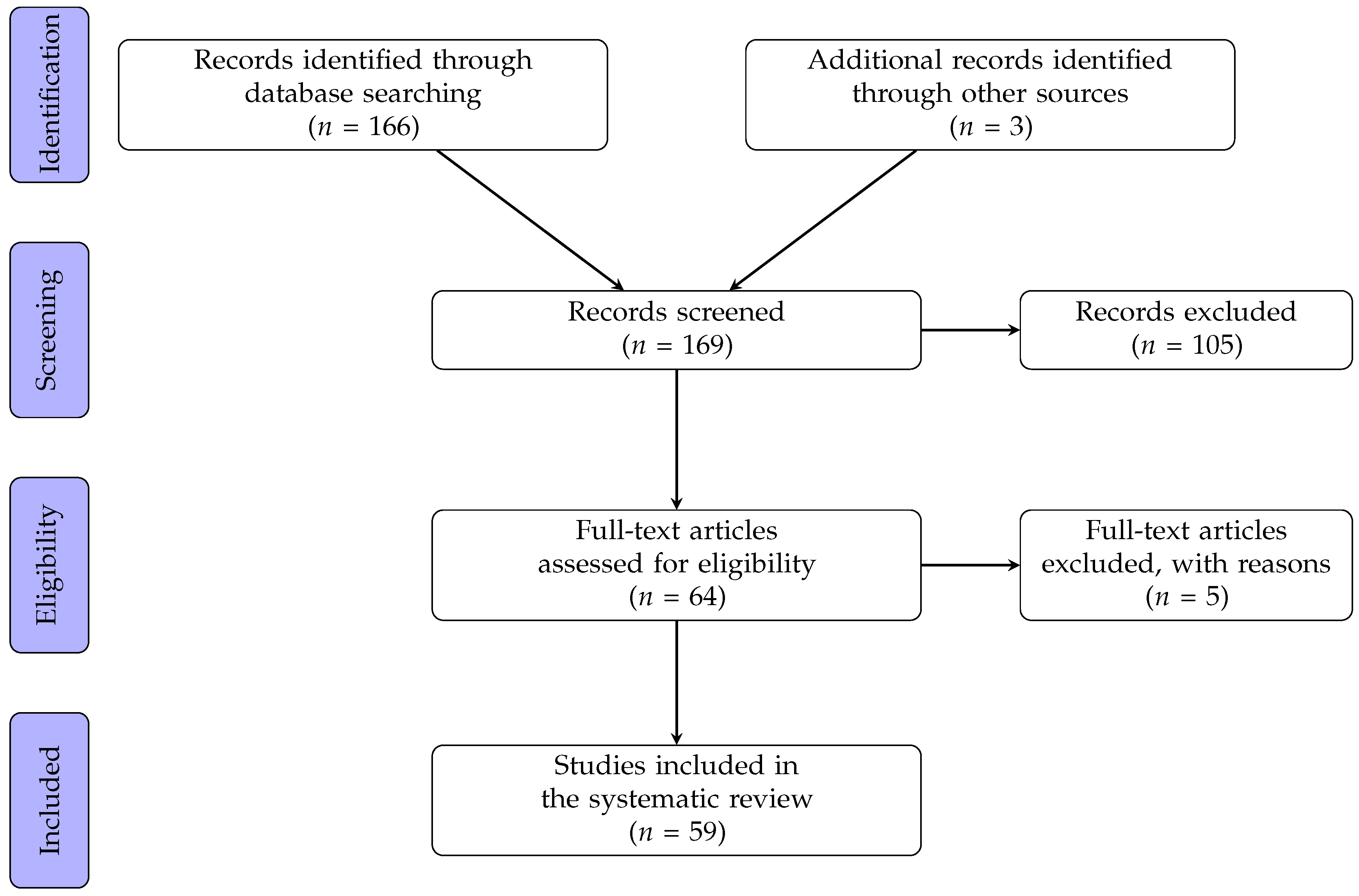
2. Methods
2.1. Search Strategy
2.2. Selection Criteria
2.3. Data Extraction
3. Results
3.1. Study Characteristics
3.2. Outcomes
3.2.1. Blood Based Biomarkers
Blood Biomarkers Associated with PCa Risk
Blood Biomarkers for Diagnosis and Staging of PCa
Blood Biomarkers for Therapy Prediction and Prognosis in PCa
3.2.2. Urine Based Biomarkers
Urine Biomarkers for Diagnosis and Prognosis of PCa
Urine Biomarkers for Risk Prediction of PCa
3.2.3. Tissue Based Biomarkers
Tissue Biomarkers for Diagnosis of PCa
Tissue Biomarkers for Staging of PCa
Tissue Biomarkers for Staging of PCa
3.2.4. Other Specimen Types
PCa Biomarkers in Urine Extracellular Vesicles
Seminal Plasma
In Situ Magnetic Resonance Spectroscopic Imaging (MRSI) Studies
3.2.5. Summary
4. Discussion
Author Contributions
Funding

Acknowledgments
Conflicts of Interest
Abbreviations
| PCa | Prostate cancer |
| BPH | Benign prstate hyperplasia |
| DRE | Digital rectal exam |
| GS | Gleason score |
| HC | Healthy controls |
| NEM | No evidence of malignancy |
| PSA | Prostate specific antigen |
| TNM | Tumour-lymphnode-metastasis classification |
| SP | Seminal plasma |
| EV | Extracellular vesicles |
| ADT | Androgen deprivation therapy |
| AJCC | American Joint Committee on Cancer |
| CRPC | Castration-resistant prostate cancer |
| PRISMA | Preferred Reporting Items for Systematic Reviews and Meta-Analyses |
| MS | Mass spectrometry |
| GC-MS | Gas chromatography mass spectrometry |
| MRIS | Magnetic resonance imaging spectroscopy |
| NMR | Nuclear magnetic resonance |
| LC-MS | Liquid chromatography mass spectrometry |
| LC-QTOF-MS | Liquid chromatography quadrupole time-of-flight mass spectrometry |
| GC-TOF-MS | Gas chromatography time-of-flight mass spectrometry |
| HPLC-ESI+QTOF-sMS | High performance liquid chromatography electrospray ionization quadrupole time-of-flight mass spectrometry |
| UHPLC-MS | Ultra high performance liquid chromatography mass spectrometry |
| HR-MAS-1H-MRS | High resolution magic angle spinning proton magnetic resonance spectroscopy |
| LC-ESI-MS | Liquid chromatography electrospray ionization mass spectrometry |
| PCA | Principal component analysis |
| PLS-DA | Partial least squares-discriminant analysis |
| RF | Random forest |
| OSC-PLS | Orthogonal signal correction for partial least square analysis |
| OPLS-DA | Orthogonal projections to latent structures-discriminant analysis |
| ROC | Receiver operator characteristics |
| SVM | Support vector machine |
| VIP | Variable importance of projection |
| LDA | Linear discrimination analysis |
| DFA | Discriminant function analysis |
| ANOVA | One-way analysis of variance |
| AUC | Area under the curve |
| DRW-GM | Directed random walk on gene metabolite pathway graph |
| (CCP/C) | Choline+creatine+polyamines over citrate |
| Cho | Choline |
| Cit | Citrate |
| Cr | Creatinine |
| GPC | Glycerophosphocholine |
| GPE | Glycerophospho-ethanolamine |
| LCFA | Long chain fatty acid |
| mI | Myo-inositol |
| NAD | Nicotinamide adenine dinucleotide (NAD+) |
| PC | Phosphocholine |
| PE | Phosphoethanolamine |
| Spm | Spermine |
| TMAO | Trimethylamine N-oxide |
References
- Heidenreich, A.; Bastian, P.J.; Bellmunt, J.; Bolla, M.; Joniau, S.; van der Kwast, T.; Mason, M.; Matveev, V.; Wiegel, T.; Zattoni, F.; et al. EAU guidelines on prostate cancer. Part 1: Screening, diagnosis, and local treatment with curative intent-update 2013. Eur. Urol. 2014, 65, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Cuzick, J.; Thorat, M.A.; Andriole, G.; Brawley, O.W.; Brown, P.H.; Culig, Z.; Eeles, R.A.; Ford, L.G.; Hamdy, F.C.; Holmberg, L.; et al. Prevention and early detection of prostate cancer. Lancet Oncol. 2014, 15, e484–e492. [Google Scholar] [CrossRef]
- Prensner, J.R.; Rubin, M.A.; Wei, J.T.; Chinnaiyan, A.M. Beyond PSA: The next generation of prostate cancer biomarkers. Sci. Transl. Med. 2012, 4, 127rv3. [Google Scholar] [CrossRef] [PubMed]
- Van den Broeck, T.; Joniau, S.; Clinckemalie, L.; Helsen, C.; Prekovic, S.; Spans, L.; Tosco, L.; Van Poppel, H.; Claessens, F. The role of single nucleotide polymorphisms in predicting prostate cancer risk and therapeutic decision making. BioMed Res. Int. 2014, 2014, 627510. [Google Scholar] [CrossRef] [PubMed]
- Silberstein, J.L.; Pal, S.K.; Lewis, B.; Sartor, O. Current clinical challenges in prostate cancer. Transl. Androl. Urol. 2013, 2, 122. [Google Scholar] [PubMed]
- Hegde, J.V.; Mulkern, R.V.; Panych, L.P.; Fennessy, F.M.; Fedorov, A.; Maier, S.E.; Tempany, C.M. Multiparametric MRI of prostate cancer: An update on state-of-the-art techniques and their performance in detecting and localizing prostate cancer. J. Magn. Reson. Imaging 2013, 37, 1035–1054. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Urrego, P.A.; Cronin, A.M.; Al-Ahmadie, H.A.; Gopalan, A.; Tickoo, S.K.; Reuter, V.E.; Fine, S.W. Interobserver and intraobserver reproducibility in digital and routine microscopic assessment of prostate needle biopsies. Hum. Pathol. 2011, 42, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Ganzer, R.; Fritsche, H.M.; Brandtner, A.; Bründl, J.; Koch, D.; Wieland, W.F.; Blana, A. Fourteen-year oncological and functional outcomes of high-intensity focused ultrasound in localized prostate cancer. BJU Int. 2013, 112, 322–329. [Google Scholar] [CrossRef]
- Karantanos, T.; Corn, P.G.; Thompson, T.C. Prostate cancer progression after androgen deprivation therapy: Mechanisms of castrate resistance and novel therapeutic approaches. Oncogene 2013, 32, 5501. [Google Scholar] [CrossRef]
- Boyd, L.K.; Mao, X.; Lu, Y.J. The complexity of prostate cancer: Genomic alterations and heterogeneity. Nat. Rev. Urol. 2012, 9, 652–664. [Google Scholar] [CrossRef]
- Welty, C.J.; Carroll, P.R. The ongoing need for improved risk stratification and monitoring for those on active surveillance for early stage prostate cancer. Eur. Urol. 2014, 65, 1032–1033. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Zhao, D.; Spring, D.J.; DePinho, R.A. Genetics and biology of prostate cancer. Genes Dev. 2018, 32, 1105–1140. [Google Scholar] [CrossRef] [PubMed]
- Di Donato, M.; Giovannelli, P.; Cernera, G.; Di Santi, A.; Marino, I.; Bilancio, A.; Galasso, G.; Auricchio, F.; Migliaccio, A.; Castoria, G. Non-genomic androgen action regulates proliferative/migratory signaling in stromal cells. Front. Endocrinol. 2015, 5, 225. [Google Scholar] [CrossRef] [PubMed]
- Mondul, A.M.; Moore, S.C.; Weinstein, S.J.; Männistö, S.; Sampson, J.N.; Albanes, D. 1-Stearoylglycerol is associated with risk of prostate cancer: results from a serum metabolomic profiling analysis. Metabolomics 2014, 10, 1036–1041. [Google Scholar] [CrossRef] [PubMed]
- Mondul, A.M.; Moore, S.C.; Weinstein, S.J.; Karoly, E.D.; Sampson, J.N.; Albanes, D. Metabolomic analysis of prostate cancer risk in a prospective cohort: The alpha-tocopherol, beta-carotene cancer prevention (ATBC) study. Int. J. Cancer 2015, 137, 2124–2132. [Google Scholar] [CrossRef]
- Huang, J.; Mondul, A.M.; Weinstein, S.J.; Karoly, E.D.; Sampson, J.N.; Albanes, D. Prospective serum metabolomic profile of prostate cancer by size and extent of primary tumor. Oncotarget 2017, 8, 45190. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Mondul, A.M.; Weinstein, S.J.; Koutros, S.; Derkach, A.; Karoly, E.; Sampson, J.N.; Moore, S.C.; Berndt, S.I.; Albanes, D. Serum metabolomic profiling of prostate cancer risk in the prostate, lung, colorectal, and ovarian cancer screening trial. Br. J. Cancer 2016, 115, 1087–1095. [Google Scholar] [CrossRef] [PubMed]
- De Vogel, S.; Ulvik, A.; Meyer, K.; Ueland, P.M.; Nygård, O.; Vollset, S.E.; Tell, G.S.; Gregory, J.F., III; Tretli, S.; Bjørge, T. Sarcosine and other metabolites along the choline oxidation pathway in relation to prostate cancer—A large nested case–control study within the JANUS cohort in Norway. Int. J. Cancer 2014, 134, 197–206. [Google Scholar] [CrossRef]
- Schmidt, J.A.; Fensom, G.K.; Rinaldi, S.; Scalbert, A.; Appleby, P.N.; Achaintre, D.; Gicquiau, A.; Gunter, M.J.; Ferrari, P.; Kaaks, R.; et al. Pre-diagnostic metabolite concentrations and prostate cancer risk in 1077 cases and 1077 matched controls in the European Prospective Investigation into Cancer and Nutrition. BMC Med. 2017, 15, 122. [Google Scholar] [CrossRef]
- Andras, I.; Crisan, N.; Vesa, S.; Rahota, R.; Romanciuc, F.; Lazar, A.; Socaciu, C.; Matei, D.V.; De Cobelli, O.; Bocsan, I.S.; et al. Serum metabolomics can predict the outcome of first systematic transrectal prostate biopsy in patients with PSA < 10 ng/mL. Future Oncol. 2017, 13, 1793–1800. [Google Scholar]
- Kumar, D.; Gupta, A.; Mandhani, A.; Sankhwar, S.N. Metabolomics-derived prostate cancer biomarkers: Fact or fiction? J. Proteome Res. 2015, 14, 1455–1464. [Google Scholar] [CrossRef] [PubMed]
- Kumar, D.; Gupta, A.; Mandhani, A.; Sankhwar, S.N. NMR spectroscopy of filtered serum of prostate cancer: A new frontier in metabolomics. Prostate 2016, 76, 1106–1119. [Google Scholar] [CrossRef]
- Giskeødegård, G.F.; Hansen, A.F.; Bertilsson, H.; Gonzalez, S.V.; Kristiansen, K.A.; Bruheim, P.; Mjøs, S.A.; Angelsen, A.; Bathen, T.F.; Tessem, M.B. Metabolic markers in blood can separate prostate cancer from benign prostatic hyperplasia. Br. J. Cancer 2015, 113, 1712. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Murphy, T.B.; Byrne, J.C.; Brennan, L.; Fitzpatrick, J.M.; Watson, R.W.G. Applying random forests to identify biomarker panels in serum 2D-DIGE data for the detection and staging of prostate cancer. J. Proteome Res. 2011, 10, 1361–1373. [Google Scholar] [CrossRef]
- Osl, M.; Dreiseitl, S.; Pfeifer, B.; Weinberger, K.; Klocker, H.; Bartsch, G.; Schäfer, G.; Tilg, B.; Graber, A.; Baumgartner, C. A new rule-based algorithm for identifying metabolic markers in prostate cancer using tandem mass spectrometry. Bioinformatics 2008, 24, 2908–2914. [Google Scholar] [CrossRef] [PubMed]
- Zang, X.; Jones, C.M.; Long, T.Q.; Monge, M.E.; Zhou, M.; Walker, L.D.; Mezencev, R.; Gray, A.; McDonald, J.F.; Fernandez, F.M. Feasibility of detecting prostate cancer by ultraperformance liquid chromatography–mass spectrometry serum metabolomics. J. Proteome Res. 2014, 13, 3444–3454. [Google Scholar] [CrossRef] [PubMed]
- Dereziński, P.; Klupczynska, A.; Sawicki, W.; Pałka, J.A.; Kokot, Z.J. Amino acid profiles of serum and urine in search for prostate cancer biomarkers: A pilot study. Int. J. Med. Sci. 2017, 14, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Sreekumar, A.; Poisson, L.M.; Rajendiran, T.M.; Khan, A.P.; Cao, Q.; Yu, J.; Laxman, B.; Mehra, R.; Lonigro, R.J.; Li, Y.; et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature 2009, 457, 910–914. [Google Scholar] [CrossRef] [PubMed]
- Thysell, E.; Surowiec, I.; Hörnberg, E.; Crnalic, S.; Widmark, A.; Johansson, A.I.; Stattin, P.; Bergh, A.; Moritz, T.; Antti, H.; et al. Metabolomic characterization of human prostate cancer bone metastases reveals increased levels of cholesterol. PLoS ONE 2010, 5, e14175. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Liu, X.; Jiao, L.; Xu, C.; Zhang, Z.; Wang, L.; Li, Y.; Yang, C.; Zhang, W.; Sun, Y. Metabolomic evaluation of the response to endocrine therapy in patients with prostate cancer. Eur. J. Pharmacol. 2014, 729, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Stabler, S.; Koyama, T.; Zhao, Z.; Martinez-Ferrer, M.; Allen, R.H.; Luka, Z.; Loukachevitch, L.V.; Clark, P.E.; Wagner, C.; Bhowmick, N.A. Serum methionine metabolites are risk factors for metastatic prostate cancer progression. PLoS ONE 2011, 6, e22486. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Watson, D.G.; Wang, L.; Abbas, M.; Murdoch, L.; Bashford, L.; Ahmad, I.; Lam, N.Y.; Ng, A.C.; Leung, H.Y. Application of holistic liquid chromatography-high resolution mass spectrometry based urinary metabolomics for prostate cancer detection and biomarker discovery. PLoS ONE 2013, 8, e65880. [Google Scholar] [CrossRef] [PubMed]
- Gkotsos, G.; Virgiliou, C.; Lagoudaki, I.; Sardeli, C.; Raikos, N.; Theodoridis, G.; Dimitriadis, G. The role of sarcosine, uracil, and kynurenic acid metabolism in urine for diagnosis and progression monitoring of prostate cancer. Metabolites 2017, 7, 9. [Google Scholar] [CrossRef] [PubMed]
- Gamagedara, S.; Kaczmarek, A.T.; Jiang, Y.; Cheng, X.; Rupasinghe, M.; Ma, Y. Validation study of urinary metabolites as potential biomarkers for prostate cancer detection. Bioanalysis 2012, 4, 1175–1183. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Peralbo, M.; Gómez-Gómez, E.; Calderón-Santiago, M.; Carrasco-Valiente, J.; Ruiz-García, J.; Requena-Tapia, M.; De Castro, M.L.; Priego-Capote, F. Prostate cancer patients–negative biopsy controls discrimination by untargeted metabolomics analysis of urine by LC-QTOF: Upstream information on other omics. Sci. Rep. 2016, 6, 38243. [Google Scholar] [CrossRef] [PubMed]
- Khalid, T.; Aggio, R.; White, P.; Costello, B.D.L.; Persad, R.; Al-Kateb, H.; Jones, P.; Probert, C.S.; Ratcliffe, N. Urinary volatile organic compounds for the detection of prostate cancer. PLoS ONE 2015, 10, e0143283. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Rambla, C.; Puchades-Carrasco, L.; García-Flores, M.; Rubio-Briones, J.; López-Guerrero, J.A.; Pineda-Lucena, A. Non-invasive urinary metabolomic profiling discriminates prostate cancer from benign prostatic hyperplasia. Metabolomics 2017, 13, 52. [Google Scholar] [CrossRef]
- Struck-Lewicka, W.; Kordalewska, M.; Bujak, R.; Mpanga, A.Y.; Markuszewski, M.; Jacyna, J.; Matuszewski, M.; Kaliszan, R.; Markuszewski, M.J. Urine metabolic fingerprinting using LC–MS and GC–MS reveals metabolite changes in prostate cancer: A pilot study. J. Pharm. Biomed. Anal. 2015, 111, 351–361. [Google Scholar] [CrossRef]
- Kosti, O.; Xu, X.; Veenstra, T.D.; Hsing, A.W.; Chu, L.W.; Goldman, L.; Bebu, I.; Collins, S.; Dritschilo, A.; Lynch, J.H.; et al. Urinary estrogen metabolites and prostate cancer risk: A pilot study. Prostate 2011, 71, 507–516. [Google Scholar] [CrossRef]
- Wu, H.; Liu, T.; Ma, C.; Xue, R.; Deng, C.; Zeng, H.; Shen, X. GC/MS-based metabolomic approach to validate the role of urinary sarcosine and target biomarkers for human prostate cancer by microwave-assisted derivatization. Anal. Bioanal. Chem. 2011, 401, 635–646. [Google Scholar] [CrossRef]
- Cao, D.L.; Ye, D.W.; Zhang, H.L.; Zhu, Y.; Wang, Y.X.; Yao, X.D. A multiplex model of combining gene-based, protein-based, and metabolite-based with positive and negative markers in urine for the early diagnosis of prostate cancer. Prostate 2011, 71, 700–710. [Google Scholar] [CrossRef] [PubMed]
- Huan, T.; Troyer, D.A.; Li, L. Metabolite Analysis and Histology on the Exact Same Tissue: Comprehensive Metabolomic Profiling and Metabolic Classification of Prostate Cancer. Sci. Rep. 2016, 6, 32272. [Google Scholar] [CrossRef] [PubMed]
- Ren, S.; Shao, Y.; Zhao, X.; Hong, C.S.; Wang, F.; Lu, X.; Li, J.; Ye, G.; Yan, M.; Zhuang, Z.; et al. Integration of metabolomics and transcriptomics reveals major metabolic pathways and potential biomarker involved in prostate cancer. Mol. Cell. Proteom. 2016, 15, 154–163. [Google Scholar] [CrossRef] [PubMed]
- McDunn, J.E.; Li, Z.; Adam, K.P.; Neri, B.P.; Wolfert, R.L.; Milburn, M.V.; Lotan, Y.; Wheeler, T.M. Metabolomic signatures of aggressive prostate cancer. Prostate 2013, 73, 1547–1560. [Google Scholar] [CrossRef] [PubMed]
- Shuster, J.R.; Lance, R.S.; Troyer, D.A. Molecular preservation by extraction and fixation, mPREF: A method for small molecule biomarker analysis and histology on exactly the same tissue. BMC Clin. Pathol. 2011, 11, 14. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Bai, X.; Liu, Y.; Wang, W.; Han, J.; Wang, Q.; Xu, Y.; Zhang, C.; Zhang, S.; Li, X.; et al. Topologically inferring pathway activity toward precise cancer classification via integrating genomic and metabolomic data: Prostate cancer as a case. Sci. Rep. 2015, 5, 13192. [Google Scholar] [CrossRef] [PubMed]
- Jung, K.; Reszka, R.; Kamlage, B.; Bethan, B.; Stephan, C.; Lein, M.; Kristiansen, G. Tissue metabolite profiling identifies differentiating and prognostic biomarkers for prostate carcinoma. Int. J. Cancer 2013, 133, 2914–2924. [Google Scholar] [CrossRef]
- Priolo, C.; Pyne, S.; Rose, J.; Regan, E.R.; Zadra, G.; Photopoulos, C.; Cacciatore, S.; Schultz, D.; Scaglia, N.; McDunn, J.E.; et al. AKT1 and MYC induce distinctive metabolic fingerprints in human prostate cancer. Cancer Res. 2014, 74, 7198–7204. [Google Scholar] [CrossRef]
- Cacciatore, S.; Zadra, G.; Bango, C.; Penney, K.L.; Tyekucheva, S.; Yanes, O.; Loda, M. Metabolic profiling in formalin-fixed and paraffin-embedded prostate cancer tissues. Mol. Cancer Res. 2017. [Google Scholar] [CrossRef]
- Wang, X.; Han, J.; Hardie, D.B.; Yang, J.; Pan, J.; Borchers, C.H. Metabolomic profiling of prostate cancer by matrix assisted laser desorption/ionization-Fourier transform ion cyclotron resonance mass spectrometry imaging using Matrix Coating Assisted by an Electric Field (MCAEF). Biochim. Biophys. Acta (BBA) Proteins Proteom. 2017, 1865, 755–767. [Google Scholar] [CrossRef]
- Brown, M.V.; McDunn, J.E.; Gunst, P.R.; Smith, E.M.; Milburn, M.V.; Troyer, D.A.; Lawton, K.A. Cancer detection and biopsy classification using concurrent histopathological and metabolomic analysis of core biopsies. Genome Med. 2012, 4, 33. [Google Scholar] [CrossRef] [PubMed]
- Giskeødegård, G.F.; Bertilsson, H.; Selnæs, K.M.; Wright, A.J.; Bathen, T.F.; Viset, T.; Halgunset, J.; Angelsen, A.; Gribbestad, I.S.; Tessem, M.B. Spermine and citrate as metabolic biomarkers for assessing prostate cancer aggressiveness. PLoS ONE 2013, 8, e62375. [Google Scholar]
- Selnæs, K.M.; Gribbestad, I.S.; Bertilsson, H.; Wright, A.; Angelsen, A.; Heerschap, A.; Tessem, M.B. Spatially matched in vivo and ex vivo MR metabolic profiles of prostate cancer–investigation of a correlation with Gleason score. NMR Biomed. 2013, 26, 600–606. [Google Scholar] [CrossRef] [PubMed]
- Sandsmark, E.; Hansen, A.F.; Selnæs, K.M.; Bertilsson, H.; Bofin, A.M.; Wright, A.J.; Viset, T.; Richardsen, E.; Drabløs, F.; Bathen, T.F.; et al. A novel non-canonical Wnt signature for prostate cancer aggressiveness. Oncotarget 2017, 8, 9572–9586. [Google Scholar] [CrossRef] [PubMed]
- Hansen, A.F.; Sandsmark, E.; Rye, M.B.; Wright, A.J.; Bertilsson, H.; Richardsen, E.; Viset, T.; Bofin, A.M.; Angelsen, A.; Selnæs, K.M.; et al. Presence of TMPRSS2-ERG is associated with alterations of the metabolic profile in human prostate cancer. Oncotarget 2016, 7, 42071. [Google Scholar] [CrossRef] [PubMed]
- Jentzmik, F.; Stephan, C.; Lein, M.; Miller, K.; Kamlage, B.; Bethan, B.; Kristiansen, G.; Jung, K. Sarcosine in prostate cancer tissue is not a differential metabolite for prostate cancer aggressiveness and biochemical progression. J. Urol. 2011, 185, 706–711. [Google Scholar] [CrossRef] [PubMed]
- Meller, S.; Meyer, H.A.; Bethan, B.; Dietrich, D.; Maldonado, S.G.; Lein, M.; Montani, M.; Reszka, R.; Schatz, P.; Peter, E.; et al. Integration of tissue metabolomics, transcriptomics and immunohistochemistry reveals ERG-and gleason score-specific metabolomic alterations in prostate cancer. Oncotarget 2016, 7, 1421. [Google Scholar] [CrossRef]
- Keshari, K.; Tsachres, H.; Iman, R.; Delos Santos, L.; Tabatabai, Z.; Shinohara, K.; Vigneron, D.; Kurhanewicz, J. Correlation of phospholipid metabolites with prostate cancer pathologic grade, proliferative status and surgical stage–impact of tissue environment. NMR Biomed. 2011, 24, 691–699. [Google Scholar] [CrossRef]
- Zhang, V.; Westphalen, A.; Delos Santos, L.; Tabatabai, Z.; Shinohara, K.; Vigneron, D.; Kurhanewicz, J. The role of metabolic imaging in radiation therapy of prostate cancer. NMR Biomed. 2014, 27, 100–111. [Google Scholar] [CrossRef]
- Maxeiner, A.; Adkins, C.B.; Zhang, Y.; Taupitz, M.; Halpern, E.F.; McDougal, W.S.; Wu, C.L.; Cheng, L.L. Retrospective analysis of prostate cancer recurrence potential with tissue metabolomic profiles. Prostate 2010, 70, 710–717. [Google Scholar] [CrossRef]
- Clos-Garcia, M.; Loizaga-Iriarte, A.; Zuñiga-Garcia, P.; Sánchez-Mosquera, P.; Rosa Cortazar, A.; González, E.; Torrano, V.; Alonso, C.; Pérez-Cormenzana, M.; Ugalde-Olano, A.; et al. Metabolic alterations in urine extracellular vesicles are associated to prostate cancer pathogenesis and progression. J. Extracell. Vesicles 2018, 7, 1470442. [Google Scholar] [CrossRef] [PubMed]
- Puhka, M.; Takatalo, M.; Nordberg, M.E.; Valkonen, S.; Nandania, J.; Aatonen, M.; Yliperttula, M.; Laitinen, S.; Velagapudi, V.; Mirtti, T.; et al. Metabolomic profiling of extracellular vesicles and alternative normalization methods reveal enriched metabolites and strategies to study prostate cancer-related changes. Theranostics 2017, 7, 3824. [Google Scholar] [CrossRef] [PubMed]
- Roberts, M.J.; Richards, R.S.; Chow, C.W.; Buck, M.; Yaxley, J.; Lavin, M.F.; Schirra, H.J.; Gardiner, R.A. Seminal plasma enables selection and monitoring of active surveillance candidates using nuclear magnetic resonance-based metabolomics: A preliminary investigation. Prostate Int. 2017, 5, 149–157. [Google Scholar] [CrossRef] [PubMed]
- D’amico, A.V.; Whittington, R.; Malkowicz, S.B.; Schultz, D.; Blank, K.; Broderick, G.A.; Tomaszewski, J.E.; Renshaw, A.A.; Kaplan, I.; Beard, C.J.; et al. Biochemical outcome after radical prostatectomy, external beam radiation therapy, or interstitial radiation therapy for clinically localized prostate cancer. JAMA 1998, 280, 969–974. [Google Scholar] [CrossRef] [PubMed]
- Weis, J.; von Below, C.; Tolf, A.; Ortiz-Nieto, F.; Wassberg, C.; Haggman, M.; Ladjevardi, S.; Ahlstrom, H. Quantification of metabolite concentrations in benign and malignant prostate tissues using 3D proton MR spectroscopic imaging. J. Magn. Reson. Imaging 2017, 45, 1232–1240. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, R.; Iqbal, Z.; Burns, B.; Wilson, N.E.; Sarma, M.K.; Margolis, D.A.; Reiter, R.E.; Raman, S.S.; Thomas, M.A. Accelerated echo planar J-resolved spectroscopic imaging in prostate cancer: A pilot validation of nonlinear reconstruction using total variation and maximum entropy. NMR Biomed. 2015, 28, 1366–1373. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, R.; Margolis, D.; Raman, S.; Sarma, M.K.; Sheng, K.; King, C.R.; Verma, G.; Sayre, J.; Reiter, R.E.; Thomas, M.A. MR spectroscopic imaging and diffusion-weighted imaging of prostate cancer with Gleason scores. J. Magn. Reson. Imaging 2012, 36, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Kobus, T.; Hambrock, T.; Hulsbergen-van de Kaa, C.A.; Wright, A.J.; Barentsz, J.O.; Heerschap, A.; Scheenen, T.W. In vivo assessment of prostate cancer aggressiveness using magnetic resonance spectroscopic imaging at 3 T with an endorectal coil. Eur. Urol. 2011, 60, 1074–1080. [Google Scholar] [CrossRef]
- Wang, X.Z.; Wang, B.; Gao, Z.Q.; Liu, J.G.; Liu, Z.Q.; Niu, Q.L.; Sun, Z.K.; Yuan, Y.X. 1H-MRSI of prostate cancer: The relationship between metabolite ratio and tumor proliferation. Eur. J. Radiol. 2010, 73, 345–351. [Google Scholar] [CrossRef]
- Kumar, R.; Nayyar, R.; Kumar, V.; Gupta, N.P.; Hemal, A.K.; Jagannathan, N.R.; Dattagupta, S.; Thulkar, S. Potential of magnetic resonance spectroscopic imaging in predicting absence of prostate cancer in men with serum prostate-specific antigen between 4 and 10 ng/ml: A follow-up study. Urology 2008, 72, 859–863. [Google Scholar] [CrossRef]
- Li, C.; Han, J.; Yao, Q.; Zou, C.; Xu, Y.; Zhang, C.; Shang, D.; Zhou, L.; Zou, C.; Sun, Z.; et al. Subpathway-GM: Identification of metabolic subpathways via joint power of interesting genes and metabolites and their topologies within pathways. Nucleic Acids Res. 2013, 41, e101. [Google Scholar] [CrossRef] [PubMed]
- Cernei, N.; Heger, Z.; Gumulec, J.; Zitka, O.; Masarik, M.; Babula, P.; Eckschlager, T.; Stiborova, M.; Kizek, R.; Adam, V. Sarcosine as a potential prostate cancer biomarker—A review. Int. J. Mol. Sci. 2013, 14, 13893–13908. [Google Scholar] [CrossRef] [PubMed]
- Garde, A.H.; Hansen, Å.M.; Kristiansen, J.; Knudsen, L.E. Comparison of uncertainties related to standardization of urine samples with volume and creatinine concentration. Ann. Occup. Hyg. 2004, 48, 171–179. [Google Scholar] [PubMed]
| Author | Year | Sample Size | Technique | Use Case | Biomarkers Candidates | Validated? | Statistical Details | Comments |
|---|---|---|---|---|---|---|---|---|
| Schmidt [19] | 2017 | PCa: 1077 Controls: 1077 | PCa risk | citrulline | No | Conditional logistic regression | Study population from European Prospective Investigation into Cancer and Nutrition (EPIC) Fasting not required | |
| Huang [16] | 2017 | n = 338 PCa: 72 (T2), 51 (T3), 15 (T4) Controls: 200 | LC-MS GC-MS | Staging PCa risk | N-acetyl-3-methylhistidine (T2)lycerophospholipid oleoyl-linoleoyl-GPI (T3) | No | logistic regression p = 0.05 | Study population from the ATBC Study cohort Overnight fasting serum Time serum collection - Dx avrg = 10 years (range 1–20) |
| Andras [20] | 2017 | Training Set: n = 59 Validation Set: n = 31 | HPLC- ESI+ QTOF - MS | Diagnosis | lisophosphatidylcholine 18:2, homocysteine-inosine, methyladenosine, lipoicacid, hydroxymelatonin and decanoilcarnitine | Yes | Mann–Whitney test PLSR-DA ROC analysis | Assessement of the predective value of metabolomic analysis for the presence of PCa at the first systematic biopsy |
| Dereziński [27] | 2017 | n = 89 PCa: 49 HC: 40 | LC-ESI-MS/MS | Diagnosis | methionine, ethanolamine, glutamine, isoleucine, arginine, leucine | No | Mann–Whitney U test Student’s t-test Welch’s F test PLS-DAROC analysisv discriminant function analysis | targeted analysis of 32 amino acids in serum urine samples were profiled as well |
| Kumar [22] | 2016 | n = 210 HC: 65 BPH: 70 PCa: 75 | NMR | Diagnosis | HC vs. PCa: alanine, sarcosine, glycine, citrate BPH vs. PCa alanine, sarcosine, glycine, citrate, creatinine HC vs. BPH + PC: aglycine, sarcosine, alanine, creatine, xanthine, and hypoxanthine HC vs. BPH: glycine, xanthine, pyruvate, methylhistidine, and creatinine | Yes | ANOVA Student–Newman–Keuls test DFA ROC analysis | Overnight fasting |
| Huang [17] | 2016 | PCa: 380 controls: 380 | UPLC-MS GC-MS | PCa risk | pyroglutamine, gamma-glutamylphenylalanine, phenylpyruvate, N-acetylcitrulline, and stearoylcarnitine | No | Conditional logistic regression p = 0.000072 | Study population from the Prostate, Lung, Colorectal and Ovarian Cancer Screening Trial (PLCO) Fasting not required |
| Giskeødegård [23] | 2015 | n = 50 PCa: 29 BPH: 21 | MRS GC-MS | Diagnosis | decanoylcarnitine (c10), tetradecenoylcarnitine (c14:1), octanoylcarnitine (c8), dimethylsulfone, phenylalanine, lysine, phosphatidylcholine diacyl C34:4, lipid signals -(CH2)n-CH2-CH2-CO | No | PCA (no discrimination) OPLS-DA and OPLS Wilcoxon rank sum testing (p ≤ 0.05) ROC analyses | Fasting serum and plasma samples Missing data for some variables were replaced by estimated values using a built-in data imputation algorithm |
| Mondul [15] | 2015 | n = 400 PCa: 200 (100 aggressive) Controls: 200 | UPLC-MS GC-MS | PCa risk | Inositol-1-phosphate oleoyl-linoleoylglycerophosphoinositol, 1-stearoylglycerophosphoglycerol, stearate and docosadienoate. Both alpha-ketoglutarate and citrate were associated with aggressivedisease risk as were elevated thyroxine and trimethylamine oxide | No | Conditional logistic regression Threshold for statistical significance p = 0.003 in the main analysis | Study population: from Alpha-Tocopherol, Beta-Carotene Cancer Prevention Study cohort Fasting serum collected up to 20 years prior to case diagnoses Missing values were assigned the minimum nonmissing value. |
| Kumar [21] | 2015 | n = 102 PCa: 70 (40 low grade PCa, 30 high grade PCa) HC: 32 | NMR | Diagnosis Staging | HC vs. PCa: alanine, pyruvate, glycine, sarcosine low grade PCa vs. high grade PCa: alanine, pyruvate, and glycine | No | Unsupervised PCA supervised OPLS-DA ANOVA Student–Newman–Keuls test ROC analysis | Fasting serum samples |
| Mondul [14] | 2014 | PCa: 74 Controls: 74 | UPLC-MS GC-MS | PCa risk | 1-stearoylglycerol Glycerolalpha-ketoglutarate | No | Logistic regression threshold for statistical significance: 0.000119 | Study population from the Alpha-Tocopherol, Beta-Carotene Cancer Prevention (ATBC)study Overnight fasting serum |
| Zang [26] | 2014 | n = 114 PCa: 64 Controls: 50 | UPLC-LS/MS | Diagnosis | fatty acids, amino acids, lysophospholipids, and bile acids | No | SVM PCA significance level 0.05 | 40 discriminant metabolites are found Only the top ranking ones are presented here |
| Huang [30] | 2014 | n = 72 newly diagnosed PCa: 18 HC: 18 good ADT responders: 18 poor ADT responders: 18 | LC-MS | Therapy prediction | deoxycholic acid (DCA), glycochenodeoxycholate (GCDC), L-tryptophan, docosapentaenoic acid (DPA), arachidonic acid, deoxycytidine triphosphate, and pyridinoline | No | PLS-DA and OPLS ANOVA Statistical significance p = 0.05 | Fasting serum (overnight fast) |
| de Vogel [18] | 2014 | PCa: 3000 Controls: 3000 | LC-MSGC-MS | PCa risk | Sarcosine; glycine | No | condiotional logistic regression | Study population within the JANUS cohort Fasting status: unknown |
| Fan [24] | 2010 | n = 56 PCa: 42 (20 GS5, 22 GS7) BPH: 14 | NMR | Diagnosis | glutamate and formate | No | ANOVA Random Forests | Article focusing on proteomics by applying RF to 2D-DIGE data NMR data were also presented but with much lower predictive performance |
| Osl [25] | 2008 | n = 320 Controls: 114 PCa: 206 | FIA-MS/MS LC-MS/MS | Diagnosis Staging | Diagnosis: PC a C16:0, PC a C18:0, Serotonin, Aspartate, OrnithineStaging: No reliable biomarkers | No | Associative Voting algorithm Logistic regression | Study population: Men participation in PCa screening One simple rejected |
| Stabler [31] | 2011 | n = 58 patients after radical prostatectomy recurrent free: 30 recurrent: 28 | GC-MS | Prognosis | homocysteine, cystathionine, cysteine | No | Wilcoxon rank sum test Logistic regression Likelihood ratio ROC analysis Kaplan Meier plots Cox proportional hazard regressio models | analysed both serum and urine samples targeted analysis of sarcosine, dimethylglycine, methionine, homocysteine, vystathionine, cysteine, methylmalonic acid, methylcitrate |
| Thysell [29] | 2010 | PCa with metastases: 7 PCa w/o metastases: 6 benign: 17 | GC-TOFMS | Staging | pseudouridine, creatinine, glucose, glutamate, taurine, phenylalanine, stearate | No | OPLS-DA Mann–Whitney U-test | main work was done on tissue extracts from fresh-frozen biopsies of bone metastases and from biopsies of primary PCa and benign prostate all patients were selected to have hihg-risk tumours (i.e., presence of bone metastases, locally advanced tumour or poorly differnitated cancer) |
| Sreekumar [28] | 2009 | n = 42 tissue samples benign adjacent: 16 localized PCa: 12 metastatic PCa: 14 | UHPLC-MS/MS GC-MS | Diagnosis | blood metabolites not reported | No | Wilcoxon rank-sum test | study analyzed tissue, blood, and urine samples but focused on tissue data only |
| Author | Year | Sample Size | Technique | Use Case | Biomarkers Candidates | Validated? | Statistical Details | Comments |
|---|---|---|---|---|---|---|---|---|
| Pérez-Rambla [37] | 2017 | n = 115 PCa: 64B BPH: 51 | 1H-NMR | Diagnosis | branched-chain amino acids (BCAA), glutamate and pseudouridine glycine, dimethylglycine, fumarate and 4-imidazole-acetate. | No | PCA analysis OPLS-DA p = 0.01 | |
| Gkotsos [33] | 2017 | PCa: 32 elevated PSA: 101 HC: 15 | UPLC-MS/MS | Diagnosis Prognosis | kynurenic acid | No | ROC analysis p = 0.05 | Ultrasonographically-guided prostatic biopsy collected before and after prostatic massage for the 101 patients Urinary concentrations of metabolites were not normalized to urinary creatinine The control group was recruited only from individuals presenting for other etiologies and these patients usually are much younger than those presenting prostate cancer. |
| Fernández- Peralbo [35] | 2016 | n = 104 PCa: 62 HC: 42 | LC-QTOF | Diagnosis | 28 significant metabolites | No | PLS-DA t-test with Benjamini–Hochberg false discovery rate p = 0.05 | Morning urine samples The controls are negative biopsy individualsCases are patients with significant PCa confirmed by prostate biopsy MSTUS was selected for normalization ofurine samples |
| Tanzeela [36] | 2015 | PCa: 59 HC: 43 | GC/MS | Diagnosis | 2,6-dimethyl-7-octen-2-ol, pentanal, 3-octanone, and 2-octanone | No | RF LDA | Urine samples were obtained at different times of the day samples were classified as prostate cancer or controls after pathological examination of the biopsy specimens |
| Struck-Lewicka [38] | 2015 | PCa: 32 HC: 32 | HPLC-TOF/MS GC-QqQ/MS | Diagnosis | metabolites involved in biochemical pathways like AA, purine and glucose metabolism as well as urea and TCA cycle | No | PCA analysis PLS-DA p = 0.05 | |
| Zhang [32] | 2013 | LC-HRMS HILICRP | Diagnosis | ureido isobutyric acid, indolylacryloyglycine, acetylvanilalinine and 2-oxoglutarate | Yes (PCa:30) | OPLS-DA ROC analysis p = 0.05 | Urine samples were stored at Three independent normalisation methodsTesting against a new cohort of patients. | |
| Gamagedara [34] | 2012 | PCa: 63 HC: 68 | LC-MS/MS | Diagnosis Prognosis | proline, kynurenine, uraciland glycerol-3-phosphate | No | Linear regression PCA analysis CART p = 0.05. | Validation study |
| Wu [40] | 2011 | PCa: 20 BPH: 8 HC: 20 | ID GC/MS MAD | Diagnosis | PCa vs. HC: Propenoic acid, Pyrimidine, Dihyroxybutanoic acid, Creatinine, Purine, Purine, Glucopyranoside, Ribofuranoside, Xylonic acid, Xylopyranose PCa vs. BPH: Dihyroxybutanoic acid, Pyrimidine, Xylonic acid, Xylopyranose, Ribofuranoside | No | PCA analysis ROC analysis p = 0.05 | |
| Cao [41] | 2010 | PCa: 86 HC: 45 | LC-MS | Diagnosis | Sarcosine | No | Logistic regression ROC analysis p = 0.05 | First voided urines after digital rectalexamination (DRE) |
| Kosti [39] | 2010 | PCa: 77 HC: 77 Biopy controls: 37 | LC-MS | Predection | 16-ketoestradiol 17-epiestriol | No | Logistic regression | The biopsy controls had elevated PSA due to BPH (n = 27) or other urologic conditions (10) |
| Dereziński [27] | 2017 | PCa: 49 HC: 40 | LC-ESI-MS/MS | Diagnosis | -amino-n-butyric acid, phosphoethanolamine, ethanolamine, homocitrulline, arginine, -hydroxylysine and asparagine | No | ROC analysis PLS-DA p = 0.05 | Evaluation of free amino acid profiles in both urine and serum samples from the same patiens Controls recruited among men subjected to the routine periodic medical examination Sample collection period over 3 months |
| Stabler [31] | 2011 | PCa: 54 | GC-MS | Reccurence | sarcosine and cysteine | No | Logistic regression p = 0.007 | Study conducted on both urine and serum Subjestcs were divided into 2 groups: - Cases who developed biochemical recurrnce within 2 years - Controls who remained recurrence-free after 5 years. |
| Author | Year | Sample Size | Technique | Use Case | Biomarkers Candidates | Validated? | Statistical Details | Comments |
|---|---|---|---|---|---|---|---|---|
| Wang [50] | 2017 | n = 3 subjects Cancer tissue: 3 Benign adjacent tissue: 3 | MALDI-FTICR-MS | Diagnosis | differential metabolites were not mentioned | No | students t-test | coated tissue slice |
| Huan [42] | 2016 | Training set: n = 16 Pca: 13; Benign: 12 Validation set 1: n = 18 Pca: 19; Benign: 17 Validation set 2: n = 12 Pca: 12; benign: 12 | LC QTOF-MS | Diagnosis | adenosine monophosphate, spermidine, uracil, ophthalmic acid + HPO3, 2,3-diaminopropionic acid + HPO3 + 2 unknown metabolites = putative identification | Yes | OPLS-DA volcano plot | Tissue extracts from core biopsies after prostatectomy |
| McDunn [44] | 2013 | Pca: 331 Benign: 178 (matched to 178 of the Pca samples) | UHPLC-MS/MS GC-MS | Staging | aggressive Pca:ADP, Glucose, 6-sialyl-N-actyllyctosamine, 2-hydroxypalmitate, 5,6 dihydrouracil, choline, fumarate, kynurenine, phophate, 2-hydrxoystearate, Ac-SDKP, choline phosphate, glycerol-3-phophate, n-acetylaspartate Gleason pattern progression:proline, malate, ADP-ribose, 6-sialyl-N-acetyllactosamineextracapsular extension:NAD+, N-acetylaspartate, putrescine, Glucose Tumor spread (regional lymph nodes/seminal vesicles):choline phosphate, Glycerol3-phophate, putrescine, 6-sialyl-N-acetyllactosamine | No | paired t-testWilcoxon test linear regression | Tissue extract from OCT embedded tissue from prostatectomies |
| Jung [47] | 2013 | n = 95 matched cancer and benign adjacent tissue | LC-MS/MS GC-MS | Diagnosis Prognosis Biological recurrence | Diagnosis of Pca:2-hdroxybehenic acid, crebronic acid, tricosanoic acid, glycerophophoethanolamine, isopentenyl pyrophosphate, 7-methylguanine, 2-aminoadipic acid, gluconic acid, maltotriose, tricosanoic acid Prediction of biological recurrence risk:2-aminoadipic acid, gluconic acid, maltotriose | No | Wilcoxon paired test ROC analysis logistic regression Kaplan–Meier curves univariate and multivariate Cox regression | tissue extracts of punch biopsy from cryosections after prostatectomy |
| Brown [51] | 2012 | n = 8 matched cancer and benign adjacent tissue | UHPLC-MS/MS GC-MS | Diagnosis | > 40 metabolites not specified further | No | Welch’s two sample t-test hierachical clustering | tissue extracts from core biopsy after prostatectomy |
| Selnæs [53] | 2012 | n = 13 subjects 40 tissue samples | HR-MAS-1H-MRS | Staging | CCS/C ratio (choline+creatine+spermine over citrate) | No | Spearman’s rank correlation Jonchheere– Terpstra test | intact tissue from needle biopsy after prostatectomy |
| Maxeiner [60] | 2010 | Pca with biological recurrence: 16 Pca without biological recurrence: 32 | HR-MAS-1H-MRS | Prognosis | spermine, glutamine, myo-inositol, phophoryl choline, scylloinositol, glutamate | Yes | PCA student’s t-test Canonical analysis ANOVA ROC analysis | intact tissue from needle biopsy after prostatectomy;training and test set with identical case group but distinct control groups |
| Sreekumar [28] | 2009 | n = 42 tissue samples benign adjacent: 16 localized Pca: 12 metastatic Pca: 14 | UHPLC-MS/MS GC-MS | Diagnosis | sarcosine, uracil, kynurenine, glycerol-3-phophate, leucine, proline | Yes | Wilcoxon rank-sum test | tissue extracts from biopsy samplesonly sarcosine was analyzed in validation set |
| Cacciatore [49] | 2017 | matched benign and Pca samples Training set: n = 8 Validation set: n = 4 | UHPLC-MS/MS GC-MS | Diagnosis | 32 metabolites reported; biomarkers included in the model not specified | Yes | Hierarchical clustering OSC-PLS | tissue extracts from tissue section after prostatectomyalso compared OCT-embedded and FFPE tissue as biospecimen |
| Sandsmark [54] | 2017 | n = 41 subjects Pca: 95 benign adjacent: 34 | HR-MAS-1H-MRS | Diagnosis | Pca with high vs. Pca with low/intermediate NCWP-EMT score: spermine and citratePca with low vs. Pca with high NCWP-EMT score taurine, phosphoethanolamine | No | t-test | intact tissue from prostatectomiesmain focus: alterations in non-canonical WNT signaling pathway (NCWP) and EMT in Pcajoint gene expression and metabolomic analyses; targeted analysis of 23 metabolites; metabolomics was performed only on the main cohort |
| Hansen [55] | 2016 | n = 41 subjects Pca: 95 benign adjacent: 34 | HR-MAS MRS | Staging | citrate, spermine (correlated with presence ERG translocation) | Yes | unsupervised multivariate analysis PLS-DA | gene expression analysis and TMPRSS2-ERG as marker for disease aggressiveness;intact tissue from tissue slices collected from prostatectomies; analyzed metabolic alterations in PCA patients positive for TMPRSS2-ERG/high ERG gene fusion; targeted analysis of 23 metabolites |
| Meller [57] | 2016 | n = 106 subjects matched cancer and benign adjacent tissue ERG-positive Pca: 27 ERG-negative Pca: 23 | GC-MS LC-MS | Staging Prognosis Biological recurrence | Gleason score:pantothenic acid, maltose, fructose-6-phosphate, gluconic acid, cholesterol ERG status:maltotriose, gluconic acid, citrate, cis-aconitate, spermine, putrescine, cerebronic acid, 2-hydroxybehenic acid, tricosanoic acid, Biological relapse:tyrosine and tryptophan | No | ANOVA PCA | tissue extracts of punch biopsy from cryosections after prostatectomy |
| Ren [43] | 2016 | Training set:25 paired PCa and adjacent benign Validation set:51 paired Pca and adjacent benign 16 BPH | LC-MS | Diagnosis | sphingosine, citicoline, choline, pantothenic acid, carnitine C4-OH, GPC, NAD, phenylacetyl-glutamine, carnitine C14:2 | Yes | PCA PLS-DA Signrank Wilcoxon signed rank two-sides test (biomarker analysis) | joint transciptomics and metabolomics to identify altered metabolic pathways in PCA tissue;tissue extracts from prostatectomies |
| Liu [46] | 2015 | n = 42 tissue samples benign adjacent: 16 localized Pca: 12 metastatic Pca: 14 | n.a. | Diagnosis | Proline, Cholesterol, sarcosine, spermidine, spermine, Putrescine, 4-Acetamidobutanoate | Yes | DRW-GM + logistic regression | joint analysis of genomic and metabolomic data and pathway topology using directed random walk on a global gene-metabolite pathway graph;used dataset established by Sreekumar et al. 2009 |
| Priolo [48] | 2014 | Discovery set: Pca: 61; benign: 25 Validation set: Pca: 40; benign: 16 | UHPLC-MS/MS GC-MS | Diagnosis Tumour subtyping | MYC-driven Pca:Oleic acid, arachidonic acid, docosahexaenoic acidsAKT1-driven Pca:2-aminoadipic acid, creatine | Yes | Mann–Whitney test | metabolomic profiling of tumors driven by MYC and AKT1 oncogenes;extracts of frozen tissue from prostatectomy;metabolomic profiling in cell lines, mice and human tissue; validation of selected markers in human tissue samples |
| Keshari [58] | 2011 | n = 49 tissue samples high-grade Pca: 13 low-grade Pca: 22 benign: 14 | 1-D and 2-D HR-MAS Spectroscopy | Staging | Benign vs. Pca tissue: choline, phosphocholine, glycerophosphocholine, phosphoethanolamine, glycerophosphoethanolamine, citrate, polyamineslow-grade vs. high-grade Pca:phosphocholine, glycerophosphocholine | No | Student’s t-test | intact tissue from core biopsies after prostatectomy;targeted analysis of phospholipid metabolites |
| Shuster [45] | 2011 | Pca: 14 benign: 14 | GC-MSLC-MS/MS | Diagnosis | cysteine, dihomo-linoleate, docosapentaenoate, N-acetylaspartate, N-acetylglucosamine, uracil, xanthine, and 1-stearoylglycerophophoinositol;uracil, kynurenine, glycerol-3-phosphate, leucine, proline; choline, lactate, alanine citrate, putrescine, spermidine, spermine | No | matched paired t-test | tissue extracts from needle biopsies after prostatectomy;description of the mPREF methodreplicated metabolites previously published by Sreekumar et al. 2009 and various in vivo studies |
| Zhang [59] | 2014 | untreated patients: benign: 58; indolent Pca: 5; aggressive Pca: 8 radiation-treated patients: benign: 32; indolent Pca: 7 (relapse); aggressive Pca: 12 (relapse) | 1-D and 2-D HR-MAS Spectroscopy | Staging Diagnosis | aggressive vs. indolent Pca:choline, phosphocholine, glycerophosphocholine, [choline + phosphocholine + glycerophosphocholine] to creatine ratio, (lactate; only in untreated)benign vs. Pca (untreated):citrate, polyamines, lactate, glutamate, alanine | No | linear mixed-effects model Wilcoxon Rank Sum Test Kruskal–Wallis Test | intact tissue from core biopsies |
| Giskeødegård [52] | 2013 | n = 158 tissue samples from 48 subjects benign: 47v low-grade Pca: 30 high-grade Pca: 81 | HR-MAS 1H MRS | Staging | Pca vs. benign: citrate, taurine, creatine, glycerohpophocholine, phosphocholine, choline, glycinelow-grade vs. high-grade Pca:spermine, citrate, CCP/C ratio | No | Linear mixed models PLS-DA models | intact tissue from biopsies |
| Jentzmik [56] | 2011 | n = 92 Matched PCa and adjacent benign tissue | GC-MS | Staging | sarcosine | No | Mann–Whitney U test Wilcoxon test Spearman rank correlation Kaplan–Meier curve Cox proportional hazards regression analysis log rank test ROC analysis | target analysis of sarcosine as biomarker for disease progression tissue extracts from punch biopsies of tissue sections collected after prostatectomy |
| Thysell [29] | 2010 | Discovery set: bone metastases: 14 (hormone-naive Pca: 7; CRPC 7) adjacent normal bone: 10 Validation set:bone metastases: 13(6 Pca, 7 other cancers) normal bone: 11 Primary tumour: with metastases: 7 w/o metastases: 6benign: 17 | GC-TOFMS | Staging | Bone tissue: metastases vs. normal: Cholesterol, myo-inositol-1-phosphate, citrate, fumarate, glycerol-3-phosphate, amino aicds Primary tumour: metastatic PCa vs. benign tissue and non-metastatic PCa:malate, dehydroascrobic acid, urea, hypoxanthine, asparagine, threonine, fumarate, linoleic acid | Yes | OPLS-DA Mann–Whitney U-test | tissue extracts from fresh-frozen biopsies of bone metastases and from biopsies of primary Pca and benign prostateall patients were selected to have hihg-risk tumours (i.e., presence of bone metastases, locally advanced tumour or poorly differnitated cancer)validation set available only for bone metastatic tissueblood plasma samples from men who underwent prostate biopsies were also analyzed |
| Author | Year | Sample Size | Technique | Use Case | Biomarkers Candidates | Validated? | Statistical Details | Comments |
|---|---|---|---|---|---|---|---|---|
| Clos-Garcia [61] | 2018 | PCa: 31 BPH: 14 | UHPLC-MS | Diagnosis Staging Prognosis | phosphathidylcholines, acyl carnitines, citrate and kynurenine steroid hormone, 3beta-hydroxyandros-5-en-17- one-3-sulphate (dehydroepiandrosterone sulphate) | NO | ROC analysis PCA PLS-DA OPLS | Sample type: Urine extracellular vesicles (EVs) |
| Roberts [63] | 2017 | n = 151 PCa: 9880 (initially diagnosed) 18 (diagnosed during follow-up period) | NMR | Risk prediction | lipids/lipoproteins (PC1)choline phosphocholinecitrate Fructose and spermine | NO | PCA PLS | Sample type: Seminal plasma Time sample collection between January 2007 and February 2013 Samples obtained prior to or at least 1 month after prostate biopsy, prior to commencement of any treatment No other specifications were provided to patients for sample collection process Glucose signals were excluded from the spectra |
| Puhka | 2017 | PCA: 3 HC: 3 | UPLC-MS | Diagnosis | glucuronate, D-ribose 5-phosphate and isobutyryl-L-carnitine | NO | Sample type: urinary and platelet EVs Urine samples and matched plasma samples were collected 0–3 days before and 5–6 weeks after the prostatectomy Control samples were from healthy <35 year-old mennormalization to EV-derived factors or with metabolite ratios, and not from the original urine samples. | |
| Weis | 2016 | PCa: 46 HC: 4 | 3D Proton MRSI | Diagnosis | choline, spermine and citrate ratios | NO | concentration referenced to water | |
| Nagarajan [66] | 2015 | PCa: 22 | EPSI 2D-JPRESS (4D) EP-JRESI | Diagnosis | choline, spermine, citrate, myo-inositol and glutamate plus glutamine (Glx) | NO | logistic regression analysis ROC analysis | Prostate cancer was histopathologically confirmed after RP |
| Nagarajan [67] | 2012 | PCa: 41 GS 3+3 (n = 12) GS 3+4 (n = 20) GS 3+4 (n = 9) | MRSI DWI | Staging | choline, creatine and citrate ratios | NO | ROC analysis | At least 6 weeks time period between biopsy and MRI |
| Kobus [68] | 2011 | PCa: 43 | MR MRSI | Staging | choline, creatine and citrate ratios | NO | 3 T | |
| Wang [69] | 2008 | PCa: 33 BPH: 33 | 1H-MRSI | Diagnosis | choline, creatine and citrate ratios | NO | 1.5 T | |
| Kumar [70] | 2008 | n = 155 | MRSI | Diagnosis | choline, creatine and citrate ratios | NO | 1.5 T TRUS-guided prostate biopsy within 1 week after the MRSI |
| Risk Prediction | Diagnosis | Prognosis | Recurrence | |
|---|---|---|---|---|
| citrate | Mondul et al. 2015 [15] (B) | Kumar et al. 2016 [22]) (B) | Giskeødegård et al. 2013 [52] (T) | |
| Huang et al. 2017 [16] (B) | Weis et al. 2017 [65] (O) | Sandsmark et al. 2017 [54] (T) | ||
| Roberts et al. 2017 [63] (O) | Nagarajan et al. 2015 [67] (O) | Selnæs et al. 2013 [53] (T) | ||
| Wang et al. 2010 [69] (O) | Hansen et al. 2016 [55] (T) | |||
| Zhang et al. 2014 [59] (T) | Meller et al. 2016 [57] (T) | |||
| Keshari et al. 2011 [58] (T) | ||||
| Thysell et al. 2010 [29] (T) | ||||
| glycine | de Vogel et al. 2014 [18] | Kumar et al. 2015 [21] (B) | Hansen et al. 2016 [55] | |
| Kumar et al. 2016 [22] (B) | ||||
| Pérez-Rambla et al. 2017 [37] (U) | ||||
| Struck-Lewicka et al. 2015 [38] (U) | ||||
| Liu et al. 2015 [46] (T) | ||||
| Jung et al. 2013 [47] (T) | ||||
| Giskeødegård et al. 2013 [52] (T) | ||||
| glycerol-3-phosphate | Andras et al. 2017 [20] (B) | McDunn et al. 2013 [44] (T) | ||
| Sreekumar et al. 2009 [28] (T) | Thysell et al. 2010 [29] (T) | |||
| glycerophosphocholine | Andras et al. 2017 [20] (B) | Giskeødegård et al. 2013 [52] (T) | ||
| Keshari et al. 2011 [58] (T) | ||||
| Zhang et al. 2014 [59] (T) | ||||
| alanine | Kumar et al. 2015 [21] (B) | Kumar et al. 2015 [21] (B) | ||
| Kumar et al. 2016 [22] (B) | Zhang et al. 2014 [59] (T) | |||
| Struck-Lewicka et al. 2015 [38] (U) | ||||
| Shuster et al. 2011 [45] (T) | ||||
| Jung et al. 2013 [47] (T) | ||||
| Zhang et al. 2014 [59] (T) | ||||
| choline | Ren et al. 2016 [43] | Selnæs et al. 2013 [53] (T) | ||
| Shuster et al. 2011 [45] | Zhang et al. 2014 [59] (T) | |||
| Giskeødegård et al. 2013 [52] (T) | Nagarajan et al. 2015 [67] (O) | |||
| Keshari et al. 2011 [58] (T) | Kobuscet al. 2011 [68] (O) | |||
| Zhang et al. 2014 [59] (T) | ||||
| Weis et al. 2017 [65] (O) | ||||
| Wang et al. 2010 [69] (O) | ||||
| phosphocholine | Giskeødegård et al. 2013 [52] (T) | Zhang et al. 2014 [59] (T) | ||
| Keshari et al. 2011 [58] (T) | ||||
| Zhang et al. 2014 [59] (T) | ||||
| glycerophosphocholine | Andras et al. 2017 [20] (B) | Zhang et al. 2014 [59] (T) | ||
| Giskeødegård et al. 2013 [52] (T) | ||||
| Keshari et al. 2011 [58] (T) | ||||
| uracil | Huan et al. 2016 [42] (T) | |||
| Sreekumar et al. 2009 [28] (T) | ||||
| Shuster et al. 2011 [45] (T) | ||||
| Jung et al. 2013 [47] (T) | ||||
| McDunn et al. 2013 [44] (T) | ||||
| proline | Dereziński et al. 2017 [27] (B) | Sreekumar et al. 2009 [28] (T) | ||
| Jung et al. 2013 [47] (T) | Liu et al. 2015 [46] (T) | |||
| McDunn et al. 2013 [44] (T) | ||||
| histidine | Fernández-Peralbo et al. 2016 [35] (U) | |||
| Jung et al. 2013 [47] (T) | ||||
| spermine | Roberts et al. 2017 [63] (O) | Nagarajan et al. 2015 (O) | Liu et al. 2015 [46] (T) | |
| Weis et al. 2017 [65] (O) | Shuster et al. 2011 [45] (T) | |||
| Giskeødegård et al. 2013 [52] (T) | ||||
| Selnæs et al. 2013 [53] (T) | ||||
| Sandsmark et al. 2017 [54] (T) | ||||
| Hansen et al. 2016 [55] (T) | ||||
| Meller et al. 2016 [57] (T) | ||||
| Maxeiner et al. 2010 [ 60] (T) | ||||
| 2-aminoadipic acid | Jung et al. 2013 [47] (T) | |||
| Shuster et al. 2011 [45] (T) | ||||
| Sreekumar et al. 2009 [28] (T) | ||||
| Priolo et al. 2014 [48] (T) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kdadra, M.; Höckner, S.; Leung, H.; Kremer, W.; Schiffer, E. Metabolomics Biomarkers of Prostate Cancer: A Systematic Review. Diagnostics 2019, 9, 21. https://doi.org/10.3390/diagnostics9010021
Kdadra M, Höckner S, Leung H, Kremer W, Schiffer E. Metabolomics Biomarkers of Prostate Cancer: A Systematic Review. Diagnostics. 2019; 9(1):21. https://doi.org/10.3390/diagnostics9010021
Chicago/Turabian StyleKdadra, Marouane, Sebastian Höckner, Hing Leung, Werner Kremer, and Eric Schiffer. 2019. "Metabolomics Biomarkers of Prostate Cancer: A Systematic Review" Diagnostics 9, no. 1: 21. https://doi.org/10.3390/diagnostics9010021
APA StyleKdadra, M., Höckner, S., Leung, H., Kremer, W., & Schiffer, E. (2019). Metabolomics Biomarkers of Prostate Cancer: A Systematic Review. Diagnostics, 9(1), 21. https://doi.org/10.3390/diagnostics9010021




