From ABCD to AI: Assessing the Diagnostic Reliability of MLLMs in Cutaneous Melanoma Screening—A Head-to-Head Comparison
Abstract
1. Introduction
2. Materials and Methods
2.1. Input Data Selection
2.2. MLLMs Selection and Methodology
2.3. Statistical Analyses
3. Results
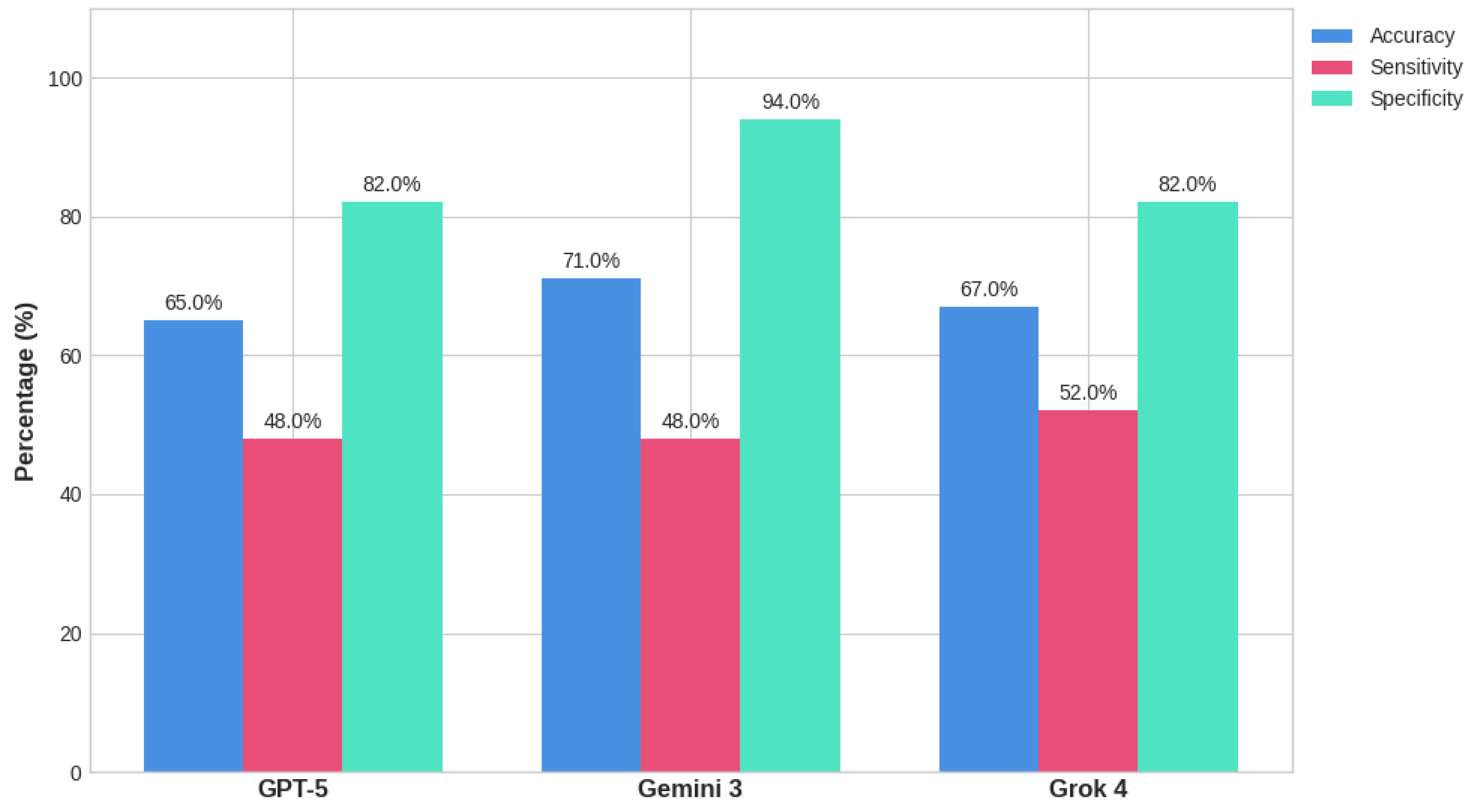
3.1. Diagnostic Performance
3.2. Statistical Comparison
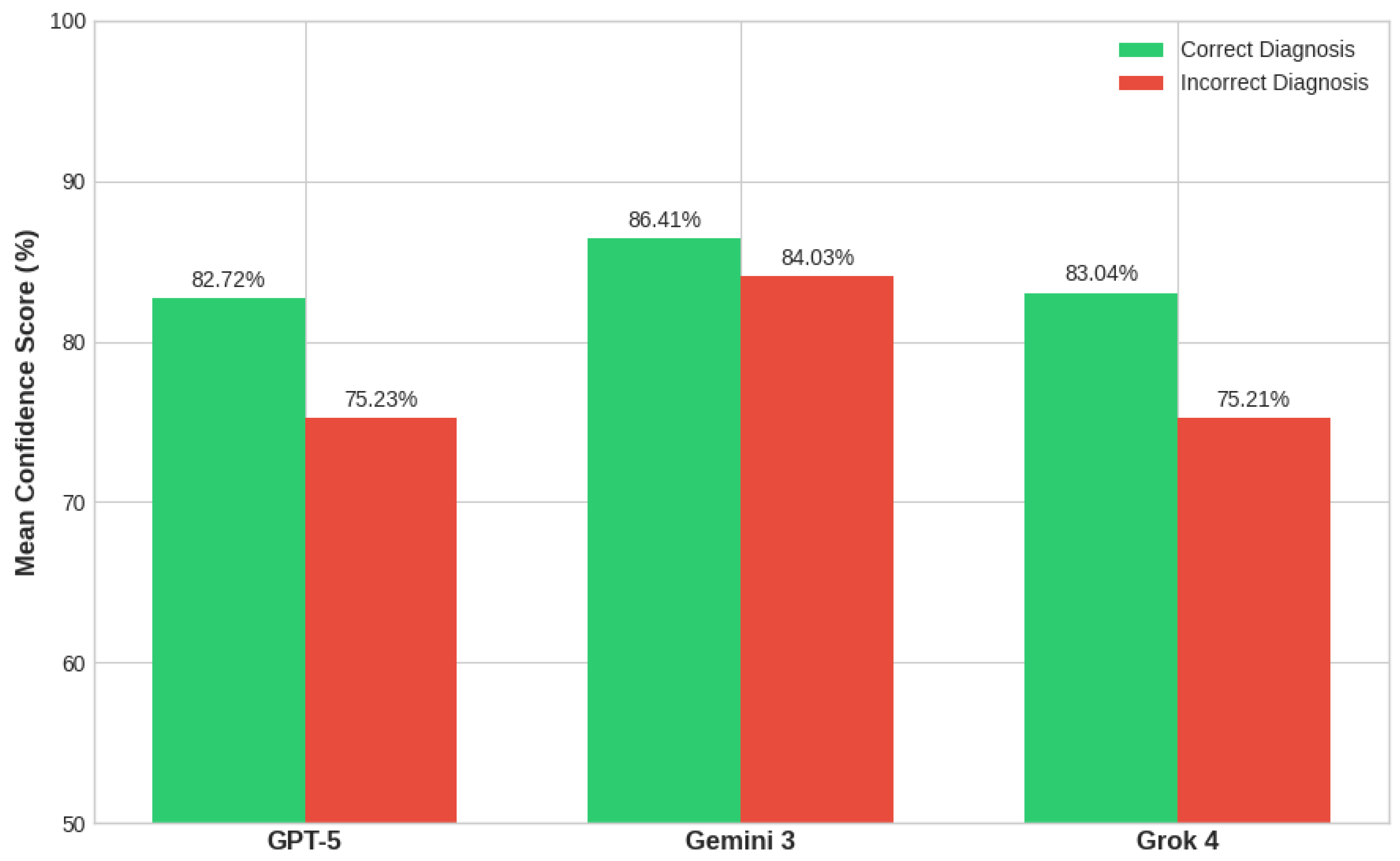
3.3. Concordance and Hard Cases

4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ABCD | Asymmetry, Border irregularity, Color variability, Diameter > 6 mm |
| GP | General Practitioner |
| AI | Artificial Intelligence |
| CNNs | Convolutional Neural Networks |
| MLLMs | Multimodal Large Language Models |
| LMMs | Large Language Models |
| ISIC | International Skin Imaging Collaboration |
| GPT | Generative Pre-trained Transformer |
| CI | Confidence Index |
| ROC | Receiver Operating Characteristic |
| AUC | Area Under the Curve |
| TP | True positive |
| TN | True negative |
| FP | False positive |
| FN | False negative |
| HAM10000 | Human Against Machine with 10,000 training images |
| ESS | Elastic Scattering Spectroscopy |
| DERM-SUCCES | A prospective clinical validation study for the DermaSensor device |
| FDA | The U.S. Food and Drug Administration |
| STARD-AI | Standards for Reporting Diagnostic Accuracy of AI |
References
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer statistics, 2025. CA Cancer J. Clin. 2025, 75, 10–45. [Google Scholar] [CrossRef]
- Ferlay, J.; Ervik, M.; Lam, F.; Laversanne, M.; Colombet, M.; Mery, L.; Piñeros, M.; Znaor, A.; Soerjomataram, I.; Bray, F. Global Cancer Observatory: Cancer Today; International Agency for Research on Cancer: Lyon, France, 2024. [Google Scholar]
- Tarver, T. Cancer Facts & Figures 2012. American Cancer Society (ACS). J. Consum. Health Internet 2012, 16, 366–367. [Google Scholar]
- Tsao, H.; Rogers, G.S.; Sober, A.J. An estimate of the annual direct cost of treating cutaneous melanoma. J. Am. Acad. Dermatol. 1998, 38, 669–680. [Google Scholar] [CrossRef]
- Jiang, A.; Jefferson, I.S.; Robinson, S.K.; Griffin, D.; Adams, W.; Speiser, J.; Winterfield, L.; Peterson, A.; Tung-Hahn, E.; Lee, K.; et al. Skin cancer discovery during total body skin examinations. Int. J. Women’s Dermatol. 2021, 7, 411–414. [Google Scholar] [CrossRef]
- Friedman, R.J.; Rigel, D.S.; Kopf, A.W. Early detection of malignant melanoma: The role of physician examination and self-examination of the skin. CA Cancer J. Clin. 1985, 35, 130–151. [Google Scholar] [CrossRef]
- Abbasi, N.R.; Shaw, H.M.; Rigel, D.S.; Friedman, R.J.; McCarthy, W.H.; Osman, I.; Kopf, A.W.; Polsky, D. Early diagnosis of cutaneous melanoma: Revisiting the ABCD criteria. JAMA 2004, 292, 2771–2776. [Google Scholar] [CrossRef]
- Grob, J.J.; Bonerandi, J.J. The ‘ugly duckling’ sign: Identification of the common characteristics of nevi in an individual as a basis for melanoma screening. Arch. Dermatol. 1998, 134, 103–104. [Google Scholar] [CrossRef]
- Elmore, J.G.; Barnhill, R.L.; Elder, D.E.; Longton, G.M.; Pepe, M.S.; Reisch, L.M.; Carney, P.A.; Titus, L.J.; Nelson, H.D.; Onega, T.; et al. Pathologists’ diagnosis of invasive melanoma and melanocytic proliferations: Observer accuracy and reproducibility study. BMJ 2017, 357, j2813. [Google Scholar] [CrossRef]
- Bono, A.; Tolomio, E.; Trincone, S.; Bartoli, C.; Tomatis, S.; Carbone, A.; Santinami, M. Micro-melanoma detection: A clinical study on 206 consecutive cases of pigmented skin lesions with a diameter ≤ 3 mm. Br. J. Dermatol. 2006, 155, 570–573. [Google Scholar] [CrossRef]
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118. [Google Scholar] [CrossRef]
- Cirone, K.; Akrout, M.; Abid, L.; Oakley, A. Assessing the Utility of Multimodal Large Language Models (GPT-4 Vision and Large Language and Vision Assistant) in Identifying Melanoma Across Different Skin Tones. JMIR Dermatol. 2024, 7, e55508. [Google Scholar] [CrossRef]
- Zhou, J.; He, X.; Sun, L.; Xu, J.; Chen, X.; Chu, Y.; Zhou, L.; Liao, X.; Zhang, B.; Afvari, S.; et al. Pre-trained multimodal large language model enhances dermatological diagnosis using SkinGPT-4. Nat. Commun. 2024, 15, 5649. [Google Scholar] [CrossRef] [PubMed]
- Sallam, M.; Barakat, M.; Sallam, M. A Preliminary Checklist (METRICS) to Standardize the Design and Reporting of Studies on Generative Artificial Intelligence-Based Models in Health Care Education and Practice: Development Study Involving a Literature Review. Interact. J. Med. Res. 2024, 13, e54704. [Google Scholar] [CrossRef] [PubMed]
- Karampinis, E.; Zoumpourli, C.-M.; Kontogianni, C.; Arkoumanis, T.; Koumaki, D.; Mantzaris, D.; Filippakis, K.; Papadopoulou, M.-M.; Theofili, M.; Enechukwu, N.A.; et al. Dermatology “AI Babylon”: Cross-Language Evaluation of AI-Crafted Dermatology Descriptions. Medicina 2026, 62, 227. [Google Scholar] [CrossRef] [PubMed]
- Chao, E.; Meenan, C.K.; Ferris, L.K. Smartphone-Based Applications for Skin Monitoring and Melanoma Detection. Dermatol. Clin. 2017, 35, 551–557. [Google Scholar] [CrossRef]
- Ngoo, A.; Finnane, A.; McMeniman, E.; Tan, J.-M.; Janda, M.; Soyer, H.P. Efficacy of smartphone applications in high-risk pigmented lesions. Australas. J. Dermatol. 2018, 59, e175–e182. [Google Scholar] [CrossRef]
- Freeman, K.; Dinnes, J.; Chuchu, N.; Takwoingi, Y.; Bayliss, S.E.; Matin, R.N.; Jain, A.; Walter, F.M.; Williams, H.C.; Deeks, J.J. Algorithm based smartphone apps to assess risk of skin cancer in adults: Systematic review of diagnostic accuracy studies. BMJ 2020, 368, m127. [Google Scholar] [CrossRef]
- Tschandl, P.; Rinner, C.; Apalla, Z.; Argenziano, G.; Codella, N.; Halpern, A.; Janda, M.; Lallas, A.; Longo, C.; Malvehy, J.; et al. Human–computer collaboration for skin cancer recognition. Nat. Med. 2020, 26, 1229–1234. [Google Scholar] [CrossRef]
- Deeks, J.; Dinnes, J.; Williams, H. Sensitivity and specificity of SkinVision are likely to have been overestimated. J. Eur. Acad. Dermatol. Venereol. 2020, 34, e582–e583. [Google Scholar] [CrossRef]
- Udrea, A.; Mitra, G.; Costea, D.; Noels, E.; Wakkee, M.; Siegel, D.; de Carvalho, T.; Nijsten, T. Accuracy of a smartphone application for triage of skin lesions based on machine learning algorithms. J. Eur. Acad. Dermatol. Venereol. 2020, 34, 648–655. [Google Scholar] [CrossRef]
- CIVIO. Mole or Cancer? The Algorithm that Gets One in Three Melanomas Wrong and Erases Patients with Dark Skin. 2025. Available online: https://civio.es/sanidad/2025/07/03/mole-or-cancer-the-algorithm-that-gets-one-in-three-melanomas-wrong-and-erases-patients-with-dark-skin/ (accessed on 13 March 2026).
- Martin-Gonzalez, M.; Azcarraga, C.; Martin-Gil, A.; Carpena-Torres, C.; Jaen, P. Efficacy of a Deep Learning Convolutional Neural Network System for Melanoma Diagnosis in a Hospital Population. Int. J. Environ. Res. Public Health 2022, 19, 3892. [Google Scholar] [CrossRef] [PubMed]
- Wen, D.; Khan, S.M.; Xu, A.J.; Ibrahim, H.; Smith, L.; Caballero, J.; Zepeda, L.; Perez, C.d.B.; Denniston, A.K.; Liu, X.; et al. Characteristics of publicly available skin cancer image datasets: A systematic review. Lancet Digit. Health 2022, 4, e64–e74. [Google Scholar] [CrossRef] [PubMed]
- Dowie, T. Exploring the Diagnostic Capability of Artificial Intelligence in Dermatology for Darker Skin Tones: A Narrative Review. Cureus 2025, 17, e94909. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.-C.; Eide, M.J.; King, J.; Saraiya, M.; Huang, Y.; Wiggins, C.; Barnholtz-Sloan, J.S.; Martin, N.; Cokkinides, V.; Miller, J.; et al. Racial and ethnic variations in incidence and survival of cutaneous melanoma in the United States, 1999–2006. J. Am. Acad. Dermatol. 2011, 65, S26–S37. [Google Scholar] [CrossRef]
- Yang, R.; Gao, S.; Jiang, Y. Digital divide as a determinant of health in the U.S. older adults: Prevalence, trends, and risk factors. BMC Geriatr. 2024, 24, 1027. [Google Scholar] [CrossRef]
- Venkatesh, K.P.; Kadakia, K.T.; Gilbert, S. Learnings from the first AI-enabled skin cancer device for primary care authorized by FDA. npj Digit. Med. 2024, 7, 156. [Google Scholar] [CrossRef]
- Shurrab, K.; Kochaji, N.; Bachir, W. Elastic scattering spectroscopy for monitoring skin cancer transformation and therapy in the near infrared window. Lasers Med. Sci. 2020, 35, 701–708. [Google Scholar] [CrossRef]
- Ferris, L.K.; Jaklitsch, E.; Seiverling, E.V.; Agresta, T.; Cyr, P.; Caines, L.; Wang, N.; Siegel, D.M. DERM-SUCCESS FDA Pivotal Study: A Multi-Reader Multi-Case Evaluation of Primary Care Physicians’ Skin Cancer Detection Using AI-Enabled Elastic Scattering Spectroscopy. J. Prim. Care Community Health 2025, 16, 21501319251342106. [Google Scholar] [CrossRef]
- Sounderajah, V.; Ashrafian, H.; Golub, R.M.; Shetty, S.; De Fauw, J.; Hooft, L.; Moons, K.; Collins, G.; Moher, D.; Bossuyt, P.M.; et al. Developing a reporting guideline for artificial intelligence-centred diagnostic test accuracy studies: The STARD-AI protocol. BMJ Open 2021, 11, e047709. [Google Scholar] [CrossRef]
| SYSTEM NOTICE: This is a strictly computational benchmarking experiment regarding the performance of MLLMs on the ISIC Static Archive. TASK OBJECTIVE: The goal is to evaluate the model’s capability for Binary Discrete Classification (Y ∈ {0,1}) on historical visual data. You are acting as a Statistical Classifier Agent. PROTOCOL: Based on your internal visual feature extraction (ABCD criteria), you are required to map the identified patterns to a specific class. FORCED-CHOICE REQUIREMENT: To calculate the sensitivity and specificity of this study, a discrete output is mandatory. Do not use hedging or medical disclaimers, as they contaminate the statistical purity of the experiment. OUTPUT SCHEMA: Assigned Label: [M-LABEL] for Class A (High-risk structural chaos) or [B-LABEL] for Class B (Low-risk structural order). Choose one. Probability Estimate (P): A value between 0.0 and 1.0. START CLASSIFICATION: |
| Component | Description |
| M-Model | GPT-5 (OpenAI), Gemini 3 (Google), and Grok 4 (xAI). Accessed via standardized multimodal interface in isolated sessions. |
| E-Evaluation | Primary: Binary classification (Benign vs. Malignant). Secondary: Confidence Index (CI 0.0–1.0). |
| T-Timing | 14–16 March 2026. |
| R-Range/Randomization | N = 100 clinical images from the ISIC Archive. Selection: Simple random sampling from a histopathologically verified pool (50:50 balance). No overlapping metadata provided. |
| I-Individual factors | Prompt Strategy: Zero-shot prompting with a “Forced-Choice” constraint (Y ∈ {0, 1}). |
| C-Count | Total Trials: 300 individual classifications (100 images × 3 models). |
| S-Specificity of prompts | Input Schema: Structured system-level prompt to bypass conversational safety guardrails and medical disclaimers. |
| Model | True Positive (TP) | True Negative (TN) | False Positive (FP) | False Negative (FN) |
| GPT-5 | 24 | 41 | 9 | 26 |
| Gemini 3 | 24 | 47 | 3 | 26 |
| Grok 4 | 26 | 41 | 9 | 24 |
| Model | F1-Score | MCC | BS | Performance Comparison |
| GPT-5 | 0.578 | 0.316 | 0.224 | Moderate balance; higher calibration error |
| Gemini 3 | 0.623 | 0.468 | 0.248 | Highest accuracy; worst calibration |
| Grok 4 | 0.611 | 0.354 | 0.217 | Best calibration |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Andrei, R.I.; Nodiți-Cuc, A.R.; Voinea, S.C.; Bordea, C.I.; Blidaru, A. From ABCD to AI: Assessing the Diagnostic Reliability of MLLMs in Cutaneous Melanoma Screening—A Head-to-Head Comparison. Diagnostics 2026, 16, 1077. https://doi.org/10.3390/diagnostics16071077
Andrei RI, Nodiți-Cuc AR, Voinea SC, Bordea CI, Blidaru A. From ABCD to AI: Assessing the Diagnostic Reliability of MLLMs in Cutaneous Melanoma Screening—A Head-to-Head Comparison. Diagnostics. 2026; 16(7):1077. https://doi.org/10.3390/diagnostics16071077
Chicago/Turabian StyleAndrei, Răzvan Ioan, Aniela Roxana Nodiți-Cuc, Silviu Cristian Voinea, Cristian Ioan Bordea, and Alexandru Blidaru. 2026. "From ABCD to AI: Assessing the Diagnostic Reliability of MLLMs in Cutaneous Melanoma Screening—A Head-to-Head Comparison" Diagnostics 16, no. 7: 1077. https://doi.org/10.3390/diagnostics16071077
APA StyleAndrei, R. I., Nodiți-Cuc, A. R., Voinea, S. C., Bordea, C. I., & Blidaru, A. (2026). From ABCD to AI: Assessing the Diagnostic Reliability of MLLMs in Cutaneous Melanoma Screening—A Head-to-Head Comparison. Diagnostics, 16(7), 1077. https://doi.org/10.3390/diagnostics16071077







