Clinical Utility of Small Extracellular Vesicles as Liquid Biopsy for Oral Mucosal Disease Diagnostics: Emerging Perspectives
Abstract
1. Introduction
2. Literature Search Methods
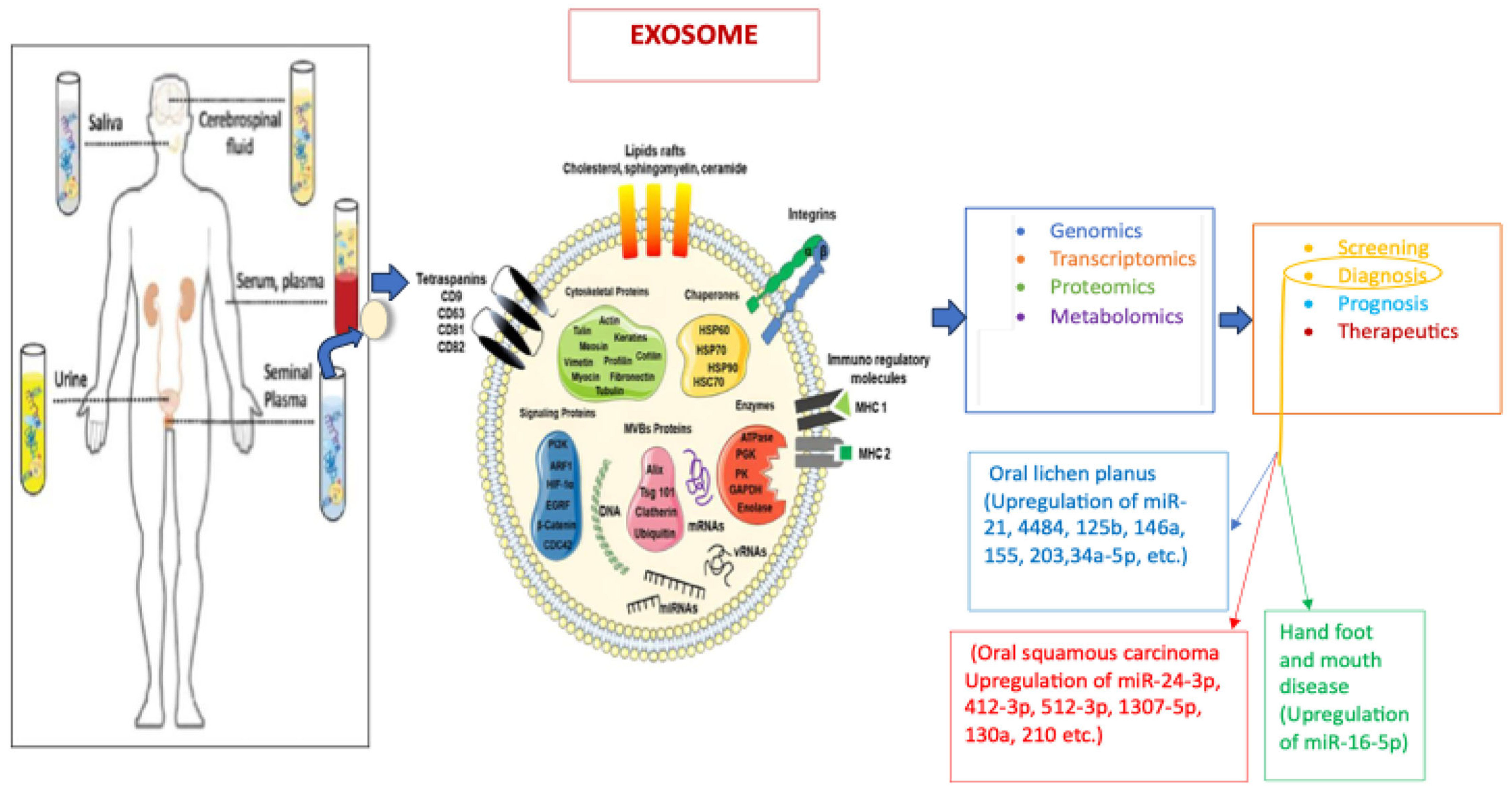
2.1. Biogenesis, Composition, and Biological Functions of sEVs
2.1.1. Biogenesis
2.1.2. sEVs Composition
2.1.3. Biological Functions of sEVs
2.2. Isolation and Characterization of sEVs
2.2.1. Isolation of sEVs
2.2.2. Characterization of sEVs
2.3. sEVs as Biomarkers for Diagnosis of Oral Mucosal Diseases
2.3.1. Hand, Foot, and Mouth Disease (HFMD)
2.3.2. Oral Lichen Planus (OLP)
2.3.3. Oral Leukoplakia (OL)
2.3.4. Oral Squamous Cell Carcinoma
3. Limitations and Future Directions
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hurjui, L.L.; Gradinaru, I.; Dorus, C.; Tanase, D.M.; Armencia, A.; Hurjui, I.; Tarniceriu, C.C.; Mitrea, M.; Lozneanu, L.; Balcos, C. Oral mucosa-pathophysiological and pharmacotherapeutic aspects. Rom. J. Oral Rehabil. 2021, 13, 108–114. [Google Scholar]
- Brizuela, M.; Winters, R. Histology, Oral Mucosa; StatPearls Publishing: Treasure Island, FL, USA, 2021. [Google Scholar]
- Akintoye, S.O.; Mupparapu, M. Clinical evaluation and anatomic variation of the oral cavity. Dermatol. Clin. 2020, 38, 399–411. [Google Scholar] [CrossRef]
- Bhalla, N.; Nudell, Y.; Thakkar, J.; Dym, H. Oral manifestation of systemic diseases. Dent. Clin. 2020, 64, 191–207. [Google Scholar] [CrossRef]
- Davidson, J. Oral mucosal lesions: Classification, diagnosis, and management. J. Oral Med. Surg. 2023, 6, 158. [Google Scholar]
- Yu, M.Y.; Liu, X.C.; Yu, Z.L.; Jia, J. The Application of Salivary Exosomes in the Diagnosis of Oral Disease. Chin. J. Dent. Res. 2024, 27, 291–301. [Google Scholar] [CrossRef]
- Lajolo, C.; Gioco, G.; Rupe, C.; Patini, R.; Rizzo, I.; Romeo, U.; Contaldo, M.; Cordaro, M. Patient perception after oral biopsies: An observational outpatient study. Clin. Oral Investig. 2021, 25, 5687–5697. [Google Scholar] [CrossRef]
- Miao, Q.; Li, S.; Lyu, W.; Zhang, J.; Han, Y. Exosomes in Oral Diseases: Mechanisms and Therapeutic Applications. Drug Des. Dev. Ther. 2025, 19, 457–469. [Google Scholar] [CrossRef] [PubMed]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef] [PubMed]
- Kourembanas, S. Exosomes: Vehicles of intercellular signaling, biomarkers, and vectors of cell therapy. Annu. Rev. Physiol. 2015, 77, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Edgar, J.R. Q&A: What are exosomes, exactly? BMC Biol. 2016, 14, 46. [Google Scholar] [CrossRef]
- Bozyk, N.; Tang, K.D.; Zhang, X.; Batstone, M.; Kenny, L.; Vasani, S.; Punyadeera, C. Salivary exosomes as biomarkers for early diagnosis of oral squamous cell carcinoma. Oral Oncol. Rep. 2023, 6, 100017. [Google Scholar] [CrossRef]
- Lu, Y.; Zheng, Z.; Yuan, Y.; Pathak, J.L.; Yang, X.; Wang, L.; Ye, Z.; Cho, W.C.; Zeng, M.; Wu, L. The emerging role of exosomes in oral squamous cell carcinoma. Front. Cell Dev. Biol. 2021, 9, 628103. [Google Scholar] [CrossRef]
- Han, Y.; Jia, L.; Zheng, Y.; Li, W. Salivary exosomes: Emerging roles in systemic disease. Int. J. Biol. Sci. 2018, 14, 633. [Google Scholar] [CrossRef] [PubMed]
- Liao, W.; Du, Y.; Zhang, C.; Pan, F.; Yao, Y.; Zhang, T.; Peng, Q. Exosomes: The next generation of endogenous nanomaterials for advanced drug delivery and therapy. Acta Biomater. 2019, 86, 1–14. [Google Scholar] [CrossRef]
- Sharma, S.; Gillespie, B.M.; Palanisamy, V.; Gimzewski, J.K. Quantitative nanostructural and single-molecule force spectroscopy biomolecular analysis of human-saliva-derived exosomes. Langmuir 2011, 27, 14394–14400. [Google Scholar] [CrossRef]
- Zlotogorski-Hurvitz, A.; Dayan, D.; Chaushu, G.; Salo, T.; Vered, M. Morphological and molecular features of oral fluid-derived exosomes: Oral cancer patients versus healthy individuals. J. Cancer Res. Clin. Oncol. 2016, 142, 101–110. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.-L.; He, C.-H.; Wang, Z.-Y.; Xu, Y.-F.; Yin, G.-Q.; Mao, L.-J.; Liu, C.-W.; Deng, L. MicroRNA expression profile in exosome discriminates extremely severe infections from mild infections for hand, foot and mouth disease. BMC Infect. Dis. 2014, 14, 506. [Google Scholar] [CrossRef] [PubMed]
- Byun, J.S.; Hong, S.H.; Choi, J.K.; Jung, J.K.; Lee, H.J. Diagnostic profiling of salivary exosomal micro RNA s in oral lichen planus patients. Oral Dis. 2015, 21, 987–993. [Google Scholar] [CrossRef]
- Mehdipour, M.; Shahidi, M.; Manifar, S.; Jafari, S.; Mashhadi Abbas, F.; Barati, M.; Mortazavi, H.; Shirkhoda, M.; Farzanegan, A.; Elmi Rankohi, Z. Diagnostic and prognostic relevance of salivary microRNA-21,-125a,-31 and-200a levels in patients with oral lichen planus-a short report. Cell. Oncol. 2018, 41, 329–334. [Google Scholar] [CrossRef]
- Stasio, D.D.; Mosca, L.; Lucchese, A.; Cave, D.D.; Kawasaki, H.; Lombardi, A.; Porcelli, M.; Caraglia, M. Salivary mir-27b expression in oral lichen planus patients: A series of cases and a narrative review of literature. Curr. Top. Med. Chem. 2019, 19, 2816–2823. [Google Scholar] [CrossRef]
- Mehdipour, M.; Shahidi, M.; Anbari, F.; Mirzaei, H.; Jafari, S.; Kholghi, A.; Lotfi, E.; Manifar, S.; Mashhadiabbas, F. Salivary level of microRNA-146a and microRNA-155 biomarkers in patients with oral lichen planus versus oral squamous cell carcinoma. BMC Oral Health 2023, 23, 433. [Google Scholar] [CrossRef] [PubMed]
- Peng, Q.; Zhang, J.; Zhou, G. Differentially circulating exosomal microRNAs expression profiling in oral lichen planus. Am. J. Transl. Res. 2018, 10, 2848. [Google Scholar]
- Čēma, I.; Dzudzilo, M.; Kleina, R.; Franckevica, I.; Svirskis, Š. Correlation of soluble CD44 expression in saliva and CD44 protein in oral leukoplakia tissues. Cancers 2021, 13, 5739. [Google Scholar] [CrossRef]
- Gai, C.; Camussi, F.; Broccoletti, R.; Gambino, A.; Cabras, M.; Molinaro, L.; Carossa, S.; Camussi, G.; Arduino, P.G. Salivary extracellular vesicle-associated miRNAs as potential biomarkers in oral squamous cell carcinoma. BMC Cancer 2018, 18, 1–11. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Ping, F.; Fan, Z.; Zhang, C.; Deng, M.; Cheng, B.; Xia, J. Salivary exosomal miR-24-3p serves as a potential detective biomarker for oral squamous cell carcinoma screening. Biomed. Pharmacother. 2020, 121, 109553. [Google Scholar] [CrossRef]
- Faur, C.I.; Roman, R.C.; Jurj, A.; Raduly, L.; Almășan, O.; Rotaru, H.; Chirilă, M.; Moldovan, M.A.; Hedeșiu, M.; Dinu, C. Salivary exosomal MicroRNA-486-5p and MicroRNA-10b-5p in oral and oropharyngeal squamous cell carcinoma. Medicina 2022, 58, 1478. [Google Scholar] [CrossRef]
- Patel, A.; Patel, S.; Patel, P.; Mandlik, D.; Patel, K.; Tanavde, V. Salivary exosomal miRNA-1307-5p predicts disease aggressiveness and poor prognosis in oral squamous cell carcinoma patients. Int. J. Mol. Sci. 2022, 23, 10639. [Google Scholar] [CrossRef]
- Bigagli, E.; Locatello, L.G.; Di Stadio, A.; Maggiore, G.; Valdarnini, F.; Bambi, F.; Gallo, O.; Luceri, C. Extracellular vesicles miR-210 as a potential biomarker for diagnosis and survival prediction of oral squamous cell carcinoma patients. J. Oral Pathol. Med. 2022, 51, 350–357. [Google Scholar] [CrossRef]
- He, T.; Guo, X.; Li, X.; Liao, C.; Wang, X.; He, K. Plasma-derived exosomal microRNA-130a serves as a noninvasive biomarker for diagnosis and prognosis of oral squamous cell carcinoma. J. Oncol. 2021, 2021, 5547911. [Google Scholar] [CrossRef]
- Yang, M.; Ding, J.; Luo, Q.; Chen, X.; Chen, F. Improving the diagnostic efficacy of squamous cell carcinoma antigen for oral squamous cell carcinoma via saponin disruption of serum extracellular vesicles. Clin. Chim. Acta 2022, 525, 40–45. [Google Scholar] [CrossRef] [PubMed]
- Hessvik, N.P.; Llorente, A. Current knowledge on exosome biogenesis and release. Cell. Mol. Life Sci. 2018, 75, 193–208. [Google Scholar] [CrossRef]
- Wolf, P. The nature and significance of platelet products in human plasma. Br. J. Haematol. 1967, 13, 269–288. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of extracellular vesicles, their origin, composition, purpose, and methods for exosome isolation and analysis. Cells 2019, 8, 727. [Google Scholar] [CrossRef]
- Bebelman, M.P.; Smit, M.J.; Pegtel, D.M.; Baglio, S.R. Biogenesis and function of extracellular vesicles in cancer. Pharmacol. Ther. 2018, 188, 1–11. [Google Scholar] [CrossRef]
- Ren, J.; Jing, X.; Liu, Y.; Liu, J.; Ning, X.; Zong, M.; Zhang, R.; Cheng, H.; Cui, J.; Li, B. Exosome-based engineering strategies for the diagnosis and treatment of oral and maxillofacial diseases. J. Nanobiotechnol. 2023, 21, 501. [Google Scholar] [CrossRef] [PubMed]
- Stuffers, S.; Sem Wegner, C.; Stenmark, H.; Brech, A. Multivesicular endosome biogenesis in the absence of ESCRTs. Traffic 2009, 10, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Perez-Hernandez, D.; Gutiérrez-Vázquez, C.; Jorge, I.; López-Martín, S.; Ursa, A.; Sánchez-Madrid, F.; Vázquez, J.; Yáñez-Mó, M. The intracellular interactome of tetraspanin-enriched microdomains reveals their function as sorting machineries toward exosomes. J. Biol. Chem. 2013, 288, 11649–11661. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, Y.; Liu, H.; Tang, W.H. Exosomes: Biogenesis, biologic function and clinical potential. Cell Biosci. 2019, 9, 1–18. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, D.; Gao, Y.; Lin, C.; An, Q.; Feng, Y.; Liu, Y.; Liu, D.; Luo, H.; Wang, D. The biology and function of extracellular vesicles in cancer development. Front. Cell Dev. Biol. 2021, 9, 777441. [Google Scholar] [CrossRef]
- Jeppesen, D.K.; Fenix, A.M.; Franklin, J.L.; Higginbotham, J.N.; Zhang, Q.; Zimmerman, L.J.; Liebler, D.C.; Ping, J.; Liu, Q.; Evans, R. Reassessment of exosome composition. Cell 2019, 177, 428–445. e418. [Google Scholar] [CrossRef]
- Li, J.; Zhang, Y.; Dong, P.-Y.; Yang, G.-M.; Gurunathan, S. A comprehensive review on the composition, biogenesis, purification, and multifunctional role of exosome as delivery vehicles for cancer therapy. Biomed. Pharmacother. 2023, 165, 115087. [Google Scholar] [CrossRef]
- Minciacchi, V.R.; Freeman, M.R.; Di Vizio, D. Extracellular vesicles in cancer: Exosomes, microvesicles and the emerging role of large oncosomes. In Seminars in Cell & Developmental Biology; Academic Press: Cambridge, MA, USA, 2015; pp. 41–51. [Google Scholar]
- Skotland, T.; Sandvig, K.; Llorente, A. Lipids in exosomes: Current knowledge and the way forward. Prog. Lipid Res. 2017, 66, 30–41. [Google Scholar] [CrossRef]
- Li, X.-X.; Yang, L.-X.; Wang, C.; Li, H.; Shi, D.-S.; Wang, J. The roles of exosomal proteins: Classification, function, and applications. Int. J. Mol. Sci. 2023, 24, 3061. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Bi, J.; Huang, J.; Tang, Y.; Du, S.; Li, P. Exosome: A review of its classification, isolation techniques, storage, diagnostic and targeted therapy applications. Int. J. Nanomed. 2020, 15, 6917–6934. [Google Scholar] [CrossRef] [PubMed]
- Dai, J.; Su, Y.; Zhong, S.; Cong, L.; Liu, B.; Yang, J.; Tao, Y.; He, Z.; Chen, C.; Jiang, Y. Exosomes: Key players in cancer and potential therapeutic strategy. Signal Transduct. Target. Ther. 2020, 5, 145. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Huang, J.; Chen, W.; Li, G.; Li, Z.; Lei, J. The updated role of exosomal proteins in the diagnosis, prognosis, and treatment of cancer. Exp. Mol. Med. 2022, 54, 1390–1400. [Google Scholar] [CrossRef]
- Ye, M.; Wang, J.; Pan, S.; Zheng, L.; Wang, Z.-W.; Zhu, X. Nucleic acids and proteins carried by exosomes of different origins as potential biomarkers for gynecologic cancers. Mol. Ther.-Oncolytics 2022, 24, 101–113. [Google Scholar] [CrossRef]
- van den Boorn, J.G.; Daßler, J.; Coch, C.; Schlee, M.; Hartmann, G. Exosomes as nucleic acid nanocarriers. Adv. Drug Deliv. Rev. 2013, 65, 331–335. [Google Scholar] [CrossRef]
- Huang, X.; Yuan, T.; Tschannen, M.; Sun, Z.; Jacob, H.; Du, M.; Liang, M.; Dittmar, R.L.; Liu, Y.; Liang, M. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genom. 2013, 14, 1–14. [Google Scholar] [CrossRef]
- Gusachenko, O.; Zenkova, M.; Vlassov, V. Nucleic acids in exosomes: Disease markers and intercellular communication molecules. Biochemistry 2013, 78, 1–7. [Google Scholar] [CrossRef]
- Waldenström, A.; Ronquist, G. Role of exosomes in myocardial remodeling. Circ. Res. 2014, 114, 315–324. [Google Scholar] [CrossRef]
- Bobrie, A.; Colombo, M.; Raposo, G.; Théry, C. Exosome secretion: Molecular mechanisms and roles in immune responses. Traffic 2011, 12, 1659–1668. [Google Scholar] [CrossRef]
- Zhang, H.-G.; Grizzle, W.E. Exosomes and cancer: A newly described pathway of immune suppression. Clin. Cancer Res. 2011, 17, 959–964. [Google Scholar] [CrossRef] [PubMed]
- Rashed, M.H.; Bayraktar, E.; Helal, G.K.; Abd-Ellah, M.F.; Amero, P.; Chavez-Reyes, A.; Rodriguez-Aguayo, C. Exosomes: From garbage bins to promising therapeutic targets. Int. J. Mol. Sci. 2017, 18, 538. [Google Scholar] [CrossRef]
- Kaur, S.; Chang, T.; Singh, S.P.; Lim, L.; Mannan, P.; Garfield, S.H.; Pendrak, M.L.; Soto-Pantoja, D.R.; Rosenberg, A.Z.; Jin, S. CD47 signaling regulates the immunosuppressive activity of VEGF in T cells. J. Immunol. 2014, 193, 3914–3924. [Google Scholar] [CrossRef]
- Rak, J. Microparticles in cancer. In Seminars in Thrombosis and Hemostasis; Thieme Medical Publishers: New York, NY, USA, 2010; pp. 888–906. [Google Scholar]
- Kujala, P.; Raymond, C.R.; Romeijn, M.; Godsave, S.F.; van Kasteren, S.I.; Wille, H.; Prusiner, S.B.; Mabbott, N.A.; Peters, P.J. Prion uptake in the gut: Identification of the first uptake and replication sites. PLoS Pathog. 2011, 7, e1002449. [Google Scholar] [CrossRef]
- Properzi, F.; Logozzi, M.; Fais, S. Exosomes: The future of biomarkers in medicine. Biomark. Med. 2013, 7, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Mukerjee, N.; Bhattacharya, A.; Maitra, S.; Kaur, M.; Ganesan, S.; Mishra, S.; Ashraf, A.; Rizwan, M.; Kesari, K.K.; Tabish, T.A.; et al. Exosome isolation and characterization for advanced diagnostic and therapeutic applications. Mater. Today Bio 2025, 31, 101613. [Google Scholar] [CrossRef]
- Peterson, M.F.; Otoc, N.; Sethi, J.K.; Gupta, A.; Antes, T.J. Integrated systems for exosome investigation. Methods 2015, 87, 31–45. [Google Scholar] [CrossRef] [PubMed]
- Lai, J.J.; Chau, Z.L.; Chen, S.Y.; Hill, J.J.; Korpany, K.V.; Liang, N.W.; Lin, L.H.; Lin, Y.H.; Liu, J.K.; Liu, Y.C. Exosome processing and characterization approaches for research and technology development. Adv. Sci. 2022, 9, 2103222. [Google Scholar] [CrossRef]
- Ding, M.; Wang, C.; Lu, X.; Zhang, C.; Zhou, Z.; Chen, X.; Zhang, C.-Y.; Zen, K.; Zhang, C. Comparison of commercial exosome isolation kits for circulating exosomal microRNA profiling. Anal. Bioanal. Chem. 2018, 410, 3805–3814. [Google Scholar] [CrossRef]
- Macías, M.; Rebmann, V.; Mateos, B.; Varo, N.; Perez-Gracia, J.L.; Alegre, E.; González, Á. Comparison of six commercial serum exosome isolation methods suitable for clinical laboratories. Effect in cytokine analysis. Clin. Chem. Lab. Med. (CCLM) 2019, 57, 1539–1545. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, A.-K.; De Miroschedji, K.; Doeppner, T.R.; Börger, V.; Ruesing, J.; Rebmann, V.; Durst, S.; Jansen, S.; Bremer, M.; Behrmann, E. Precipitation with polyethylene glycol followed by washing and pelleting by ultracentrifugation enriches extracellular vesicles from tissue culture supernatants in small and large scales. J. Extracell. Vesicles 2018, 7, 1528109. [Google Scholar] [CrossRef] [PubMed]
- Lobb, R.J.; Becker, M.; Wen Wen, S.; Wong, C.S.; Wiegmans, A.P.; Leimgruber, A.; Möller, A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J. Extracell. Vesicles 2015, 4, 27031. [Google Scholar] [CrossRef]
- Patel, G.K.; Khan, M.A.; Zubair, H.; Srivastava, S.K.; Khushman, M.d.; Singh, S.; Singh, A.P. Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci. Rep. 2019, 9, 5335. [Google Scholar] [CrossRef]
- Miron, R.J.; Zhang, Y. Understanding exosomes: Part 1—Characterization, quantification and isolation techniques. Periodontology 2000 2024, 94, 231–256. [Google Scholar] [CrossRef]
- Kaddour, H.; Lyu, Y.; Welch, J.L.; Paromov, V.; Mandape, S.N.; Sakhare, S.S.; Pandhare, J.; Stapleton, J.T.; Pratap, S.; Dash, C. Proteomics profiling of autologous blood and semen exosomes from HIV-infected and uninfected individuals reveals compositional and functional variabilities. Mol. Cell. Proteom. 2020, 19, 78–100. [Google Scholar] [CrossRef]
- Cao, Z.; Wu, Y.; Liu, G.; Jiang, Y.; Wang, X.; Wang, Z.; Feng, T. α-Synuclein in salivary extracellular vesicles as a potential biomarker of Parkinson’s disease. Neurosci. Lett. 2019, 696, 114–120. [Google Scholar] [CrossRef]
- Shi, M.; Kovac, A.; Korff, A.; Cook, T.J.; Ginghina, C.; Bullock, K.M.; Yang, L.; Stewart, T.; Zheng, D.; Aro, P. CNS tau efflux via exosomes is likely increased in Parkinson’s disease but not in Alzheimer’s disease. Alzheimer’s Dement. 2016, 12, 1125–1131. [Google Scholar] [CrossRef]
- Welch, J.L.; Stapleton, J.T.; Okeoma, C.M. Vehicles of intercellular communication: Exosomes and HIV-1. J. Gen. Virol. 2019, 100, 350–366. [Google Scholar] [CrossRef] [PubMed]
- Aswathyraj, S.; Arunkumar, G.; Alidjinou, E.; Hober, D. Hand, foot and mouth disease (HFMD): Emerging epidemiology and the need for a vaccine strategy. Med. Microbiol. Immunol. 2016, 205, 397–407. [Google Scholar] [CrossRef]
- Zhu, P.; Ji, W.; Li, D.; Li, Z.; Chen, Y.; Dai, B.; Han, S.; Chen, S.; Jin, Y.; Duan, G. Current status of hand-foot-and-mouth disease. J. Biomed. Sci. 2023, 30, 15. [Google Scholar] [CrossRef]
- Robinson, C.; Doane, F.W.; Rhodes, A.J. Report of an outbreak of febrile illness with pharyngeal lesions and exanthem: Toronto, summer 1957—Isolation of group A coxsackie virus. Can. Med. Assoc. J. 1958, 79, 615. [Google Scholar]
- Solomon, T.; Lewthwaite, P.; Perera, D.; Cardosa, M.J.; McMinn, P.; Ooi, M.H. Virology, epidemiology, pathogenesis, and control of enterovirus 71. Lancet Infect. Dis. 2010, 10, 778–790. [Google Scholar] [CrossRef]
- Li, X.-W.; Ni, X.; Qian, S.-Y.; Wang, Q.; Jiang, R.-M.; Xu, W.-B.; Zhang, Y.-C.; Yu, G.-J.; Chen, Q.; Shang, Y.-X. Chinese guidelines for the diagnosis and treatment of hand, foot and mouth disease (2018 edition). World J. Pediatr. 2018, 14, 437–447. [Google Scholar] [CrossRef]
- Gupta, S.; Jawanda, M.K. Oral lichen planus: An update on etiology, pathogenesis, clinical presentation, diagnosis and management. Indian J. Dermatol. 2015, 60, 222–229. [Google Scholar] [CrossRef] [PubMed]
- Manchanda, Y.; Rathi, S.K.; Joshi, A.; Das, S. Oral lichen planus: An updated review of etiopathogenesis, clinical presentation, and management. Indian Dermatol. Online J. 2024, 15, 8–23. [Google Scholar] [CrossRef] [PubMed]
- Tziotzios, C.; Lee, J.Y.; Brier, T.; Saito, R.; Hsu, C.-K.; Bhargava, K.; Stefanato, C.M.; Fenton, D.A.; McGrath, J.A. Lichen planus and lichenoid dermatoses: Clinical overview and molecular basis. J. Am. Acad. Dermatol. 2018, 79, 789–804. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.-P.; Chang, J.Y.-F.; Wang, Y.-P.; Wu, Y.-H.; Lu, S.-Y.; Sun, A. Oral lichen planus–differential diagnoses, serum autoantibodies, hematinic deficiencies, and management. J. Formos. Med. Assoc. 2018, 117, 756–765. [Google Scholar] [CrossRef]
- González-Moles, M.Á.; Ruiz-Avila, I.; Gonzalez-Ruiz, L.; Ayen, A.; Gil-Montoya, J.A.; Ramos-Garcia, P. Malignant transformation risk of oral lichen planus: A systematic review and comprehensive meta-analysis. Oral Oncol. 2019, 96, 121–130. [Google Scholar] [CrossRef]
- Tao, Y.; Ai, R.; Hao, Y.; Jiang, L.; Dan, H.; Ji, N.; Zeng, X.; Zhou, Y.; Chen, Q. Role of miR-155 in immune regulation and its relevance in oral lichen planus. Exp. Ther. Med. 2019, 17, 575–586. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; He, Y.; Xiang, J.; Feng, L.; Wang, Y.; Chen, R. The functional mechanism of MicroRNA in oral lichen planus. J. Inflamm. Res. 2022, 15, 4261–4274. [Google Scholar] [CrossRef]
- Seif, S.; Afra, N.; Dadgar, E.; Enteghad, S.; Argani, P.; Aghdasi, N.; Masouleh, S.S.; Barati, G. The expression of salivary microRNAs in oral lichen planus: Searching for a prognostic biomarker. Pathol.-Res. Pract. 2022, 234, 153923. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.Y.; Zhang, J.; Lu, R.; Tan, Y.Q.; Du, G.F.; Zhou, G. T cell–derived exosomes induced macrophage inflammatory protein-1α/β drive the trafficking of CD8+ T cells in oral lichen planus. J. Cell. Mol. Med. 2020, 24, 14086–14098. [Google Scholar] [CrossRef] [PubMed]
- Maloney, B.; Galvin, S.; Healy, C. Oral leukoplakia: An update for dental practitioners. J. Ir. Dent. Assoc. 2025, 71, 34–40. [Google Scholar] [CrossRef]
- Warnakulasuriya, S.; Kujan, O.; Aguirre-Urizar, J.M.; Bagan, J.V.; González-Moles, M.Á.; Kerr, A.R.; Lodi, G.; Mello, F.W.; Monteiro, L.; Ogden, G.R. Oral potentially malignant disorders: A consensus report from an international seminar on nomenclature and classification, convened by the WHO Collaborating Centre for Oral Cancer. Oral Dis. 2021, 27, 1862–1880. [Google Scholar] [CrossRef]
- Maserejian, N.N.; Joshipura, K.J.; Rosner, B.A.; Giovannucci, E.; Zavras, A.I. Prospective study of alcohol consumption and risk of oral premalignant lesions in men. Cancer Epidemiol. Biomark. Prev. 2006, 15, 774–781. [Google Scholar] [CrossRef]
- Khan, Z.; Khan, S.; Christianson, L.; Rehman, S.; Ekwunife, O.; Samkange-Zeeb, F. Smokeless tobacco and oral potentially malignant disorders in South Asia: A systematic review and meta-analysis. Nicotine Tob. Res. 2017, 20, 12–21. [Google Scholar] [CrossRef]
- Observatory, G.C. Cancer Today (version 1.1); International Agency for Research on Cancer: Lyon, France, 16 June 2025. [Google Scholar]
- Coletta, R.D.; Yeudall, W.A.; Salo, T. Current trends on prevalence, risk factors and prevention of oral cancer. Front. Oral Health 2024, 5, 1505833. [Google Scholar] [CrossRef]
- Coletta, R.D.; Yeudall, W.A.; Salo, T. Grand challenges in oral cancers. Front. Oral Health 2020, 1, 3. [Google Scholar] [CrossRef]
- Shah, J.P.; Gil, Z. Current concepts in management of oral cancer–surgery. Oral Oncol. 2009, 45, 394–401. [Google Scholar] [CrossRef]
- Thomson, P. Perspectives on oral squamous cell carcinoma prevention—Proliferation, position, progression and prediction. J. Oral Pathol. Med. 2018, 47, 803–807. [Google Scholar] [CrossRef]
- Dineja, R.; Ashwini, R. Journey of Exosomes in Oral Squamous Cell Carcinoma (OSCC). Oral Oncol. Rep. 2024, 12, 100674. [Google Scholar] [CrossRef]
- Nakamichi, E.; Sakakura, H.; Mii, S.; Yamamoto, N.; Hibi, H.; Asai, M.; Takahashi, M. Detection of serum/salivary exosomal Alix in patients with oral squamous cell carcinoma. Oral Dis. 2021, 27, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, L.; Zhu, G. Role of extracellular vesicles on cancer lymphangiogenesis and lymph node metastasis. Front. Oncol. 2021, 11, 721785. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhou, Y.; Liu, J.; Su, X.; Qin, H.; Huang, S.; Huang, X.; Zhou, N. Potential markers from serum-purified exosomes for detecting oral squamous cell carcinoma metastasis. Cancer Epidemiol. Biomark. Prev. 2019, 28, 1668–1681. [Google Scholar] [CrossRef]
- Liu, C.J.; Lin, S.C.; Yang, C.C.; Cheng, H.W.; Chang, K.W. Exploiting salivary miR-31 as a clinical biomarker of oral squamous cell carcinoma. Head Neck 2012, 34, 219–224. [Google Scholar] [CrossRef]
| Country | Type of Oral Mucosa Diseases | Source of Small Extracellular Vesicles | Analytical Techniques | Findings | Exosomal Biomarker | Sample Size | Author (Year) |
|---|---|---|---|---|---|---|---|
| China | Hand, foot, and mouth disease (HFMD) | Serum | Microarray method | Upregulation in HFMD patients compared with controls | miR-16-5p | 54 (18 ESHFMD, 18 MHFMD, and 18 healthy controls) | Jia et al. 2014 [18] |
| Downregulation in HFMD patients compared with controls | miR-150-3p and miR-671-5p | ||||||
| South Korea | Oral lichen planus (OLP) | Saliva | miRNA microarray analysis and TaqMan quantitative polymerase chain reaction | Upregulation in OLP patients compared with controls | miR-4484 | 24 (16 OLP patients and 8 healthy controls) | Byun et al. 2015 [19] |
| Iran | Saliva | Quantitative RT-PCR | Upregulation in OLP patients compared with controls | miR-21 | 60 (30 OLP patients, 15 OSCC patients, and 15 healthy controls) | Mehdipour et al. 2018 [20] | |
| Downregulation in OLP patients compared with controls | miR-125a | ||||||
| Italy | Saliva | Low-density microarray analysis and qRT-PCR | Upregulation in OLP patients compared with controls | miR-21, miR-125b, miR-203, and miR15b | A systematic review of 6 research articles | Stasio et al. 2019 [21] | |
| Down regulation in OLP patients compared with controls | miR-27b | ||||||
| Iran | RT-qPCR | Upregulation in OLP patients compared with controls | miR-146a, miR-155 | 60 patients (15 patients each with and without dysplastic OLP, 15 OSCC patients, and 15 healthy controls) | Mehdipour et al. 2023 [22] | ||
| China | Plasma | Exosomal miRNA microarray analysis and quantitative real-time RT-PCR confirmation | Upregulation in OLP patients compared with controls | miR-34a-5p and miR- 130b-5p | 30 (19 OLP patients and 11 age- and sex-matched healthy controls) | Peng et al. 2018 [23] | |
| Downregulation in OLP patients compared with controls | miR-301b-3p | ||||||
| Latvia | Oral Leukoplakia (OL) | Saliva | OncAlert® oral cancer rapid test | Increased level in OL patients compared to healthy controls and increasing level with severity | SolCD44 and total protein | 70 (50 OL patients and 20 controls with benign lesions) | Čēma et al. 2021 [24] |
| USA | Oral squamous cell carcinoma (OSCC) | Saliva | Irregular in OSCC patients | Morphology of sEVs | 10 (5 OSCC patients and 5 healthy volunteers) | Sharma et al. 2011 [16] | |
| Increased in OSCC patients compared with healthy controls | Size of sEVs | ||||||
| Increased in OSCC patients compared with healthy controls | Particle aggregation of sEVs | ||||||
| Israel | Saliva | Transmission electron microscopy, atomic force microscopy (AFM), and nanoparticle tracking analysis (NTA) | Increased value in OSCC patients compared with controls | Morphology, size, and concentration of sEVs | 61 (36 OSCC patients and 25 healthy individuals) | Zlotogorski- Hurvitz et al. 2016 [17] | |
| ELISA and Western blotting | Higher concentration in OSCC patients compared with healthy controls | CD 63 | |||||
| Lower concentration in OSCC patients compared with healthy controls | CD 9, 81, 82 | ||||||
| Italy | Saliva | qRT-PCR | Overexpression in OSCC patients compared with controls | miR-27a-3p, miR-373-3p, miR-494-3p | 32 (21 OSCC patients and 11 healthy controls) | Gai et al. 2018 [25] | |
| Higher levels in OSCC patients compared with healthy individuals | miR-412-3p and miR-512-3p | ||||||
| Expressed only in OSCC patients | miR-302b-3p and miR-517b-3p | ||||||
| China | Saliva | miRNA microarray analysis and qRT-PCR | Higher levels in OSCC patients compared with healthy individuals | miR-24-3p | 8 (4 OSCC patients and 4 healthy controls) | He et al. 2020 [26] | |
| Romania | Saliva | qRT-PCR | Higher levels in OSCC patients compared with healthy individuals | miR-486-5p | 50 (25 OSCC patients and 25 healthy controls) | Faur et al. 2022 [27] | |
| Lower levels in OSCC patients compared with healthy individuals | miR-10b-5p | ||||||
| India | Saliva | Real-time PCR | Overexpression in OSCC patients compared with controls | miR-1307-5p | 17 (12 OSCC patients and 5 healthy controls) | Patel et al. 2022 [28] | |
| Italy | Plasma | qRT-PCR | Overexpression in OSCC patients compared with controls | miR-210 | 44 (30 OSCC patients and 14 healthy controls) | Bigagli et al. 2022 [29] | |
| China | Plasma | qRT-PCR | Higher in OSCC patients compared to controls | miR-130a | 380 (184 OSCC patients and 196 healthy controls) | He et al. 2021 [30] | |
| China | Serum | Increased expression in OSCC patients | Squamous cell carcinoma antigen (SCCA) | 186 (73 OSCC patients and 113 healthy controls | Yang et al. 2022 [31] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Adebayo, O.F.; Temilola, D.O.; Owotade, F.J.; Chetty, M. Clinical Utility of Small Extracellular Vesicles as Liquid Biopsy for Oral Mucosal Disease Diagnostics: Emerging Perspectives. Diagnostics 2026, 16, 1044. https://doi.org/10.3390/diagnostics16071044
Adebayo OF, Temilola DO, Owotade FJ, Chetty M. Clinical Utility of Small Extracellular Vesicles as Liquid Biopsy for Oral Mucosal Disease Diagnostics: Emerging Perspectives. Diagnostics. 2026; 16(7):1044. https://doi.org/10.3390/diagnostics16071044
Chicago/Turabian StyleAdebayo, Olawande Funmilola, Dada Oluwaseyi Temilola, Foluso John Owotade, and Manogari Chetty. 2026. "Clinical Utility of Small Extracellular Vesicles as Liquid Biopsy for Oral Mucosal Disease Diagnostics: Emerging Perspectives" Diagnostics 16, no. 7: 1044. https://doi.org/10.3390/diagnostics16071044
APA StyleAdebayo, O. F., Temilola, D. O., Owotade, F. J., & Chetty, M. (2026). Clinical Utility of Small Extracellular Vesicles as Liquid Biopsy for Oral Mucosal Disease Diagnostics: Emerging Perspectives. Diagnostics, 16(7), 1044. https://doi.org/10.3390/diagnostics16071044






