A Novel Approach Using Microarray Testing as a Screening Method with Clinical Validation Using Whole-Genome Sequencing and Karyotyping for Identifying 46,XX Testicular Differences of Sex Development
Abstract
1. Introduction
2. Case Presentation
2.1. Subject Profile
2.2. Discovery of Genetic Abnormality
2.3. Microarray Analysis
2.4. Whole-Genome Sequencing (WGS) Analysis
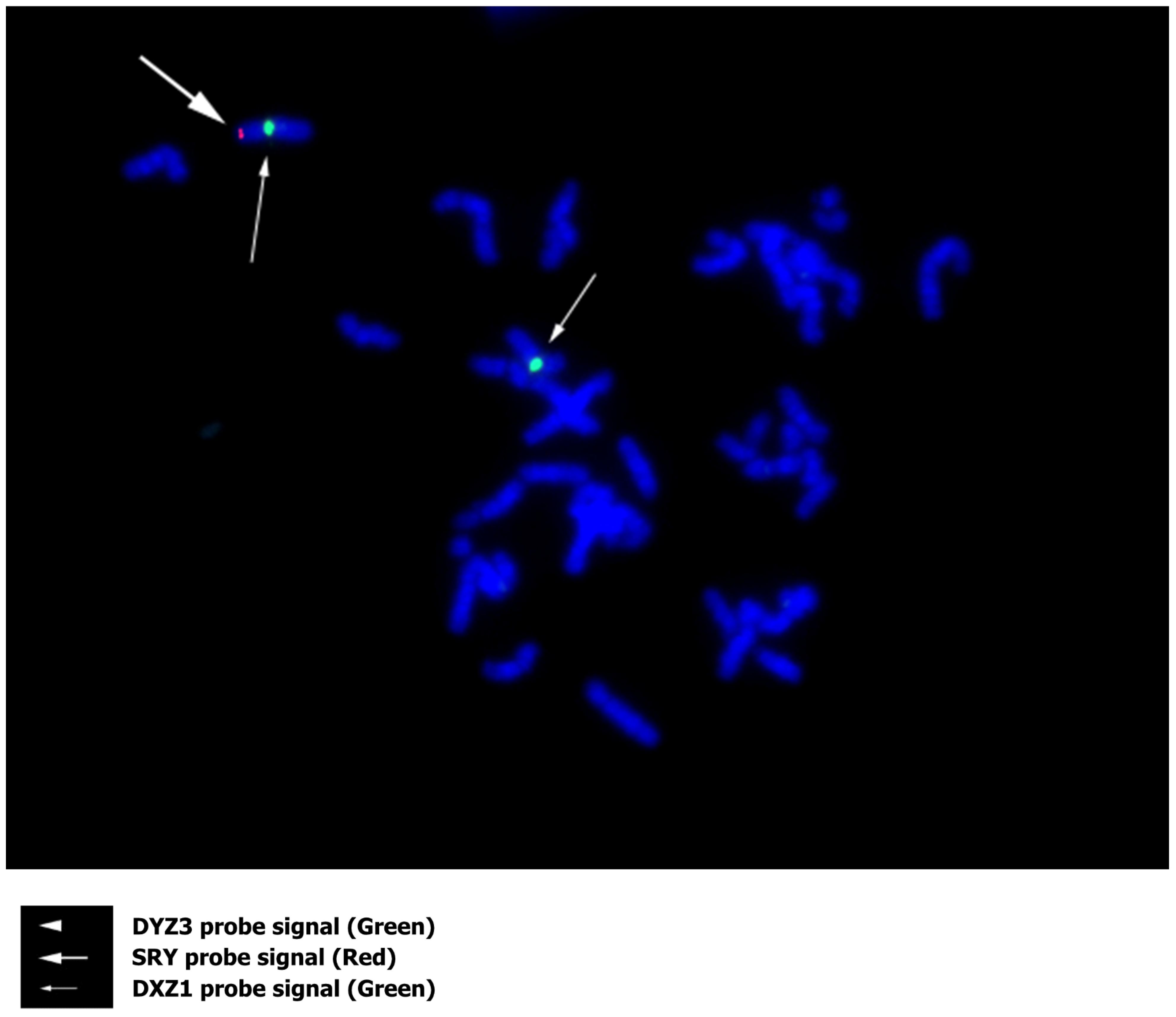
2.5. G-Banding, Fluorescence In Situ Hybridization (FISH), and Endocrine Testing
3. Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| GWAS | Genome-wide association study |
| DTC | Direct to consumer |
| DSD | Differences of sex development |
| WGS | Whole-genome sequencing |
| SRY | Sex-determining Region Y |
| GH | Genesis Healthcare Co. |
| SNP | Single-nucleotide polymorphism |
| PAR | Pseudo autosomal region |
| FISH | Fluorescence in situ hybridization |
| DHEA-S | Dehydroepiandrosterone sulfate |
| GH | Growth hormone |
| LH | Luteinizing hormone |
| FSH | Follicle-stimulating hormone |
References
- Frisova, V. Prenatal Screening for Chromosomal Defects. Reprod. Med. 2025, 6, 15. [Google Scholar] [CrossRef]
- Hochstenbach, R.; Van Binsbergen, E.; Schuring-Blom, H.; Buijs, A.; Ploos Van Amstel, H.K. A Survey of Undetected, Clinically Relevant Chromosome Abnormalities When Replacing Postnatal Karyotyping by Whole Genome Sequencing. Eur. J. Med. Genet. 2019, 62, 103543. [Google Scholar] [CrossRef]
- Guttmacher, A.E.; Collins, F.S. Genomic Medicine—A Primer. N. Engl. J. Med. 2002, 347, 1512–1520. [Google Scholar] [CrossRef] [PubMed]
- Feero, W.G.; Guttmacher, A.E.; Collins, F.S. Genomic Medicine—An Updated Primer. N. Engl. J. Med. 2010, 362, 2001–2011. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.; Barapatre, A.R.; Babar, N.; Doshi, J.; Ghaly, M.; Patel, K.G.; Nawaz, S.; Hasana, U.; Khatri, S.P.; Pathange, S.; et al. Genomic Medicine and Personalized Treatment: A Narrative Review. Ann. Med. Surg. 2025, 87, 1406–1414. [Google Scholar] [CrossRef] [PubMed]
- Davies, K. The Era of Genomic Medicine. Clin. Med. 2013, 13, 594–601. [Google Scholar] [CrossRef]
- Manolio, T.A.; Chisholm, R.L.; Ozenberger, B.; Roden, D.M.; Williams, M.S.; Wilson, R.; Bick, D.; Bottinger, E.P.; Brilliant, M.H.; Eng, C.; et al. Implementing Genomic Medicine in the Clinic: The Future Is Here. Genet. Med. 2013, 15, 258–267. [Google Scholar] [CrossRef]
- Milko, L.V.; Khoury, M.J. Editorial: DNA-Based Population Screening for Precision Public Health. Front. Genet. 2022, 13, 1061329. [Google Scholar] [CrossRef]
- Martins, M.F.; Murry, L.T.; Telford, L.; Moriarty, F. Direct-to-Consumer Genetic Testing: An Updated Systematic Review of Healthcare Professionals’ Knowledge and Views, and Ethical and Legal Concerns. Eur. J. Hum. Genet. 2022, 30, 1331–1343. [Google Scholar] [CrossRef]
- Roberts, J.S.; Ostergren, J. Direct-to-Consumer Genetic Testing and Personal Genomics Services: A Review of Recent Empirical Studies. Curr. Genet. Med. Rep. 2013, 1, 182–200. [Google Scholar] [CrossRef]
- Kilbride, M.K.; Kessler, L.J.; Cronier, B.; Park, J.J.; Cacioppo, C.N.; Beem, J.; Bradbury, A.R. Test-Takers’ Perspectives on Consumer Genetic Testing for Hereditary Cancer Risk. Front. Genet. 2024, 15, 1374602. [Google Scholar] [CrossRef]
- Horton, R.; Crawford, G.; Freeman, L.; Fenwick, A.; Lucassen, A. Direct-to-Consumer Genetic Testing with Third Party Interpretation: Beware of Spurious Results. Emerg. Top. Life Sci. 2019, 3, 669–674. [Google Scholar] [CrossRef]
- Malgorzata, M.; Maria, S.; Michał, W. Genetic Testing—Whether to Allow Complete Freedom? Direct to Consumer Tests versus Genetic Tests for Medical Purposes. J. Appl. Genet. 2022, 63, 119–126. [Google Scholar] [CrossRef]
- Nolan, J.J.; Ormondroyd, E. Direct-to-consumer Genetic Tests Providing Health Risk Information: A Systematic Review of Consequences for Consumers and Health Services. Clin. Genet. 2023, 104, 3–21. [Google Scholar] [CrossRef]
- Ozaki, K.; Ohnishi, Y.; Iida, A.; Sekine, A.; Yamada, R.; Tsunoda, T.; Sato, H.; Sato, H.; Hori, M.; Nakamura, Y.; et al. Functional SNPs in the Lymphotoxin-α Gene That Are Associated with Susceptibility to Myocardial Infarction. Nat. Genet. 2002, 32, 650–654. [Google Scholar] [CrossRef] [PubMed]
- Abdellaoui, A.; Yengo, L.; Verweij, K.J.H.; Visscher, P.M. 15 Years of GWAS Discovery: Realizing the Promise. Am. J. Hum. Genet. 2023, 110, 179–194. [Google Scholar] [CrossRef] [PubMed]
- Levy, B.; Wapner, R. Prenatal Diagnosis by Chromosomal Microarray Analysis. Fertil. Steril. 2018, 109, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Miller, D.T.; Adam, M.P.; Aradhya, S.; Biesecker, L.G.; Brothman, A.R.; Carter, N.P.; Church, D.M.; Crolla, J.A.; Eichler, E.E.; Epstein, C.J.; et al. Consensus Statement: Chromosomal Microarray Is a First-Tier Clinical Diagnostic Test for Individuals with Developmental Disabilities or Congenital Anomalies. Am. J. Hum. Genet. 2010, 86, 749–764. [Google Scholar] [CrossRef]
- Hughes, I.A.; Houk, C.P.; Ahmed, S.F.; Lee, P.A.; ESPE Consensus Group; Group EC. Consensus statement on management of intersex disorders. Arc. Dis. Child. 2006, 91, 554–563. [Google Scholar] [CrossRef]
- Lee, P.A.; Nordenström, A.; Houk, C.P.; Ahmed, S.F.; Auchus, R.; Baratz, A.; Baratz Dalke, K.; Liao, L.M.; Lin-Su, K.; Mazur, T.; et al. Global disorders of sex development update since 2006: Perceptions, approach, and care. Horm. Res. Paediatr. 2016, 85, 158–180. [Google Scholar] [CrossRef]
- Cools, M.; Nordenström, A.; Robeva, R.; Hall, J.; Westerveld, P.; Flück, C.; Köhler, B.; Berra, M.; Springer, A.; Schweizer, K.; et al. Caring for individuals with a difference of sex development (DSD): A Consensus Statement. Nat. Rev. Endocrinol. 2018, 14, 415–429. [Google Scholar] [CrossRef] [PubMed]
- De La Chapelle, A.; Hortling, H.; Niemi, M.; Wennström, J. XX Sex Chromosomes in a Human Male: First Case. J. Int. Med. 1964, 175, 25–38. [Google Scholar] [CrossRef] [PubMed]
- Berglund, A.; Johannsen, T.H.; Stochholm, K.; Aksglaede, L.; Fedder, J.; Viuff, M.H.; Main, K.M.; Gravholt, C.H. Incidence, Prevalence, Diagnostic Delay, Morbidity, Mortality and Socioeconomic Status in Males with 46,XX Disorders of Sex Development: A Nationwide Study. Hum. Reprod. 2017, 32, 1751–1760. [Google Scholar] [CrossRef] [PubMed]
- Okashita, N.; Tachibana, M. Transcriptional Regulation of the Y-Linked Mammalian Testis-Determining Gene SRY. Sex. Dev. 2021, 15, 351–359. [Google Scholar] [CrossRef]
- Monteiro, B.; Arenas, M.; Prata, M.J.; Amorim, A. Evolutionary Dynamics of the Human Pseudoautosomal Regions. PLoS Genet. 2021, 17, e1009532. [Google Scholar] [CrossRef]
- Li, H.; He, J.; Leong, I. A Rare Case of 46, XX (SRY Positive) Testicular Disorder of Sex Development with Growth Hormone Deficiency: Case Report. Medicine 2021, 100, e24641. [Google Scholar] [CrossRef]
- Adrião, M.; Ferreira, S.; Silva, R.S.; Garcia, M.; Dória, S.; Costa, C.; Castro-Correia, C.; Fontoura, M. 46,XX Male Disorder of Sexual Development. Clin. Pediatr. Endocrinol. 2020, 29, 43–45. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative Genomics Viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef]
- Thorvaldsdottir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-Performance Genomics Data Visualization and Exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Wenger, A.M.; Zehir, A.; Mesirov, J.P. Variant Review with the Integrative Genomics Viewer. Cancer Res. 2017, 77, e31–e34. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdottir, H.; Turner, D.; Mesirov, J.P. Igv.Js: An Embeddable JavaScript Implementation of the Integrative Genomics Viewer (IGV). Bioinformatics 2023, 39, btac830. [Google Scholar] [CrossRef] [PubMed]
- Mensah, M.A.; Hestand, M.S.; Larmuseau, M.H.D.; Isrie, M.; Vanderheyden, N.; Declercq, M.; Souche, E.L.; Van Houdt, J.; Stoeva, R.; Van Esch, H.; et al. Pseudoautosomal Region 1 Length Polymorphism in the Human Population. PLoS Genet. 2014, 10, e1004578. [Google Scholar] [CrossRef] [PubMed]
- Kirsch, S.; Weiss, B.; De Rosa, M.; Ogata, T.; Lombardi, G.; Rappold, G.A. FISH deletion mapping defines a single location for the Y chromosome stature gene, GCY. J. Med. Genet. 2000, 37, 593–599. [Google Scholar] [CrossRef][Green Version]
- Iwamoto, T.; Yanase, T.; Koh, E.; Horie, H.; Baba, K.; Namiki, M.; Nawata, H. Reference ranges of serum total and free testosterone in Japanese male adults. Jpn. J. Urol. 2004, 95, 751–760. [Google Scholar] [CrossRef]
- Chen, T.; Tian, L.; Wu, F.; Xuan, X.; Ma, G.; Tang, R.; Lu, J. Clinical and Genetic Analysis in Males with 46,XX Disorders of Sex Development: A Reproductive Centre Experience of 144 Cases. Andrologia 2019, 51, e13232. [Google Scholar] [CrossRef] [PubMed]
- Akinsal, E.C.; Baydilli, N.; Demirtas, A.; Saatci, C.; Ekmekcioglu, O. Ten Cases with 46,XX Testicular Disorder of Sex Development: Single Center Experience. Int. Braz. J. Urol. 2017, 43, 770–775. [Google Scholar] [CrossRef]
- Chin, R.T.; Mok, S.F. 46,XX Testicular Disorder of Sex Development (DSD) Presenting with Male Hypogonadism. JCEM Case Rep. 2025, 3, luae237. [Google Scholar] [CrossRef]
- Howard, S.R. Interpretation of Reproductive Hormones Before, During and After the Pubertal Transition—Identifying Health and Disordered Puberty. Clin. Endocrinol. 2021, 95, 702–715. [Google Scholar] [CrossRef]
- Aksglaede, L.; Jensen, R.B.; Carlsen, E.; Kok, P.; Keenan, D.M.; Veldhuis, J.; Skakkebæk, N.E.; Juul, A. Increased Basal and Pulsatile Secretion of FSH and LH in Young Men with 47,XXY or 46,XX Karyotypes. Eur. J. Endocrinol. 2008, 158, 803–810. [Google Scholar] [CrossRef]



| Entry | Male (Mean ± SD) (n = 27,181) | Female (Mean ± SD) (n = 40,509) | Subject |
|---|---|---|---|
| Y chromosome call rate 1 | 71.9 ± 1.2% | 26.4 ± 2.5% | 39.1% |
| X chromosome heterozygosity 2 | 0.9 ± 0.1% | 16.2 ± 0.8% | 16.2% |
| Deduced sex | Male | Female | Female? |
| Result in accordance with ISCN 2024 | arr[GRCh37] (X,Y)×1,(1–2)×2 | arr[GRCh37] (X,1–22) × 2 | arr[GRCh37] Xp22.33p22.3(181,779_3,555,302) × 1, Yp11.32p11.1(181,779_9,314,387) × 1 |
| Test Item 1 | Result (Unit) | Reference Value (Unit) |
|---|---|---|
| Free testosterone | 5.6 pg/mL | 4.7–21.6 pg/mL |
| Testosterone | 4.76 ng/mL | 1.92–8.84 ng/mL |
| DHEA-S | 201 μg/dL | 123–422 μg/dL |
| GH | 0.78 ng/mL | <2.47 ng/mL |
| LH | 21.61 mIU/mL | 0.79–5.72 mIU/mL |
| FSH | 51.19 mIU/mL | 2.00–8.30 mIU/mL |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ozeki, T.; Nakano, Y.; Ishigaki, A.; Kawashima, Y.; Crawford, C.W.; Ordinario, D.D.; Sato-Baran, I.; Higashida, T. A Novel Approach Using Microarray Testing as a Screening Method with Clinical Validation Using Whole-Genome Sequencing and Karyotyping for Identifying 46,XX Testicular Differences of Sex Development. Diagnostics 2026, 16, 706. https://doi.org/10.3390/diagnostics16050706
Ozeki T, Nakano Y, Ishigaki A, Kawashima Y, Crawford CW, Ordinario DD, Sato-Baran I, Higashida T. A Novel Approach Using Microarray Testing as a Screening Method with Clinical Validation Using Whole-Genome Sequencing and Karyotyping for Identifying 46,XX Testicular Differences of Sex Development. Diagnostics. 2026; 16(5):706. https://doi.org/10.3390/diagnostics16050706
Chicago/Turabian StyleOzeki, Takeshi, Yuka Nakano, Ayumu Ishigaki, Yoichi Kawashima, Charles W. Crawford, David D. Ordinario, Iri Sato-Baran, and Toshihiko Higashida. 2026. "A Novel Approach Using Microarray Testing as a Screening Method with Clinical Validation Using Whole-Genome Sequencing and Karyotyping for Identifying 46,XX Testicular Differences of Sex Development" Diagnostics 16, no. 5: 706. https://doi.org/10.3390/diagnostics16050706
APA StyleOzeki, T., Nakano, Y., Ishigaki, A., Kawashima, Y., Crawford, C. W., Ordinario, D. D., Sato-Baran, I., & Higashida, T. (2026). A Novel Approach Using Microarray Testing as a Screening Method with Clinical Validation Using Whole-Genome Sequencing and Karyotyping for Identifying 46,XX Testicular Differences of Sex Development. Diagnostics, 16(5), 706. https://doi.org/10.3390/diagnostics16050706





