Progression-Aware and Explainable CNN–Transformer Framework for Multiclass Alzheimer’s Disease Staging Using MRI
Abstract
1. Introduction
- We use progression-aware ordinal learning to directly model the ordering of severity of Alzheimer disease.
- We apply consistency regularization to enhance the robustness of the slice-level predictions of MRI images.
- We provide token-level interpretability analysis to improve transparency and trust in model decisions.
- We design an efficient CNN–Transformer architecture suitable for real-world MRI imaging applications.
2. Related Work
3. Methodology
3.1. Dataset Description
3.2. The Proposed Framework
3.3. Data Preparation and Leakage-Free Evaluation Protocol
3.4. Convolutional Feature Extraction
3.5. Transformer-Based Global Context Modeling
3.6. Progression-Aware Ordinal Learning
3.7. Consistency Regularization for Robust Prediction
3.8. Training Objective
3.9. Model Interpretability via Token Importance
3.10. Implementation Details
4. Results
4.1. Ordinal Error Analysis and Clinical Consistency
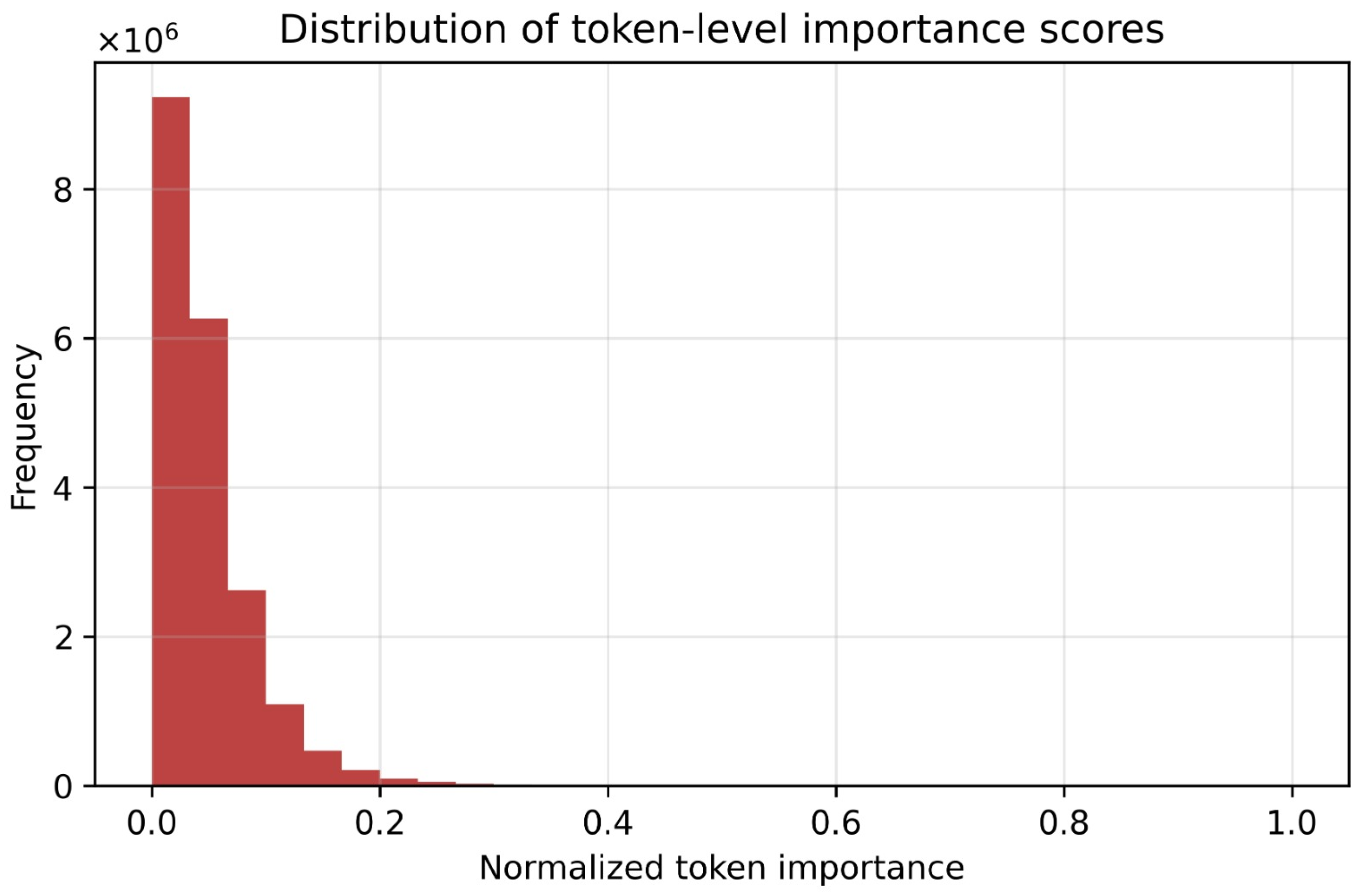
4.2. Token-Level Interpretability and Disease Progression Analysis
4.3. Ablation Study
5. Discussion
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jack, C.R., Jr.; Bennett, D.A.; Blennow, K.; Carrillo, M.C.; Dunn, B.; Haeberlein, S.B.; Holtzman, D.M.; Jagust, W.; Jessen, F.; Karlawish, J.; et al. NIA-AA Research Framework: Toward a biological definition of Alzheimer’s disease. Alzheimer’s Dement. 2018, 14, 535–562. [Google Scholar] [CrossRef] [PubMed]
- Alzheimer’s Disease International. World Alzheimer Report 2021: Journey Through the Diagnosis of Dementia; Alzheimer’s Disease International: London, UK, 2021; Available online: https://www.alzint.org/resource/world-alzheimer-report-2021/ (accessed on 9 January 2026).
- Scheltens, P.; Strooper, B.D.; Kivipelto, M.; Holstege, H.; Chételat, G.; Teunissen, C.E.; Cummings, J.; Van der Flier, W.M. Alzheimer’s disease. Lancet 2021, 397, 1577–1590. [Google Scholar] [CrossRef] [PubMed]
- Frisoni, G.B.; Fox, N.C.; Jack, C.R.; Scheltens, P.; Thompson, P.M. The clinical use of structural MRI in Alzheimer disease. Nat. Rev. Neurol. 2010, 6, 67–77. [Google Scholar] [CrossRef]
- Jack, C.R., Jr.; Petersen, R.C.; Xu, Y.C.; Waring, S.C.; O’Brien, P.C.; Tangalos, E.G.; Smith, G.E.; Ivnik, R.J.; Kokmen, E. Medial temporal atrophy on MRI in normal aging and very mild Alzheimer’s disease. Neurology 1997, 49, 786–794. [Google Scholar] [CrossRef]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; Van Der Laak, J.A.; Van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Shen, D.; Wu, G.; Suk, H.-I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef]
- Wen, J.; Thibeau-Sutre, E.; Diaz-Melo, M.; Samper-González, J.; Routier, A.; Bottani, S.; Dormont, D.; Durrleman, S.; Burgos, N.; Colliot, O. Convolutional neural networks for classification of Alzheimer’s disease: Overview and reproducible evaluation. Med. Image Anal. 2020, 63, 101694. [Google Scholar] [CrossRef]
- Zhuang, X.; Li, L.; Payer, C.; Štern, D.; Urschler, M.; Heinrich, M.P.; Oster, J.; Wang, C.; Smedby, Ö; Bian, C.; et al. Evaluation of algorithms for multi-modality whole heart segmentation: An open-access grand challenge. Med. Image Anal. 2019, 58, 101537. [Google Scholar] [CrossRef]
- Suk, H.-I.; Lee, S.-W.; Shen, D. Deep ensemble learning of sparse regression models for brain disease diagnosis. Med. Image Anal. 2017, 37, 101–113. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An Image is Worth 16 × 16 Words: Transformers for Image Recognition at Scale. In Proceedings of the International Conference on Learning Representations (ICLR); ICLR: Appleton, WI, USA, 2021. [Google Scholar]
- Yao, Z.; Xie, W.; Chen, J.; Zhan, Y.; Wu, X.; Dai, Y.; Pei, Y.; Wang, Z.; Zhang, G. IT: An interpretable transformer model for Alzheimer’s disease prediction based on PET/MR images. NeuroImage 2025, 311, 121210. [Google Scholar] [CrossRef]
- Singhal, R. Alzheimer Multiclass MRI Dataset (4-Class). Kaggle. 2022. Available online: https://www.kaggle.com/datasets/uraninjo/augmented-alzheimer-mri-dataset-v2 (accessed on 9 January 2026).
- Suk, H.-I.; Lee, S.-W.; Shen, D. The ADNI. Hierarchical feature representation and multimodal fusion with deep learning for AD/MCI diagnosis. NeuroImage 2014, 101, 569–582. [Google Scholar] [CrossRef]
- Hosseini-Asl, E.; Keynton, R.; El-Baz, A. Alzheimer’s disease diagnostics by adaptation of 3D convolutional network. In 2016 IEEE International Conference on Image Processing (ICIP); IEEE: Piscataway, NJ, USA, 2016; pp. 126–130. [Google Scholar] [CrossRef]
- Korolev, S.; Safiullin, A.; Belyaev, M.; Dodonova, Y. Residual and plain convolutional neural networks for 3D brain MRI classification. In 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017); IEEE: Piscataway, NJ, USA, 2017. [Google Scholar]
- Islam, J.; Zhang, Y. Brain MRI analysis for Alzheimer’s disease diagnosis using an ensemble system of deep convolutional neural networks. Brain Inform. 2018, 5, 2. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; He, X.; Liu, Y.; Cai, Q.; Chen, H.; Qing, L. Multi-modal cross-attention network for Alzheimer’s disease diagnosis with multi-modality data. Comput. Biol. Med. 2023, 162, 107203. [Google Scholar] [CrossRef] [PubMed]
- Falahati, F.; Westman, E.; Simmons, A. Multivariate data analysis and machine learning in Alzheimer’s disease with a focus on structural magnetic resonance imaging. J. Alzheimer’s Dis. 2014, 41, 685–708. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.F.J.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015; Springer: Cham, Switzerland, 2015. [Google Scholar]
- Dickerson, B.C.; Bakkour, A.; Salat, D.H.; Feczko, E.; Pacheco, J.; Greve, D.N.; Grodstein, F.; Wright, C.I.; Blacker, D.; Rosas, H.D.; et al. The Cortical Signature of Alzheimer’s Disease: Regionally Specific Cortical Thinning Relates to Symptom Severity in Very Mild to Mild AD Dementia and is Detectable in Asymptomatic Amyloid-Positive Individuals. Cereb. Cortex 2009, 19, 497–510. [Google Scholar] [CrossRef]
- Basaia, S.; Agosta, F.; Wagner, L.; Canu, E.; Magnani, G.; Santangelo, R.; Filippi, M. Alzheimer’s Disease Neuroimaging Initiative. Automated classification of Alzheimer’s disease and mild cognitive impairment using a single MRI and deep neural networks. NeuroImage Clin. 2019, 21, 101645. [Google Scholar] [CrossRef]
- Li, X.; Gong, B.; Chen, X.; Li, H.; Yuan, G. Alzheimer’s disease image classification based on enhanced residual attention network. PLoS ONE 2025, 20, e0317376. [Google Scholar] [CrossRef]
- Ibrar, W.; Khan, M.A.; Hamza, A.; Rubab, S.; Alqahtani, O.; Alouane, M.T.H.; Teng, S.; Nam, Y. A novel interpreted deep network for Alzheimer’s disease prediction based on inverted self attention and vision transformer. Sci. Rep. 2025, 15, 29974. [Google Scholar] [CrossRef]
- Ul Haq, E.; Qin, Y.; Zhou, Y.; Xu, H.; Ul Haq, R. Multimodal fusion diagnosis of Alzheimer’s disease via lightweight CNN–LSTM model using magnetic resonance imaging (MRI). Biomed. Signal Process. Control 2023, 85, 104985. [Google Scholar]
- Venkatraman, S.; Pandiyaraju, V.; Abeshek, A.; Kumar, S.P.; Aravintakshan, S.A. Leveraging bi-focal perspectives and granular feature integration for accurate reliable early Alzheimer’s detection. IEEE Access 2025, 13, 28678–28692. [Google Scholar] [CrossRef]
- Alorf, A. Transformer and convolutional neural network: A hybrid model for multimodal data in multiclass classification of Alzheimer’s disease. Mathematics 2025, 13, 1548. [Google Scholar] [CrossRef]
- Zolfaghari, S.; Joudaki, A.; Sarbaz, Y. A hybrid learning approach for MRI-based detection of Alzheimer’s disease stages using dual CNNs and ensemble classifier. Sci. Rep. 2025, 15, 25342. [Google Scholar] [CrossRef]
- Hu, Z.; Wang, Y.; Li, Y. A transformer-based hybrid network for Alzheimer’s disease diagnosis via MRI. Digit. Signal Process. 2026, 168, 105562. [Google Scholar] [CrossRef]







| Fold | Accuracy | Precision | Recall | F1-Score | AUROC | AUPR | Cohen’s Kappa |
|---|---|---|---|---|---|---|---|
| Fold 1 | 0.9901 | 0.9912 | 0.9915 | 0.9914 | 0.9998 | 0.9995 | 0.9866 |
| Fold 2 | 0.9926 | 0.9932 | 0.9936 | 0.9934 | 0.9999 | 0.9998 | 0.9899 |
| Fold 3 | 0.9908 | 0.9919 | 0.9920 | 0.9920 | 0.9998 | 0.9996 | 0.9876 |
| Fold 4 | 0.9840 | 0.9854 | 0.9869 | 0.9861 | 0.9994 | 0.9988 | 0.9783 |
| Fold 5 | 0.9912 | 0.9919 | 0.9927 | 0.9923 | 0.9998 | 0.9996 | 0.9881 |
| Mean | 0.9897 | 0.9907 | 0.9913 | 0.9910 | 0.9998 | 0.9994 | 0.9861 |
| Std | 0.0033 | 0.0031 | 0.0026 | 0.0028 | 0.0002 | 0.0004 | 0.0040 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Alsalem, K.; Elbashir, M.K.; Alzahrani, A.O.; Mohammed, M.; Mahmood, M.A.; Abd El Fattah, T. Progression-Aware and Explainable CNN–Transformer Framework for Multiclass Alzheimer’s Disease Staging Using MRI. Diagnostics 2026, 16, 593. https://doi.org/10.3390/diagnostics16040593
Alsalem K, Elbashir MK, Alzahrani AO, Mohammed M, Mahmood MA, Abd El Fattah T. Progression-Aware and Explainable CNN–Transformer Framework for Multiclass Alzheimer’s Disease Staging Using MRI. Diagnostics. 2026; 16(4):593. https://doi.org/10.3390/diagnostics16040593
Chicago/Turabian StyleAlsalem, Khalaf, Murtada K. Elbashir, Ahmed Omar Alzahrani, Mohanad Mohammed, Mahmood A. Mahmood, and Tarek Abd El Fattah. 2026. "Progression-Aware and Explainable CNN–Transformer Framework for Multiclass Alzheimer’s Disease Staging Using MRI" Diagnostics 16, no. 4: 593. https://doi.org/10.3390/diagnostics16040593
APA StyleAlsalem, K., Elbashir, M. K., Alzahrani, A. O., Mohammed, M., Mahmood, M. A., & Abd El Fattah, T. (2026). Progression-Aware and Explainable CNN–Transformer Framework for Multiclass Alzheimer’s Disease Staging Using MRI. Diagnostics, 16(4), 593. https://doi.org/10.3390/diagnostics16040593





