Benchmark-Driven Clinical Decision Framework for Multi-Class Middle Ear Disease Diagnosis: Superiority of Swin Transformer in Accuracy and Stability
Abstract
1. Introduction
- To establish an accurate performance benchmark on a standardized, multicenter dataset comprising five diagnostic categories.
- To perform an extensive and fair comparison between representative CNN models (e.g., VGG16, ResNet50, Inception-v3) and Transformer variants (e.g., ViT, DeiT, Swin Transformer) under consistent experimental conditions to determine their relative performance.
- To conduct an in-depth analysis of the optimal model’s robustness to class imbalance and, based on its output characteristics, develop a practical, probability-guided Top-K clinical decision-support framework to provide a solution for real-world applications.
2. Materials and Methods
2.1. Dataset Construction
2.1.1. Data Source and Collection
2.1.2. Image Data Annotation and Characteristics
2.2. Data Preprocessing
2.2.1. Automatic Removal of Interference Information
2.2.2. Class Imbalance Handling
2.3. Swin Transformer Architecture and Model Setup
2.4. CNN Models for Comparison
2.5. Training Parameters and Strategy
2.6. Comprehensive Evaluation Strategy
2.7. Probability Output and Top-K Clinical Decision Mechanism
2.8. Performance Metrics
3. Results
3.1. Dataset Characteristics and Model Performance Overview
3.2. Benchmark Performance: Swin Transformer Outperforms CNNs
3.3. Model Robustness: Swin Transformer Exhibits Superior Stability
3.4. Disease-Specific Analysis: Superiority in Complex and Rare Conditions
3.5. Ablation Studies: Preprocessing Essential; Architectural Robustness Evident
3.6. Interpretability and Clinical Decision Support
4. Discussion
4.1. Performance Convergence and the Need for a Multi-Dimensional Evaluation Paradigm
4.2. Model Stability as a Primary Criterion for Clinical Deployment
4.3. Performance on Critical Diseases: Translating Architectural Advantage into Clinical Impact
4.4. Methodological Refinements and Their Impact
4.5. Clinical Workflow Integration and Implications
4.6. Advancements over the State of the Art: A Multi-Center Benchmark and Clinical Framework
4.7. Analysis of Clinically Meaningful Misclassification
4.8. Limitations and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CNNs | Convolutional Neural Networks |
| CAD | Computer-aided diagnosis |
| TM | Tympanic membrane |
| CSOM | Chronic Suppurative Otitis Media |
| OME | Otitis Media with Effusion |
| GM | Granular Myringitis |
| W-MSA | Window-based Multi-head Self-Attention |
| FC | Fully Connected Laye |
| LN | Layer Normalization |
| SW-MSA | Shifted Window-based Multi-head Self-Attention |
| MLP | Multi-Layer Perceptron |
| CI | Confidence Interval |
| SD | Standard Deviation |
| LDAM (DRW) | Label-Distribution-Aware Margin Loss with Deferred Re-Weighting training strategy |
| CB-Sampling | Class-Balanced Sampling |
| CB-Loss (Focal) | Class-Balanced Loss (based on Focal Loss) |
| Grad-CAM | Gradient-weighted Class Activation Mapping |
| STMC | Stable Tympanic Membrane Condition |
References
- Wang, H.; Zeng, X.; Miao, X.; Yang, B.; Zhang, S.; Fu, Q.; Zhang, Q.; Tang, M. Global, regional, and national epidemiology of otitis media in children from 1990 to 2021. Front. Pediatr. 2025, 13, 1513629. [Google Scholar] [CrossRef] [PubMed]
- Sirota, S.B.; Doxey, M.C.; Dominguez, R.V.; Bender, R.G.; Vongpradith, A.; Albertson, S.B.; Novotney, A.; Burkart, K.; Carter, A.; Abdi, P.; et al. Global, regional, and national burden of upper respiratory infections and otitis media, 1990–2021: A systematic analysis from the Global Burden of Disease Study 2021. Lancet Infect. Dis. 2025, 25, 36–51. [Google Scholar] [CrossRef] [PubMed]
- Dong, L.; Jin, Y.; Dong, W.; Jiang, Y.; Li, Z.; Su, K.; Yu, D. Trends in the incidence and burden of otitis media in children: A global analysis from 1990 to 2021. Eur. Arch. Oto-Rhino-Laryngol. 2025, 282, 2959–2970. [Google Scholar] [CrossRef] [PubMed]
- Monasta, L.; Ronfani, L.; Marchetti, F.; Montico, M.; Vecchi, B.L.; Bavcar, A.; Grasso, D.; Barbiero, C.; Tamburlini, G. Burden of disease caused by otitis media: Systematic review and global estimates. PLoS ONE 2012, 7, e36226. [Google Scholar] [CrossRef]
- Rosenfeld, R.M.; Shin, J.J.; Schwartz, S.R.; Coggins, R.; Gagnon, L.; Hackell, J.M.; Hoelting, D.; Hunter, L.L.; Kummer, A.W.; Payne, S.C.; et al. Clinical Practice Guideline: Otitis Media with Effusion (Update). Otolaryngol.-Head Neck Surg. Off. J. Am. Acad. Otolaryngol.-Head Neck Surg. 2016, 154, S1–S41. [Google Scholar] [CrossRef]
- Oyewumi, M.; Brandt, M.G.; Carrillo, B.; Atkinson, A.; Iglar, K.; Forte, V.; Campisi, P. Objective Evaluation of Otoscopy Skills Among Family and Community Medicine, Pediatric, and Otolaryngology Residents. J. Surg. Educ. 2016, 73, 129–135. [Google Scholar] [CrossRef]
- Pichichero, M.E.; Poole, M.D. Assessing diagnostic accuracy and tympanocentesis skills in the management of otitis media. Arch. Pediatr. Adolesc. Med. 2001, 155, 1137–1142. [Google Scholar] [CrossRef]
- Leung, A.K.C.; Wong, A.H.C. Acute Otitis Media in Children. Recent Pat. Inflamm. Allergy Drug Discov. 2017, 11, 32–40. [Google Scholar]
- Habib, A.R.; Kajbafzadeh, M.; Hasan, Z.; Wong, E.; Gunasekera, H.; Perry, C.; Sacks, R.; Kumar, A.; Singh, N. Artificial intelligence to classify ear disease from otoscopy: A systematic review and meta-analysis. Clin. Otolaryngol. 2022, 47, 401–413. [Google Scholar] [CrossRef]
- Basaran, E.; Cmert, Z.; Celik, Y. Convolutional neural network approach for automatic tympanic membrane detection and classification. Biomed. Signal Proces. 2020, 56, 101734. [Google Scholar] [CrossRef]
- Tsutsumi, K.; Goshtasbi, K.; Risbud, A.; Khosravi, P.; Pang, J.C.; Lin, H.W.; Djalilian, H.R.; Abouzari, M. A Web-Based Deep Learning Model for Automated Diagnosis of Otoscopic Images. Otol. Neurotol. 2021, 42, e1382–e1388. [Google Scholar] [CrossRef]
- Pham, V.T.; Tran, T.T.; Wang, P.C.; Chen, P.Y.; Lo, M.T. EAR-UNet: A deep learning-based approach for segmentation of tympanic membranes from otoscopic images. Artif. Intell. Med. 2021, 115, 102065. [Google Scholar] [CrossRef]
- Livingstone, D.; Chau, J. Otoscopic diagnosis using computer vision: An automated machine learning approach. Laryngoscope 2020, 130, 1408–1413. [Google Scholar] [CrossRef]
- Cha, D.; Pae, C.; Seong, S.B.; Choi, J.Y.; Park, H.J. Automated diagnosis of ear disease using ensemble deep learning with a big otoendoscopy image database. eBiomedicine 2019, 45, 606–614. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Flores, W.; de Albuquerque Pereira, W.C. A comparative study of pre-trained convolutional neural networks for semantic segmentation of breast tumors in ultrasound. Comput. Biol. Med. 2020, 126, 104036. [Google Scholar] [CrossRef]
- Oloruntoba, A.I.; Vestergaard, T.; Nguyen, T.D.; Yu, Z.; Sashindranath, M.; Betz-Stablein, B.; Soyer, H.P.; Ge, Z.; Mar, V. Assessing the Generalizability of Deep Learning Models Trained on Standardized and Nonstandardized Images and Their Performance Against Teledermatologists: Retrospective Comparative Study. JMIR Dermatol. 2022, 5, e35150. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, S.; Qureshi, A.N.; Alhussein, M.; Aurangzeb, K.; Anwar, M.S. AD-CAM: Enhancing Interpretability of Convolutional Neural Networks with a Lightweight Framework-From Black Box to Glass Box. IEEE J. Biomed. Health 2023, 28, 514–525. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Yang, E.; Qi, X.; Xie, R.; Bai, Z. High-performance medicine: The convergence of human and artificial intelligence. J. Alloys Compd. 2019. [Google Scholar]
- Zhang, H.; Li, H.; Ali, R.; Jia, W.; Pan, W.; Reeder, S.B.; Harris, D.; Masch, W.; Aslam, A.; Shanbhogue, K.; et al. Development and Validation of a Modality-Invariant 3D Swin U-Net Transformer for Liver and Spleen Segmentation on Multi-Site Clinical Bi-parametric MR Images. J. Imaging Inform. Med. 2025, 38, 2688–2699. [Google Scholar] [CrossRef]
- Li, M.; Ma, J.; Zhao, J. LKDA-Net: Hierarchical transformer with large Kernel depthwise convolution attention for 3D medical image segmentation. PLoS ONE 2025, 20, e0329806. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. In Proceedings of the International Conference on Learning Representations, Virtual, 3–7 May 2021. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Paris, France, 2–6 October 2023. [Google Scholar]
- Papanastasiou, G.; Dikaios, N.; Huang, J.; Wang, C.; Yang, G. Is Attention all You Need in Medical Image Analysis? A Review. IEEE J. Biomed. Health 2024, 28, 1398–1411. [Google Scholar]
- Pang, Y.; Liang, J.; Huang, T.; Chen, H.; Li, Y.; Li, D.; Huang, L.; Wang, Q. Slim UNETR: Scale Hybrid Transformers to Efficient 3D Medical Image Segmentation Under Limited Computational Resources. IEEE Trans. Med. Imaging 2024, 43, 994–1005. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Kwon, S.; Choo, J.; Hong, S.M.; Kang, S.H.; Park, I.H.; Kim, S.K.; Hong, S.J. Automatic detection of tympanic membrane and middle ear infection from oto-endoscopic images via convolutional neural networks. Neural Netw. 2020, 126, 384–394. [Google Scholar] [CrossRef] [PubMed]
- Xie, S.; Tu, Z. Holistically-Nested Edge Detection. Int. J. Comput. Vision 2015, 125, 3–18. [Google Scholar] [CrossRef]
- Banerji, A. An introduction to image analysis using mathematical morphology. IEEE Eng. Med. Biol. Mag. Q. Mag. Eng. Med. Biol. Soc. 2000, 19, 13–14. [Google Scholar]
- Shen, L.; Lin, Z.; Huang, Q. Relay Backpropagation for Effective Learning of Deep Convolutional Neural Networks. In Computer Vision–ECCV 2016; Lecture Notes in Computer Science; Springer: Cham, Switzerland, 2016; Volume 9911, pp. 467–482. [Google Scholar]
- Mahajan, D.; Girshick, R.; Ramanathan, V.; He, K.; Maaten, L.V.D. Exploring the Limits of Weakly Supervised Pretraining. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; Springer: Cham, Switzerland, 2018; pp. 433–451. [Google Scholar]
- Lin, T.Y.; Goyal, P.; Girshick, R.; He, K.; Dollár, P. Focal Loss for Dense Object Detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2999–3007. [Google Scholar]
- Cui, Y.; Jia, M.; Lin, T.Y.; Song, Y.; Belongie, S. Class-balanced loss based on effective number of samples. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 16–20 June 2019; pp. 9268–9277. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 26 June–1 July 2016; pp. 2818–2826. [Google Scholar]
- Tan, M.; Le, Q.V. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. In Proceedings of the International Conference on Machine Learning (ICML), Long Beach, CA, USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Touvron, H.; Cord, M.; Douze, M.; Massa, F.; Jégou, H. Training Data-Efficient Image Transformers & Distillation through Attention. In Proceedings of the International Conference on Machine Learning, Virtual, 18–24 July 2021; pp. 10347–10357. [Google Scholar]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Sánchez, C.I. A Survey on Deep Learning in Medical Image Analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Selvaraju, R.R.; Cogswell, M.; Das, A.; Vedantam, R.; Parikh, D.; Batra, D. Visual Explanations from Deep Networks via Gradient-Based Localization. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 618–626. [Google Scholar]
- Kumar, R.R.; Shankar, S.V.; Jaiswal, R.; Ray, M.; Budhlakoti, N.; Singh, K.N. Advances in Deep Learning for Medical Image Analysis: A Comprehensive Investigation. J. Stat. Theory Pract. 2025, 19, 23. [Google Scholar] [CrossRef]
- Choi, Y.; Chae, J.; Park, K.; Hur, J.; Kweon, J.; Ahn, J.H. Automated multi-class classification for prediction of tympanic membrane changes with deep learning models. PLoS ONE 2022, 17, e0275846. [Google Scholar] [CrossRef]
- Sandström, J.; Myburgh, H.; Laurent, C.; Swanepoel, W.; Lundberg, T. A Machine Learning Approach to Screen for Otitis Media Using Digital Otoscope Images Labelled by an Expert Panel. Diagnostics 2022, 12, 1318. [Google Scholar] [CrossRef]
- Canares, T.L.; Wang, W.; Unberath, M.; Clark, J.H. Artificial intelligence to diagnose ear disease using otoscopic image analysis: A review. J. Investig. Med. 2022, 70, 354–362. [Google Scholar] [CrossRef]
- Koyama, H.; Kashio, A.; Yamasoba, T. Application of Artificial Intelligence in Otology: Past, Present, and Future. J. Clin. Med. 2024, 13, 7577. [Google Scholar] [CrossRef]
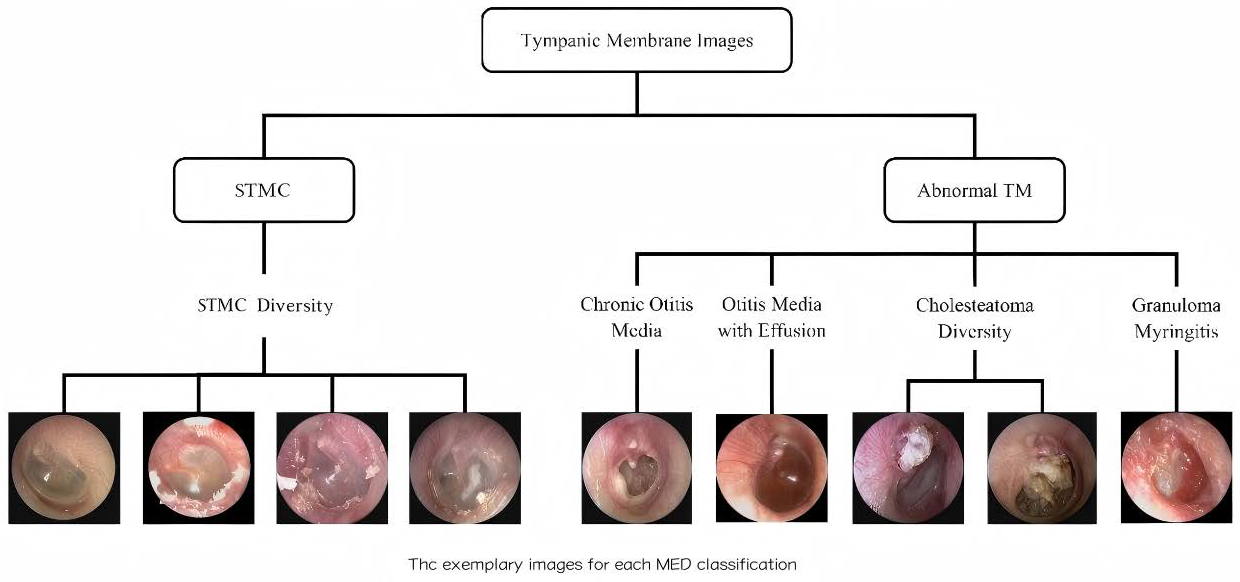
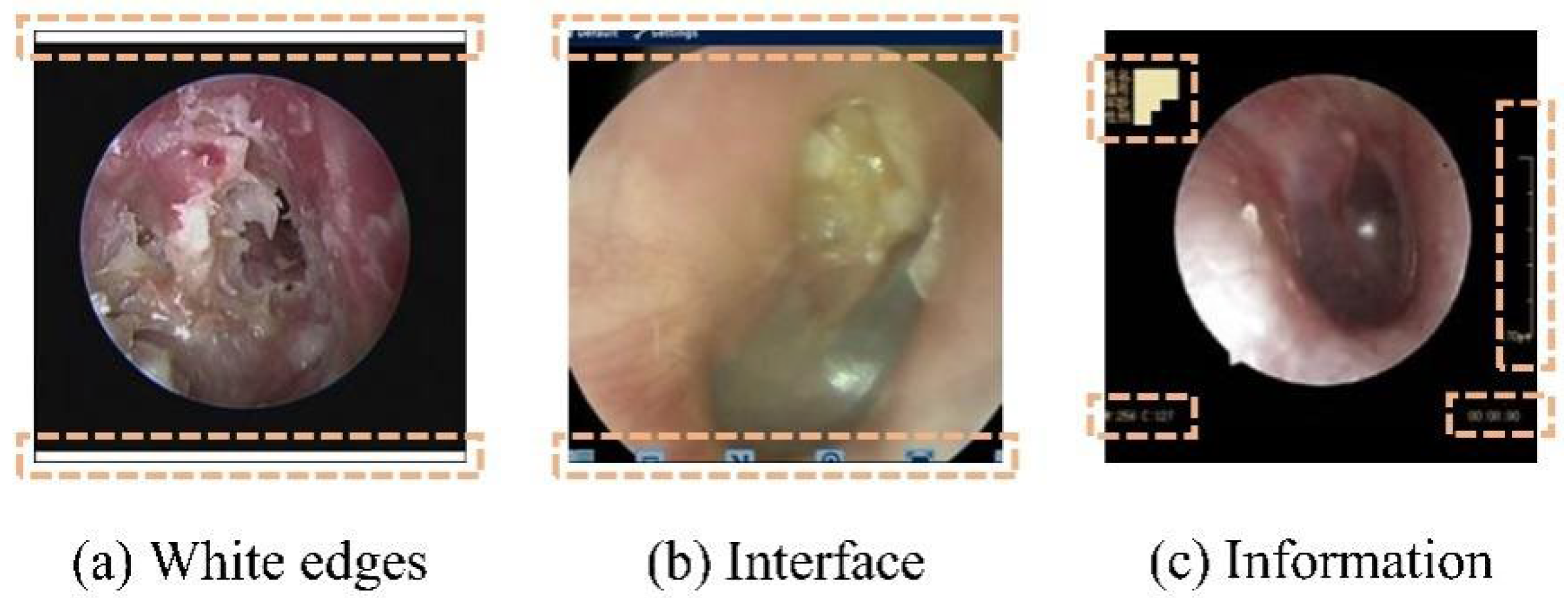




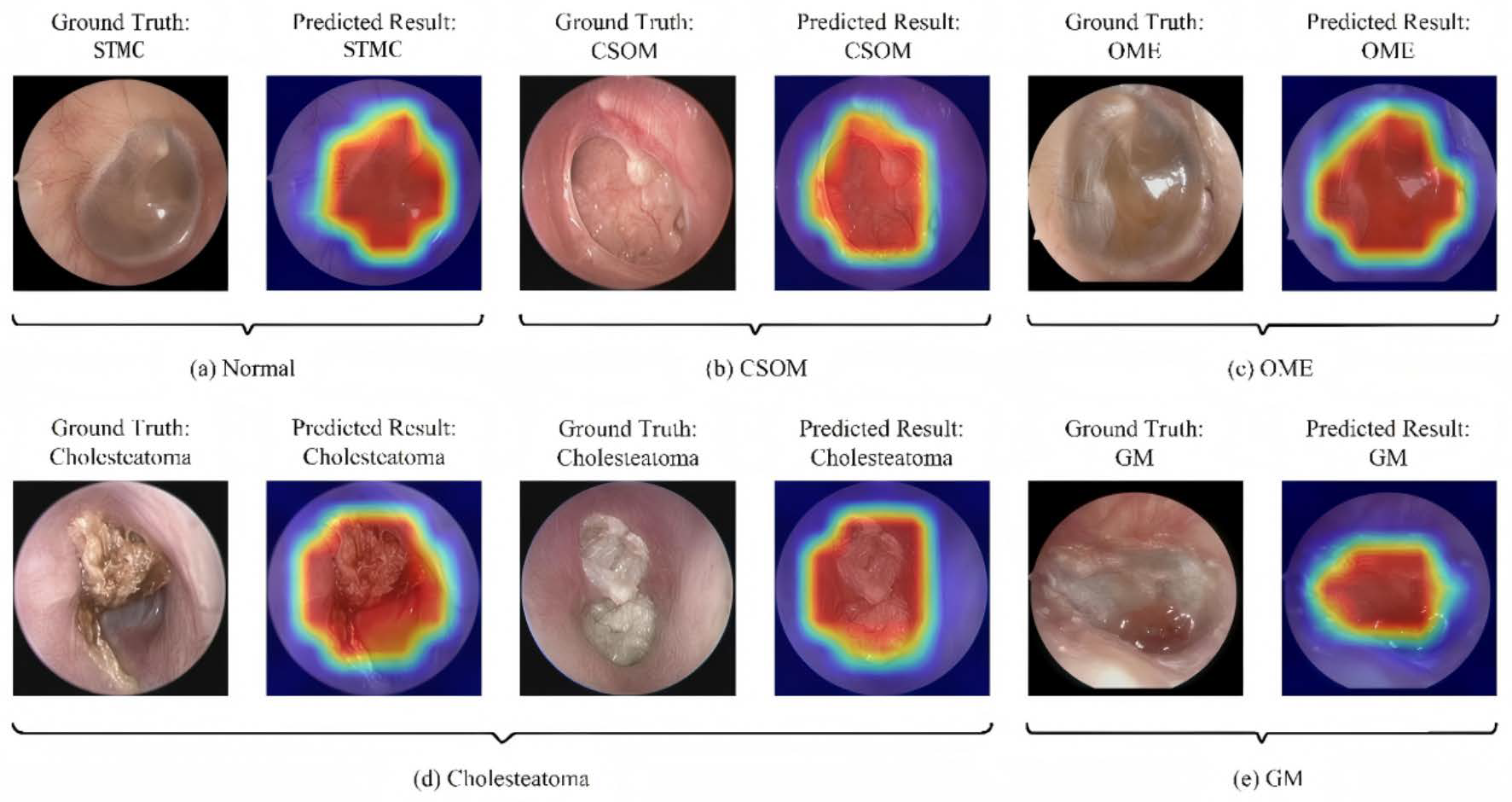

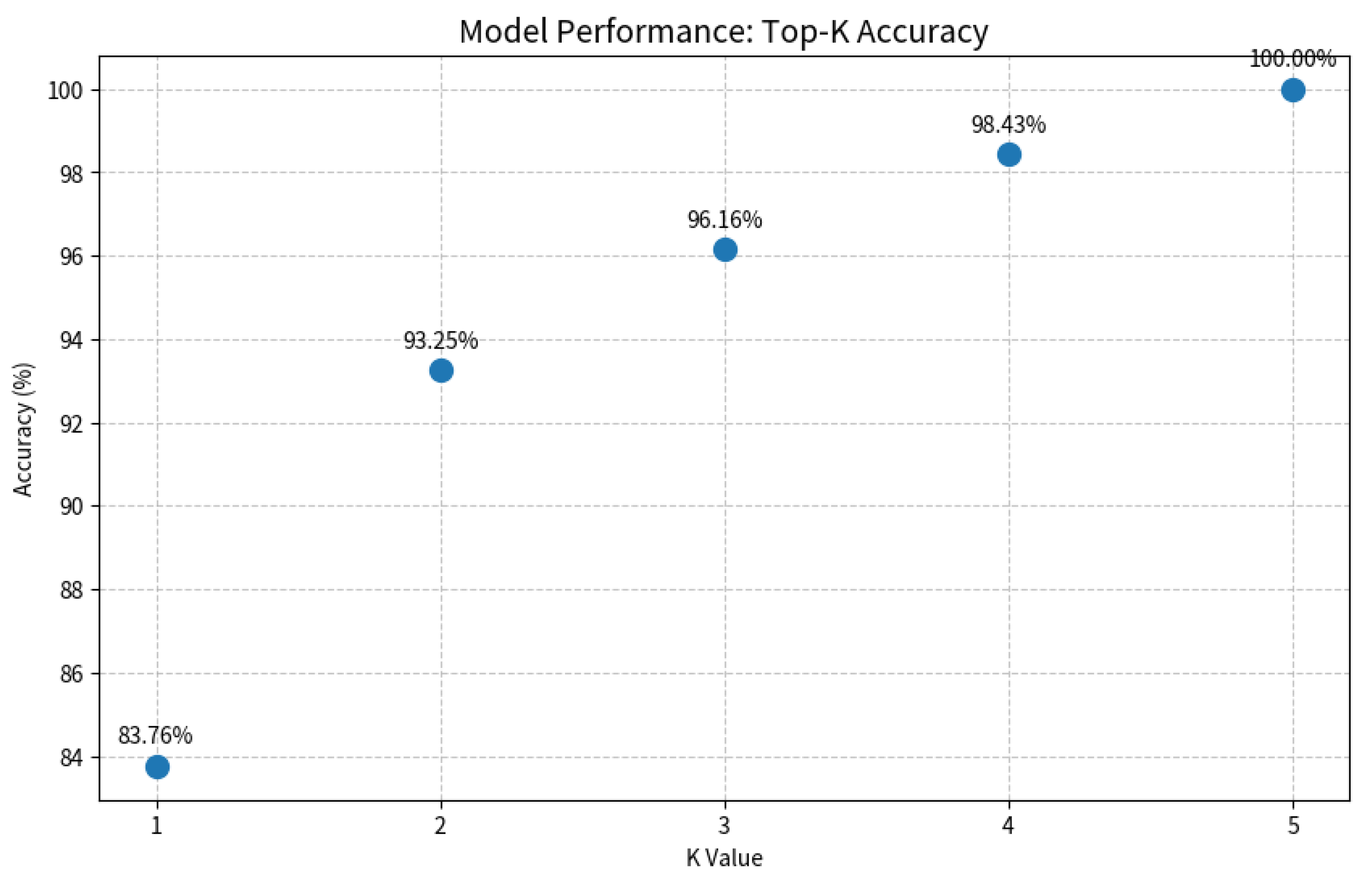
| Disease Category | Sample Size | Percentage (%) | 95% CI | Annotation Consistency (Kappa) |
|---|---|---|---|---|
| STMC | 786 | 12.6 | [11.8–13.4] | 0.89 |
| CSOM | 2488 | 39.9 | [38.7–41.1] | 0.92 |
| OME | 2276 | 36.5 | [35.3–37.7] | 0.91 |
| Cholesteatoma | 472 | 7.6 | [6.9–8.3] | 0.87 |
| GM | 339 | 5.4 | [4.8–6.0] | 0.85 |
| Model | Accuracy (Mean ± SD) | 95% CI | Macro-F1 (Mean ± SD) | 95% CI | T-Value (Acc) | T-Value (Macro-F1) F1 | p-Value (Acc) | p-Value (Macro-F1) | Accuracy Cohen’s d | Macro-F1 Cohen’s d |
|---|---|---|---|---|---|---|---|---|---|---|
| Swin-B | 0.8745 ± 0.0107 | [0.868–0.881] | 0.8391 ± 0.0107 | [0.832–0.846] | Reference | Reference | Reference | Reference | - | - |
| VGG16 | 0.8549 ± 0.0125 | [0.847–0.863] | 0.7923 ± 0.0142 | [0.783–0.802] | −3.89 | −7.82 | p < 0.001 | p < 0.001 | −1.30 | −1.42 |
| ResNet50 | 0.8698 ± 0.0089 | [0.864–0.876] | 0.8311 ± 0.0098 | [0.825–0.837] | −1.08 | −1.82 | p = 0.092 | p = 0.085 | −0.34 | 0.58 |
| Inception-v3 | 0.8596 ± 0.0112 | [0.852–0.867] | 0.8233 ± 0.0105 | [0.817–0.830] | −3.15 | −3.32 | p < 0.001 | p = 0.103 | −0.39 | −0.47 |
| EfficientNet-B4 | 0.7435 ± 0.0187 | [0.732–0.755] | 0.6968 ± 0.0163 | [0.686–0.708] | −19.23 | −20.87 | p < 0.001 | p < 0.001 | −1.58 | −1.64 |
| ViT-S/16 | 0.8220 ± 0.0134 | [0.814–0.830] | 0.7831 ± 0.0128 | [0.775–0.791] | −9.87 | −10.52 | p < 0.001 | p < 0.001 | −1.11 | −1.2 |
| ViT-B/16 | 0.8047 ± 0.0156 | [0.795–0.814] | 0.7592 ± 0.0141 | [0.750–0.768] | −11.76 | −13.45 | p < 0.001 | p < 0.001 | −1.26 | −1.33 |
| DeiT-B | 0.8071 ± 0.0148 | [0.798–0.816] | 0.7711 ± 0.0135 | [0.763–0.779] | −11.34 | −11.45 | p < 0.001 | p < 0.001 | −1.33 | −1.39 |
| Model | Accuracy (Mean ± SD) | 95% CI | Macro-F1 (Mean ± SD) | 95% CI | T-Value (Acc) | T-Value (Macro-F1) | p-Value (Acc) | p-Value (Macro-F) | Selection Status | Accuracy Cohen’s d | Macro-F1 Cohen’s d |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Swin-B | 0.9553 ± 0.0085 | [0.9470–0.9640] | 0.9337 ± 0.0078 | [0.9290–0.9380] | - | - | Reference | Reference | Advanced | - | - |
| Inception-v3 | 0.9506 ± 0.0081 | [0.9460–0.9550] | 0.9351 ± 0.0072 | [0.9310–0.9390] | −1.23 | +0.45 | p = 0.25 | p = 0.103 | Advanced | −0.389 | +0.45 |
| VGG16 | 0.9418 ± 0.0092 | [0.9440–0.9560] | 0.9208 ± 0.0085 | [0.9230–0.9330] | −3.3 | −4.15 | p < 0.001 | p < 0.001 | Excluded | −0.436 | −1.44 |
| ResNet50 | 0.9404 ± 0.0076 | [0.9360–0.9450] | 0.9185 ± 0.0069 | [0.9140–0.9230] | −4.12 | −4.21 | p < 0.001 | p < 0.001 | Excluded | −1.303 | −4.21 |
| EfficientNet-B4 | 0.9302 ± 0.0103 | [0.9240–0.9360] | 0.9017 ± 0.0091 | [0.8950–0.9070] | −6.12 | −0.45 | p < 0.001 | p < 0.001 | Excluded | −1.935 | −8.45 |
| ViT-s/16 | 0.9383 ± 0.0088 | [0.9330–0.9440] | 0.9181 ± 0.0075 | [0.9140–0.9220] | −4.33 | −4.48 | p < 0.001 | p < 0.001 | Excluded | −1.369 | −4.48 |
| ViT-B/16 | 0.9427 ± 0.0082 | [0.9380–0.9470] | 0.9150 ± 0.0067 | [0.9110–0.9190] | −3.28 | −5.12 | p < 0.001 | p < 0.001 | Excluded | 1.037 | −5.12 |
| DeiT-B | 0.9349 ± 0.0095 | [0.9290–0.9410] | 0.9142 ± 0.0073 | [0.9100–0.9180] | −5.01 | −5.23 | p < 0.001 | p < 0.001 | Excluded | 1.584 | −5.43 |
| Metric | Swin-B | Inception-v3 | Performance Gap | Effect Size (Cohen’s d) |
|---|---|---|---|---|
| Accuracy | 95.61 ± 0.38% | 95.18 ± 0.49% | +0.43% | 0.62 |
| Macro-F1 | 93.45 ± 0.34% | 92.91 ± 0.42% | +0.54% | 0.58 |
| Fold Variation | 0.40% | 0.51% | −21.6% | - |
| Disease Category | Swin-B Accuracy (Mean ± SD%) | (Mean ± SD, [95% CI]) | Swin-B F1-Score (Mean ± SD%) | (Mean ± SD, [95% CI]) | Inception-v3 Accuracy (Mean ± SD%) | (Mean ± SD, [95% CI]) | Inception-v3 F1-Score (Mean ± SD%) | (Mean ± SD, [95% CI]) | p-Value (Accuracy) | p-Value (F1-Score) | Effect Size (Acc, Cohen’s d) | Effect Size (F1, Cohen’s d) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| STMC | 94.5238 ± 0.8123 | [93.52, 95.53] | 94.1176 ± 0.7342 | [93.21, 95.03] | 94.0476 ± 1.0235 | [92.78, 95.32] | 93.7500 ± 1.0237 | [92.48, 95.02] | 0.0970 | 0.1100 | +0.15 | +0.12 |
| CSOM | 96.9231 ± 0.6238 | [96.15, 97.70] | 96.8254 ± 0.6238 | [96.05, 97.60] | 96.6420 ± 0.8123 | [95.63, 97.65] | 96.4912 ± 0.8123 | [95.48, 97.50] | 0.0380 | 0.0420 | +0.40 | +0.45 |
| OME | 95.3488 ± 0.5231 | [94.70, 96.00] | 95.2381 ± 0.5231 | [94.59, 95.89] | 95.0000 ± 0.9342 | [93.84, 96.16] | 94.9367 ± 0.9342 | [93.78, 96.10] | 0.0890 | 0.0950 | +0.44 | +0.40 |
| Cholesteatoma | 92.5373 ± 1.2345 | [91.00, 94.07] | 92.3077 ± 1.1234 | [90.91, 93.70] | 91.9403 ± 1.8123 | [89.69, 94.19] | 91.8367 ± 1.7342 | [89.68, 93.99] | 0.0250 | 0.0300 | +0.39 | +0.30 |
| GM | 87.8358 ± 1.5231 | [85.94, 89.73] | 87.6238 ± 1.4238 | [85.86, 89.39] | 87.1343 ± 2.2342 | [84.36, 89.91] | 86.9450 ± 2.1234 | [84.31, 89.58] | 0.0180 | 0.0220 | +0.36 | +0.35 |
| Training–Testing Set | Accuracy (Mean ± SD) | Macro-F1 (Mean ± SD) | Acc. Comparison | Macro-F1 Comparison | Acc. Cohen’s d | Macro-F1 Cohen’s d |
|---|---|---|---|---|---|---|
| Removed–Removed | 0.9553 ± 0.0028 | 0.9337 ± 0.0025 | (Baseline) | (Baseline) | - | - |
| Normal–Normal | 0.9427 ± 0.0035 | 0.9337 ± 0.0026 | t = 4.32, p < 0.01 | t = 0.00, p = 1.000 | 1.366 | 0.000 |
| Removed–Normal | 0.9231 ± 0.0032 | 0.8905 ± 0.0035 | t = 25.67, p < 0.001 | t = 19.84, p < 0.001 | 8.117 | 6.273 |
| Normal–Removed | 0.8659 ± 0.0042 | 0.8206 ± 0.0048 | t = 105.21, p < 0.001 | t = 98.15, p < 0.001 | 33.266 | 31.037 |
| Strategy | GM | STMC | OME | CSOM | Macro-F1 (Mean ± SD) | Effect Size (Cohen’s d) | Significance (vs. Baseline) |
|---|---|---|---|---|---|---|---|
| Baseline | 0.8702 ± 0.0243 | 0.9430 ± 0.0136 | 0.9640 ± 0.0098 | 0.9681 ± 0.0079 | 0.9337 ± 0.0107 | - | - |
| CB-Sampling | 0.9265 ± 0.0186 ** | 0.9216 ± 0.0152 * | 0.9581 ± 0.0109 * | 0.9669 ± 0.0088 | 0.9400 ± 0.0098 * | +0.614 | p < 0.05 |
| Focal Loss | 0.8986 ± 0.0257 | 0.9148 ± 0.0169 ** | 0.9481 ± 0.0118 * | 0.9582 ± 0.0107 * | 0.9203 ± 0.0126 ** | −1.146 | p < 0.01 |
| CB-Loss (Focal) | 0.8872 ± 0.0279 | 0.8970 ± 0.0205 ** | 0.9447 ± 0.0129 ** | 0.9649 ± 0.0098 | 0.9151 ± 0.0138 ** | −1.506 | p < 0.01 |
| LDAM (DRW) | 0.8906 ± 0.0248 | 0.9125 ± 0.0178 ** | 0.9527 ± 0.0108 | 0.9678 ± 0.0081 | 0.9260 ± 0.0117 | −0.687 | p > 0.05 |
| Confidence Grade | Prob. Threshold (Pth) | Sample Count | Correct Predictions | Correct Prediction (%) | Accuracy (%) | Kappa (κ) | Clinical Suggestion |
|---|---|---|---|---|---|---|---|
| Preliminary Screening | 0.4 | 1121 | 1082 | 90.54% | 96.52% | 0.96 | Suitable for preliminary screening |
| Routine Diagnosis | 0.5 | 859 | 846 | 70.79% | 98.49% | 0.98 | Suitable for routine diagnosis |
| High-Confidence Diagnosis | 0.6 | 418 | 413 | 34.56% | 98.80% | 0.98 | Suitable for high-confidence diagnosis |
| Confirmed Diagnosis | 0.7 | 229 | 228 | 19.08% | 99.56% | 0.99 | Suitable for confirmed diagnosis |
| Gold Standard Diagnosis | 0.75 | 172 | 172 | 14.39% | 100.00% | 1 | Suitable as gold standard |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Chen, G.; Zhang, H.; Zeng, J.; Cai, Y.; Huang, D.; Chen, Y.; Li, P.; Zheng, Y. Benchmark-Driven Clinical Decision Framework for Multi-Class Middle Ear Disease Diagnosis: Superiority of Swin Transformer in Accuracy and Stability. Diagnostics 2026, 16, 482. https://doi.org/10.3390/diagnostics16030482
Chen G, Zhang H, Zeng J, Cai Y, Huang D, Chen Y, Li P, Zheng Y. Benchmark-Driven Clinical Decision Framework for Multi-Class Middle Ear Disease Diagnosis: Superiority of Swin Transformer in Accuracy and Stability. Diagnostics. 2026; 16(3):482. https://doi.org/10.3390/diagnostics16030482
Chicago/Turabian StyleChen, Guoping, Haoyi Zhang, Junbo Zeng, Yuexin Cai, Dong Huang, Yubin Chen, Peng Li, and Yiqing Zheng. 2026. "Benchmark-Driven Clinical Decision Framework for Multi-Class Middle Ear Disease Diagnosis: Superiority of Swin Transformer in Accuracy and Stability" Diagnostics 16, no. 3: 482. https://doi.org/10.3390/diagnostics16030482
APA StyleChen, G., Zhang, H., Zeng, J., Cai, Y., Huang, D., Chen, Y., Li, P., & Zheng, Y. (2026). Benchmark-Driven Clinical Decision Framework for Multi-Class Middle Ear Disease Diagnosis: Superiority of Swin Transformer in Accuracy and Stability. Diagnostics, 16(3), 482. https://doi.org/10.3390/diagnostics16030482






