The Diagnostic Performance of the Cellavision DC-1 Digital Morphology Analyser on Leukaemia Samples
Abstract
1. Introduction
2. Materials and Methods
2.1. Peripheral Blood Smears
2.2. Cellavision DC-1 Digital Morphology Analyser
2.3. Statistical Analysis: Specificity, Sensitivity, and Cohen’s Kappa
2.4. Statistical Analysis: Correlation, Accuracy, and Bias
3. Results
3.1. DC-1 Pre-Classification Performance
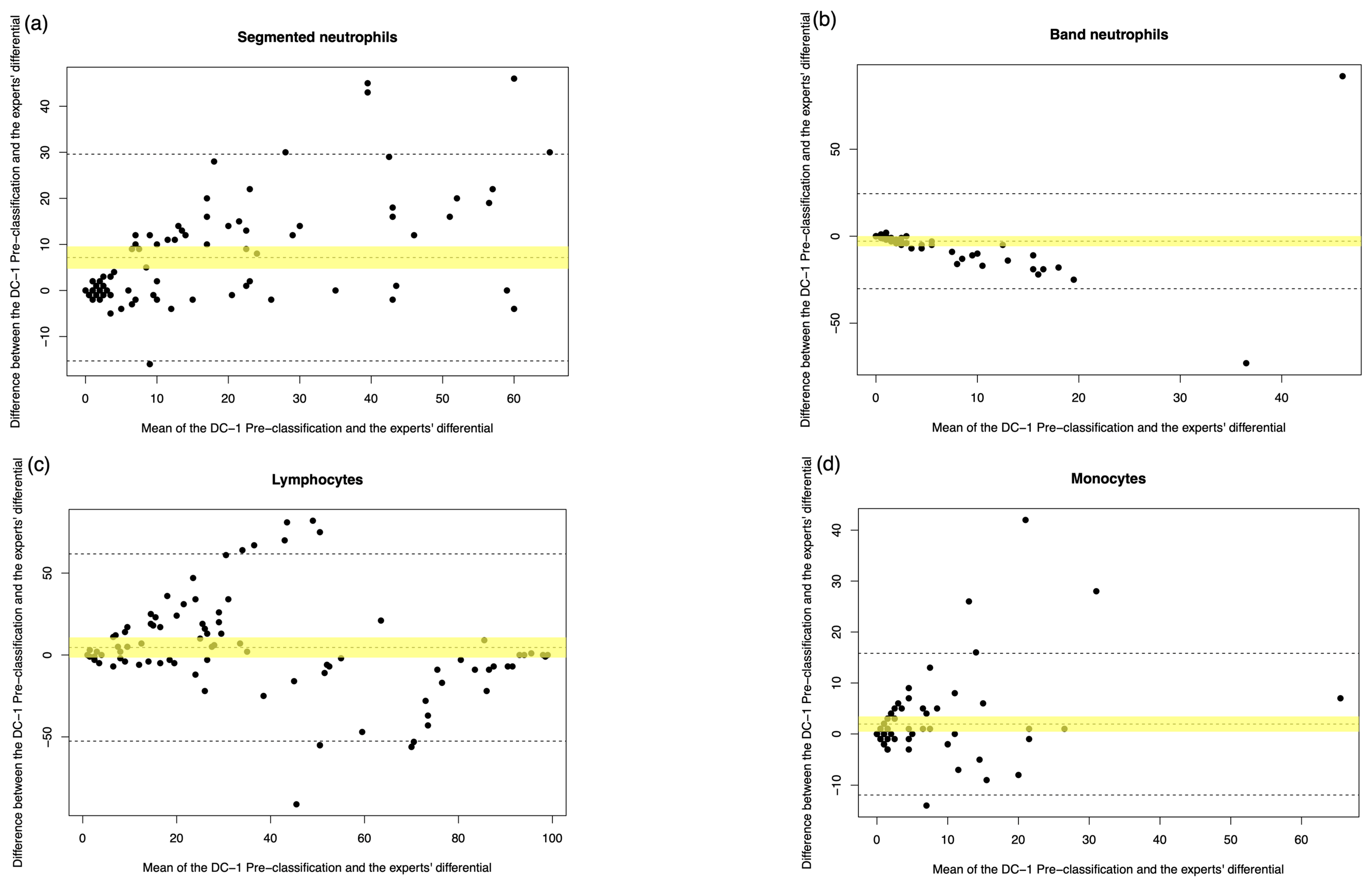
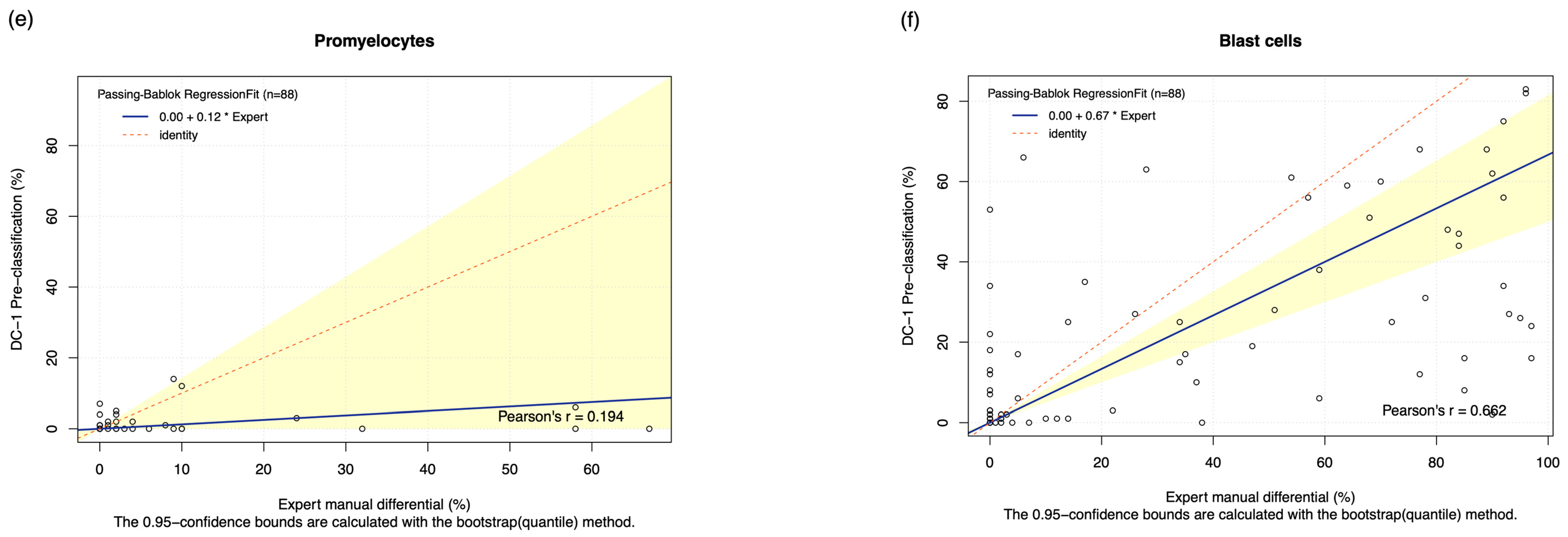
3.2. Comparison of Methods: DC-1 Pre-Classification
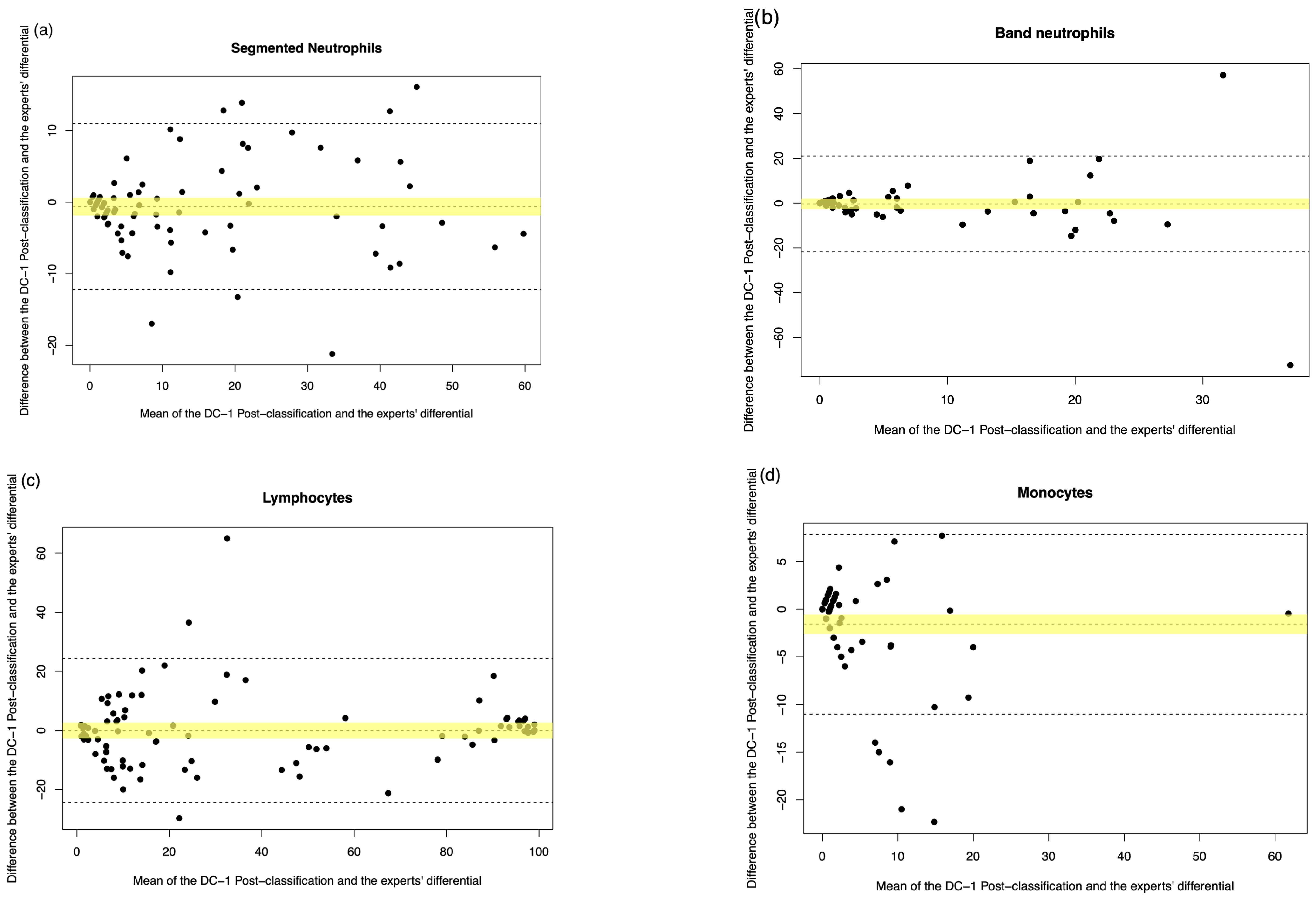
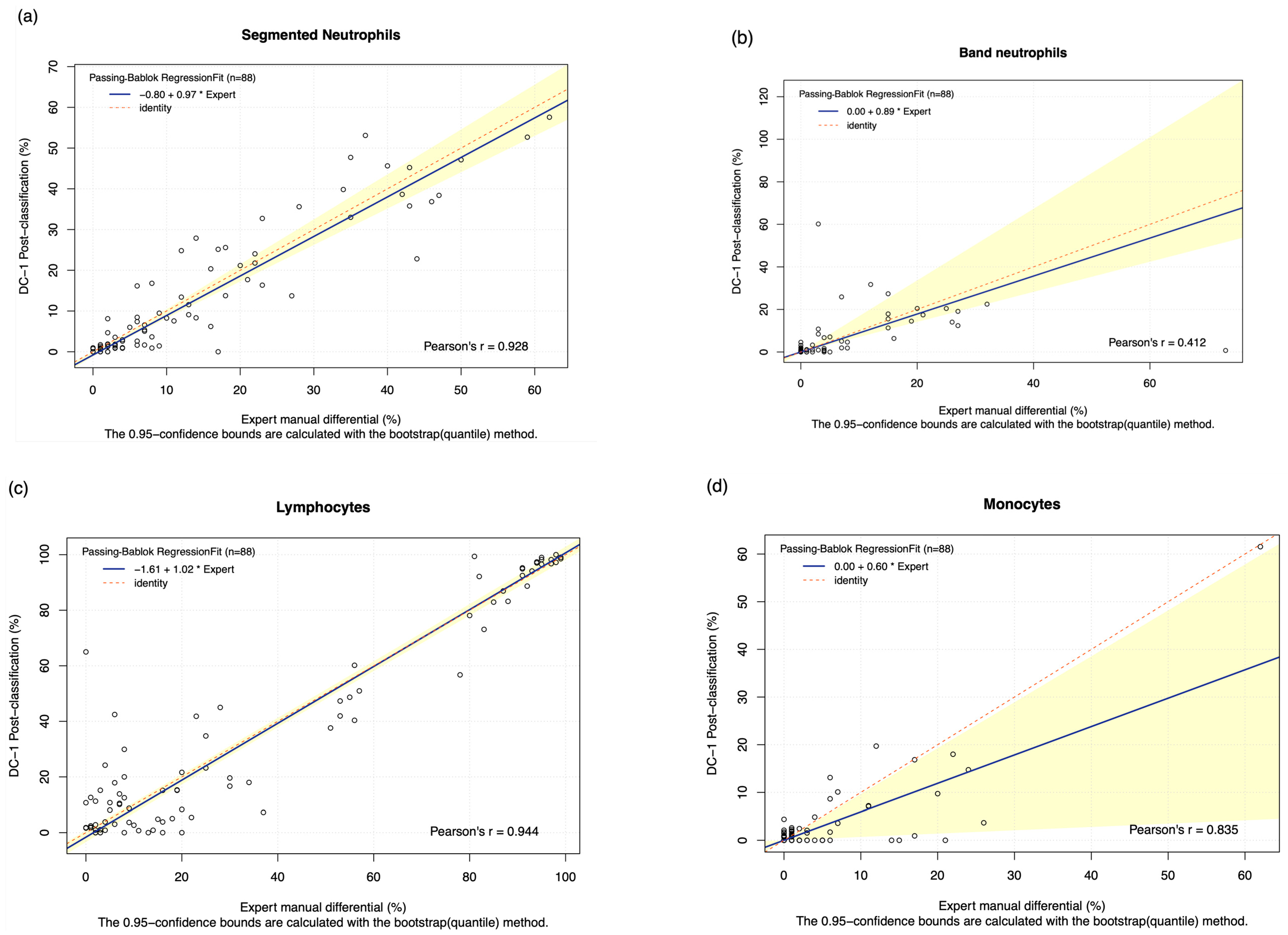
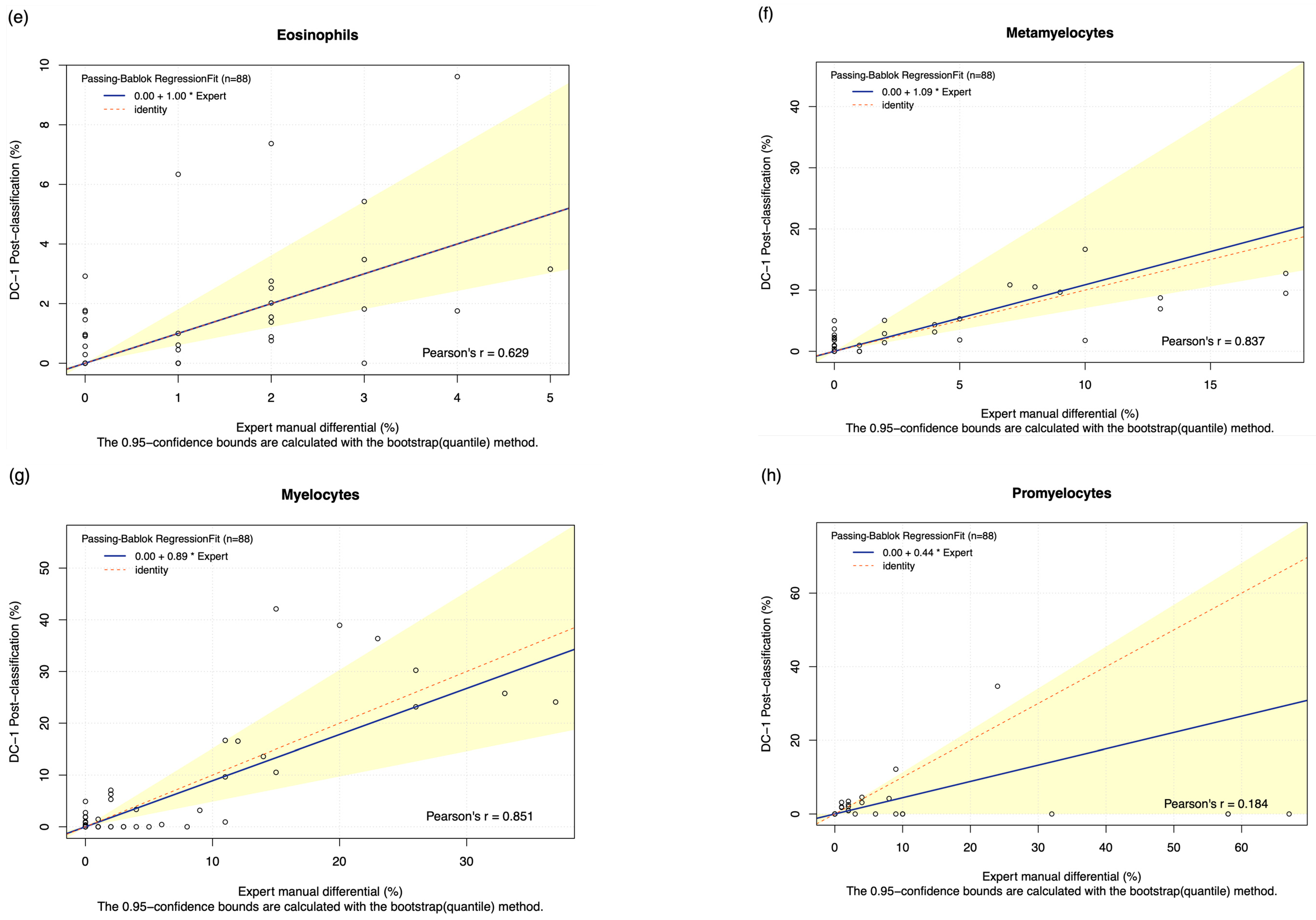
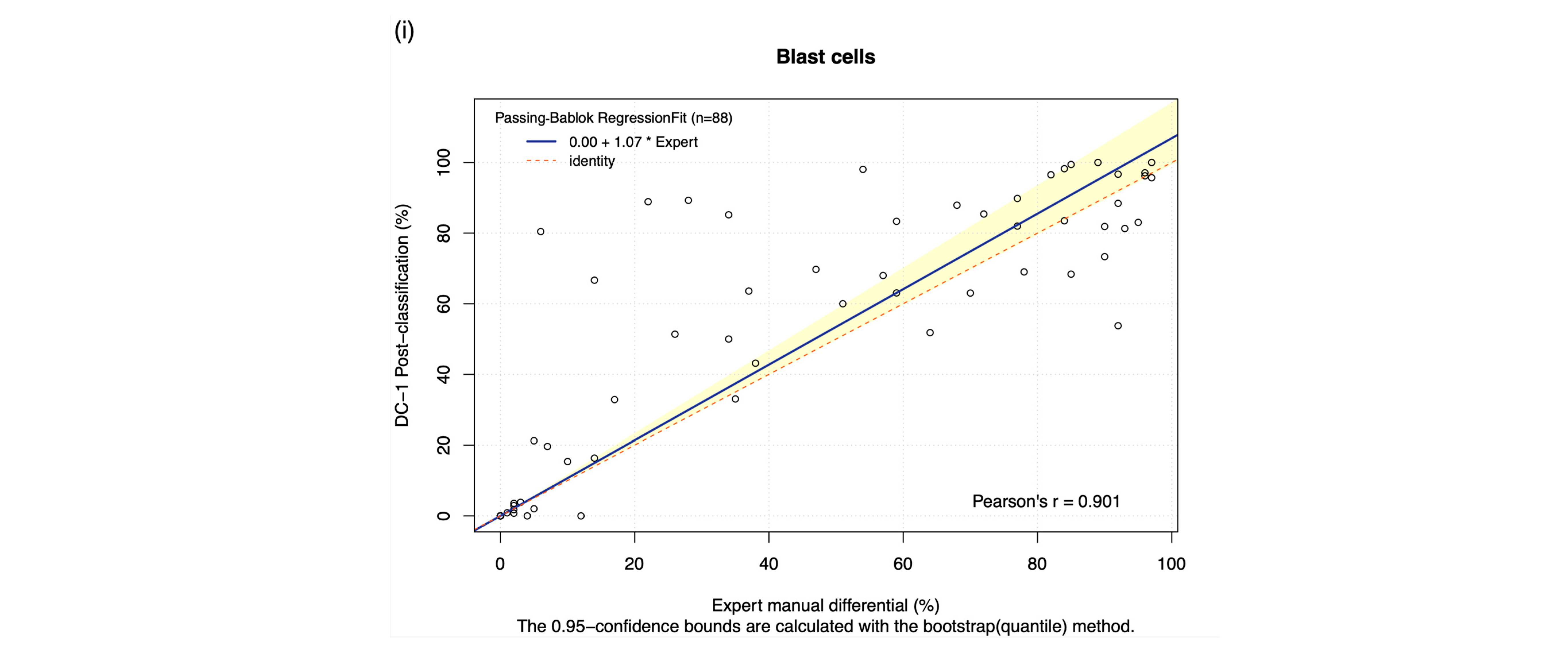
3.3. Comparison of Methods: DC-1 Post-Classification
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CNN | Convolutional Neural Network |
| ANN | Artificial Neural Network |
| RMIT | Royal Melbourne Institute of Technology |
| CML | Chronic Myeloid Leukaemia |
| CLL | Chronic Lymphocytic Leukaemia |
| JMML | Juvenile Myelomonocytic Leukaemia |
| PLL | Prolymphocytic Leukaemia |
| HCL | Hairy Cell Leukaemia |
| CMML | Chronic Myelomonocytic Leukaemia |
| ALL | Acute Lymphoblastic Leukaemia |
| AML | Acute Myeloid Leukaemia |
| APML | Acute Promyelocytic Leukaemia |
| AMML | Acute Myelomonocytic Leukaemia |
| AMoL | Acute Monoblastic/cytic Leukaemia |
Appendix A
| Leukaemia Cases | Sample Size (n = 88) | Proportion (%) in Total |
|---|---|---|
| CML | 15 | 16.3 |
| CLL | 18 | 20.7 |
| JMML | 1 | 1.1 |
| PLL | 6 | 6.5 |
| HCL | 2 | 3.3 |
| CMML | 3 | 5.4 |
| ALL | 18 | 19.6 |
| AML M0 | 1 | 1.1 |
| AML M1 | 4 | 4.3 |
| AML M2 | 3 | 3.3 |
| APML M3 | 5 | 5.4 |
| AMML M4 | 4 | 4.3 |
| AMoL M5 | 2 | 2.2 |
| AML M6 | 2 | 2.2 |
| AML M7 | 2 | 2.2 |
| AML recurrent | 2 | 2.2 |
| Leukocytes | Total Cell Number Taken for Account (n = 8799) | Proportion (%) in Total |
|---|---|---|
| Segmented Neutrophils | 1177 | 13.4 |
| Band neutrophils | 449 | 5.1 |
| Lymphocytes | 3182 | 36.2 |
| Monocytes | 377 | 4.3 |
| Eosinophils | 54 | 0.6 |
| Basophils | 55 | 0.6 |
| Metamyelocytes | 136 | 1.5 |
| Myelocytes | 312 | 3.5 |
| Promyelocytes | 325 | 3.7 |
| Blast cells | 2732 | 31.0 |
Appendix B
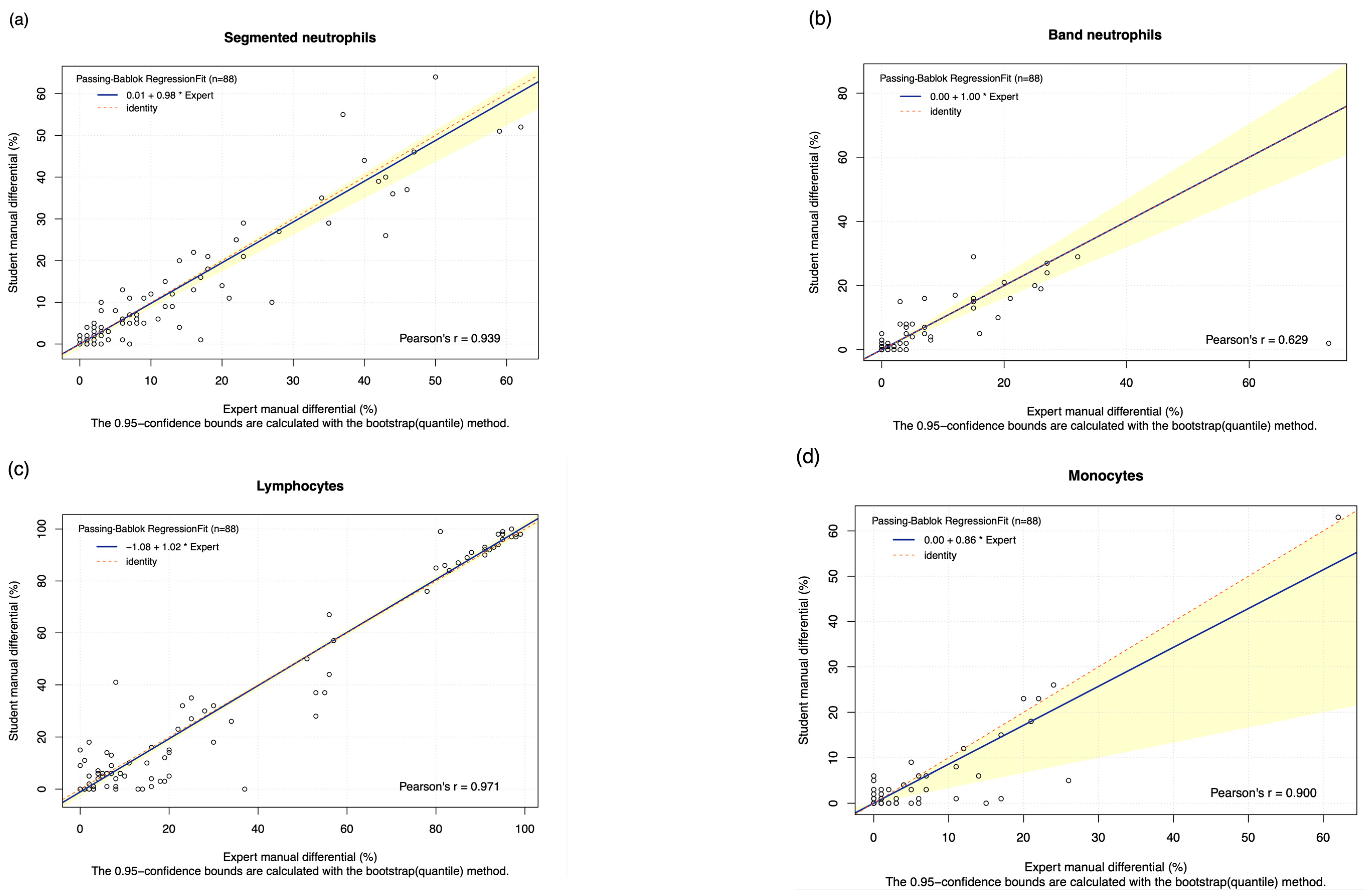
| Leukocytes | Estimated Accuracy | Pearson’s r Coefficient Value (95% CI) | Slope (95% CI) | Intercept (95% CI) | Mean Difference (95% CI) |
|---|---|---|---|---|---|
| Neutrophils | 86.268% | 0.939 (p < 0.001) | 0.975 (0.875 to 1.032) | 0.013 (−1.000 to 0.133) | −0.932 (−2.065 to 0.202) |
| Band neutrophils | 73.165% | 0.629 (p < 0.001) | 1.000 (0.800 to 1.200) | 0.000 (0.000 to 0.000) | −2.515 (−0.761 to 0.992) |
| Lymphocytes | 91.813% | 0.971 (p < 0.001) | 1.021 (1.000 to 1.038) | −1.083 (−2.591 to 0.000) | −1.25 (−3.184 to 0.684) |
| Monocytes | 74.504% | 0.900 (p < 0.001) | 0.857 (0.332 to 1.000) | 0.000 (0.000 to 0.000) | −1.125 (−1.937 to −0.313) |
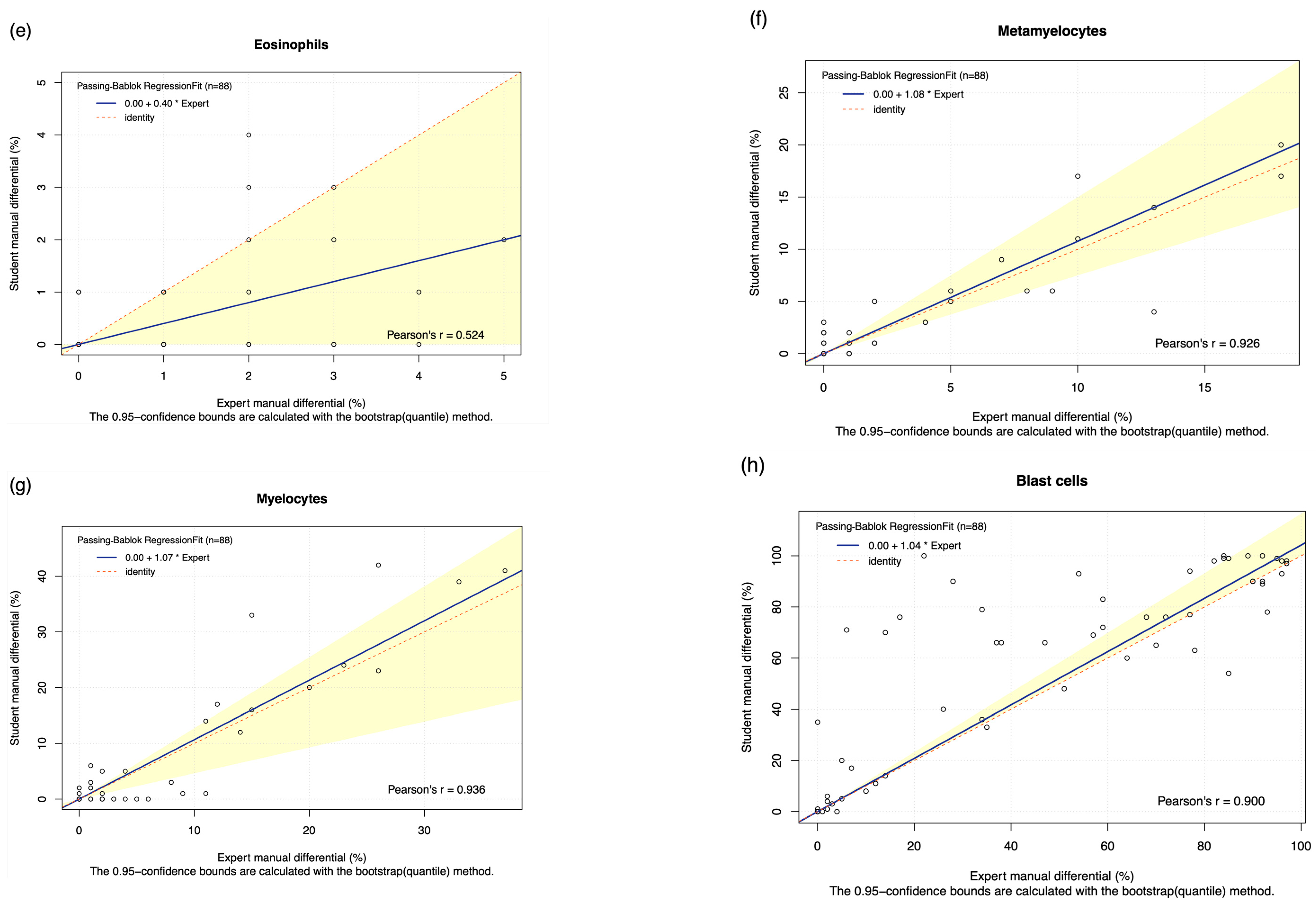
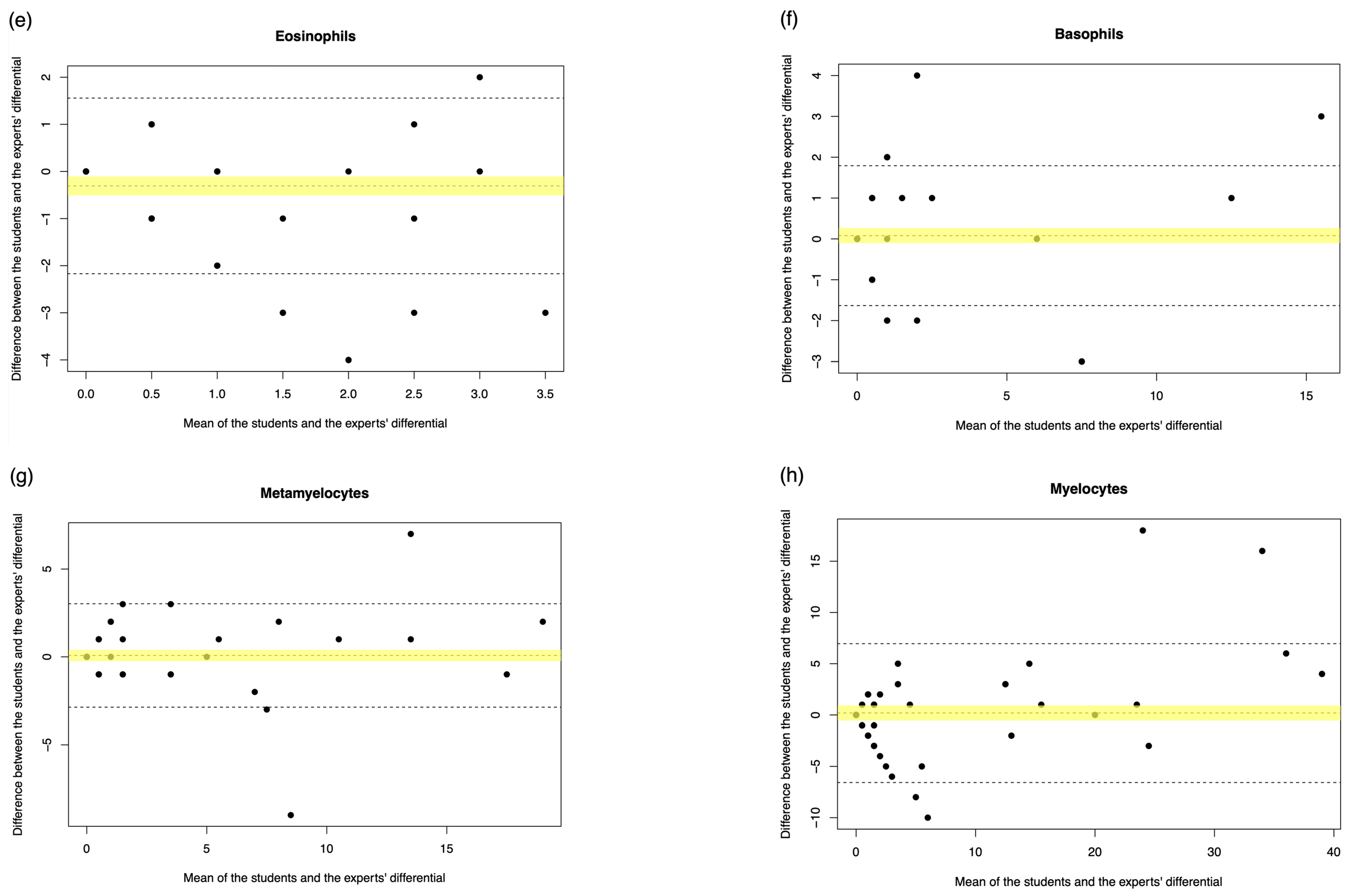
| Eosinophils | 49.383% | 0.524 (p < 0.001) | 0.400 (0.000 to 1.000) | 0.000 (0.000 to 0.000) | −0.307 (−0.508 to −0.105) |
| Basophils | 73.504% | * 0.542 (p < 0.001) | N.A. | N.A. | 0.080 (−0.106 to 0.265) |
| Metamyelocytes | 81.004% | 0.926 (p < 0.001) | 1.077 (0.750 to 1.500) | 0.000 (0.000 to 0.000) | 0.080 (−0.239 to 0.398) |
| Myelocytes | 79.251% | 0.936 (p < 0.001) | 1.067 (0.375 to 1.273) | 0.00 (0.000 to 0.000) | 0.193 (−0.538 to 0.924) |
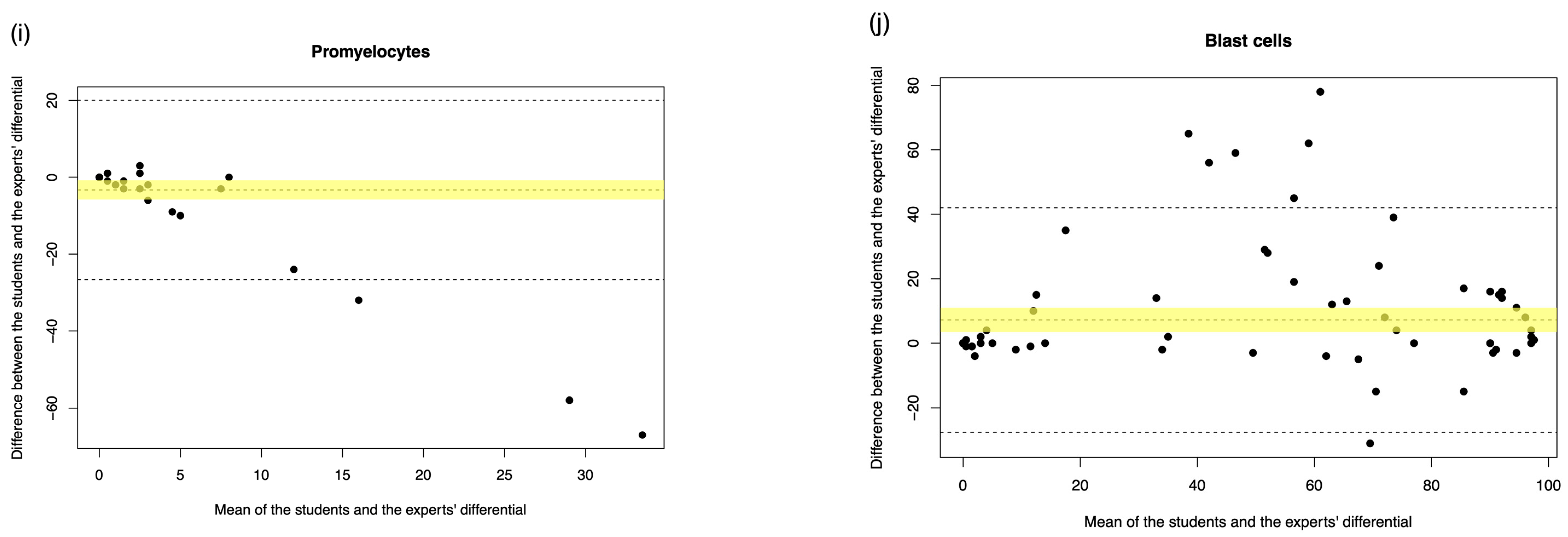
| Promyelocytes | 13.370% | * 0.417 (p < 0.001) | N.A. | N.A. | −3.307 (−5.830 to −0.784) |
| Blast cells | 86.539% | 0.900 (p < 0.001) | 1.042 (1.000 to 1.159) | 0.000 (0.000 to 0.000) | 7.216 (3.456 to 10.976) |

Appendix C




References
- Lee, G.H.; Yoon, S.; Nam, M.; Kim, H.; Hur, M. Performance of digital morphology analyzer CellaVision DC-1. CCLM 2023, 61, 133–141. [Google Scholar] [CrossRef]
- Lapic, I.; Milos, M.; Dorotic, M.; Drenski, V.; Coen Herak, D.; Rogic, D. Analytical validation of white blood cell differential and platelet assessment on the Sysmex DI-60 digital morphology analyzer. Int. J. Lab. Hematol. 2023, 45, 668–677. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Fang, L.; Chen, Y.; Tong, J.; Ning, X.; Feng, L.; Xu, Y.; Yang, D. Performance comparison of two automated digital mor phology analyzers for leukocyte differential in patients with malignant hematological diseases: Mindray MC -80 and Sysmex DI-60. Int. J. Lab. Hematol. 2024, 46, 457–465. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Park, C.J.; Choi, M.O.; Kim, M.J.; Cho, Y.U.; Jang, S.; Chi, H.S. Automated digital cell morphology identification system (CellaVision DM96) is very useful for leukocyte differentials inspecimens with qualitative or quantitative abnormalit ies. Int. J. Lab. Hematol. 2013, 35, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Kufel, J.; Bargiel-Laczek, K.; Kocot, S.; Kozlik, M.; Bartnikowska, W.; Janik, M.; Czogalic, L.; Dudek, P.; Magiera, M.; Lis, A.; et al. What Is Machine Learning, Artificial Neural Networks and Deep Learning?-Examples of Practical Applications in Medicine. Diagnostics 2023, 13, 2582. [Google Scholar] [CrossRef]
- Zhao, Y.; Diao, Y.; Zheng, J.; Li, X.; Luan, H. Performance evaluation of the digital morphology analyser Sysmex DI-60 for white blood cell differentials in abnormal samples. Sci. Rep. 2024, 14, 14344. [Google Scholar] [CrossRef]
- Kim, H.N.; Hur, M.; Kim, H.; Kim, S.W.; Moon, H.W.; Yun, Y.M. Performance of automated digital cell imaging analyzer Sysmex DI-60. CCLM 2017, 56, 94–102. [Google Scholar] [CrossRef]
- Christiansen, M.; Abildgaard, A.; Larsen, J.B.; Tindbæk, G.; Vestergaard, E.M. Diagnostic performance of the CellaVision preclassification neutrophil count-time to bypass the reclassification? Scand. J. Clin. Lab. Investig. 2024, 84, 278–284. [Google Scholar] [CrossRef]
- Kweon, O.J.; Lim, Y.K.; Lee, M.K.; Kim, H.R. Red and white blood cell morphology characterization and hands-on time analysis by the digital cell imaging analyzer DI-60. PLoS ONE 2022, 17, e0267638. [Google Scholar] [CrossRef]
- Tabe, Y.; Yamamoto, T.; Maenou, I.; Nakai, R.; Idei, M.; Horii, T.; Miida, T.; Ohsaka, A. Performance evaluation of the digital cell imaging analyzer DI-60 integrated into the fully automated Sysmex XN hematology analyzer system. Clin. Chem. Lab. Med. 2015, 53, 281–289. [Google Scholar] [CrossRef]
- Zini, G.; Mancini, F.; Rossi, E.; Landucci, S.; D’Onofrio, G. Artificial intelligence and the blood film: Performance of the MC-80 digital morphology analyzer in samples with neoplastic and reactive cell types. Int. J. Lab. Hematol. 2023, 45, 881–889. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.H.; Mansoor, A.; Wood, B.; Nelson, H.; Higa, D.; Naugler, C. Performance of CellaVision DM96 in leukocyte classification. J. Pathol. Inform. 2013, 4, 14. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.; Hur, M.; Lee, G.H.; Nam, M.; Kim, H. How Reproducible Is the Data from Sysmex DI-60 in Leukopenic Samples? Diagnostics 2021, 11, 2173. [Google Scholar] [CrossRef] [PubMed]
- Van Der Vorm, L.N.; Hendriks, H.A.; Smits, S.M. Performance of the CellaVision DC-1 digital cell imaging analyser for differential counting and morphological classification of blood cells. J. Clin. Pathol. 2023, 76, 194–201. [Google Scholar] [CrossRef] [PubMed]
- Mayes, C.; Gwilliam, T.; Mahe, E.R. Improving turn-around times in low-throughput distributed hematology laboratory se tings with the CellaVision® DC-1 instrument. J. Lab. Med. 2025, 49, 71–75. [Google Scholar] [CrossRef]
- Alsalem, M.A.; Zaidan, A.A.; Zaidan, B.B.; Hashim, M.; Madhloom, H.T.; Azeez, N.D.; Alsyisuf, S. A review of the automated detection and classification of acute leukaemia: Coherent taxonomy, datasets, validation and performance measurements, motivation, open challenges and recommendations. Comput. Methods Programs Biomed. 2018, 158, 93–112. [Google Scholar] [CrossRef]
- Xing, Y.; Liu, X.; Dai, J.; Ge, X.; Wang, Q.; Hu, Z.; Wu, Z.; Zeng, X.; Xu, D.; Qu, C. Artificial intelligence of digital morphology analyzers improves the efficiency of manual leukocyte differentiation of peripheral blood. BMC Med. Inform. Decis. Mak. 2023, 23, 50. [Google Scholar] [CrossRef]
- McHugh, M.L. Interrater reliability: The kappa statistic. Biochem. Medica 2012, 22, 276–282. [Google Scholar] [CrossRef]
- IBM Corp. IBM SPSS Statistics 29, Statistics for Windows; IBM Corp: Armonk, NY, USA, 2023.
- Miot, H.A. Correlation analysis in clinical and experimental studies. J. Vasc. Bras. 2018, 17, 275–279. [Google Scholar] [CrossRef]
- PBC. RStudio, Version 4.4.2. Integrated development environment for R. PBC: Boston, MA, USA, 2024.
- NATA. Available online: https://nata.com.au/ (accessed on 27 July 2025).
- Zhang, S.; He, Y.; Wu, W.; Tan, H.; Xie, S.; Liu, M.; Chen, W.; Sun, D. Comparison of the performance of two automatic cell morphology analyzers for peripheral-blood leukocyte morphology analysis: Mindray MC-100i and Sysmex DI-60. Int. J. Lab. Hematol. 2023, 45, 860–868. [Google Scholar] [CrossRef]
- Rollins-Raval, M.A.; Raval, J.S.; Contis, L. Experience with CellaVision DM96 for peripheral blood differentials in a large multicenter academic hospital system. J. Pathol. Inform. 2012, 3, 29. [Google Scholar] [CrossRef] [PubMed]
- George, T.I.; Bajel, A. Diagnosis of rare subtypes of acute myeloid leukaemia and related neoplasms. Pathology 2021, 53, 312–327. [Google Scholar] [CrossRef]
- Kratz, A.; Lee, S.H.; Zini, G.; Riedl, J.A.; Hur, M.; Machin, S. Digital morphology analyzers in hematology: ICSH review and recommendations. Int. J. Lab. Hematol. 2019, 41, 437–447. [Google Scholar] [CrossRef]
- Merino, A.; Laguna, J.; Rodríguez-García, M.; Julian, J.; Casanova, A.; Molina, A. Performance of the new MC-80 automated digital cell morphology analyser in detection of normal and abnormal blood cells: Comparison with the CellaVision DM9600. Int. J. Lab. Hematol. 2024, 46, 72–82. [Google Scholar] [CrossRef]
- Eilertsen, H.; Saether, P.C.; Henriksson, C.E.; Petersen, A.S.; Hagve, T.A. Evaluation of the detection of blasts by Sysmex hematology instruments, CellaVision DM96, and manual microscopy using flow cytometry as the confirmatory method. Int. J. Lab. Hematol. 2019, 41, 338–344. [Google Scholar] [CrossRef]
- Vergara-Lluri, M.; Kovach, A.E.; Nakashima, M.O.; Bradley, K.T.; Mahe, E.; Tsao, L.; Savage, N.M.; Salansky, S.A.; Long, T.; Perkins, S.L.; et al. Significant Variability in the Identification and Reporting of Band Neutrophils by Participants Enrolled in the College of American Pathologists Proficiency Testing Program: Time for a Change. Arch. Pathol. Lab. Med. 2024, 148, 666–676. [Google Scholar] [CrossRef] [PubMed]
- Biljak, V.R.; Juresa, V.; Vidranski, V.; Vuga, I.; Tomic, F.; Smaic, F.; Horvat, M.; Kresic, B.; Simac, B.; Lapic, I. The band count imprecision—A Croatian multicentric pilot study. Biochem. Medica 2024, 34, 335–339. [Google Scholar]
- Van der Meer, W.; Van Gelder, W.; de Keijzer, R.; Willems, H. Does the band cell survive the 21st century? Eur. J. Haematol. 2006, 76, 251–254. [Google Scholar] [CrossRef]
- Cellavision. Available online: https://www.cellavision.com (accessed on 27 July 2025).

| Pre-Classification | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Post- Classification | Segmented Neutrophils | Band Neutrophils | Lymphocytes | Monocytes | Eosinophils | Basophils | Metamyelocytes | Myelocytes | Promyelocytes | Blasts | Post- Classification Total |
| Segmented Neutrophils | 997 | 4 | 3 | 2 | 3 | 5 | 0 | 1 | 0 | 0 | 1015 |
| Band neutrophils | 209 | 70 | 5 | 0 | 6 | 0 | 0 | 1 | 0 | 0 | 291 |
| Lymphocytes | 46 | 0 | 2101 | 43 | 1 | 8 | 4 | 44 | 0 | 180 | 2427 |
| Monocytes | 8 | 0 | 3 | 189 | 0 | 0 | 0 | 0 | 0 | 0 | 200 |
| Eosinophils | 2 | 1 | 0 | 0 | 55 | 2 | 0 | 2 | 0 | 0 | 62 |
| Basophils | 0 | 0 | 10 | 0 | 0 | 42 | 0 | 2 | 1 | 1 | 56 |
| Metamyelocytes | 45 | 6 | 16 | 5 | 1 | 0 | 50 | 2 | 0 | 1 | 126 |
| Myelocytes | 17 | 0 | 137 | 7 | 3 | 2 | 5 | 79 | 0 | 1 | 251 |
| Promyelocytes | 22 | 0 | 4 | 6 | 2 | 12 | 0 | 24 | 15 | 33 | 98 |
| Blasts | 53 | 0 | 1142 | 279 | 81 | 88 | 25 | 103 | 54 | 1375 | 3200 |
| Pre- classification Total | 1379 | 81 | 3421 | 531 | 152 | 159 | 84 | 258 | 70 | 1591 | 7726 |
| True Positive | 997 | 70 | 2101 | 189 | 55 | 42 | 50 | 79 | 15 | 1375 | - |
| False Negative | 382 | 11 | 1320 | 342 | 97 | 117 | 34 | 179 | 55 | 216 | - |
| True Negative | 6711 | 7435 | 5299 | 7526 | 7664 | 7670 | 7600 | 7475 | 7628 | 4526 | - |
| False Positive | 18 | 221 | 326 | 11 | 7 | 14 | 76 | 172 | 83 | 1825 | - |
| Specificity (%) | 99.7 | 97.1 | 94.2 | 99.9 | 99.9 | 99.8 | 99 | 97.8 | 98.9 | 71.3 | - |
| Sensitivity (%) | 72.3 | 86.4 | 61.4 | 35.6 | 36.2 | 26.4 | 59.5 | 30.6 | 21.4 | 86.4 | - |
| Leukocytes | Pre-Classification | Post-Classification | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Estimated Accuracy | r (95% CI) | Slope (95% CI) | Intercept (95% CI) | Mean Difference (95% CI) | Estimated Accuracy | r (95% CI) | Slope (95% CI) | Intercept (95% CI) | Mean Difference (95% CI) | |
| Segmented neutrophils | 74.757% | 0.867 (p < 0.001) | 1.511 (1.353 to 1.868) | −0.511 (−1.568 to 0.711) | 7.148 (4.721 to 9.574) | 84.823% | 0.928 (p < 0.001) | 0.970 (0.897 to 1.117) | −0.799 (−1.147 to −0.037) | −0.613 (−1.865 to 0.640) |
| Band neutrophils | 30.341% | 0.083 (p = 0.444) | 0.250 (0.167 to 0.333) | 0.000 (0.000 to 0.000) | −2.864 (−5.815 to 0.088) | 60.697% | 0.412 (p < 0.001) | 0.892 (0.707 to 1.683) | 0.000 (0.000 to 0.000) | 0.355 (−2.669 to 1.958) |
| Lymphocytes | 74.786% | 0.635 (p < 0.001) | 0.909 (0.800 to 1.000) | 4.773 (0.000 to 10.600) | 4.568 (−1.610 to 10.747) | 88.786% | 0.944 (p < 0.001) | 1.023 (0.990 to 1.054) | −1.614 (−3.460 to 0.808) | −0.046 (−2.683 to 2.591) |
| Monocytes | 65.297% | 0.762 (p < 0.001) | 1.417 (1.000 to 2.591) | 0.000 (−0.265 to 0.400) | 1.943 (0.443 to 3.443) | 63.10% | 0.835 (p < 0.001) | 0.595 (0.058 to 0.961) | 0.000 (0.000 to 0.000) | −1.573 (−2.593 to −0.554) |
| Eosinophils | 29.508% | * 0.448 (p < 0.001) | N.A. | N.A. | 1.545 (0.428 to 2.663) | 54.226% | 0.629 (p < 0.001) | 1.000 (0.606 to 2.020) | 0.000 (0.000 to 0.000) | 0.129 (−0.146 to 0.404) |
| Basophils | 38.222% | * 0.336. (p = 0.001) | N.A. | N.A. | 1.307 (0.514 to 2.100) | 61.67% | * 0.567 (p < 0.001) | N.A. | N.A. | −0.062 (−0.366 to 0.241) |
| Metamyelocytes | 56.911% | * 0.592 (p < 0.001) | N.A. | N.A. | −0.295 (−0.846 to 0.256) | 69.203% | 0.837 (p < 0.001) | 1.087 (0.707 to 2.944) | 0.000 (0.000 to 0.000) | −0.002 (−0.446 to 0.441) |
| Myelocytes | 55.241% | * 0.542 (p < 0.001) | N.A. | N.A. | 0.932 (−0.652 to 2.515) | 71.297% | 0.851 (p < 0.001) | 0.891 (0.438 to 1.515) | 0.000 (0.000 to 0.000) | 0.198 (−0.814 to 1.211) |
| Promyelocytes | 19.90% | 0.194 (p = 0.070) | 0.125 (0.000 to 1.200) | 0.000 (0.000 to 0.000) | −2.932 (−5.397 to −0.466) | 27.56% | 0.184 (p = 0.09) | 0.442 (0.000 to 1.136) | 0.000 (0.000 to 0.000) | −2.836 (−5.534 to −0.337) |
| Blast cells | 62.920% | 0.662 (p < 0.001) | 0.667 (0.500 to 0.815) | 0.000 (0.000 to 0.261) | −11.341 (−17.218 to − 5.464) | 86.193% | 0.901 (p < 0.001) | 1.069 (0.994 to 1.170) | 0.000 (0.000 to 0.000) | 5.744 (2.064 to 9.424) |
| Accuracy and Correlation (with Bias Analysis) | ||
|---|---|---|
| DC-1 Pre-Classification | DC-1 Post-Classification | |
| Segmented neutrophils |
|
|
| Band neutrophils |
|
|
| Lymphocytes |
|
|
| Monocytes |
|
|
| Eosinophils |
|
|
| Basophils |
|
|
| Metamyelocytes |
|
|
| Myelocytes |
|
|
| Promyelocytes |
|
|
| Blast cells |
|
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kowald, A.; Fung, C.H.; Moon, J.; Shibeeb, S. The Diagnostic Performance of the Cellavision DC-1 Digital Morphology Analyser on Leukaemia Samples. Diagnostics 2025, 15, 2029. https://doi.org/10.3390/diagnostics15162029
Kowald A, Fung CH, Moon J, Shibeeb S. The Diagnostic Performance of the Cellavision DC-1 Digital Morphology Analyser on Leukaemia Samples. Diagnostics. 2025; 15(16):2029. https://doi.org/10.3390/diagnostics15162029
Chicago/Turabian StyleKowald, Annabel, Chun Ho Fung, Jane Moon, and Sapha Shibeeb. 2025. "The Diagnostic Performance of the Cellavision DC-1 Digital Morphology Analyser on Leukaemia Samples" Diagnostics 15, no. 16: 2029. https://doi.org/10.3390/diagnostics15162029
APA StyleKowald, A., Fung, C. H., Moon, J., & Shibeeb, S. (2025). The Diagnostic Performance of the Cellavision DC-1 Digital Morphology Analyser on Leukaemia Samples. Diagnostics, 15(16), 2029. https://doi.org/10.3390/diagnostics15162029





