Early Diagnosis of COVID-19 Images Using Optimal CNN Hyperparameters
Abstract
1. Introduction
- The number of convolutional layers.
- The number of filters.
- The filter size.
- The stride is thus the number of steps the filter takes as it slides across the input image.
- The kind of padding might be the same or a valid convolution.
- The batch size.
- The number of training epochs
- The learning rate, momentum, and dropout probability.
- The CNN hyperparameters are optimized using the grid search approach to reduce model losses and achieve the best degree of COVID-19 diagnostic accuracy.
- The grid search optimization algorithm chooses the optimum CNN hyperparameters from a provided list of parameter possibilities, automating the “trial-and-error” process to obtain the optimized parameters with the greatest diagnosis accuracy.
- The optimization approach is tested and compared using three different well-known CNN structures (GoogleNet, VGG16, and ResNet50).
- The optimized CNN architectures were used to classify CT and CXR images for COVID-19 patients to increase the diagnostic sensitivity for identifying COVID-19.
- Simulation results confirm the efficiency of the CNN architectures with optimized hyperparameters regarding disease categorization. Hence, our envisioned approach outperforms unoptimized CNN methods.
- Furthermore, our envisioned technique attains the highest accuracy compared to previous techniques using both X-ray and CT images.
2. Related Work
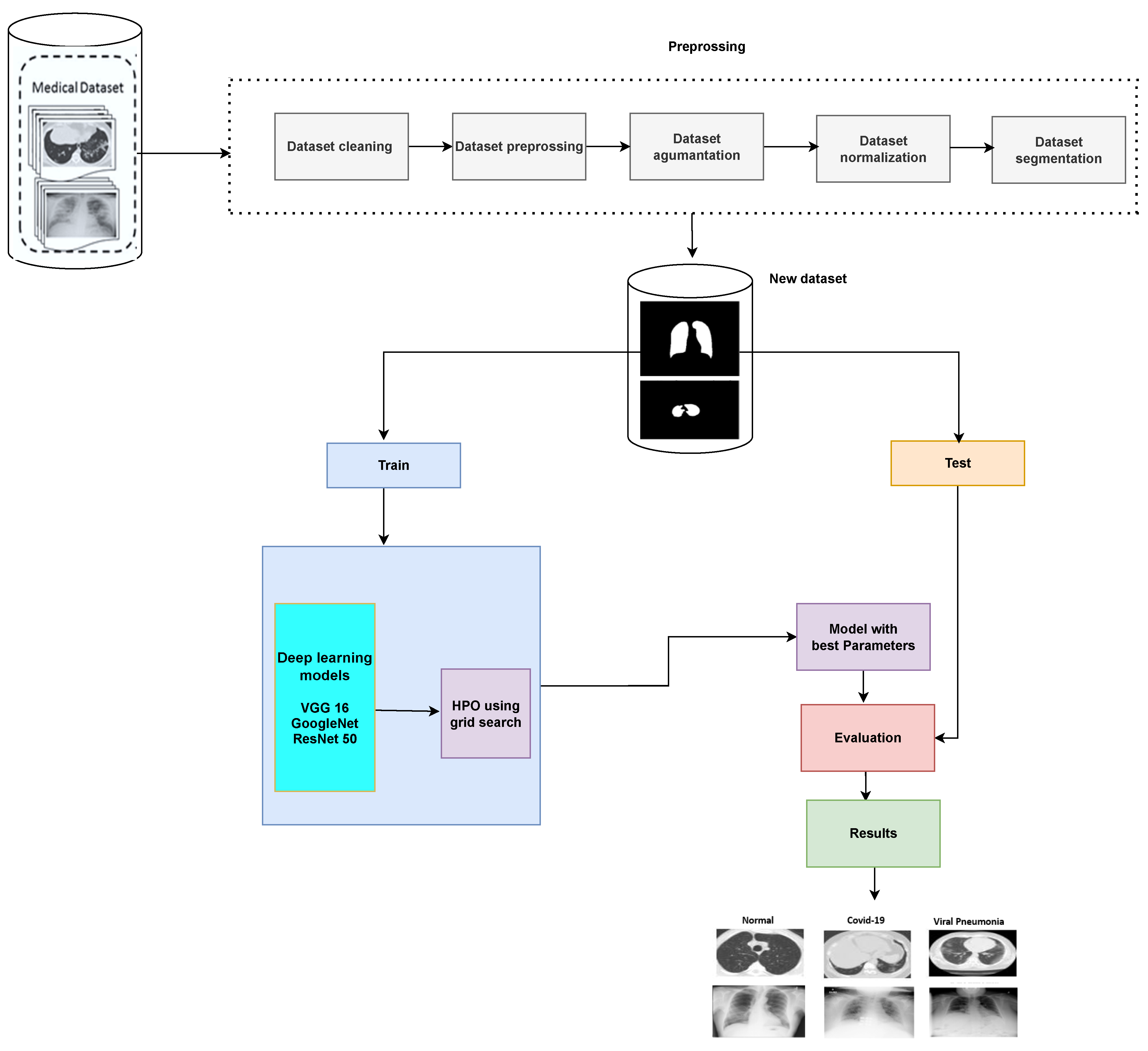
3. Proposed COVID-19 Detection System
- Load the images and prepare the training and test images.
- Create an image data augmenter that configures a set of preprocessing options for image augmentation, such as resizing, rotation, and reflection.
- Resize the training and testing images to the size required by the network.
- Implement the grid search method to get the optimal learning rate and momentum.
- Finally, train the model with the optimal learning rate and momentum based on grid search.
3.1. Preprocessing Phase
3.1.1. Image Augmentation
- Gaussian blur: a Gaussian filter may be used to remove high-frequency elements, resulting in a blurred image version.
- Rotation: a rotation of between 10° and 180° is applied to the picture.
- Shear: using rotation and the imitation factor for the third dimension, picture shearing may be done.
3.1.2. Image Processing
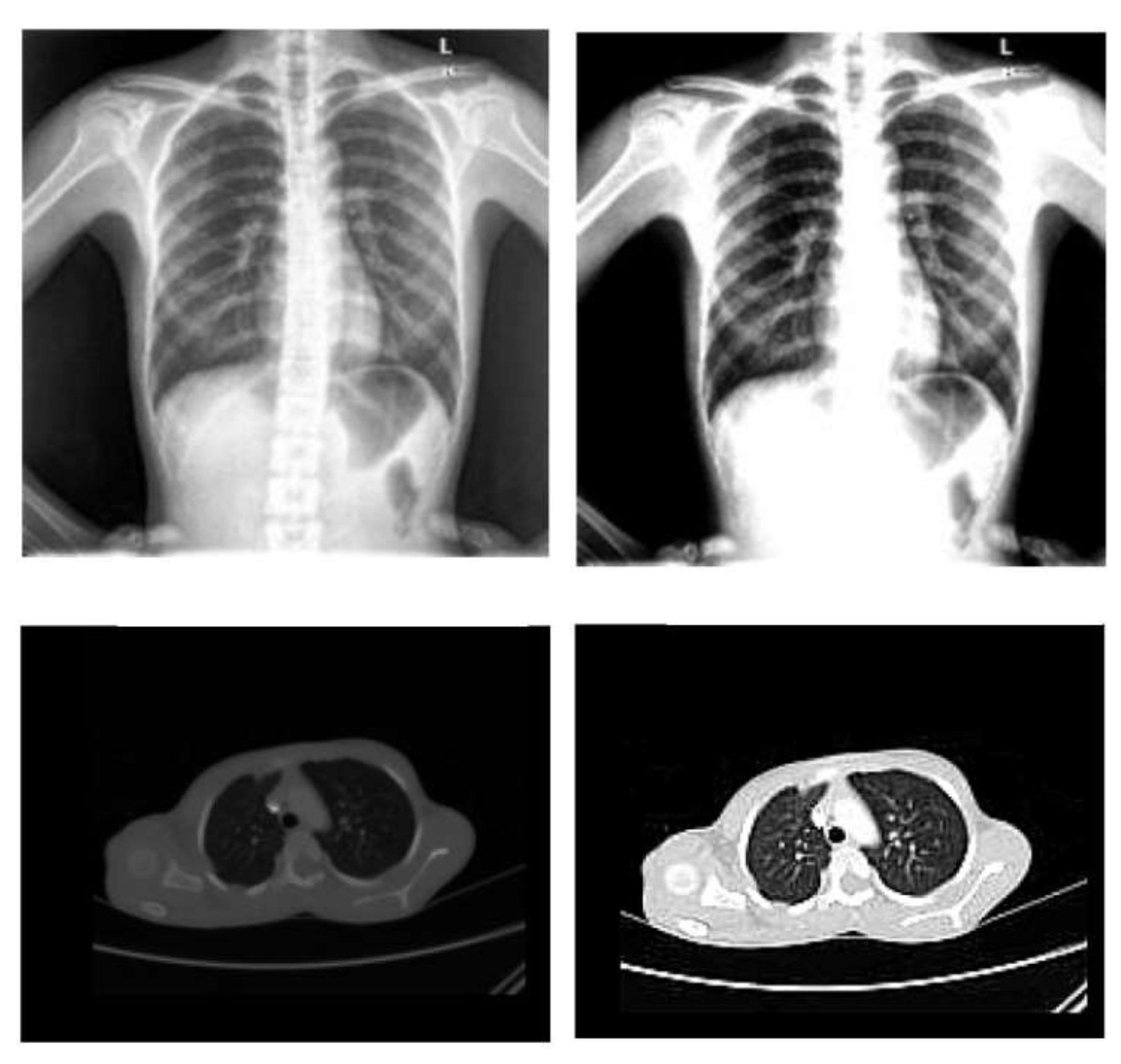
- Image standardization: it is necessary for CNNs because they deal with images. As a result, the images must first be resized into distinct dimensions and a square form, which is the typical shape used in neural networks (NNs).
- Normalization: To improve the convergence of the training phase, input pixels to any AI system must have a normalized data distribution. To normalize an image, the distribution’s mean value is first subtracted from each pixel, then divided by the result by the standard deviation. Sample X-ray and CT images are shown in Figure 2 before (on the left) and after (on the right) the preprocessing steps.
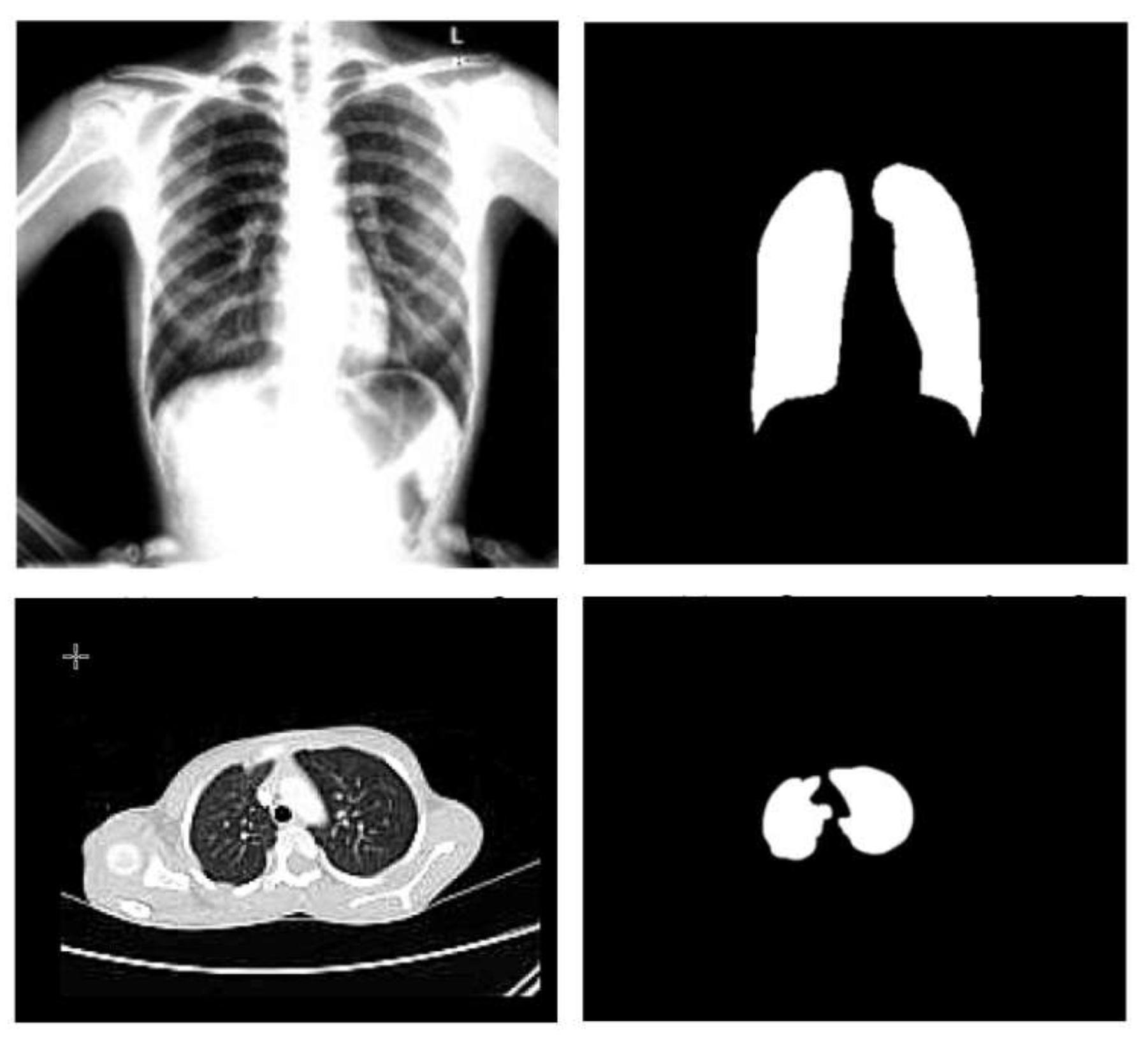
3.1.3. Segmentation
3.2. Reference CNN Models for Transfer Learning
- VGG16: The foundation for the 2014 ImageNet competition entry was the VGG network design (VGG16), which has 16 layers. Five blocks of convolutional layers and three fully linked layers make up VGG16. Convolution uses a filter of size 3 × 3 with stride 1 and padding 1. After each convolution, the ReLU activation function is applied, and the spatial dimensions are decreased by max-pooling with a 2 × 2 filter, stride 2, and no padding.
- GoogleNet: Using inception modules, GoogleNet conducts convolutions with various filter sizes. ImageNet’s 2014 large-scale visual recognition competition (ILSVRC) was known as GoogleNet thanks to its superior performance. It employs four million parameters and has 2 × 2 layers. Layers are deeper with concurrent use of various field widths, and a 6.67% error rate was attained. The ReLu activation function is utilized for all convolutions, including those inside the inception module, and a 1 × 1 filter is applied before the and convolutions.
- ResNet: ResNet, a 2015 ILSVRC winner, is another name for the residual network. In order to increase the classification accuracy of challenging tasks, very deep models are employed for visual recognition tasks. However, the training procedure becomes more challenging, and accuracy begins to decline as the network depth increases. Skip connections were utilized to add residual learning to solve this issue. In general, layers are placed for training and network learn features at the end of these layers in a convolutional-based deep NN. A residual-based network has a residual link that spans two or more network levels. ResNet accomplishes this goal by linking the nth layer to the th layer. The 34-layered ResNet solves the problem of accuracy loss in a deeper convolutional network and is simple to train.
3.2.1. Hyperparameters
3.2.2. Grid Search Scheme
- Start with a large search space and phase scale, then limit them based on prior observations of hyperparameter settings that performed well.
- Repeat multiple times until the best result is obtained.
3.2.3. Optimized CNN Model
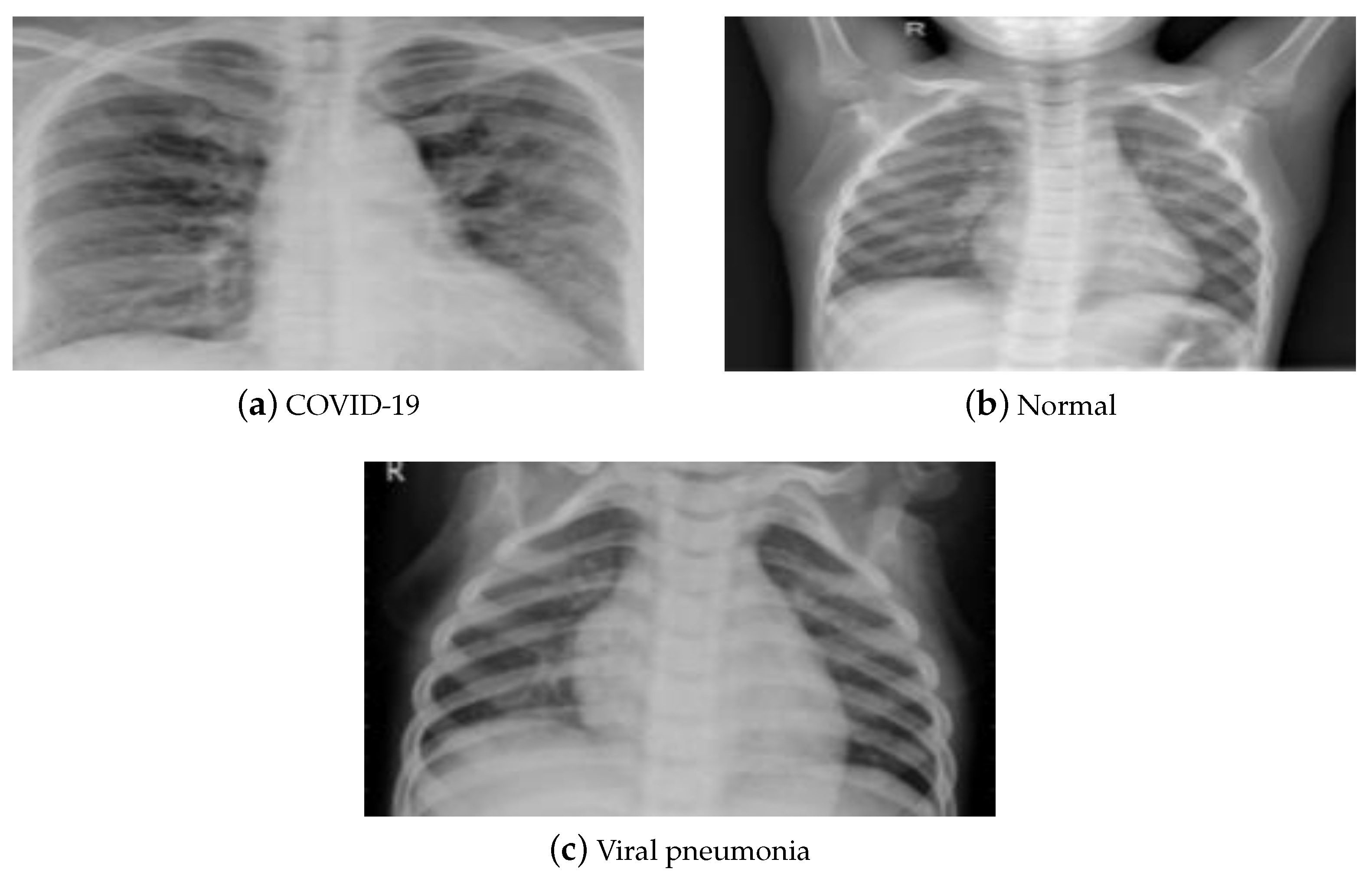
4. Utilized Data Sets
5. Results and Discussions
5.1. Evaluation Metrics
- The sensitivity or recallIt is the accuracy of positive examples. It refers to how many examples of the positive classes were labeled correctly. This is shown in Equation (4), where is the true positives, which are the number of instances that are correctly identified, and FN is the false negatives, which are the number of positive cases that are classified as negative by mistake [54].
- SpecificityIt refers to the conditional probability of true negatives given a secondary class. It approximates the probability of the negative label being true as in Equation (5), where is the number of true negatives classified as negative and is the number of false positives, defined by the negative instances that are classified incorrectly as positive cases. In general, sensitivity and specificity evaluate the positive and negative effectiveness of the algorithm on a single class, respectively [55].
- Accuracy
- PrecisionIt is calculated by the number of true positives divided by the number of true positives plus the number of false positives as in Equation (7). It evaluates the predictive power of the algorithm. Precision is how “precise” the model is out of those predicted positive and how many of them are actually positive [55].
- F-score
- False-positive rate and false-negative rateFalse-positive rate (FPR) is the portion of negative cases identified improperly as positive instances to the total number of negative instances. FPR is an error in classification in which a test result incorrectly indicates the presence of a condition (such as a disease when the disease is not present), while a false-negative rate (FNR) is the opposite error, where the test result incorrectly indicates the absence of a condition when it is actually present. These are the two kinds of errors in a test, in contrast to the two kinds of correct results (a true positive and a true negative). They are also known in medicine as a false-positive (or false-negative) diagnosis, and in statistical classification as a false-positive (or false-negative) error.The error rate is the performance statistic that informs of incorrect predictions without classifying positive and negative forecasts. It can evaluate by
5.2. Results
6. Conclusions
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zeyaullah, M.; AlShahrani, A.M.; Muzammil, K.; Ahmad, I.; Alam, S.M.; Khan, W.H.; Ahmad, R. COVID-19 and SARS-CoV-2 Variants: Current Challenges and Health Concern. Front. Genet. 2021, 12, 693916. [Google Scholar] [CrossRef] [PubMed]
- Jebril, N.M.T. World Health Organization Declared a Pandemic Public Health Menace: A Systematic Review of the Coronavirus Disease 2019 “COVID-19”. 2020. Available online: https://ssrn.com/abstract=3566298 (accessed on 23 November 2022).
- Mair, M.; Hussain, M.; Siddiqui, S.; Das, S.; Baker, A.J.; Conboy, P.J.; Valsamakis, T.; Uddin, J.; Rae, P.D. A systematic review and meta-analysis comparing the diagnostic accuracy of initial RT-PCR and CT scan in suspected COVID-19 patients. Br. J. Radiol. 2020, 94, 20201039. [Google Scholar] [CrossRef] [PubMed]
- Karimi, F.; Vaezi, A.A.; Qorbani, M.; Moghadasi, F.; Gelsfid, S.H.; Maghoul, A.; Mahmoodi, N.; Eskandari, Z.; Gholami, H.; Mokhames, Z.; et al. Clinical and laboratory findings in COVID-19 adult hospitalized patients from Alborz province/Iran: Comparison of rRT-PCR positive and negative. BMC Infect. Dis. 2021, 21, 05948-5. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.C.; Chatterjee, D. COVID-19, Older Adults and the Ageing Society; Taylor & Francis Group: Abingdon, UK, 2022. [Google Scholar] [CrossRef]
- Zhang, Y.D.; Zhang, Z.; Zhang, X.; Wang, S. MIDCAN: A multiple input deep convolutional attention network for COVID-19 diagnosis based on chest CT and chest X-ray. Pattern Recognit. Lett. 2021, 150, 8–16. [Google Scholar] [CrossRef]
- Hosny, A.; Parmar, C.; Quackenbush, J.; Schwartz, L.H.; Aerts, H.J. Artificial intelligence in radiology. Nat. Rev. Cancer 2018, 18, 500–510. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Chabert, G.L.; Carriero, A.; Paschè, A.; Danna, P.S.C.; Saba, L.; Mehmedović, A.; Faa, G.; Singh, I.M.; et al. COVLIAS 1.0Lesion vs. MedSeg: An Artificial Intelligence Framework for Automated Lesion Segmentation in COVID-19 Lung Computed Tomography Scans. Diagnostics 2022, 12, 1283. [Google Scholar] [CrossRef]
- Suri, J.S.; Agarwal, S.; Chabert, G.L.; Carriero, A.; Paschè, A.; Danna, P.S.C.; Saba, L.; Mehmedović, A.; Faa, G.; Singh, I.M.; et al. COVLIAS 2.0-cXAI: Cloud-Based Explainable Deep Learning System for COVID-19 Lesion Localization in Computed Tomography Scans. Diagnostics 2022, 12, 1482. [Google Scholar] [CrossRef]
- Nillmani; Sharma, N.; Saba, L.; Khanna, N.N.; Kalra, M.K.; Fouda, M.M.; Suri, J.S. Segmentation-Based Classification Deep Learning Model Embedded with Explainable AI for COVID-19 Detection in Chest X-ray Scans. Diagnostics 2022, 12, 2132. [Google Scholar] [CrossRef]
- Ahmad, R.W.; Salah, K.; Jayaraman, R.; Yaqoob, I.; Ellahham, S.; Omar, M.A. Blockchain and COVID-19 Pandemic: Applications and Challenges. IEEE TechRxiv 2020. [Google Scholar] [CrossRef]
- Alom, M.Z.; Taha, T.M.; Yakopcic, C.; Westberg, S.; Sidike, P.; Nasrin, M.S.; Hasan, M.; Essen, B.C.V.; Awwal, A.A.S.; Asari, V.K. A State-of-the-Art Survey on Deep Learning Theory and Architectures. Electronics 2019, 8, 292. [Google Scholar] [CrossRef]
- Liang, S.; Liu, H.; Gu, Y.; Guo, X.; Li, H.; Li, L.; Wu, Z.; Liu, M.; Tao, L. Fast automated detection of COVID-19 from medical images using convolutional neural networks. Commun. Biol. 2021, 4, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Sakib, S.; Tazrin, T.; Fouda, M.M.; Fadlullah, Z.M.; Guizani, M. DL-CRC: Deep Learning-Based Chest Radiograph Classification for COVID-19 Detection: A Novel Approach. IEEE Access 2020, 8, 171575–171589. [Google Scholar] [CrossRef] [PubMed]
- Munjral, S.; Maindarkar, M.; Ahluwalia, P.; Puvvula, A.; Jamthikar, A.; Jujaray, T.; Suri, N.; Paul, S.; Pathak, R.; Saba, L.; et al. Cardiovascular Risk Stratification in Diabetic Retinopathy via Atherosclerotic Pathway in COVID-19/Non-COVID-19 Frameworks Using Artificial Intelligence Paradigm: A Narrative Review. Diagnostics 2022, 12, 1234. [Google Scholar] [CrossRef] [PubMed]
- Saxena, S.; Jena, B.; Gupta, N.; Das, S.; Sarmah, D.; Bhattacharya, P.; Nath, T.; Paul, S.; Fouda, M.M.; Kalra, M.; et al. Role of Artificial Intelligence in Radiogenomics for Cancers in the Era of Precision Medicine. Cancers 2022, 14, 2860. [Google Scholar] [CrossRef]
- Rajpurkar, P.; Irvin, J.A.; Ball, R.L.; Zhu, K.; Yang, B.; Mehta, H.; Duan, T.; Ding, D.Y.; Bagul, A.; Langlotz, C.; et al. Deep learning for chest radiography diagnosis: A retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med. 2018, 15, e1002686. [Google Scholar] [CrossRef]
- Castiglioni, I.; Rundo, L.; Codari, M.; Leo, G.D.; Salvatore, C.; Interlenghi, M.; Gallivanone, F.; Cozzi, A.; D’Amico, N.C.; Sardanelli, F. AI applications to medical images: From machine learning to deep learning. Phys. Med. 2021, 83, 9–24. [Google Scholar] [CrossRef]
- El-Shazly, E.H.; Zhang, X.; Jiang, J. Improved appearance loss for deep estimation of image depth. Electron. Lett. 2019, 55, 264–266. [Google Scholar] [CrossRef]
- Jiang, J.; El-Shazly, E.H.; Zhang, X. Gaussian weighted deep modeling for improved depth estimation in monocular images. IEEE Access 2019, 7, 134718–134729. [Google Scholar] [CrossRef]
- Jin, C.; Netrapalli, P.; Ge, R.; Kakade, S.M.; Jordan, M. On Nonconvex Optimization for Machine Learning. J. ACM 2021, 68, 1–29. [Google Scholar] [CrossRef]
- Lu, T.C. CNN Convolutional layer optimisation based on quantum evolutionary algorithm. Connect. Sci. 2021, 33, 482–494. [Google Scholar] [CrossRef]
- Singh, P.; Chaudhury, S.; Panigrahi, B.K. Hybrid MPSO-CNN: Multi-level Particle Swarm optimized hyperparameters of Convolutional Neural Network. Swarm Evol. Comput. 2021, 63, 100863. [Google Scholar] [CrossRef]
- Al-qaness, M.A.A.; Ewees, A.A.; Fan, H.; Aziz, M.A.A.E. Optimization Method for Forecasting Confirmed Cases of COVID-19 in China. J. Clin. Med. 2020, 9, 674. [Google Scholar] [CrossRef] [PubMed]
- Apostolopoulos, I.D.; Bessiana, T. COVID-19: Automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys. Eng. Sci. Med. 2020, 43, 635–640. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xie, Y.; Li, Y.; Shen, C.; Xia, Y. COVID-19 Screening on Chest X-ray Images Using Deep Learning based Anomaly Detection. arXiv 2020, arXiv:2003.12338. [Google Scholar]
- Singh, D.; Kumar, V.; Vaishali; Kaur, M. Classification of COVID-19 patients from chest CT images using multi-objective differential evolution–based convolutional neural networks. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 1379–1389. [Google Scholar] [CrossRef]
- Jaiswal, A.; Gianchandani, N.; Singh, D.; Kumar, V.; Kaur, M. Classification of the COVID-19 infected patients using DenseNet201 based deep transfer learning. J. Biomol. Struct. Dyn. 2020, 39, 5682–5689. [Google Scholar] [CrossRef]
- Chen, X.; Yao, L.; Zhang, Y. Residual Attention U-Net for Automated Multi-Class Segmentation of COVID-19 Chest CT Images. arXiv 2020, arXiv:2004.05645. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention; Springer: Cham, Switzerland, 2015; pp. 234–241. [Google Scholar] [CrossRef]
- Adhikari, N.C.D. Infection Severity Detection of CoVID19 from X-rays and CT Scans Using Artificial Intelligence. Int. J. Comput. 2020, 38, 73–92. [Google Scholar]
- Khan, A.I.; Shah, J.L.; Bhat, M. CoroNet: A deep neural network for detection and diagnosis of COVID-19 from chest X-ray images. Comput. Methods Programs Biomed. 2020, 196, 105581. [Google Scholar] [CrossRef]
- Ghoshal, B.; Tucker, A. Estimating Uncertainty and Interpretability in Deep Learning for Coronavirus (COVID-19) Detection. arXiv 2020, arXiv:2003.10769. [Google Scholar]
- Hemdan, E.E.D.; Shouman, M.A.; Karar, M.E. COVIDX-Net: A Framework of Deep Learning Classifiers to Diagnose COVID-19 in X-ray Images. arXiv 2020, arXiv:2003.11055. [Google Scholar]
- Reshi, A.A.; Rustam, F.; Mehmood, A.; Alhossan, A.; Alrabiah, Z.; Ahmad, A.; Alsuwailem, H.; Choi, G.S. An Efficient CNN Model for COVID-19 Disease Detection Based on X-ray Image Classification. Complexity 2021, 2021, 6621607:1–6621607:12. [Google Scholar] [CrossRef]
- Hota, S.; Satapathy, P.; Acharya, B.M. Performance Analysis of Hyperparameters of Convolutional Neural Networks for COVID-19 X-ray Image Classification. In Ambient Intelligence in Health Care; Springer: Singapore, 2022. [Google Scholar] [CrossRef]
- Zubair, S.; Singha, A.K. Parameter optimization in convolutional neural networks using gradient descent. In Microservices in Big Data Analytics; Springer: Cham, Switzerland, 2020; pp. 87–94. [Google Scholar]
- Lacerda, P.; Barros, B.; Albuquerque, C.; Conci, A. Hyperparameter Optimization for COVID-19 Pneumonia Diagnosis Based on Chest CT. Sensors 2021, 21, 2174. [Google Scholar] [CrossRef] [PubMed]
- Singha, A.K.; Pathak, N.; Sharma, N.; Gandhar, A.; Urooj, S.; Zubair, S.; Sultana, J.; Nagalaxmi, G. An Experimental Approach to Diagnose COVID-19 Using Optimized CNN. Intell. Autom. Soft Comput. 2022, 34, 1066–1080. [Google Scholar] [CrossRef]
- Aslan, M.F.; Sabanci, K.; Durdu, A.; Unlersen, M.F. COVID-19 diagnosis using state-of-the-art CNN architecture features and Bayesian Optimization. Comput. Biol. Med. 2022, 142, 105244. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.K.; Greenspan, H.; Davatzikos, C.; Duncan, J.S.; van Ginneken, B.; Madabhushi, A.; Prince, J.L.; Rueckert, D.; Summers, R.M. A Review of Deep Learning in Medical Imaging: Imaging Traits, Technology Trends, Case Studies with Progress Highlights, and Future Promises. Proc. IEEE 2021, 109, 820–838. [Google Scholar] [CrossRef]
- Chatzimparmpas, A.; Martins, R.M.; Kucher, K.; Kerren, A. VisEvol: Visual Analytics to Support Hyperparameter Search through Evolutionary Optimization. Comput. Graph. Forum 2021, 40, 201–214. [Google Scholar] [CrossRef]
- Wu, Y.; Cheng, M.; Huang, S.; Pei, Z.; Zuo, Y.; Liu, J.; Yang, K.; Zhu, Q.; Zhang, J.; Hong, H.; et al. Recent Advances of Deep Learning for Computational Histopathology: Principles and Applications. Cancers 2022, 14, 1199. [Google Scholar] [CrossRef]
- Gaspar, A.; Oliva, D.; Cuevas, E.; Zaldívar, D.; Pérez, M.; Pajares, G. Hyperparameter Optimization in a Convolutional Neural Network Using Metaheuristic Algorithms. In Metaheuristics in Machine Learning: Theory and Applications; Springer: Cham, Switzerland, 2021; pp. 37–59. [Google Scholar]
- Bacanin, N.; Bezdan, T.; Tuba, E.; Strumberger, I.; Tuba, M. Optimizing Convolutional Neural Network Hyperparameters by Enhanced Swarm Intelligence Metaheuristics. Algorithms 2020, 13, 67. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, H.; Zhang, G. cPSO-CNN: An efficient PSO-based algorithm for fine-tuning hyper-parameters of convolutional neural networks. Swarm Evol. Comput. 2019, 49, 114–123. [Google Scholar] [CrossRef]
- Uyar, K.; Tasdemir, S.; Özkan, İ.A. The analysis and optimization of CNN Hyperparameters with fuzzy tree modelfor image classification. Turk. J. Electr. Eng. Comput. Sci. 2022, 30, 961–977. [Google Scholar] [CrossRef]
- Belete, D.M.; Huchaiah, M.D. Grid search in hyperparameter optimization of machine learning models for prediction of HIV/AIDS test results. Int. J. Comput. Appl. 2021, 44, 875–886. [Google Scholar] [CrossRef]
- Zahedi, L.; Mohammadi, F.; Rezapour, S.; Ohland, M.W.; Amini, M.H. Search Algorithms for Automated Hyper-Parameter Tuning. arXiv 2021, arXiv:2104.14677. [Google Scholar]
- Mustapha, A.; Mohamed, L.; Ali, K. Comparative study of optimization techniques in deep learning: Application in the ophthalmology field. J. Phys. Conf. Ser. 2021, 1743, 012002. [Google Scholar] [CrossRef]
- Sajjad, U.; Hussain, I.; Hamid, K.; Ali, H.M.; Wang, C.C.; Yan, W.M. Liquid-to-vapor phase change heat transfer evaluation and parameter sensitivity analysis of nanoporous surface coatings. Int. J. Heat Mass Transf. 2022, 194, 123088. [Google Scholar] [CrossRef]
- Cengil, E.; Cinar, A.C. The effect of deep feature concatenation in the classification problem: An approach on COVID-19 disease detection. Int. J. Imaging Syst. Technol. 2022, 32, 26–40. [Google Scholar] [CrossRef]
- Mahanty, C.; Kumar, R.; Patro, S.G.K. Internet of Medical Things-Based COVID-19 Detection in CT Images Fused with Fuzzy Ensemble and Transfer Learning Models. New Gener. Comput. 2022, 40, 1125–1141. [Google Scholar] [CrossRef]
- Zoabi, Y.; Deri-Rozov, S.; Shomron, N. Machine learning-based prediction of COVID-19 diagnosis based on symptoms. npj Digit. Med. 2021, 4, 1–5. [Google Scholar] [CrossRef]
- Soleymani, R.; Granger, É.; Fumera, G. F-measure curves: A tool to visualize classifier performance under imbalance. Pattern Recognit. 2020, 100, 107146. [Google Scholar] [CrossRef]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (COVID-19) using X-ray images and deep convolutional neural networks. Pattern Anal. Appl. 2021, 24, 1207–1220. [Google Scholar] [CrossRef]
- Salman, F.M.; Abu-Naser, S.S.; Alajrami, E.; Abu-Nasser, B.S.; Alashqar, B.A.M. COVID-19 Detection Using Artificial Intelligence. 2020. Available online: http://dstore.alazhar.edu.ps/xmlui/handle/123456789/587 (accessed on 23 November 2022).
- Matsuyama, Eri, Haruyuki Watanabe, and Noriyuki Takahashi, Explainable Analysis of Deep Learning Models for Coronavirus Disease (COVID-19) Classification with Chest X-Ray Images: Towards Practical Applications. Open J. Med. Imaging 2022, 12, 83–102. [CrossRef]
- Xu, X.; Jiang, X.; Ma, C.; Du, P.; Li, X.; Lv, S.; Yu, L.; Ni, Q.; Chen, Y.; Su, J.; et al. A Deep Learning System to Screen Novel Coronavirus Disease 2019 Pneumonia. Engineering 2020, 6, 1122–1129. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Fang, Y.; Li, W.; Pan, C.; Qin, P.; Zhong, Y.; Liu, X.; Huang, M.; Liao, Y.; Li, S. CT image visual quantitative evaluation and clinical classification of coronavirus disease (COVID-19). Eur. Radiol. 2020, 30, 4407–4416. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Zhao, W.; Xie, X.; Zhong, Z.; Shi, F.; Liu, J.; Shen, D. Severity Assessment of Coronavirus Disease 2019 (COVID-19) Using Quantitative Features from Chest CT Images. arXiv 2020, arXiv:2003.11988. [Google Scholar]
- Ozkaya, U.; Öztürk, Ş.; Barstugan, M. Coronavirus (COVID-19) Classification Using Deep Features Fusion and Ranking Technique. In Big Data Analytics and Artificial Intelligence against COVID-19: Innovation Vision and Approach; Springer: Cham, Switzerland, 2020; Volume 78, pp. 281–295. [Google Scholar]
| Hyperparameters | Description |
|---|---|
| Learning Rate | The learning rate defines how quickly a network updates its parameters. For the classification problem, it is important to choose the optimal learning rate to minimize the loss function. A low learning rate slows down the learning process but converges smoothly. A larger learning rate speeds up the learning but may not converge. |
| Momentum | Momentum helps to know the direction of the next step with the knowledge of the previous steps. It helps to prevent oscillations. |
| Number of Epochs | The number of epochs is the number of times the whole training data are introduced to the network. It is important to determine an ideal epoch number to prevent overfitting. |
| MiniBatch Size | The larger minibatch size causes running of the model for a long period of time with constant weights that causes overall performance loses and increases the memory requirements. Carrying out the experiments with small minibatch sizes can be more beneficial. |
| Hyperparameters | Range |
|---|---|
| Learning Rate | [0.09, 0.07, 0.05, 0.03, 0.01] |
| Momentum | [0.9, 0.7, 0.5, 0.3, 0.1] |
| Number of Epochs | 10 to 100 with step 5 |
| Hyperparameters | Resnet | Google Net | VGG16 |
|---|---|---|---|
| Learning Rate | 0.01 | 0.01 | 0.02 |
| Momentum | 0.1 | 0.1 | 0.3 |
| Number of Epochs | 25 | 30 | 45 |
| Activation Function | Relu | Relu | Relu |
| Classifier | Softmax | Softmax | Softmax |
| Loss Function | Cross entropy | Cross entropy | Cross entropy |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 94.86 | 94.91 | 93.52 | 94.24 | 93.88 | 0.0509 | 0.0576 | 0.0514 |
| COVID-19 | 93.52 | 93.36 | 92.67 | 94.38 | 93.52 | 0.0664 | 0.0562 | 0.0648 |
| Viral Pneumonia | 92.89 | 94.82 | 94.83 | 90.14 | 92.43 | 0.0518 | 0.0986 | 0.0711 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 98.91 | 98.65 | 97.67 | 97.59 | 97.63 | 0.0253 | 0.0648 | 0.0218 |
| COVID-19 | 97.69 | 96.25 | 96.57 | 96.78 | 96.67 | 0.0135 | 0.0241 | 0.0109 |
| Viral Pneumonia | 97.82 | 97.47 | 98.18 | 93.52 | 95.79 | 0.0375 | 0.0322 | 0.0231 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 96.86 | 96.91 | 95.52 | 96.24 | 95.88 | 0.0309 | 0.0376 | 0.0314 |
| COVID-19 | 96.52 | 96.51 | 96.67 | 96.38 | 96.52 | 0.0349 | 0.0362 | 0.0348 |
| Viral Pneumonia | 96.39 | 96.82 | 96.83 | 96.14 | 96.48 | 0.0318 | 0.0386 | 0.0361 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 98.93 | 98.88 | 98.93 | 98.54 | 98.73 | 0.0112 | 0.0146 | 0.0107 |
| COVID-19 | 98.49 | 98.29 | 98.78 | 98.88 | 98.83 | 0.0171 | 0.0112 | 0.0151 |
| Viral Pneumonia | 98.18 | 98.28 | 98.58 | 97.12 | 97.84 | 0.0172 | 0.0288 | 0.0182 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 97.58 | 97.56 | 97.52 | 97.47 | 97.49 | 0.0265 | 0.0365 | 0.0313 |
| COVID-19 | 96.84 | 96.92 | 96.95 | 96.87 | 96.91 | 0.0244 | 0.0253 | 0.0242 |
| Viral Pneumonia | 96.87 | 97.35 | 97.27 | 96.35 | 96.81 | 0.0308 | 0.0313 | 0.0316 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 99.84 | 99.79 | 99.48 | 99.57 | 99.52 | 0.0021 | 0.0043 | 0.0016 |
| COVID-19 | 98.98 | 98.88 | 98.78 | 98.88 | 98.83 | 0.0112 | 0.0112 | 0.0102 |
| Viral Pneumonia | 98.75 | 98.95 | 98.58 | 98.49 | 98.53 | 0.0105 | 0.0151 | 0.0125 |
| Category | Accuracy (%) | Specificity (%) | Precision (%) | Recall (%) | F1-Score (%) | FPR | FNR | Error Rate |
|---|---|---|---|---|---|---|---|---|
| Normal | 99.79 | 99.65 | 99.25 | 99.36 | 99.30 | 0.0118 | 0.0162 | 0.0147 |
| COVID-19 | 98.78 | 98.75 | 98.63 | 98.59 | 98.61 | 0.0035 | 0.0064 | 0.0021 |
| Viral Pneumonia | 98.53 | 98.82 | 98.58 | 98.38 | 98.48 | 0.0125 | 0.0141 | 0.0122 |
| Reference | Technique | Accuracy (%) |
|---|---|---|
| [56] | Resnet50 | 92.74 |
| [57] | CNN | 93.37 |
| [25] | Transfer Learning | 95.23 |
| [58] | CNN | 95.92 |
| This paper | Optimal CNN Hyperparameters | 98.98 |
| Reference | Technique | Accuracy (%) |
|---|---|---|
| [59] | Random forest | 88.6 |
| [60] | 3D-CNN | 90.7 |
| [61] | Statistical Analysis | 89.9 |
| [62] | CNN Future Fusion | 94.8 |
| This paper | Optimal CNN Hyperparameters | 98.78 |
| Data Type | Model with Optimal Parameters | Time (s) |
|---|---|---|
| X-ray | VGG16 | 26.451 |
| X-ray | Google Net | 22.372 |
| X-ray | ResNet | 20.003 |
| CT | ResNet | 21.145 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saad, M.H.; Hashima, S.; Sayed, W.; El-Shazly, E.H.; Madian, A.H.; Fouda, M.M. Early Diagnosis of COVID-19 Images Using Optimal CNN Hyperparameters. Diagnostics 2023, 13, 76. https://doi.org/10.3390/diagnostics13010076
Saad MH, Hashima S, Sayed W, El-Shazly EH, Madian AH, Fouda MM. Early Diagnosis of COVID-19 Images Using Optimal CNN Hyperparameters. Diagnostics. 2023; 13(1):76. https://doi.org/10.3390/diagnostics13010076
Chicago/Turabian StyleSaad, Mohamed H., Sherief Hashima, Wessam Sayed, Ehab H. El-Shazly, Ahmed H. Madian, and Mostafa M. Fouda. 2023. "Early Diagnosis of COVID-19 Images Using Optimal CNN Hyperparameters" Diagnostics 13, no. 1: 76. https://doi.org/10.3390/diagnostics13010076
APA StyleSaad, M. H., Hashima, S., Sayed, W., El-Shazly, E. H., Madian, A. H., & Fouda, M. M. (2023). Early Diagnosis of COVID-19 Images Using Optimal CNN Hyperparameters. Diagnostics, 13(1), 76. https://doi.org/10.3390/diagnostics13010076








