Dermatopathology of Malignant Melanoma in the Era of Artificial Intelligence: A Single Institutional Experience
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data Acquisition
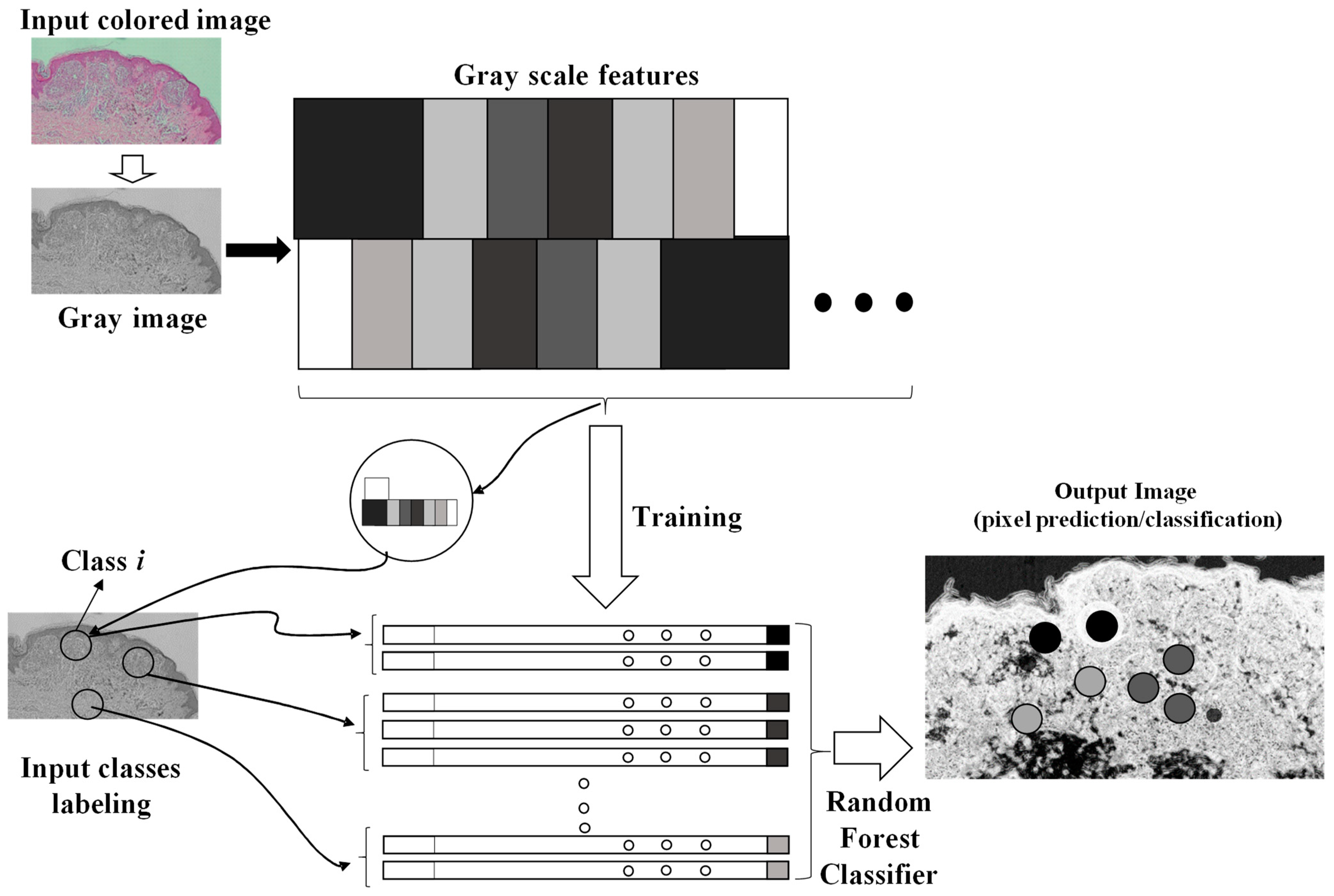
2.2. Fast Random Forest (FRF) Image Classification
- Different clusters of pixels (features) are used for the training model;
- A marked region is distinguished as the features to find in the same image to process;
- The RF is executed by finding similar features in the same image (similar features of similar clusters having a similar gray pixel intensity distribution).
- Gaussian blur filtering, obtaining a blurred image to process;
- Sobel filtering, which is able to approximate the image by a gradient of the intensity;
- Hessian filtering, defined as:
- Difference of Gaussian functions;
- Membrane projections due to the rotation of the original image kernel;
- Main pixel parameters (mean, minimum, maximum, etc.);
- Anisotropic diffusion filtering preserving sharp edges;
- Bilateral filtering preserving edges;
- Lipschitz filtering (preserving edges and decreasing noise);
- Gabor filtering (mainly adopted for texture analysis);
- Derivative filtering estimating high order derivatives;
- Structured filtering estimating the eigenvalues;
- Shifting of the image in different directions.
- -
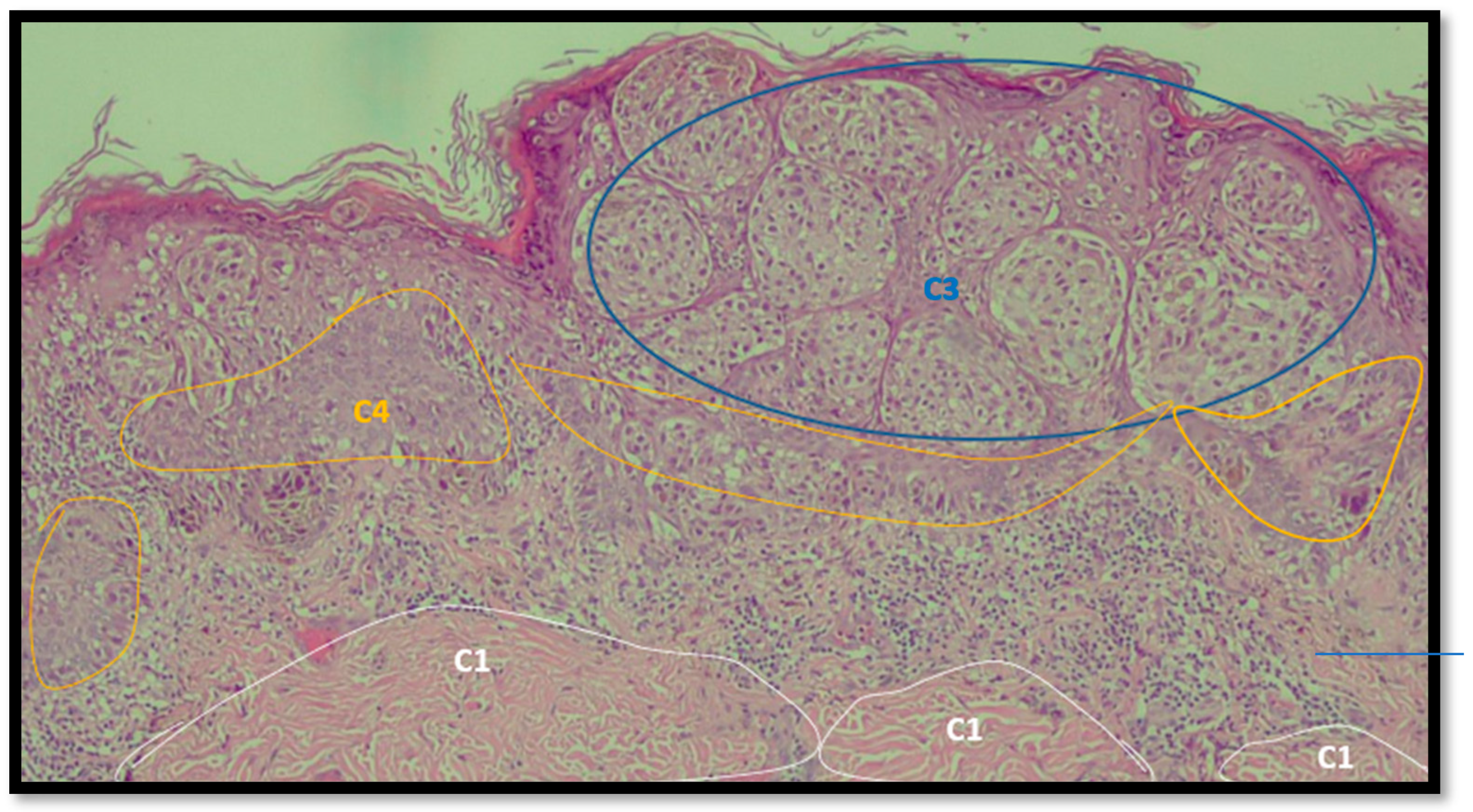
- A preliminary selection of images focusing the attention on the characteristics of dysplastic nevi and malignant melanoma (validation of the training model focusing on this classification);
- -
- Performing of a training of the selected images (FRF training model based on the classification of anomalous image areas embedding features of dysplastic nevi and MM, and the identification of other no-risk areas as structures of clusters of grayscale pixels);
- -
- Setting the optimization of the FRF algorithm parameters for the best identification of classes of the testing dataset;
- -
- Testing the execution of the FRF algorithm’s detection and estimation of anomalous areas by applying, after the analysis, image threshold filters (for the calculus, all the images have the same dimension of 1000 pixels × 2000 pixels);
- -
- Verification of the algorithm performance by estimating its precision.
3. Results
4. Discussion
Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A

Appendix B

References
- Turing, A.M. I.—Computing Machinery and Intelligence. Mind 1950, LIX, 433–460. [Google Scholar] [CrossRef]
- Bjola, C. AI for development: Implications for theory and practice. Oxf. Dev. Stud. 2022, 50, 78–90. [Google Scholar] [CrossRef]
- Venerito, V.; Angelini, O.; Cazzato, G.; Lopalco, G.; Maiorano, E.; Cimmino, A.; Iannone, F. A convolutional neural network with transfer learning for automatic discrimination between low and high-grade synovitis: A pilot study. Intern. Emerg. Med. 2021, 16, 1457–1465. [Google Scholar] [CrossRef] [PubMed]
- Onega, T.; Reisch, L.M.; Frederick, P.D.; Geller, B.M.; Nelson, H.D.; Lott, J.P.; Radick, A.C.; Elder, D.E.; Barnhill, R.L.; Piepkorn, M.W.; et al. Use of digital whole slide imaging in dermatopathology. J. Digit. Imaging 2016, 29, 243–253. [Google Scholar] [CrossRef] [PubMed]
- Massaro, A. Electronic in Advanced Research Industry: From Industry 4.0 to Industry 5.0 Advances; Wiley: Hoboken, NJ, USA; IEEE: New York, NY, USA, 2021; ISBN 9781119716877. [Google Scholar]
- Allen, A.C.; Spitz, S. Malignant melanoma; a clinicopathological analysis of the criteria for diagnosis and prognosis. Cancer 1953, 6, 1–45. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Mahapatra, D. Analyzing training information from random forests for Improved image segmentation. IEEE Trans. Image Process. 2014, 23, 1504–1512. [Google Scholar] [CrossRef] [PubMed]
- Belgiu, M.; Drăguţ, L. Random forest in remote sensing: A review of applications and future directions. ISPRS J. Photogramm. Remote Sens. 2016, 114, 24–31. [Google Scholar] [CrossRef]
- Arganda-Carreras, I.; Kaynig, V.; Rueden, C.; Eliceiri, K.W.; Schindelin, J.; Cardona, A.; Seung, H.S. Trainable Weka segmentation: A machine learning tool for microscopy pixel classification. Bioinformatics 2017, 33, 2424–2426. [Google Scholar] [CrossRef] [PubMed]
- Lormand, C.; Zellmer, G.F.; Németh, K.; Kilgour, G.; Measd, S.; Palmer, A.S.; Sakamoto, N.; Yurimoto, H.; Moebis, A. Weka trainable segmentation plugin in ImageJ: A semi-automatic tool applied to crystal size distributions of microlites in volcanic rocks. Microsc. Microanal. 2018, 24, 667–675. [Google Scholar] [CrossRef] [PubMed]
- Alessandro, M. Detection of Anomalies in Images by Machine Learning Fast Random Forest—FRF-Algorithm; Zenodo: Geneva, Switzerland, 2022. [Google Scholar] [CrossRef]
- Lay-Ekuakille, A.; Spano, F.; Mvemba, P.K.; Massaro, A.; Galiano, A.; Casciaro, S.; Conversano, F. Conductivity image characterization of gold nanoparticles based-device through atomic force microscopy. In Proceedings of the IEEE 2018 Nanotechnology for Instrumentation and Measurement (NANOfIM), Mexico City, Mexico, 7–8 November 2018; IEEE: Piscataway, NJ, USA, 2019. [Google Scholar]
- Roush, G.C.; Kirkwood, J.M.; Ernstoff, M.; Somma, S.J.; Duray, P.H.; Klaus, S.N.; Stenn, K.S.; Titus, L.J.; Lerner, A.B. Reproducibility and validity in the clinical diagnosis of the nonfamilial dysplastic nevus: Work in progress. Recent Results Cancer Res. 1986, 102, 154–158. [Google Scholar] [CrossRef] [PubMed]
- Cazzato, G.; Colagrande, A.; Cimmino, A.; Demarco, A.; Lospalluti, L.; Arezzo, F.; Resta, L.; Ingravallo, G. The Great Mime: Three Cases of Melanoma with Carcinoid-Like and Paraganglioma-Like Pattern with Emphasis on Differential Diagnosis. Dermatopathology 2021, 8, 130–134. [Google Scholar] [CrossRef] [PubMed]
- Elmore, J.G.; Barnhill, R.L.; Elder, D.E.; Longton, G.M.; Pepe, M.S.; Reisch, L.M.; Carney, P.A.; Titus, L.J.; Nelson, H.D.; Onega, T.; et al. Pathologists’ diagnosis of invasive melanoma and melanocytic proliferations: Observer accuracy and reproducibility study. BMJ 2017, 357, j2813. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.H.; Gilbertson, J.R.; Hanna, M.G.; Olson, N.H.; Seheult, J.N.; Sorace, J.M.; Stram, M.N. Introduction to Artificial Intelligence and Machine Learning for Pathology. Arch. Pathol. Lab. Med. 2021, 145, 1228–1254. [Google Scholar] [CrossRef] [PubMed]
- Hekler, A.; Utikal, J.S.; Enk, A.H.; Berking, C.; Klode, J.; Schadendorf, D.; Jansen, P.; Franklin, C.; Holland-Letz, T.; Krahl, D.; et al. Pathologist-level classification of histopathological melanoma images with deep neural networks. Eur. J. Cancer 2019, 115, 79–83. [Google Scholar] [CrossRef] [PubMed]
- Hart, S.N.; Flotte, W.; Norgan, A.P.; Shah, K.K.; Buchan, Z.R.; Mounajjed, T.; Flotte, T.J. Classification of Melanocytic Lesions in Selected and Whole-Slide Images via Convolutional Neural Networks. J. Pathol. Inform. 2019, 10, 5. [Google Scholar] [CrossRef] [PubMed]
- Witten, I.H.; Frank, E.; Hall, M.A.; Pal, C.J. Data Mining, 4th ed.; Morgan Kaufmann: Burlington, MA, USA, 2017; ISBN 9780128042915. [Google Scholar] [CrossRef]
- Raza, R.; Zulfiqar, F.; Tariq, S.; Anwar, G.B.; Sargano, A.B.; Habib, Z. Melanoma Classification from Dermoscopy Images Using Ensemble of Convolutional Neural Networks. Mathematics 2022, 10, 26. [Google Scholar] [CrossRef]
- Argenziano, G.; Soyer, H.P.; Chimenti, S.; Talamini, R.; Corona, R.; Sera, F.; Binder, M.; Cerroni, L.; De Rosa, G.; Ferrara, G.; et al. Dermoscopy of pigmented skin lesions: Results of a consensus meeting via the Internet. J. Am. Acad. Dermatol. 2003, 48, 679–693. [Google Scholar] [CrossRef] [PubMed]
- Kłóś, A.; Płonka, P.M. Growth of mixed cancer cell population—In silico the size matters. Acta Biochim. Pol. 2020, 67, 453–463. [Google Scholar] [CrossRef] [PubMed]

| Dysplastic Nevus | Malignant Melanoma |
|---|---|
| Architectural criteria (mandatory, major) | |
| Lentiginous or contiguous melanocytic hyperplasia | Poor circumscription of the intraepidermal melanocytic component of the lesion |
| Focal melanocytic atypia | Increased number of melanocytes, solitary and in nests, within and above the epidermal basal cell layer and within adnexal epithelia (pagetoid spreading) |
| Marked variation in size and shape of the melanocytic nests | |
| Absence of maturation of melanocytes with descent into the dermis | |
| Melanocytes in mitosis | |
| Architectural criteria (minor, at least 2) | |
| “Shoulder phenomenon” | Melanocytes with nuclear atypia |
| Fusion of epithelial cones | Necrosis or degeneration of melanocytes |
| Subepidermal concentric lamellar fibrosis |
| Original Image | Defect Type (Name) | Defect Cluster (Enhanced Probability Image) | Percentage Presence on the Whole Image | Extension [mm2] |
|---|---|---|---|---|
| IMG00131 EE | // |  | 5.3% | 0.106 |
| IMG00132 EE | // |  | 4.1% | 0.082 |
| Original Image | Defect Type (Name) | Defect Cluster (Enhanced Probability Image) | Percentage Presence on the Whole Image | Extension [mm2] |
|---|---|---|---|---|
| IMG00150 | // |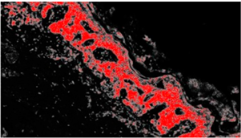  | 6.6% | 0.132 |
| IMG00151 | // |  | 8.8% | 0.176 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cazzato, G.; Massaro, A.; Colagrande, A.; Lettini, T.; Cicco, S.; Parente, P.; Nacchiero, E.; Lospalluti, L.; Cascardi, E.; Giudice, G.; et al. Dermatopathology of Malignant Melanoma in the Era of Artificial Intelligence: A Single Institutional Experience. Diagnostics 2022, 12, 1972. https://doi.org/10.3390/diagnostics12081972
Cazzato G, Massaro A, Colagrande A, Lettini T, Cicco S, Parente P, Nacchiero E, Lospalluti L, Cascardi E, Giudice G, et al. Dermatopathology of Malignant Melanoma in the Era of Artificial Intelligence: A Single Institutional Experience. Diagnostics. 2022; 12(8):1972. https://doi.org/10.3390/diagnostics12081972
Chicago/Turabian StyleCazzato, Gerardo, Alessandro Massaro, Anna Colagrande, Teresa Lettini, Sebastiano Cicco, Paola Parente, Eleonora Nacchiero, Lucia Lospalluti, Eliano Cascardi, Giuseppe Giudice, and et al. 2022. "Dermatopathology of Malignant Melanoma in the Era of Artificial Intelligence: A Single Institutional Experience" Diagnostics 12, no. 8: 1972. https://doi.org/10.3390/diagnostics12081972
APA StyleCazzato, G., Massaro, A., Colagrande, A., Lettini, T., Cicco, S., Parente, P., Nacchiero, E., Lospalluti, L., Cascardi, E., Giudice, G., Ingravallo, G., Resta, L., Maiorano, E., & Vacca, A. (2022). Dermatopathology of Malignant Melanoma in the Era of Artificial Intelligence: A Single Institutional Experience. Diagnostics, 12(8), 1972. https://doi.org/10.3390/diagnostics12081972














