An Embedded System Using Convolutional Neural Network Model for Online and Real-Time ECG Signal Classification and Prediction
Abstract
:1. Introduction
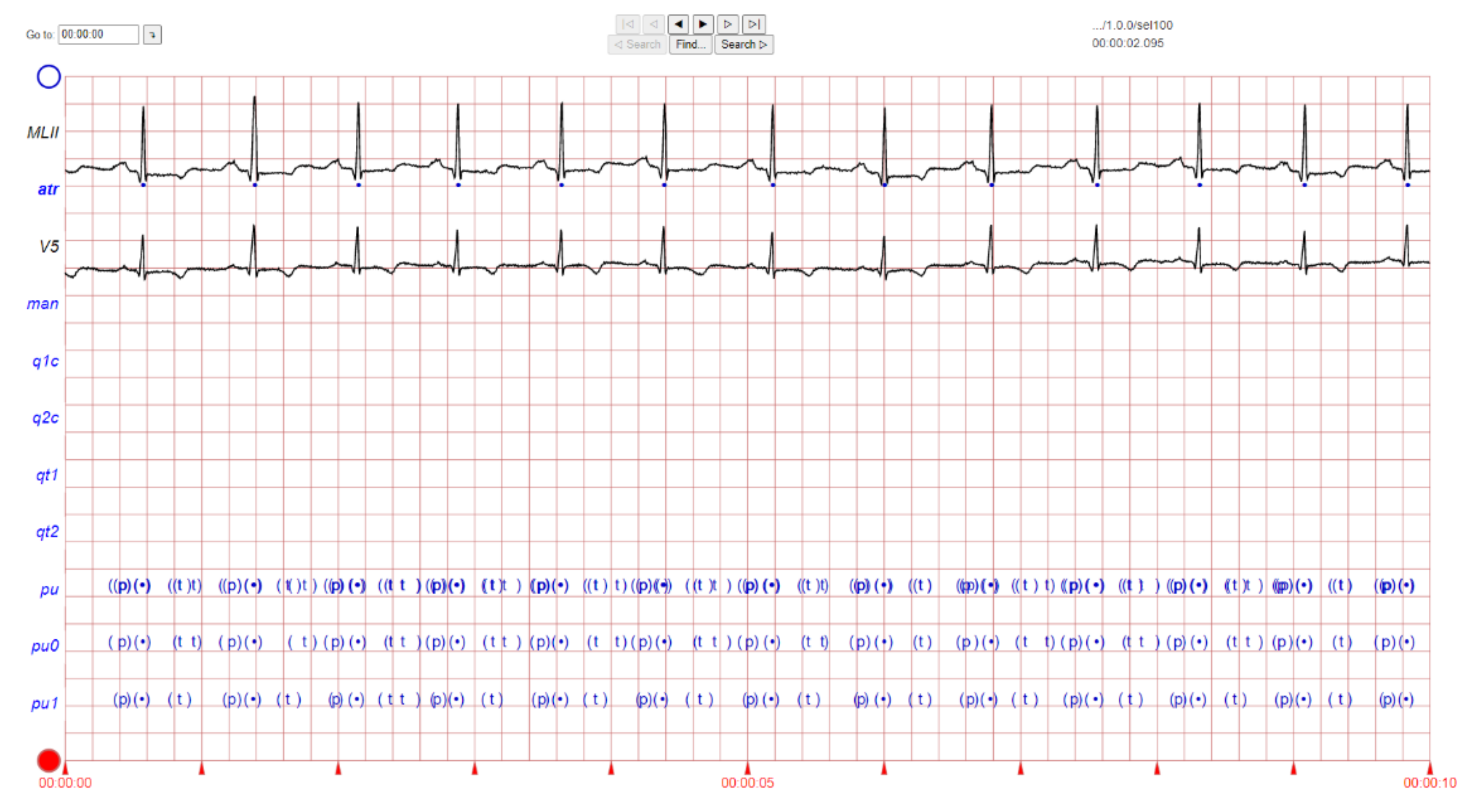
2. ECG Image Data and Preprocessing Method
3. Convolutional Neural Network (CNN) Method and Data Processing Stage
4. Results and Discussion
4.1. Hyperparameter Settings
4.2. Training and Validation Performance
4.3. CNN Prediction
5. Development of Online and Real-Time ECG Image Classification and Prediction
6. Limitations
7. Future Work
8. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Dutta, A.; Batabyal, T.; Basu, M.; Acton, S.T. An efficient convolutional neural network for coronary heart disease prediction. Expert Syst. Appl. 2020, 159, 113408. [Google Scholar] [CrossRef]
- Pursnani, S.; Merchant, M. South Asian ethnicity as a risk factor for coronary heart disease. Atherosclerosis 2020, 315, 126–130. [Google Scholar] [CrossRef]
- Venkatesan, C.; Karthigaikumar, P.; Paul, A.; Satheeskumaran, S.; Kumar, R. ECG Signal Preprocessing and SVM Classifier-Based Abnormality Detection in Remote Healthcare Applications. IEEE Access 2018, 6, 9767–9773. [Google Scholar] [CrossRef]
- Hagiwara, Y.; Fujita, H.; Oh, S.L.; Tan, J.H.; Tan, R.S.; Ciaccio, E.J.; Acharya, U.R. Computer-aided diagnosis of atrial fibrillation based on ECG Signals: A review. Inf. Sci. 2018, 467, 99–114. [Google Scholar] [CrossRef]
- Banerjee, R.; Ghose, A.; Khandelwal, S. A Novel Recurrent Neural Network Architecture for Classification of Atrial Fibrillation Using Single-lead ECG. In Proceedings of the 2019 27th European Signal Processing Conference (EUSIPCO), A Coruna, Spain, 2–6 September 2019; pp. 1–5. [Google Scholar]
- Faust, O.; Shenfield, A.; Kareem, M.; San, T.R.; Fujita, H.; Acharya, U.R. Automated detection of atrial fibrillation using long short-term memory network with RR interval signals. Comput. Biol. Med. 2018, 102, 327–335. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, C.; Yan, Y.; Zhou, L.; Bai, J.; Wang, L. Automated atrial fibrillation detection based on deep learning network. In Proceedings of the 2016 IEEE International Conference on Information and Automation (ICIA), Ningbo, China, 1–3 August 2016; pp. 1159–1164. [Google Scholar]
- Gawali, D.H.; Wadhai, V.M. Implementation of ECG sensor for real time signal processing applications. In Proceedings of the 2014 International Conference on Advances in Electronics Computers and Communications, Bangalore, India, 10–11 October 2014; pp. 1–3. [Google Scholar]
- Jeon, T.; Kim, B.; Jeon, M.; Lee, B.-G. Implementation of a portable device for real-time ECG signal analysis. Biomed. Eng. Online 2014, 13, 160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sadhukhan, D.; Pal, S.; Mitra, M. Automated Identification of Myocardial Infarction Using Harmonic Phase Distribution Pattern of ECG Data. IEEE Trans. Instrum. Meas. 2018, 67, 2303–2313. [Google Scholar] [CrossRef]
- Celin, S.; Vasanth, K. ECG Signal Classification Using Various Machine Learning Techniques. J. Med. Syst. 2018, 42, 241. [Google Scholar] [CrossRef]
- Shameer, K.; Johnson, K.; Glicksberg, B.; Dudley, J.T.; Sengupta, P.P. Machine learning in cardiovascular medicine: Are we there yet? Heart 2018, 104, 1156–1164. [Google Scholar] [CrossRef]
- Sameni, R.; Shamsollahi, M.B.; Jutten, C.; Clifford, G.D. A Nonlinear Bayesian Filtering Framework for ECG Denoising. IEEE Trans. Biomed. Eng. 2007, 54, 2172–2185. [Google Scholar] [CrossRef] [Green Version]
- Asl, B.M.; Setarehdan, S.K.; Mohebbi, M. Support vector machine-based arrhythmia classification using reduced features of heart rate variability signal. Artif. Intell. Med. 2008, 44, 51–64. [Google Scholar] [CrossRef]
- Oyeleye, M.; Chen, T.; Titarenko, S.; Antoniou, G. A Predictive Analysis of Heart Rates Using Machine Learning Techniques. Int. J. Environ. Res. Public Health 2022, 19, 2417. [Google Scholar] [CrossRef] [PubMed]
- Krittanawong, C.; Zhang, H.; Wang, Z.; Aydar, M.; Kitai, T. Artificial Intelligence in Precision Cardiovascular Medicine. J. Am. Coll. Cardiol. 2017, 69, 2657–2664. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.B.; Nguyen, M.N.; San, P.P.; Li, X.L.; Krishnaswamy, S. Deep convolutional neural networks on multichannel time series for human activity recognition. In Proceedings of the Twenty-Fourth International Joint Conference on Artificial Intelligence, Buenos Aires, Argentina, 25–31 July 2015. [Google Scholar]
- Khan, S.; Yairi, T. A review on the application of deep learning in system health management. Mech. Syst. Signal Process. 2018, 107, 241–265. [Google Scholar] [CrossRef]
- Darmawahyuni, A.; Nurmaini, S.; Sukemi; Caesarendra, W.; Bhayyu, V.; Rachmatullah, M.N. Firdaus Deep Learning with a Recurrent Network Structure in the Sequence Modeling of Imbalanced Data for ECG-Rhythm Classifier. Algorithms 2019, 12, 118. [Google Scholar] [CrossRef] [Green Version]
- Ni, A.; Azarang, A.; Kehtarnavaz, N. A Review of Deep Learning-Based Contactless Heart Rate Measurement Methods. Sensors 2021, 21, 3719. [Google Scholar] [CrossRef]
- Staffini, A.; Svensson, T.; Chung, U.-I.; Svensson, A.K. Heart Rate Modeling and Prediction Using Autoregressive Models and Deep Learning. Sensors 2021, 22, 34. [Google Scholar] [CrossRef]
- Wang, J.; Chen, Y.; Hao, S.; Peng, X.; Hu, L. Deep learning for sensor-based activity recognition: A survey. Pattern Recognit. Lett. 2019, 119, 3–11. [Google Scholar] [CrossRef] [Green Version]
- Cheng, C.-H.; Wong, K.-L.; Chin, J.-W.; Chan, T.-T.; So, R.H.Y. Deep Learning Methods for Remote Heart Rate Measurement: A Review and Future Research Agenda. Sensors 2021, 21, 6296. [Google Scholar] [CrossRef]
- Massaro, A.; Maritati, V.; Savino, N.; Galiano, A. Neural Networks for Automated Smart Health Platforms oriented on Heart Predictive Diagnostic Big Data Systems. In Proceedings of the 2018 AEIT International Annual Conference, Bari, Italy, 3–5 October 2018. [Google Scholar]
- Massaro, A.; Ricci, G.; Selicato, S.; Raminelli, S.; Galiano, A. Decisional Support System with Artificial Intelligence oriented on Health Prediction using a Wearable Device and Big Data. In Proceedings of the 2020 IEEE International Workshop on Metrology for Industry 4.0 & IoT, Rome, Italy, 3–5 June 2020. [Google Scholar]
- LeCun, Y.; Bengio, Y. Convolutional networks for images, speech, and time series. Handb. Brain Theory Neural Netw. 2019, 3361, 1995. [Google Scholar]
- Acharya, U.R.; Fujita, H.; Lih, O.S.; Hagiwara, Y.; Tan, J.H.; Adam, M. Automated detection of arrhythmias using different intervals of tachycardia ECG segments with convolutional neural network. Inf. Sci. 2017, 405, 81–90. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farrukh, F.U.D.; Xie, T.; Zhang, C.; Wang, Z. Optimization for Efficient Hardware Implementation of CNN on FPGA. In Proceedings of the 2018 IEEE International Conference on Integrated Circuits, Technologies and Applications (ICTA), Beijing, China, 21–23 November 2018; pp. 88–89. [Google Scholar]
- Li, Q.; Cai, W.; Wang, X.; Zhou, Y.; Feng, D.D.; Chen, M. Medical image classification with convolutional neural network. In Proceedings of the 2014 13th International Conference on Control Automation Robotics & Vision (ICARCV), Singapore, 10–12 December 2014; pp. 844–848. [Google Scholar]
- Caesarendra, W.; Triwiyanto, T.; Pandiyan, V.; Glowacz, A.; Permana, S.; Tjahjowidodo, T. A CNN Prediction Method for Belt Grinding Tool Wear in a Polishing Process Utilizing 3-Axes Force and Vibration Data. Electronics 2021, 10, 1429. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. In Proceedings of the 2015 3rd International Conference on Learning Representations (ICLR), San Diego, CA, USA, 7–9 May 2015; pp. 1–14. [Google Scholar]
- Kulathunga, N.; Ranasinghe, N.; Vrinceanu, D.; Kinsman, Z.; Huang, L.; Wang, Y. Effects of Nonlinearity and Network Architecture on the Performance of Supervised Neural Networks. Algorithms 2021, 14, 51. [Google Scholar] [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning, 1st ed.; MIT Press: Cambridge, MA, USA, 2016; pp. 255–265. [Google Scholar]
- Lam, C.S. Heart failure in Southeast Asia: Facts and numbers. ESC Hear. Fail. 2015, 2, 46–49. [Google Scholar] [CrossRef]
- Pan, Q.; Li, X.; Fang, L. Data augmentation for deep learning-based ECG analysis. In Feature Engineering and Computational Intelligence in ECG Monitoring; Liu, C., Ed.; Springer: Singapore, 2020; pp. 91–111. [Google Scholar]
- Vella, M.; Mota, J.F.C. Overcoming Measurement Inconsistency in Deep Learning for Linear Inverse Problems: Applications in Medical Imaging. In Proceedings of the 2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Toronto, ON, Canada, 6–11 June 2021. [Google Scholar]
- Guha, I.; Nadeem, S.A.; You, C.; Zhang, X.; Levy, S.M.; Wang, G.; Torner, J.C.; Saha, P.K. Deep learning based high-resolution reconstruction of trabecular bone microstructures from low-resolution CT scans using GAN-CIRCLE. In Proceedings of the SPIE-the International Society for Optical Engineering, Houston, TX, USA, 28 February 2020; Volume 11317. [Google Scholar]
- Oksuz, I.; Clough, J.R.; Ruijsink, B.; Anton, E.P.; Bustin, A.; Cruz, G.; Prieto, C.; King, A.P.; Schnabel, J.A. Deep Learning-Based Detection and Correction of Cardiac MR Motion Artefacts During Reconstruction for High-Quality Segmentation. IEEE Trans. Med Imaging 2020, 39, 4001–4010. [Google Scholar] [CrossRef]
- Pham, D.L.; Xu, C.; Prince, J. Current Methods in Medical Image Segmentation. Annu. Rev. Biomed. Eng. 2000, 2, 315–337. [Google Scholar] [CrossRef]
- Wu, J.; Chen, X.; Zhang, H.; Xiong, L.D.; Lei, H.; Deng, S. Hyperparameter Optimization for Machine Learning Models Based on Bayesian Optimization. J. Electron. Sci. Technol. 2019, 17, 26–40. [Google Scholar]
- Howard, A.; Sandler, M.; Chen, B.; Wang, W.; Chen, L.-C.; Tan, M.; Chu, G.; Vasudevan, V.; Zhu, Y.; Pang, R.; et al. Searching for MobileNetV3. In Proceedings of the 2019 IEEE/CVF International Conference on Computer Vision (ICCV), Seoul, Korea, 27 October–2 November 2019; pp. 1314–1324. [Google Scholar] [CrossRef]

















| Layer (Type) | Output Shape | Parameter # |
|---|---|---|
| Flatten | (None, 25,088) | 0 |
| Fully Connected Layer | (None, 256) | 6,422,784 |
| LeakyReLU | (None, 256) | 0 |
| Dropout Layer (p = 0.5) | (None, 256) | 0 |
| Fully Connected Layer | (None, 256) | 65,792 |
| LeakyReLU | (None, 256) | 0 |
| Dropout Layer (p = 0.3) | (None, 256) | 0 |
| Fully Connected Layer | (None, 4) | 1028 |
| Total parameters: 6,489,604 | ||
| Trainable parameters: 6,489,604 | ||
| Non-trainable parameters: 0 | ||
| Training Parameters | Description/Values |
|---|---|
| Optimizer | ‘Adam’ |
| Loss | cross entropy |
| Mini-Batch Size | 16 |
| Epochs | 100 |
| Training dataset | 80% |
| Validation dataset | 20% |
| Class | Precision | Recall | F1-Score | Accuracy | Loss |
|---|---|---|---|---|---|
| Arrhythmia | 0.99 | 0.95 | 0.97 | 0.9596 | 0.0859 |
| Normal sinus | 0.99 | 0.93 | 0.95 | ||
| Sudden death | 1.00 | 1.00 | 1.00 | ||
| Supraventricular arrhythmia | 0.91 | 0.96 | 0.93 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caesarendra, W.; Hishamuddin, T.A.; Lai, D.T.C.; Husaini, A.; Nurhasanah, L.; Glowacz, A.; Alfarisy, G.A.F. An Embedded System Using Convolutional Neural Network Model for Online and Real-Time ECG Signal Classification and Prediction. Diagnostics 2022, 12, 795. https://doi.org/10.3390/diagnostics12040795
Caesarendra W, Hishamuddin TA, Lai DTC, Husaini A, Nurhasanah L, Glowacz A, Alfarisy GAF. An Embedded System Using Convolutional Neural Network Model for Online and Real-Time ECG Signal Classification and Prediction. Diagnostics. 2022; 12(4):795. https://doi.org/10.3390/diagnostics12040795
Chicago/Turabian StyleCaesarendra, Wahyu, Taufiq Aiman Hishamuddin, Daphne Teck Ching Lai, Asmah Husaini, Lisa Nurhasanah, Adam Glowacz, and Gusti Ahmad Fanshuri Alfarisy. 2022. "An Embedded System Using Convolutional Neural Network Model for Online and Real-Time ECG Signal Classification and Prediction" Diagnostics 12, no. 4: 795. https://doi.org/10.3390/diagnostics12040795
APA StyleCaesarendra, W., Hishamuddin, T. A., Lai, D. T. C., Husaini, A., Nurhasanah, L., Glowacz, A., & Alfarisy, G. A. F. (2022). An Embedded System Using Convolutional Neural Network Model for Online and Real-Time ECG Signal Classification and Prediction. Diagnostics, 12(4), 795. https://doi.org/10.3390/diagnostics12040795







