Efficient Staining-Invariant Nuclei Segmentation Approach Using Self-Supervised Deep Contrastive Network
Abstract
1. Introduction
- Proposing an efficient nuclei segmentation method for hematoxylin and eosin (H&E) WSI images using a deep staining-invariant self-supervised contrastive network. This method eliminates the need for a stain normalization step;
- Proposing an effective weighted hybrid dilated convolutional (WHDC) block that helps extract multi-scale nuclei-relevant representations;
- Achieving accurate nuclei segmentation on unseen single-organ and multi-organ datasets collected from different laboratories without employing stain color normalization or fine-tuning that demonstrate the proposed method’s generalization capabilities.
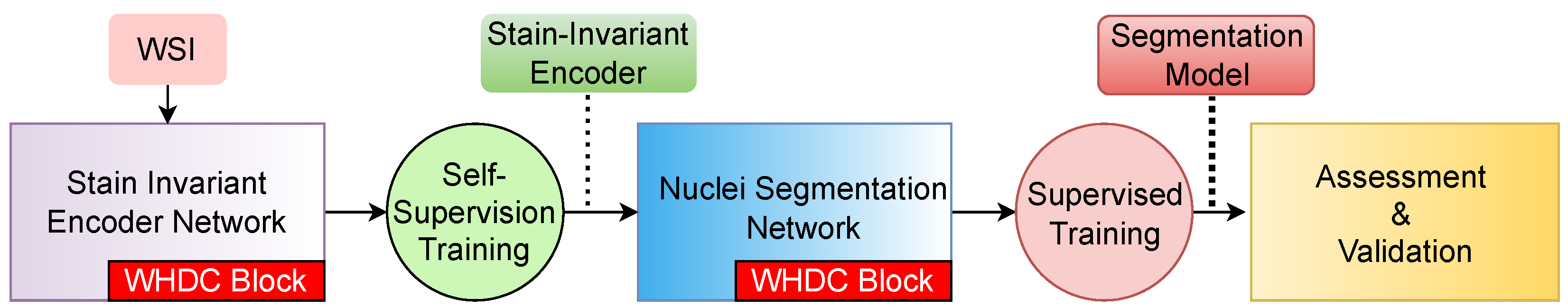
2. Proposed Method
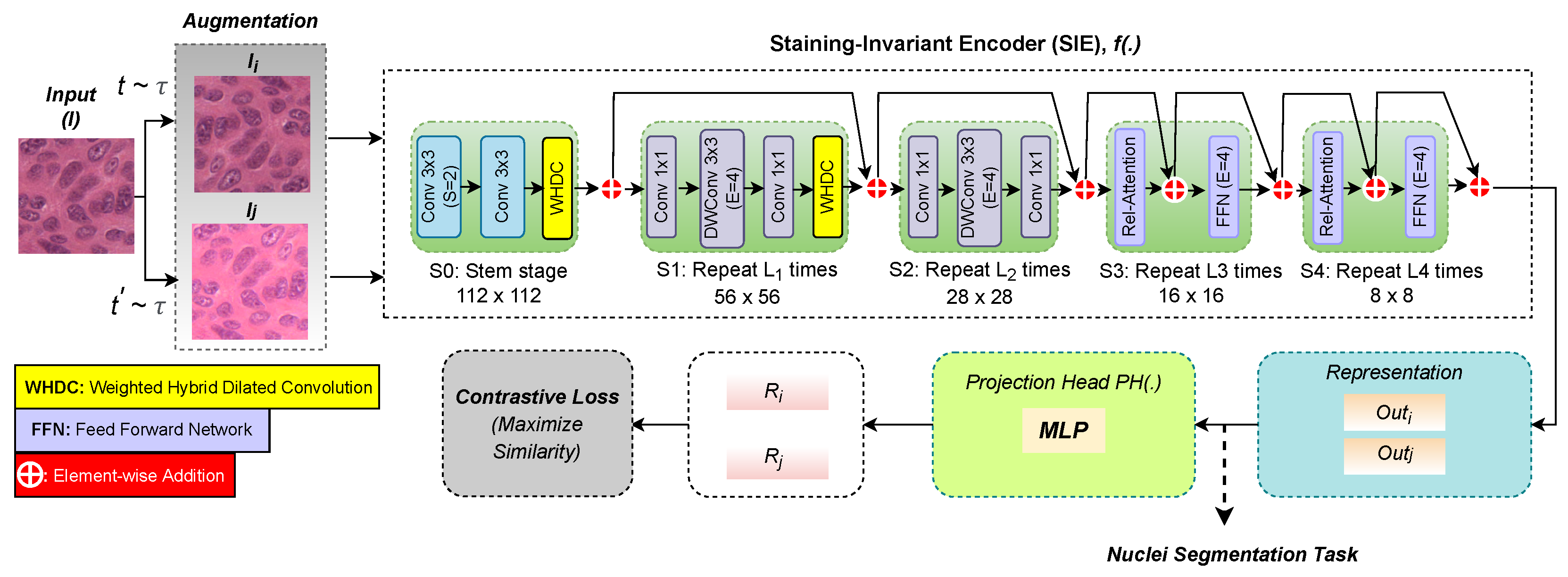
2.1. Staining-Invariant Encoder
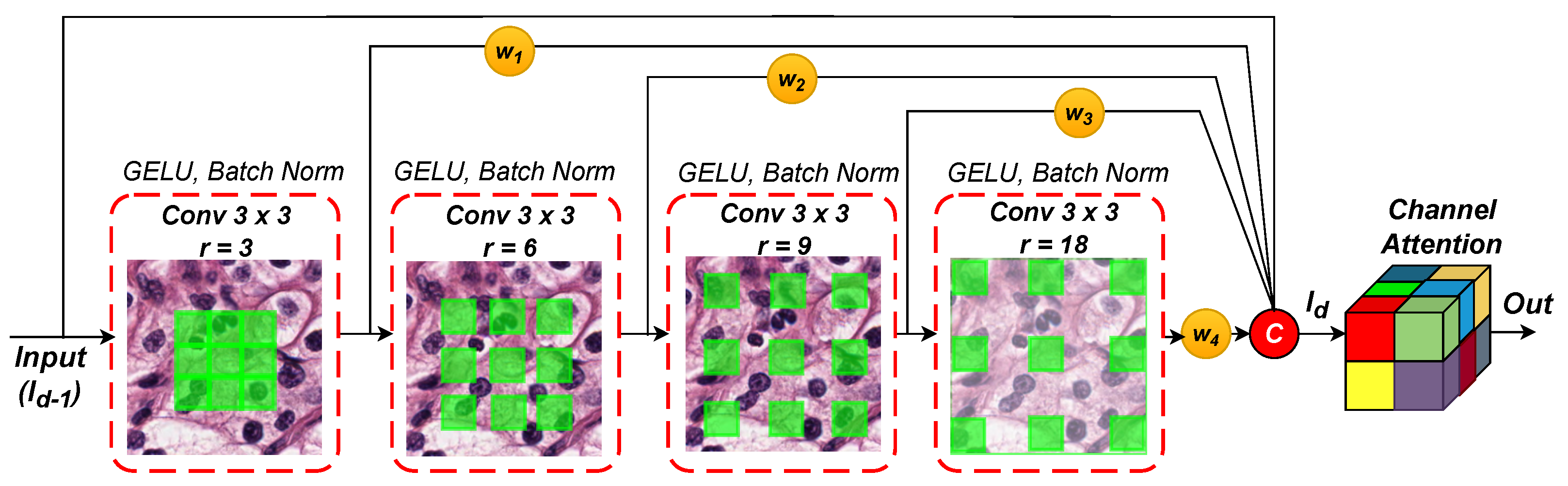
2.2. Weighted Hybrid Dilated Convolution (WHDC) Block
2.3. Nuclei Segmentation Network
3. Results and Discussion
3.1. Datasets
3.2. Implementation Details
3.3. Evaluation Metrics
3.4. Ablation Study
3.4.1. Analysis of Various Configurations
3.4.2. Analysis of the Loss Function
3.5. Comparison with Existing Methods
3.6. Evaluating the Proposed Method on Other Datasets
3.7. Discussion and Limitations
4. Conclusions and Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hanna, M.G.; Parwani, A.; Sirintrapun, S.J. Whole slide imaging: Technology and applications. Adv. Anat. Pathol. 2020, 27, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Roullier, V.; Lézoray, O.; Ta, V.T.; Elmoataz, A. Multi-resolution graph-based analysis of histopathological whole slide images: Application to mitotic cell extraction and visualization. Comput. Med. Imaging Graph. 2011, 35, 603–615. [Google Scholar] [CrossRef] [PubMed]
- Doyle, S.; Madabhushi, A.; Feldman, M.; Tomaszeweski, J. A boosting cascade for automated detection of prostate cancer from digitized histology. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Copenhagen, Denmark, 1–6 October 2006; Springer: Berlin/Heidelberg, Germany, 2006; pp. 504–511. [Google Scholar]
- Janowski, P.; Strzelecki, M.; Brzezinska-Blaszczyk, E.; Zalewska, A. Computer analysis of normal and basal cell carcinoma mast cells. Med. Sci. Monit. 2001, 7, 260–265. [Google Scholar] [PubMed]
- Lu, C.; Ma, Z.; Mandal, M. Automated segmentation of the epidermis area in skin whole slide histopathological images. IET Image Process. 2015, 9, 735–742. [Google Scholar] [CrossRef]
- Kleczek, P.; Jaworek-Korjakowska, J.; Gorgon, M. A novel method for tissue segmentation in high-resolution H&E-stained histopathological whole-slide images. Comput. Med. Imaging Graph. 2020, 79, 101686. [Google Scholar]
- Kłeczek, P.; Dyduch, G.; Jaworek-Korjakowska, J.; Tadeusiewicz, R. Automated epidermis segmentation in histopathological images of human skin stained with hematoxylin and eosin. In Proceedings of the Medical Imaging 2017: Digital Pathology, Orlando, FL, USA, 11–16 February 2017; Volume 10140, pp. 140–158. [Google Scholar]
- Wu, Y.; Cheng, M.; Huang, S.; Pei, Z.; Zuo, Y.; Liu, J.; Yang, K.; Zhu, Q.; Zhang, J.; Hong, H.; et al. Recent Advances of Deep Learning for Computational Histopathology: Principles and Applications. Cancers 2022, 14, 1199. [Google Scholar] [CrossRef]
- Bándi, P.; van de Loo, R.; Intezar, M.; Geijs, D.; Ciompi, F.; van Ginneken, B.; van der Laak, J.; Litjens, G. Comparison of different methods for tissue segmentation in histopathological whole-slide images. In Proceedings of the 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, Australia, 18–21 April 2017; pp. 591–595. [Google Scholar]
- Wang, Z. A new approach for segmentation and quantification of cells or nanoparticles. IEEE Trans. Ind. Inform. 2016, 12, 962–971. [Google Scholar] [CrossRef]
- Zhao, M.; Wang, H.; Han, Y.; Wang, X.; Dai, H.N.; Sun, X.; Zhang, J.; Pedersen, M. Seens: Nuclei segmentation in pap smear images with selective edge enhancement. Future Gener. Comput. Syst. 2021, 114, 185–194. [Google Scholar] [CrossRef]
- Lagree, A.; Mohebpour, M.; Meti, N.; Saednia, K.; Lu, F.I.; Slodkowska, E.; Gandhi, S.; Rakovitch, E.; Shenfield, A.; Sadeghi-Naini, A.; et al. A review and comparison of breast tumor cell nuclei segmentation performances using deep convolutional neural networks. Sci. Rep. 2021, 11, 8025. [Google Scholar] [CrossRef]
- Cui, Y.; Zhang, G.; Liu, Z.; Xiong, Z.; Hu, J. A deep learning algorithm for one-step contour aware nuclei segmentation of histopathology images. Med. Biol. Eng. Comput. 2019, 57, 2027–2043. [Google Scholar] [CrossRef]
- Xie, L.; Qi, J.; Pan, L.; Wali, S. Integrating deep convolutional neural networks with marker-controlled watershed for overlapping nuclei segmentation in histopathology images. Neurocomputing 2020, 376, 166–179. [Google Scholar] [CrossRef]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. UNet++: Redesigning Skip Connections to Exploit Multiscale Features in Image Segmentation. IEEE Trans. Med. Imaging 2020, 39, 1856–1867. [Google Scholar] [CrossRef] [PubMed]
- Ilyas, T.; Mannan, Z.I.; Khan, A.; Azam, S.; Kim, H.; De Boer, F. TSFD-Net: Tissue specific feature distillation network for nuclei segmentation and classification. Neural Netw. 2022, 115, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Kiran, I.; Raza, B.; Ijaz, A.; Khan, M.A. DenseRes-Unet: Segmentation of overlapped/clustered nuclei from multi organ histopathology images. Comput. Biol. Med. 2022, 143, 105267. [Google Scholar] [CrossRef] [PubMed]
- Rączkowski, Ł.; Możejko, M.; Zambonelli, J.; Szczurek, E. ARA: Accurate, reliable and active histopathological image classification framework with Bayesian deep learning. Sci. Rep. 2019, 9, 14347. [Google Scholar] [CrossRef] [PubMed]
- Hassan, L.; Abdel-Nasser, M.; Saleh, A.; A Omer, O.; Puig, D. Efficient Stain-Aware Nuclei Segmentation Deep Learning Framework for Multi-Center Histopathological Images. Electronics 2021, 10, 954. [Google Scholar] [CrossRef]
- Ciga, O.; Xu, T.; Martel, A.L. Self supervised contrastive learning for digital histopathology. Mach. Learn. Appl. 2022, 7, 100198. [Google Scholar] [CrossRef]
- Ye, H.L.; Wang, D.H. Stain-Adaptive Self-Supervised Learning for Histopathology Image Analysis. arXiv 2022, arXiv:2208.04017. [Google Scholar]
- Boserup, N.; Selvan, R. Efficient Self-Supervision using Patch-based Contrastive Learning for Histopathology Image Segmentation. arXiv 2022, arXiv:2208.10779. [Google Scholar]
- Dai, Z.; Liu, H.; Le, Q.; Tan, M. Coatnet: Marrying convolution and attention for all data sizes. Adv. Neural Inf. Process. Syst. 2021, 34, 3965–3977. [Google Scholar]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the International Conference on Machine Learning, PMLR, Online, 26–28 August 2020; pp. 1597–1607. [Google Scholar]
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7132–7141. [Google Scholar]
- Oktay, O.; Schlemper, J.; Folgoc, L.L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.Y.; Kainz, B.; et al. Attention U-Net: Learning where to look for the pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar]
- Graham, S.; Jahanifar, M.; Azam, A.; Nimir, M.; Tsang, Y.W.; Dodd, K.; Hero, E.; Sahota, H.; Tank, A.; Benes, K.; et al. Lizard: A Large-Scale Dataset for Colonic Nuclear Instance Segmentation and Classification. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada, 10–17 October 2021; pp. 684–693. [Google Scholar]
- Wei, J.; Suriawinata, A.; Ren, B.; Liu, X.; Lisovsky, M.; Vaickus, L.; Brown, C.; Baker, M.; Tomita, N.; Torresani, L.; et al. A Petri Dish for Histopathology Image Analysis. In Proceedings of the International Conference on Artificial Intelligence in Medicine, Virtual Event, 15–18 June 2021; Springer: Berlin/Heidelberg, Germany, 2021; pp. 11–24. [Google Scholar]
- Aksac, A.; Demetrick, D.J.; Ozyer, T.; Alhajj, R. BreCaHAD: A dataset for breast cancer histopathological annotation and diagnosis. BMC Res. Notes 2019, 12, 82. [Google Scholar] [CrossRef] [PubMed]
- Petrick, N.; Akbar, S.; Cha, K.H.; Nofech-Mozes, S.; Sahiner, B.; Gavrielides, M.A.; Kalpathy-Cramer, J.; Drukker, K.; Martel, A.L.; BreastPathQ Challenge Group. SPIE-AAPM-NCI BreastPathQ Challenge: An image analysis challenge for quantitative tumor cellularity assessment in breast cancer histology images following neoadjuvant treatment. J. Med. Imaging 2021, 8, 034501. [Google Scholar] [CrossRef] [PubMed]
- Kather, J.N.; Krisam, J.; Charoentong, P.; Luedde, T.; Herpel, E.; Weis, C.A.; Gaiser, T.; Marx, A.; Valous, N.A.; Ferber, D.; et al. Predicting survival from colorectal cancer histology slides using deep learning: A retrospective multicenter study. PLoS Med. 2019, 16, e1002730. [Google Scholar] [CrossRef]
- Kumar, N.; Verma, R.; Anand, D.; Zhou, Y.; Onder, O.F.; Tsougenis, E.; Chen, H.; Heng, P.A.; Li, J.; Hu, Z.; et al. A multi-organ nucleus segmentation challenge. IEEE Trans. Med. Imaging 2019, 39, 1380–1391. [Google Scholar] [CrossRef] [PubMed]
- Mahbod, A.; Schaefer, G.; Bancher, B.; Löw, C.; Dorffner, G.; Ecker, R.; Ellinger, I. CryoNuSeg: A dataset for nuclei instance segmentation of cryosectioned H&E-stained histological images. Comput. Biol. Med. 2021, 132, 104349. [Google Scholar]
- Naylor, P.; Laé, M.; Reyal, F.; Walter, T. Nuclei segmentation in histopathology images using deep neural networks. In Proceedings of the 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), Melbourne, Australia, 18–21 April 2017; pp. 933–936. [Google Scholar]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer: Berlin/Heidelberg, Germany, 2018; pp. 3–11. [Google Scholar]
- Mahmood, F.; Borders, D.; Chen, R.J.; McKay, G.N.; Salimian, K.J.; Baras, A.; Durr, N.J. Deep adversarial training for multi-organ nuclei segmentation in histopathology images. IEEE Trans. Med. Imaging 2019, 39, 3257–3267. [Google Scholar] [CrossRef]
- Zeng, Z.; Xie, W.; Zhang, Y.; Lu, Y. RIC-Unet: An Improved Neural Network Based on Unet for Nuclei Segmentation in Histology Images. IEEE Access 2019, 7, 21420–21428. [Google Scholar] [CrossRef]
- Naylor, P.; Laé, M.; Reyal, F.; Walter, T. Segmentation of nuclei in histopathology images by deep regression of the distance map. IEEE Trans. Med. Imaging 2018, 38, 448–459. [Google Scholar] [CrossRef]
- Valanarasu, J.M.J.; Oza, P.; Hacihaliloglu, I.; Patel, V.M. Medical transformer: Gated axial-attention for medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Virtual Event, 27 September–1 October 2021; Springer: Berlin/Heidelberg, Germany, 2021; pp. 36–46. [Google Scholar]
- Chanchal, A.K.; Lal, S.; Kini, J. Deep structured residual encoder-decoder network with a novel loss function for nuclei segmentation of kidney and breast histopathology images. Multimed. Tools Appl. 2022, 81, 9201–9224. [Google Scholar] [CrossRef]
- Xiang, T.; Zhang, C.; Liu, D.; Song, Y.; Huang, H.; Cai, W. BiO-Net: Learning recurrent bi-directional connections for encoder-decoder architecture. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Lima, Peru, 4–8 October 2020; Springer: Berlin/Heidelberg, Germany, 2020; pp. 74–84. [Google Scholar]
- Ali, H.; Cui, L.; Feng, J. MSAL-Net: Improve accurate segmentation of nuclei in histopathology images by multiscale attention learning network. BMC Med. Inform. Decis. Mak. 2022, 22, 22. [Google Scholar] [CrossRef] [PubMed]

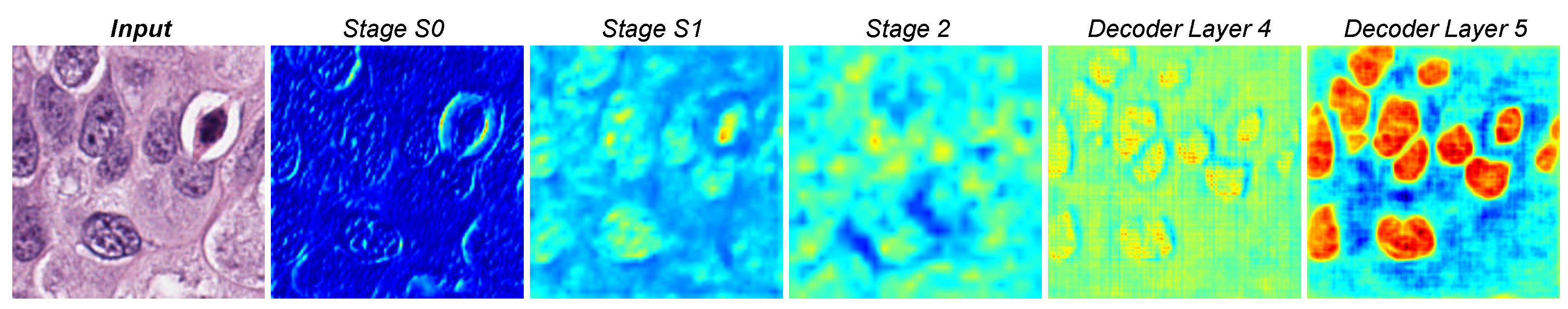
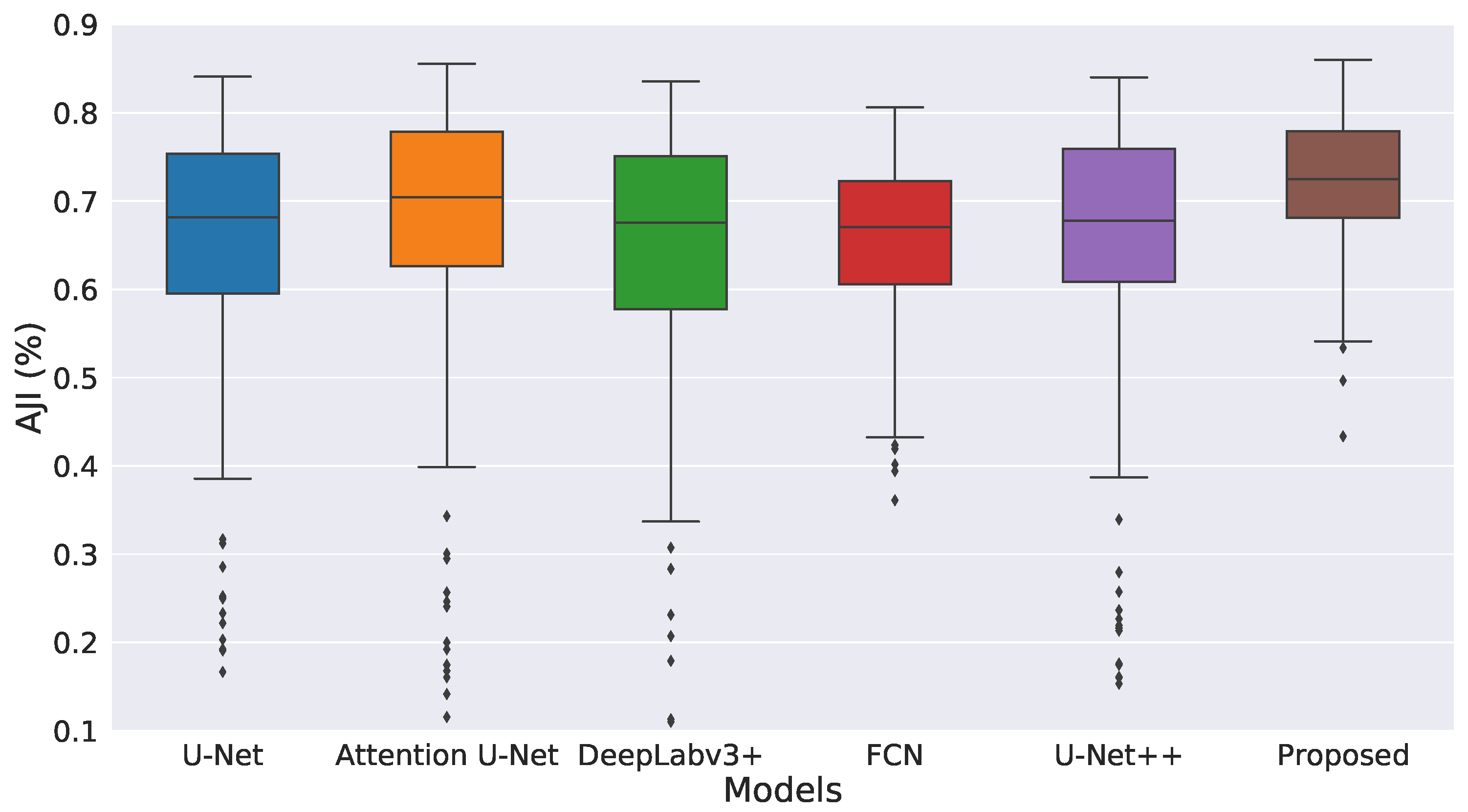

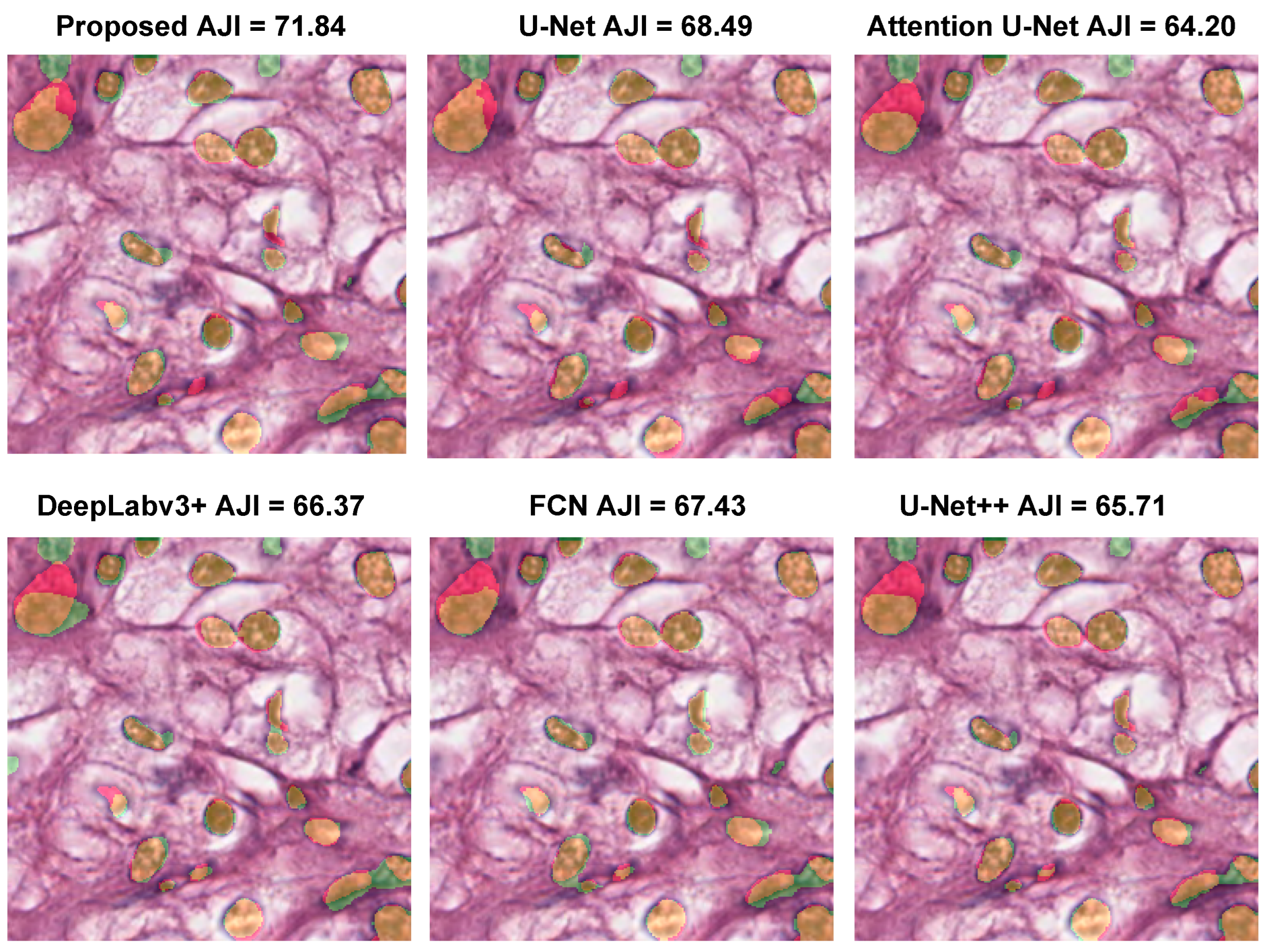
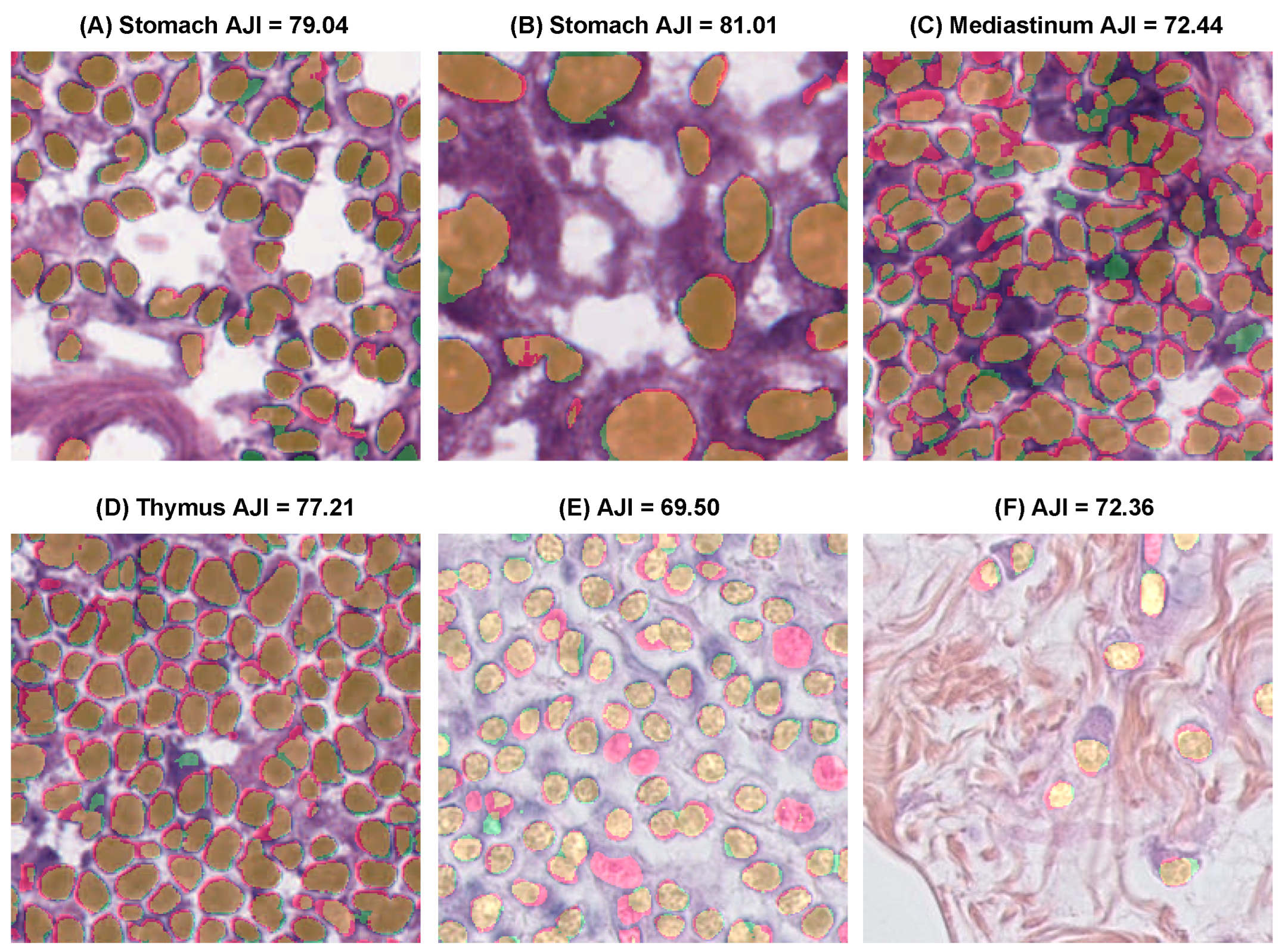
| Model | Dice (%) ↑ | AJI (%) ↑ | Precision (%) ↑ | Recall (%) ↑ |
|---|---|---|---|---|
| BL | ||||
| BL + WHDC | ||||
| Proposed w/o CL | ||||
| Proposed |
| Loss Function | Dice (%) ↑ | AJI (%) ↑ | Precision (%) ↑ | Recall (%) ↑ |
|---|---|---|---|---|
| + |
| Model | Dice (%) ↑ | AJI (%) ↑ | Precision (%) ↑ | Recall (%) ↑ |
|---|---|---|---|---|
| U-Net | ||||
| Attention U-Net | ||||
| DeepLabv3+ | ||||
| FCN | ||||
| U-Net++ | ||||
| RIC-UNet [37] | − | − | ||
| DIST [38] | − | − | ||
| Chanchal et al. [40] | − | − | ||
| cGANs [36] | ||||
| MedT [39] | − | − | ||
| MSAL-Net [42] | ||||
| BiO-Net [41] | − | − | ||
| Proposed |
| Model | Dice (%) ↑ | AJI (%) ↑ | Precision (%) ↑ | Recall (%) ↑ |
|---|---|---|---|---|
| U-Net | ||||
| Attention U-Net | ||||
| DeepLabv3+ | ||||
| FCN | ||||
| U-Net++ | ||||
| Hassan et al. [19] | ||||
| Proposed |
| Model | Dice (%) ↑ | AJI (%) ↑ | Precision (%) ↑ | Recall (%) ↑ |
|---|---|---|---|---|
| U-Net | ||||
| Attention U-Net | ||||
| DeepLabv3+ | ||||
| FCN | ||||
| U-Net++ | ||||
| Proposed |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdel-Nasser, M.; Singh, V.K.; Mohamed, E.M. Efficient Staining-Invariant Nuclei Segmentation Approach Using Self-Supervised Deep Contrastive Network. Diagnostics 2022, 12, 3024. https://doi.org/10.3390/diagnostics12123024
Abdel-Nasser M, Singh VK, Mohamed EM. Efficient Staining-Invariant Nuclei Segmentation Approach Using Self-Supervised Deep Contrastive Network. Diagnostics. 2022; 12(12):3024. https://doi.org/10.3390/diagnostics12123024
Chicago/Turabian StyleAbdel-Nasser, Mohamed, Vivek Kumar Singh, and Ehab Mahmoud Mohamed. 2022. "Efficient Staining-Invariant Nuclei Segmentation Approach Using Self-Supervised Deep Contrastive Network" Diagnostics 12, no. 12: 3024. https://doi.org/10.3390/diagnostics12123024
APA StyleAbdel-Nasser, M., Singh, V. K., & Mohamed, E. M. (2022). Efficient Staining-Invariant Nuclei Segmentation Approach Using Self-Supervised Deep Contrastive Network. Diagnostics, 12(12), 3024. https://doi.org/10.3390/diagnostics12123024








