Urinary Protein Profiling for Potential Biomarkers of Chronic Kidney Disease: A Pilot Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Study Settings
2.2. Laboratory Tests and Data Collection
2.3. Mass Spectrometry Analysis
2.4. Reactome and GO Analysis
2.5. Statistical Analyses
3. Results
3.1. General Characteristics of the Study Population
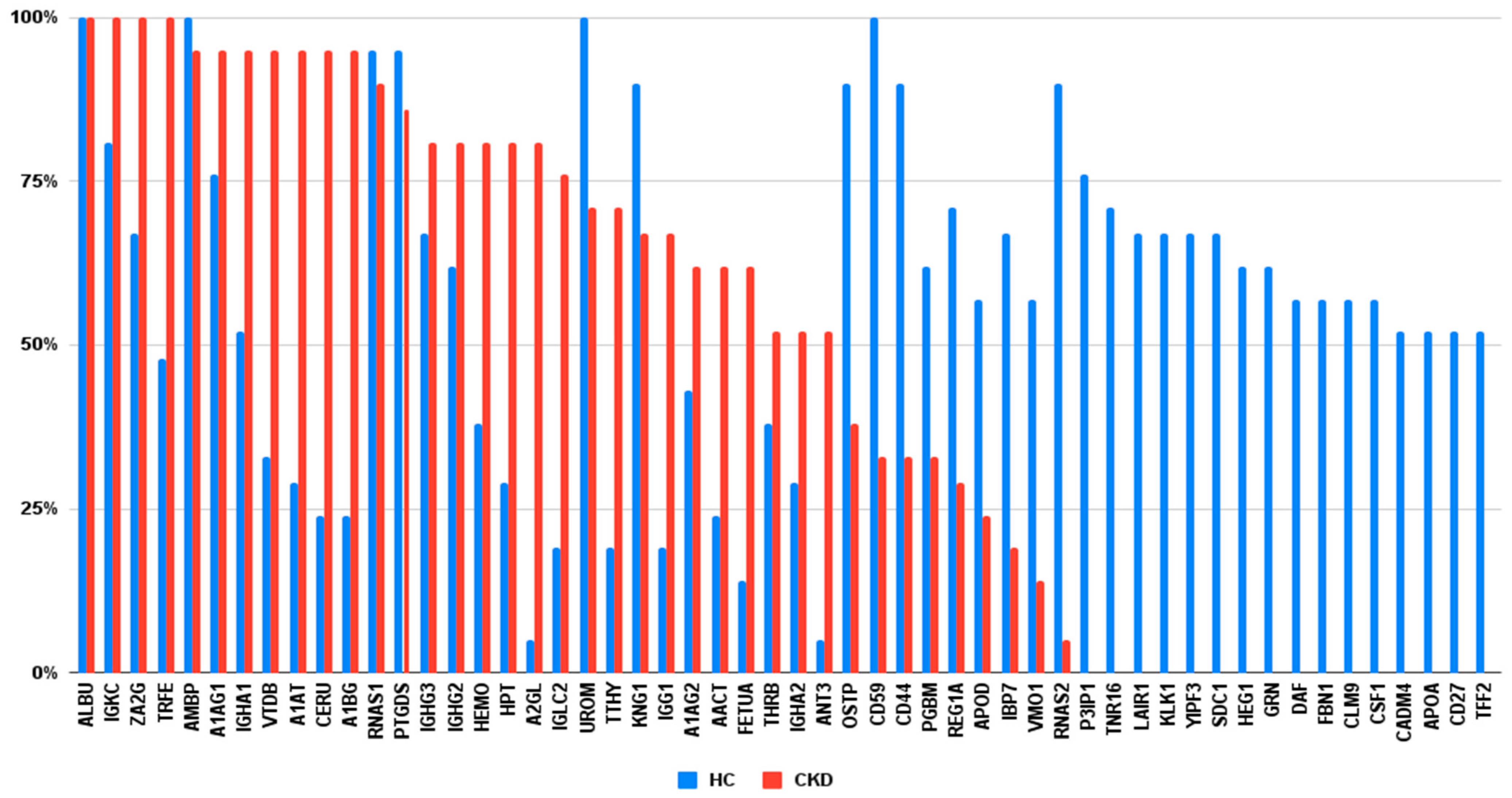
3.2. Urinary Protein Profiles of Study Population
3.3. Associations between Renal Function and Protein Composition
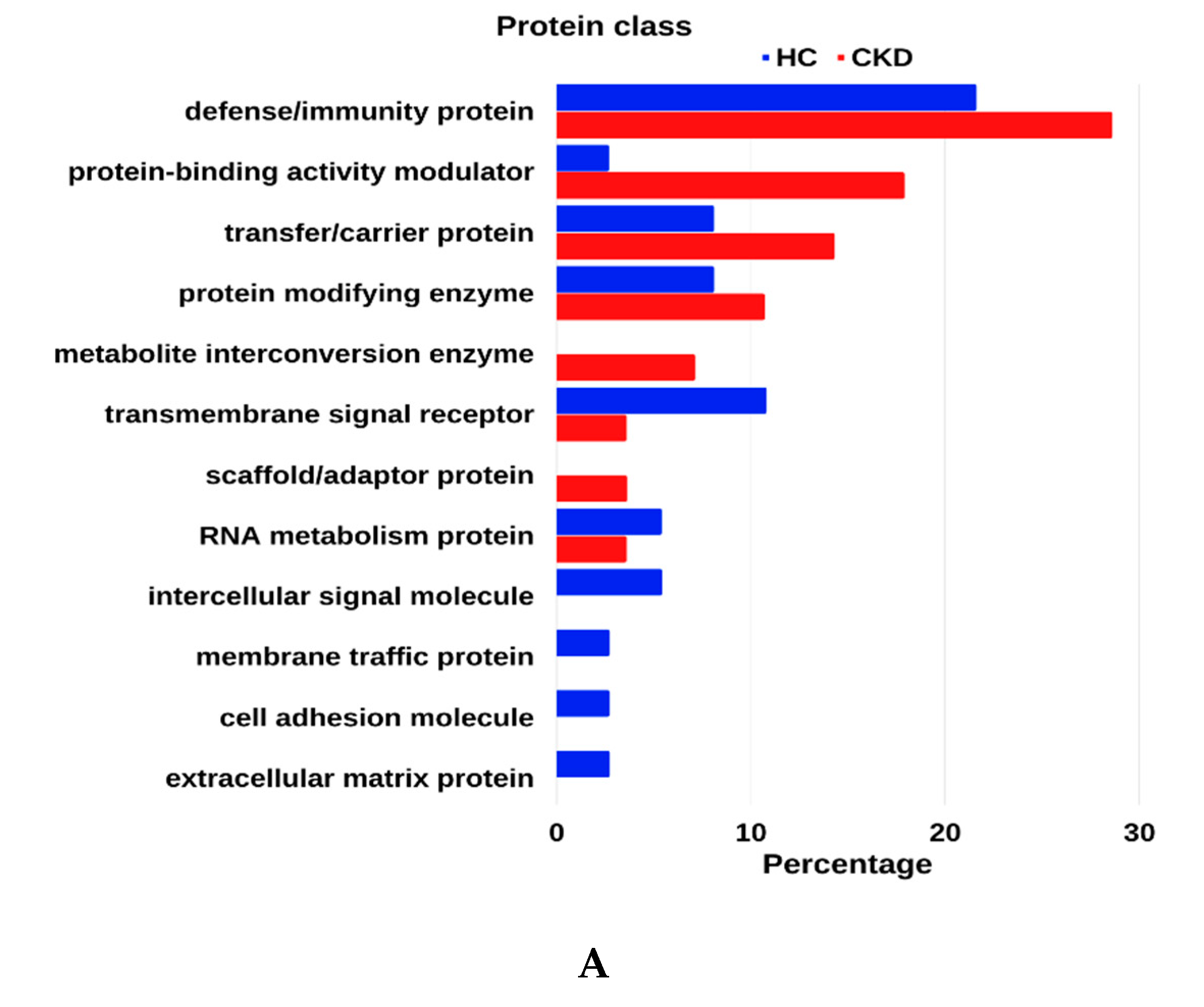
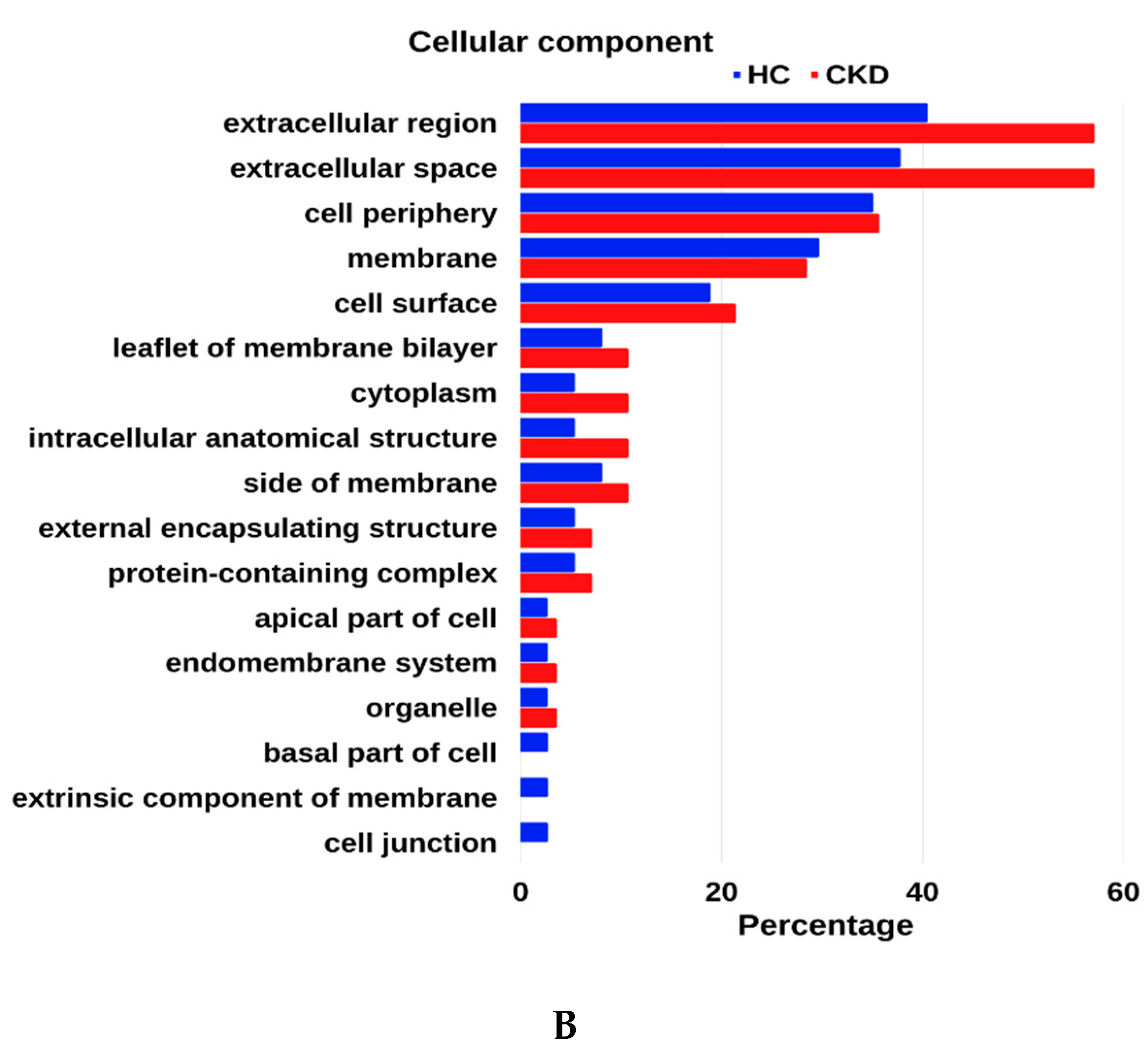
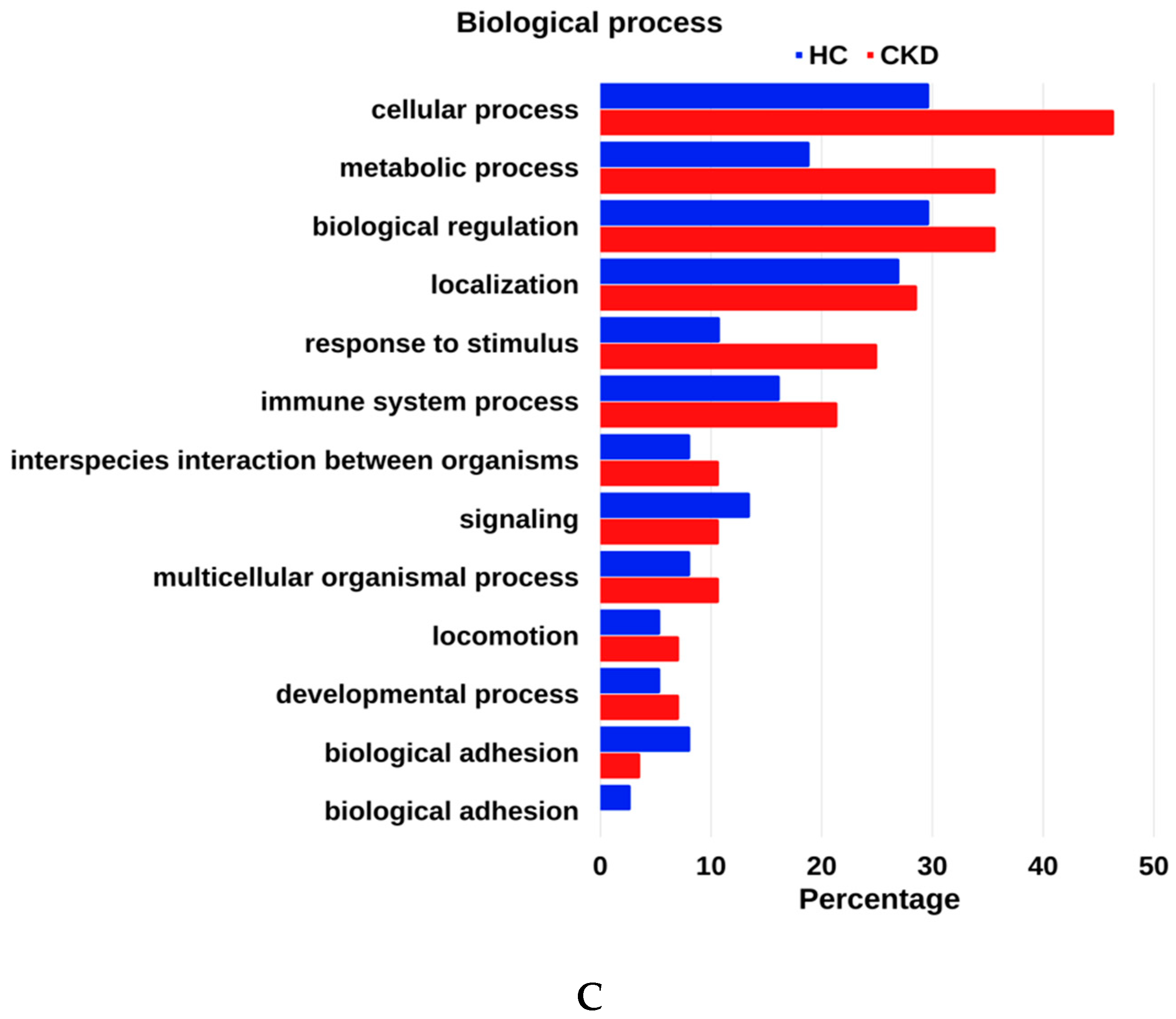
3.4. GO Enrichment Analysis of Urinary Proteome
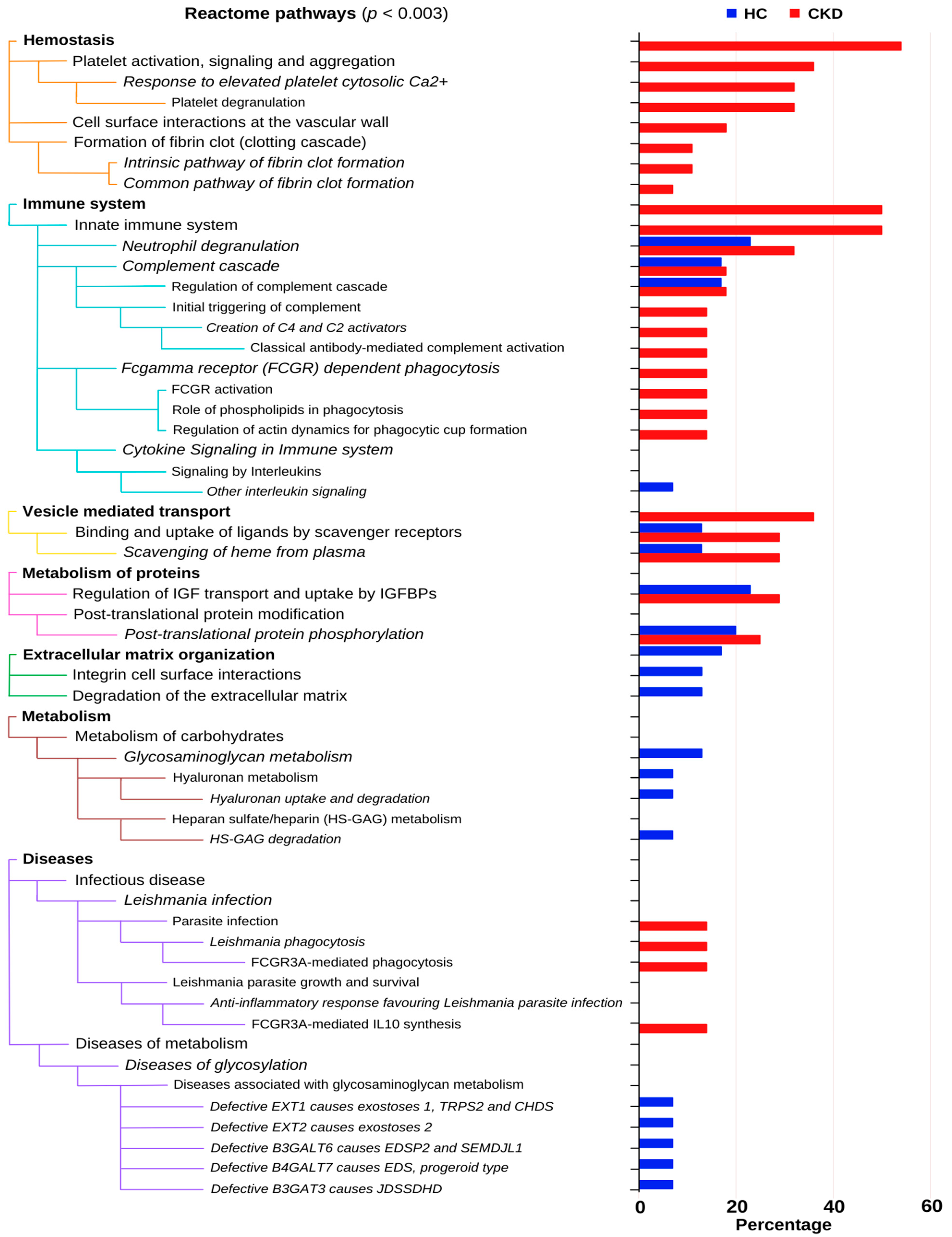
3.5. Pathway Enrichment Analysis of Proteins
4. Discussion
5. Conclusions
6. Trial Registration
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jager, K.J.; Kovesdy, C.; Langham, R.; Rosenberg, M.; Jha, V.; Zoccali, C. A Single Number for Advocacy and Communication—Worldwide More than 850 Million Individuals Have Kidney Diseases; Oxford University Press: Oxford, UK, 2019; pp. 1803–1805. [Google Scholar]
- Carney, E.F. The impact of chronic kidney disease on global health. Nat. Rev. Nephrol. 2020, 16, 251. [Google Scholar] [CrossRef] [PubMed]
- Foreman, K.J.; Marquez, N.; Dolgert, A.; Fukutaki, K.; Fullman, N.; McGaughey, M.; Pletcher, M.A.; Smith, A.E.; Tang, K.; Yuan, C.-W.; et al. Forecasting life expectancy, years of life lost, and all-cause and cause-specific mortality for 250 causes of death: Reference and alternative scenarios for 2016–40 for 195 countries and territories. Lancet 2018, 392, 2052–2090. [Google Scholar] [CrossRef]
- Cravedi, P.; Remuzzi, G. Pathophysiology of proteinuria and its value as an outcome measure in chronic kidney disease. Br. J. Clin. Pharmacol. 2013, 76, 516–523. [Google Scholar]
- Liu, D.; Lv, L.-L. New understanding on the role of proteinuria in progression of chronic kidney disease. In Renal Fibrosis: Mechanisms and Therapies; Springer: Berlin/Heidelberg, Germany, 2019; pp. 487–500. [Google Scholar]
- Kasiske, B.; Wheeler, D. Kidney Disease: Improving Global Outcomes CKD Work Group. KDIGO 2012 clinical practice guideline for the evaluation and management of chronic kidney disease. Kidney Int. Suppl. 2013, 3, 1–150. [Google Scholar]
- Perico, N.; Benigni, A.; Remuzzi, G. Proteinuria and tubulotoxicity. In Diabetic Nephropathy; Springer: Berlin/Heidelberg, Germany, 2019; pp. 197–214. [Google Scholar]
- Ledingham, J. Tubular toxicity of filtered proteins. Am. J. Nephrol. 1990, 10 (Suppl. S1), 52–57. [Google Scholar] [CrossRef]
- Meng, Z.; Bustamante Lopez, S.C.; Meissner, K.E.; Yakovlev, V.V. Subcellular measurements of mechanical and chemical properties using dual Raman-Brillouin microspectroscopy. J. Biophotonics 2016, 9, 201–207. [Google Scholar] [CrossRef]
- Gaipov, A.; Utegulov, Z.; Bukasov, R.; Turebekov, D.; Tarlykov, P.; Markhametova, Z.; Nurekeyev, Z.; Kunushpayeva, Z.; Sultangaziyev, A. Development and validation of hybrid Brillouin-Raman spectroscopy for non-contact assessment of mechano-chemical properties of urine proteins as biomarkers of kidney diseases. BMC Nephrol. 2020, 21, 1–9. [Google Scholar] [CrossRef]
- Levey, A.S.; Eckardt, K.-U.; Tsukamoto, Y.; Levin, A.; Coresh, J.; Rossert, J.; Eknoyan, G. Definition and classification of chronic kidney disease: A position statement from Kidney Disease: Improving Global Outcomes (KDIGO). Kidney Int. 2005, 67, 2089–2100. [Google Scholar] [CrossRef]
- Aitekenov, S.; Gaipov, A.; Bukasov, R. Detection and quantification of proteins in human urine. Talanta 2021, 223, 121718. [Google Scholar] [CrossRef]
- Sun, W.; Gao, Y. Liquid Chromatography Coupled to Mass Spectrometry for Analysis of the Urinary Proteome. In Renal and Urinary Proteomics: Methods and Protocols; Wiley: Hoboken, NJ, USA, 2009; pp. 271–279. [Google Scholar]
- Ishihama, Y.; Oda, Y.; Tabata, T.; Sato, T.; Nagasu, T.; Rappsilber, J.; Mann, M. Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein* s. Mol. Cell. Proteom. 2005, 4, 1265–1272. [Google Scholar] [CrossRef]
- The Gene Ontology Consortium. The Gene Ontology resource: Enriching a GOld mine. Nucleic Acids Res. 2021, 49, D325–D334. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef] [PubMed]
- Jassal, B.; Matthews, L.; Viteri, G.; Gong, C.; Lorente, P.; Fabregat, A.; D’Eustachio, P. The reactome pathway knowledgebase. Nucleic Acids Res. 2020, 48, D498–D503. [Google Scholar] [CrossRef]
- D’amico, G.; Bazzi, C. Pathophysiology of proteinuria. Kidney Int. 2003, 63, 809–825. [Google Scholar] [CrossRef]
- Tofte, N.; Lindhardt, M.; Adamova, K.; Bakker, S.J.L.; Beige, J.; Beulens, J.W.J.; Kilic, C. Early detection of diabetic kidney disease by urinary proteomics and subsequent intervention with spironolactone to delay progression (PRIORITY): A prospective observational study and embedded randomised placebo-controlled trial. Lancet Diabetes Endocrinol. 2020, 8, 301–312. [Google Scholar] [CrossRef]
- Rodríguez-Ortiz, M.E.; Pontillo, C.; Rodríguez, M.; Zürbig, P.; Mischak, H.; Ortiz, A. Novel Urinary Biomarkers for Improved Prediction of Progressive Egfr Loss in Early Chronic Kidney Disease Stages And In High Risk Individuals Without Chronic Kidney Disease. Sci. Rep. 2018, 8, 15940. [Google Scholar] [CrossRef]
- Captur, G.; Moon, J.C.; Topriceanu, C.C.; Joy, G.; Swadling, L.; Hallqvist, J.; Doykov, I.; Patel, N.; Spiewak, J.; Baldwin, T.; et al. Plasma proteomic signature predicts who will get persistent symptoms following SARS-CoV-2 infection. eBioMedicine 2022, 2022, 104293. [Google Scholar] [CrossRef]
- Martin, H. Laboratory measurement of urine albumin and urine total protein in screening for proteinuria in chronic kidney disease. Clin. Biochem. Rev. 2011, 32, 97. [Google Scholar] [PubMed]
- Methven, S.; MacGregor, M.S.; Traynor, J.P.; Hair, M.; O’Reilly, D.S.J.; Deighan, C.J. Comparison of urinary albumin and urinary total protein as predictors of patient outcomes in CKD. Am. J. Kidney Dis. 2011, 57, 21–28. [Google Scholar] [CrossRef]
- Thongboonkerd, V.; Malasit, P. Renal and urinary proteomics: Current applications and challenges. Proteomics 2005, 5, 1033–1042. [Google Scholar] [CrossRef]
- Scolari, F.; Izzi, C.; Ghiggeri, G.M. Uromodulin: From monogenic to multifactorial diseases. Nephrol. Dial. Transplant. 2015, 30, 1250–1256. [Google Scholar] [CrossRef] [PubMed]
- Molitoris, B.A.; Sandoval, R.M.; Yadav, S.P.S.; Wagner, M.C. Albumin Uptake and Processing by the Proximal Tubule: Physiologic, Pathologic and Therapeutic Implications. Physiol. Rev. 2022, 102, 1625–1667. [Google Scholar] [CrossRef] [PubMed]
- Jassil, N.K.; Sharma, A.; Bikle, D.; Wang, X. Vitamin D binding protein and 25-hydroxyvitamin D levels: Emerging clinical applications. Endocr. Pract. 2017, 23, 605–613. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Riera, E.; García-Arguinzonis, M.; López, L.; Garcia-Moll, X.; Badimon, L.; Padró, T. Vitamin D Binding Protein and Renal Injury in Acute Decompensated Heart Failure. Front. Cardiovasc. Med. 2022, 9, 829490. [Google Scholar] [CrossRef]
- Suresh, C.; Saha, A.; Kaur, M.; Kumar, R.; Dubey, N.; Basak, T.; Upadhyay, A.D. Differentially expressed urinary biomarkers in children with idiopathic nephrotic syndrome. Clin. Exp. Nephrol. 2016, 20, 273–283. [Google Scholar] [CrossRef]
- Zhou, X.-H.; Liu, S.-Y.; Feng, L.; Yang, B.; Li, Y.-F.; Wang, X.-Q.; Chen, J.; Wang, H.H. Urinary orosomucoid and retinol binding protein levels as early diagnostic markers for diabetic nephropathy. Res. Sq. 2020. [Google Scholar]
- Alhazmi, S.; Basingab, F.; Alrahimi, J.; Alharbi, M.; Mufawwaz, A.; Alotaibi, K. The Role of Alpha-1-acid Glycoprotein 2 Protein and the Underlying Orosomucoid 2 Gene in Different Diseases. J. Pharm. Res. Int. 2022, 34, 15–29. [Google Scholar]
- Jin, Y.; Wang, W.; Wang, Q.; Zhang, Y.; Zahid, K.R.; Raza, U.; Gong, Y. Alpha-1-antichymotrypsin as a novel biomarker for diagnosis, prognosis, and therapy prediction in human diseases. Cancer Cell Int. 2022, 22, 156. [Google Scholar] [CrossRef]
- Sánchez-Navarro, A.; Mejía-Vilet, J.M.; Pérez-Villalva, R.; Carrillo-Pérez, D.L.; Marquina-Castillo, B.; Gamba, G.; Bobadilla, N.A. SerpinA3 in the Early Recognition of Acute Kidney Injury to Chronic Kidney Disease (CKD) transition in the rat and its Potentiality in the Recognition of Patients with CKD. Sci. Rep. 2019, 9, 10350. [Google Scholar] [CrossRef]
- López-Guisa, J.M.; Rassa, A.C.; Cai, X.; Collins, S.J.; Eddy, A.A. Vitronectin accumulates in the interstitium but minimally impacts fibrogenesis in experimental chronic kidney disease. Am. J. Physiol.-Ren. Physiol. 2011, 300, F1244–F1254. [Google Scholar] [CrossRef]
- Carreras-Planella, L.; Cucchiari, D.; Cañas, L.; Juega, J.; Franquesa, M.; Bonet, J.; Borràs, F.E. Urinary vitronectin identifies patients with high levels of fibrosis in kidney grafts. J. Nephrol. 2021, 34, 861–874. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.; Gingras, D.; Bendayan, M. Alterations of vitronectin and its receptor αv integrin in the rat renal glomerular wall during diabetes. Am. J. Kidney Dis. 2001, 38, 1298–1306. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Chen, X.; Chima, A.; Zhang, Y.; George, J.; Cobbs, A.; Emmett, N. Albumin induces CD44 expression in glomerular parietal epithelial cells by activating extracellular signal-regulated kinase 1/2 pathway. J. Cell. Physiol. 2019, 234, 7224–7235. [Google Scholar] [CrossRef] [PubMed]
- Rouschop, K.; Roelofs, J.; Sylva, M.; Rowshani, A.; Ten Berge, I.; Weening, J.; Florquin, S. Renal expression of CD44 correlates with acute renal allograft rejection. Kidney Int. 2006, 70, 1127–1134. [Google Scholar] [CrossRef] [PubMed]
- Froes, B.P.; de Almeida Araújo, S.; Bambirra, E.A.; Oliveira, E.A.; Simoes e Silva, A.C.; Pinheiro, S.V.B. Is CD44 in glomerular parietal epithelial cells a pathological marker of renal function deterioration in primary focal segmental glomerulosclerosis? Pediatr. Nephrol. 2017, 32, 2165–2169. [Google Scholar] [CrossRef] [PubMed]
- Weening, J.J.; Ronco, P.; Remuzzi, G. Advances in the pathology of glomerular diseases. New Insights Glomerulonephritis 2013, 181, 12–21. [Google Scholar]
- Gros, A.; Ollivier, V.; Ho-Tin-Noé, B. Platelets in inflammation: Regulation of leukocyte activities and vascular repair. Front. Immunol. 2015, 5, 678. [Google Scholar] [CrossRef]
- Finsterbusch, M.; Schrottmaier, W.C.; Kral-Pointner, J.B.; Salzmann, M.; Assinger, A. Measuring and interpreting platelet-leukocyte aggregates. Platelets 2018, 29, 677–685. [Google Scholar] [CrossRef]
- Mayadas, T.N.; Rosetti, F.; Ernandez, T.; Sethi, S. Neutrophils: Game changers in glomerulonephritis? Trends Mol. Med. 2010, 16, 368–378. [Google Scholar] [CrossRef]
- Chen, S.-F.; Chen, M. Complement activation in progression of chronic kidney disease. In Renal Fibrosis: Mechanisms and Therapies; Springer: Berlin/Heidelberg, Germany, 2019; pp. 423–441. [Google Scholar]
- Barcellini, W. Immune hemolysis: Diagnosis and treatment recommendations. Semin. Hematol. 2015, 52, 304–312. [Google Scholar] [CrossRef]
- Zager, R.A.; Johnson, A.C.; Becker, K. Renal cortical hemopexin accumulation in response to acute kidney injury. Am. J. Physiol.-Ren. Physiol. 2012, 303, F1460–F1472. [Google Scholar] [CrossRef] [PubMed]
- Moreno, J.A.; Martín-Cleary, C.; Gutiérrez, E.; Rubio-Navarro, A.; Ortiz, A.; Praga, M.; Egido, J. Haematuria: The forgotten CKD factor? Nephrol. Dial. Transplant. 2012, 27, 28–34. [Google Scholar] [CrossRef] [PubMed]
- Moreno, J.A.; Martín-Cleary, C.; Gutiérrez, E.; Toldos, O.; Blanco-Colio, L.M.; Praga, M.; Egido, J. AKI associated with macroscopic glomerular hematuria: Clinical and pathophysiologic consequences. Clin. J. Am. Soc. Nephrol. 2012, 7, 175–184. [Google Scholar] [CrossRef] [PubMed]
- Stephan, J.-P.; Mao, W.; Filvaroff, E.; Cai, L.; Rabkin, R.; Pan, G. Albumin stimulates the accumulation of extracellular matrix in renal tubular epithelial cells. Am. J. Nephrol. 2004, 24, 14–19. [Google Scholar] [CrossRef]
| Parameters | Healthy Controls, n = 21 | CKD Group, n = 21 | p-Value |
|---|---|---|---|
| Demographics and clinical data | |||
| Gender, female, % | 15 (71%) | 12 (57%) | 0.334 |
| Age, year | 35.6 ± 8.3 | 38.5 ± 13.7 | 0.419 |
| eGFR, mL/min/1.73 m2 | 109.1 (103.3–120.1) | 111 (72.6–138.1) | 0.835 |
| Primary diagnosis, n (%) | |||
| Glomerular disease | N/A | 19 (90%) | |
| Post-transplant CKD | N/A | 1 (4,76%) | |
| CKD of unknown etiology | N/A | 1 (4.76%) | |
| Biochemistry | |||
| Serum creatinine, µmol/L | 65.1 (59.6–71.4) | 69.6 (57.4–127.9) | 0.285 |
| Serum urea, mmol/L | 4.0 (3.5–4.6) | 5.2 (4.0–8.6) | 0.014 |
| Serum glucose, mmol/L | 5.1 (4.9–5.5) | 5.0 (4.4–5.3) | 0.973 |
| Serum uric acid, mcmol/L | 277.5 (234.7–321.0) | 359.6 (283.0–436.4) | 0.018 |
| Serum sodium, mmol/L | 139.5 (138–141) | 140 (139–141) | 0.863 |
| Serum potassium, mmol/L | 4.4 (4.1–4.8) | 4.2 (4.0–4.6) | 0.232 |
| Ca ionized, mmol/L | 1.3 (1.3–1.3) | 1.3 (1.3–1.3) | 0.071 |
| Total bilirubin, mcmol/L | 7.1 (5.0–11.5) | 5.0 (3.1–7.7) | 0.042 |
| ALT, mckat/L | 0.2 (0.1–0.3) | 0.3 (0.2–0.5) | 0.111 |
| ACT, mckat/L | 0.3 (0.2–0.3) | 0.3 (0.2–0.4) | 0.396 |
| Total cholesterol, mmol/L | 4.4 (4.0–4.7) | 6.5 (5.5–7.9) | <0.0001 |
| HDL, mmol/L | 1.5 (1.2–1.9) | 1.4 (1.1–1.7) | 0.514 |
| LDL, mmol/L | 2.7 (2.2–3.2) | 4.1 (3.3–5.6) | 0.0001 |
| Triglycerides, mmol/L | 0.9 (0.6–1.3) | 2.3 (1.4–3.0) | <0.0001 |
| Serum total protein, g/L | 68.9 (66.5–72.0) | 50.8 (44.8–64.6) | <0.0001 |
| Protein fractions, % | |||
| Albumin | 68.4 (66.1–69.7) | 58.0 (43.9–63.1) | <0.0001 |
| Globulin alpa-1 | 1.9 (1.8–2.1) | 2.9 (2.3–3.4) | <0.0001 |
| Globulin alpa-2 | 8.5 (8.0–8.9) | 14.3 (10.9–19.9) | <0.0001 |
| Globulin beta-1 | 6.0 (5.6–6.4) | 8.5 (7.1–9.7) | <0.0001 |
| Globulin beta-2 | 3.2 (3.1–3.7) | 5.4 (3.9–7.3) | 0.0001 |
| Gamma globulin | 12.3 (10.9–14.4) | 10.3 (8.5–13.8) | 0.039 |
| CBC | |||
| WBC 10 × 109/L | 5.4 (5.0–6.2) | 7.3 (6.8–8.9) | 0.0003 |
| RBC 10 × 1012/L | 4.7 (4.3–5.2) | 4.4 (4.0–4.7) | 0.043 |
| HGB g/L | 129 (121–155) | 125 (115–138) | 0.220 |
| PLT 10 × 109/L | 252 (228–298) | 284 (228–329) | 0.410 |
| ESR, mm/h | 9 (5–15) | 27 (12–37) | 0.002 |
| Urinalysis, 24 h | |||
| Proteinuria, g/24 h | 0.1 (0.1–0.2) | 2.0 (1.4–6.9) | <0.0001 |
| Sodium, mmol/24 h | 119 (71.0–164.9) | 206.4 (145.0–317.1) | 0.009 |
| Potassium, mmol/24 h | 40.8 (33.8–49.5) | 42.2 (33.5–51.1) | 0.697 |
| Uric acid, mmol/24 h | 2.4 (2.0–3.3) | 2.6 (1.6–3.4) | 0.990 |
| Creatinine, mmol/24 h | 9.6 (8.5–10.5) | 9.7 (8.0–13.5) | 0.464 |
| Proteomic data | |||
| Total # of defined proteins, n | 250 | 142 | |
| emPAI for total proteins | 39.1 (19–53) | 67.8 (49–117) | 0.003 |
| # of selected proteins, n | 45 | 32 | |
| emPAI for selected proteins | 43.4 (24–71) | 45.1 (18–80) | 0.0001 |
| Urinary Proteome | Protein Name | Spearman’s Rho | p-Value | N of Obs | Sum of emPAI | emPAI, Median (IQR) | Molecular Weight (Da) |
|---|---|---|---|---|---|---|---|
| ALBU | Albumin | 0.6287 | <0.0001 | 42 | 1388.9 | 21.5 (5.0–44.1) | 69,367 |
| ZA2G | Zinc-alpha-2-glycoprotein | 0.6254 | 0.0001 | 35 | 37.32 | 1 (0.4–1.85) | 34,259 |
| IGKC | Immunoglobulin kappa constant | 0.4199 | 0.0087 | 38 | 245.0 | 3.3 (1.1–6.2) | 11,765 |
| FETUA | Alpha-2-HS-glycoprotein | 0.5347 | 0.0328 | 16 | 4.06 | 0.2 (0.1–0.4) | 39,341 |
| VTDB | Vitamin D-binding protein | 0.4742 | 0.0125 | 27 | 13.67 | 0.3 (0.2–0.8) | 52,918 |
| A1BG | Alpha-1B-glycoprotein | 0.4296 | 0.0321 | 25 | 25.85 | 1.1 (0.7–1.3) | 54,254 |
| TRFE | Serotransferrin | 0.3815 | 0.0342 | 31 | 83.71 | 2.5 (0.7–4.0) | 77,064 |
| IGHA2 | Immunoglobulin heavy constant alpha 2 | 0.4833 | 0.0494 | 17 | 11.13 | 0.6 (0.5–0.8) | 36,591 |
| OSTP | Osteopontin | −0.8017 | <0.0001 | 27 | 9.47 | 0.3 (0.2–0.4) | 35,423 |
| CD59 | CD59 glycoprotein | −0.7633 | <0.0001 | 28 | 85.42 | 2.4 (1.3–4.1) | 14,177 |
| UROM | Uromodulin | −0.7589 | <0.0001 | 36 | 56.16 | 1.1 (0.2–2.7) | 69,761 |
| KNG1 | Kininogen-1 | −0.7269 | <0.0001 | 33 | 21.01 | 0.6 (0.1–1.0) | 71,957 |
| RNAS1 | Ribonuclease pancreatic | −0.6924 | <0.0001 | 39 | 38.62 | 0.6 (0.4–1.7) | 17,644 |
| CD44 | CD44 antigen | −0.6803 | 0.0001 | 26 | 2.56 | 0.1 (0.04–0.1) | 81,538 |
| AMBP | Protein AMBP | −0.5899 | <0.0001 | 41 | 160.57 | 1.9 (0.8–4.4) | 38,999 |
| A2GL | Leucine-rich alpha-2-glycoprotein | −0.5478 | 0.0186 | 18 | 4.60 | 0.2 (0.2–0.3) | 38,178 |
| PTGDS | Prostaglandin-H2 D-isomerase | −0.3365 | 0.0389 | 38 | 44.05 | 1.2 (0.5–1.3) | 21,029 |
| Information about Proteins | Serum Creatinine | eGFR | |||||
|---|---|---|---|---|---|---|---|
| Urinary Proteome | Protein Name | Molecular Weight (Da) | N of Obs | Spearman’s Rho | p-Value | Spearman’s Rho | p-Value |
| VTDB | Vitamin D-binding protein | 52,918 | 26 | 0.6172 | 0.0008 | −0.3944 | 0.0461 |
| AACT | Alpha-1-antichymotrypsin | 47,651 | 17 | −0.6271 | 0.0071 | 0.3611 | 0.1544 |
| A1AG2 | Alpha-1-acid glycoprotein 2 | 23,603 | 21 | −0.4637 | 0.0342 | 0.2023 | 0.3791 |
| VTNC | Vitronectin | 54,306 | 12 | −0.5999 | 0.0392 | 0.4888 | 0.1068 |
| CD44 | CD44 antigen | 81,538 | 25 | −0.1365 | 0.5154 | 0.4696 | 0.0179 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gaipov, A.; Makhammajanov, Z.; Dauyey, Z.; Markhametova, Z.; Mussina, K.; Nogaibayeva, A.; Kozina, L.; Auganova, D.; Tarlykov, P.; Bukasov, R.; et al. Urinary Protein Profiling for Potential Biomarkers of Chronic Kidney Disease: A Pilot Study. Diagnostics 2022, 12, 2583. https://doi.org/10.3390/diagnostics12112583
Gaipov A, Makhammajanov Z, Dauyey Z, Markhametova Z, Mussina K, Nogaibayeva A, Kozina L, Auganova D, Tarlykov P, Bukasov R, et al. Urinary Protein Profiling for Potential Biomarkers of Chronic Kidney Disease: A Pilot Study. Diagnostics. 2022; 12(11):2583. https://doi.org/10.3390/diagnostics12112583
Chicago/Turabian StyleGaipov, Abduzhappar, Zhalaliddin Makhammajanov, Zhanna Dauyey, Zhannur Markhametova, Kamilla Mussina, Assem Nogaibayeva, Larissa Kozina, Dana Auganova, Pavel Tarlykov, Rostislav Bukasov, and et al. 2022. "Urinary Protein Profiling for Potential Biomarkers of Chronic Kidney Disease: A Pilot Study" Diagnostics 12, no. 11: 2583. https://doi.org/10.3390/diagnostics12112583
APA StyleGaipov, A., Makhammajanov, Z., Dauyey, Z., Markhametova, Z., Mussina, K., Nogaibayeva, A., Kozina, L., Auganova, D., Tarlykov, P., Bukasov, R., Utegulov, Z., Turebekov, D., Soler, M. J., Ortiz, A., & Kanbay, M. (2022). Urinary Protein Profiling for Potential Biomarkers of Chronic Kidney Disease: A Pilot Study. Diagnostics, 12(11), 2583. https://doi.org/10.3390/diagnostics12112583





