Cancer Diagnosis of Microscopic Biopsy Images Using a Social Spider Optimisation-Tuned Neural Network
Abstract
:1. Introduction
- The proposed design for an automated model to diagnose the microscopic biopsy images as benign or malignant.
- The optimal tuning of the weights of the NN classifier by the SSO algorithm plays a vital role in enhancing the performance of the proposed classification model.
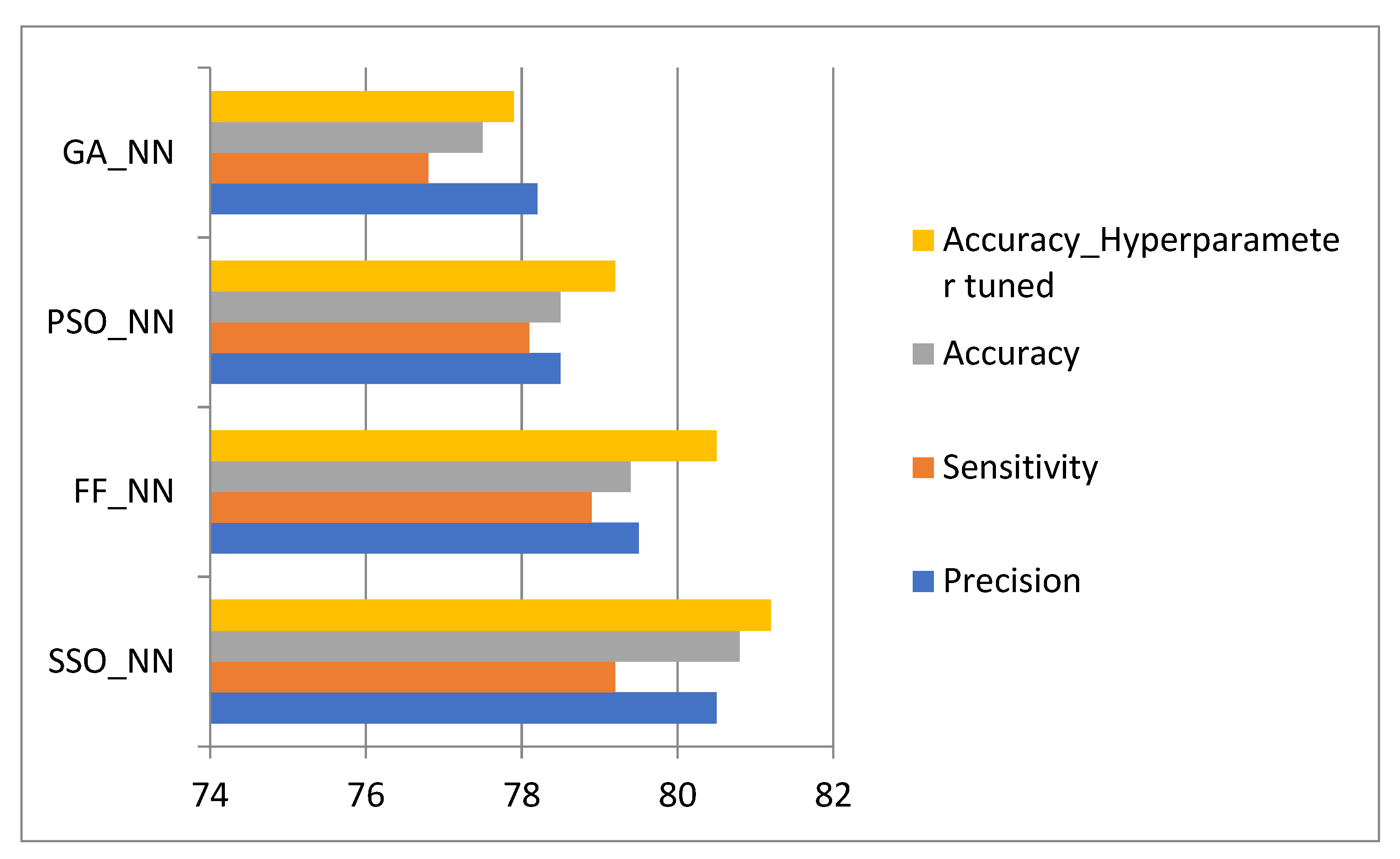
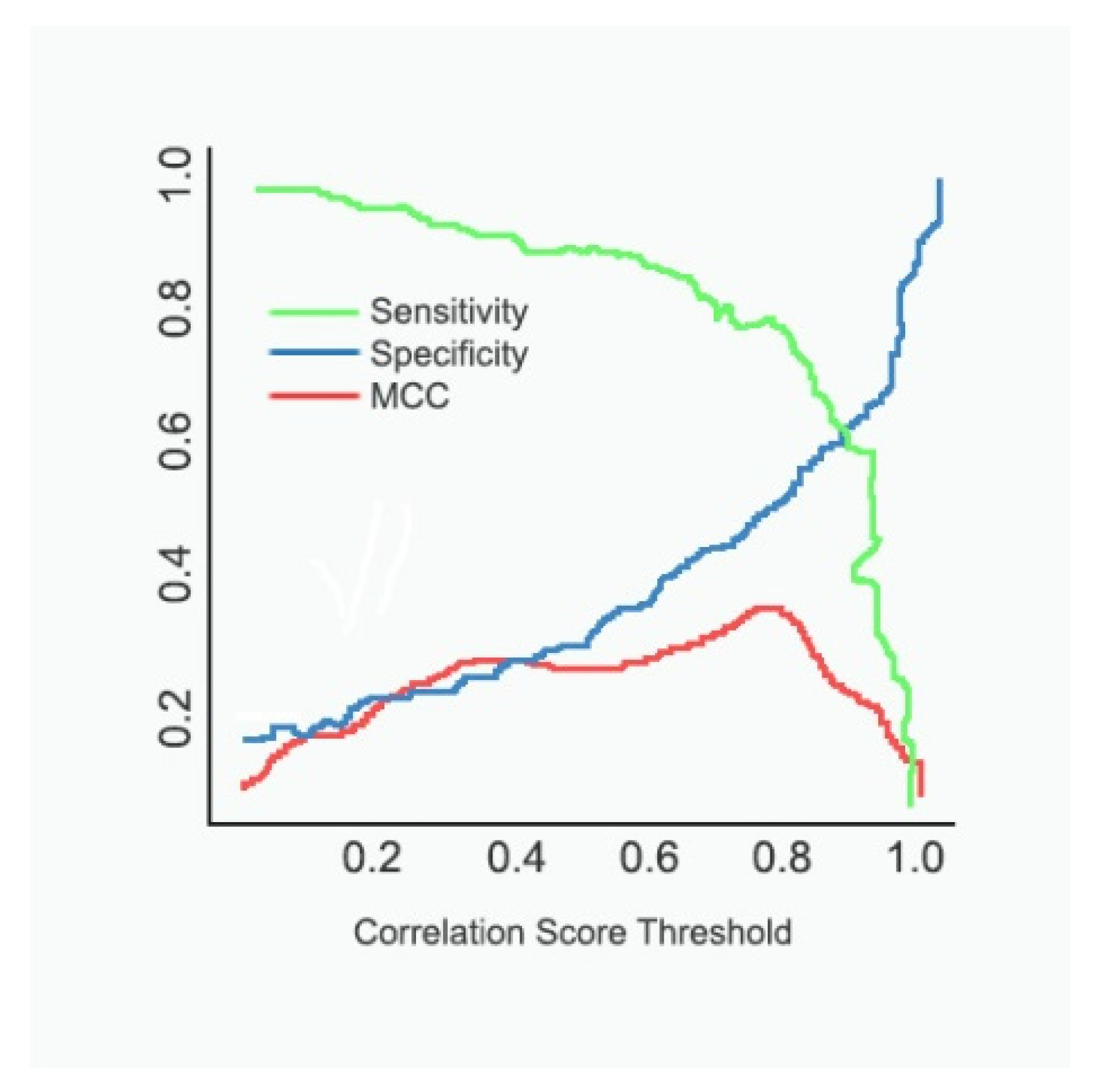
- The proposed model is analysed with conventional methods in terms of performance measures such as accuracy, sensitivity, specificity, and Matthew’s correlation coefficient (MCC) to validate the effectiveness of the proposed strategy.
2. Literature Survey
2.1. Related Works
2.2. Challenges
- The DNN-based strategies, particularly the CNN-based methods, have solved the handcrafted extraction of features. However, when this model is trained from scratch, it needs more annotated images and requires very high resources [3].
- In breast biopsy, the breast cancer samples are taken and preserved into microscopic slides for manual evaluation. An expert pathologist carries out the microscopic analysis, and the conclusion is made after the agreement of more than two pathologists for enhanced diagnosis. However, it may need increased time for diagnosis, and there may be a disagreement of opinion among two pathologists [3].
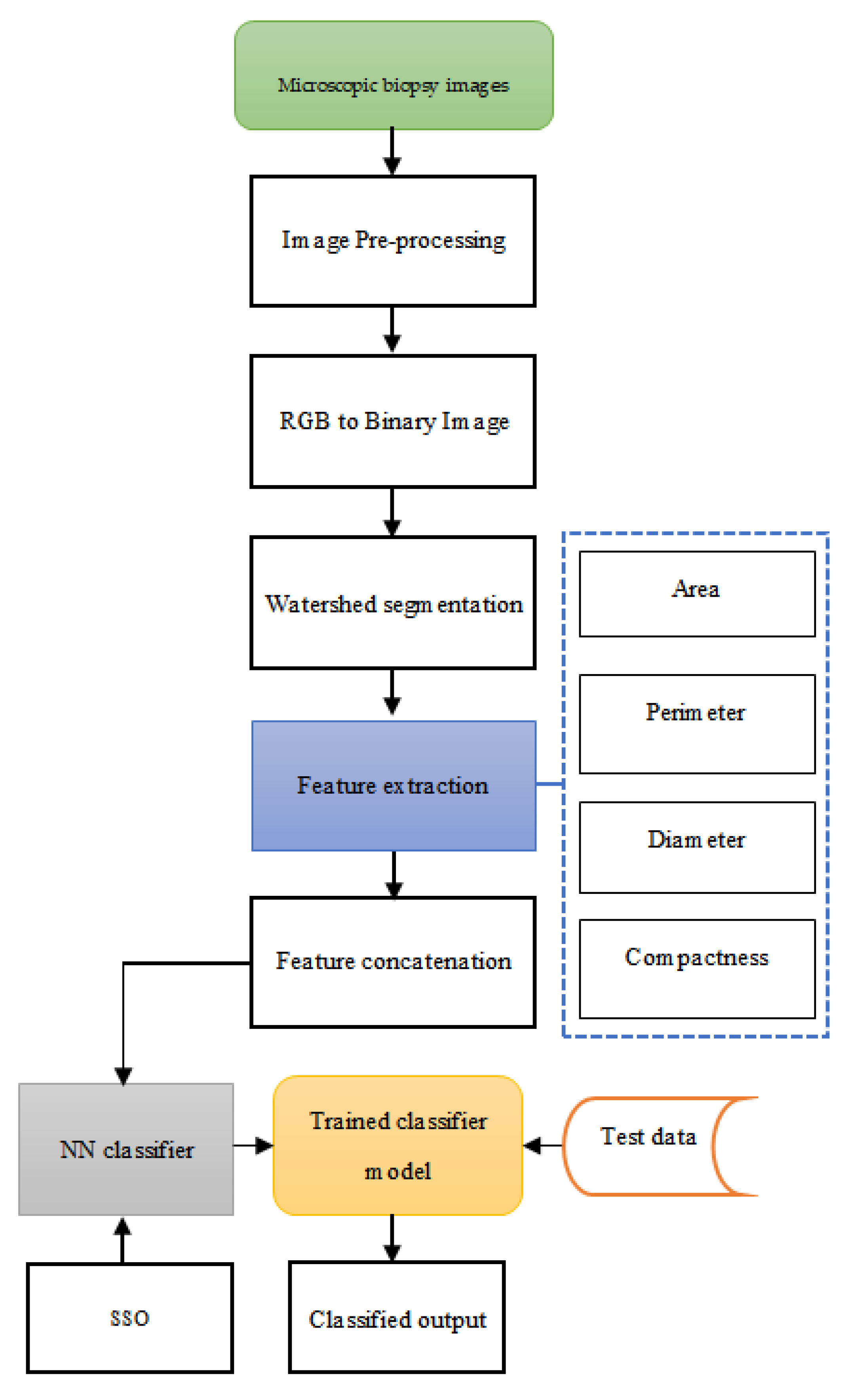
3. Proposed Method of Cancer Diagnosis
3.1. Image Pre-Processing
3.2. Watershed Segmentation
3.3. Feature Extraction and Concatenation
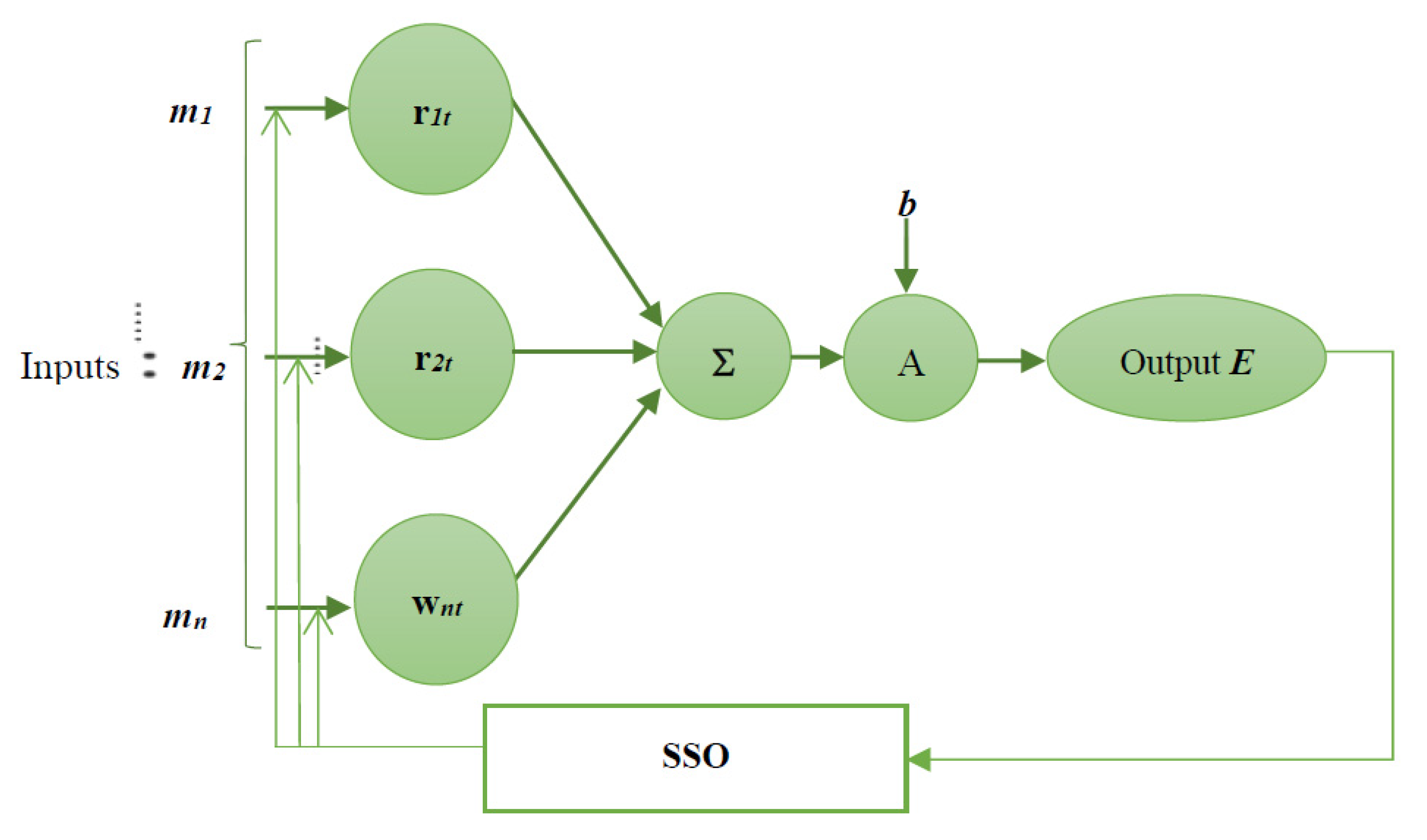
3.4. Proposed Social Spider Optimisation Tuned Neural Network Classifier in Cancer Diagnosis
- The ANN is initialised with a sequential layerann = tf.keras.models.Sequential()
- The fully connected input and the first hidden layer is added as a dense layer to the sequential layer with a uniform initialisation layer with a ‘relu’ activation function.ann.add(tf.keras.layers.Dense(units = 6, activation = ‘relu’))
- The second fully connected layer is added to the existing dense layer with the same ‘relu’ activation function.ann.add(tf.keras.layers.Dense(units = 6, activation = ‘relu’))
- Finally a fully connected output layer is added to the existing dense layer with the ‘sigmoid’ activation function.ann.add(tf.keras.layers.Dense(units = 1, activation = ‘sigmoid’))
3.5. Social Spider Optimisation Algorithm in an Update of NN Weights
| Algorithm 1. Pseudocode of SSO algorithm |
| Initialise the population of social spiders |
| Initialise the target vibration |
| Initialise the parameters, and |
| Evaluate the fitness measure for all social spiders |
| For all social spiders, |
| { |
| Calculate the vibration intensity |
| Select the strongest vibration among all |
| { |
| Store position of foraging spider as the best solution |
| } |
| While |
| { |
| Update the position of the foraging spider as per Equation (8) |
| } |
| End For |
| Update the parameters, and |
| Evaluate fitness of all social spiders |
| Sort the positions as per fitness measure (accuracy) |
| } |
| End For |
| Return |
4. Results and Discussions
4.1. Experimental Setup
4.2. Evaluation Metrics
4.3. Comparative Analysis of Methods Involved in the Diagnosis of Cancer
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Alix-Panabières, C.; Pantel, K. Circulating Tumor Cells: Liquid Biopsy of Cancer. Clin. Chem. 2013, 59, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Fowler, J.E.; Bigler, S.A.; Miles, D.; Yalkut, D.A. Predictors of first repeat biopsy cancer detection with suspected local stage prostate cancer. J. Urol. 2000, 163, 813–818. [Google Scholar] [CrossRef]
- Murtaza, G.; Shuib, L.; Wahab, A.W.A.; Mujtaba, G.; Raza, G.; Azmi, N.A. Breast cancer classification using digital biopsy histopathology images through transfer learning. J. Phys. Conf. Ser. 2019, 1339, 012035. [Google Scholar] [CrossRef]
- Kasivisvanathan, V.; Rannikko, A.S.; Borghi, M.; Panebianco, V.; Mynderse, L.A.; Vaarala, M.H.; Briganti, A.; Budäus, L.; Hellawell, G.; Hindley, R.G.; et al. MRI-Targeted or Standard Biopsy for Prostate-Cancer Diagnosis. N. Engl. J. Med. 2018, 378, 1767–1777. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, O.; Kanavati, F.; Kato, K.; Rambeau, M.; Arihiro, K.; Tsuneki, M. Deep Learning Models for Histopathological Classification of Gastric and Colonic Epithelial Tumours. Sci. Rep. 2020, 10, 1504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhattacharjee, S.; Park, H.-G.; Kim, C.-H.; Prakash, D.; Madusanka, N.; So, J.-H.; Cho, N.-H.; Choi, H.-K. Quantitative Analysis of Benign and Malignant Tumors in Histopathology: Predicting Prostate Cancer Grading Using SVM. Appl. Sci. 2019, 9, 2969. [Google Scholar] [CrossRef] [Green Version]
- Khan, I.U.; Aslam, N.; Alshehri, R.; Alzahrani, S.; Alghamdi, M.; Almalki, A.; Balabeed, M. Cervical Cancer Diagnosis Model Using Extreme Gradient Boosting and Bioinspired Firefly Optimization. Sci. Program. 2021, 2021, 5540024. [Google Scholar] [CrossRef]
- Alheejawi, S.; Mandal, M.; Berendt, R.; Jha, N. Automated melanoma staging in lymph node biopsy image using deep learning. In Proceedings of the 2019 IEEE Canadian Conference of Electrical and Computer Engineering (CCECE), Edmonton, AB, Canada, 5–8 May 2019; pp. 1–4. [Google Scholar]
- Bacanin, N.; Tuba, M. Artificial Bee Colony (ABC) Algorithm for Constrained Optimisation Improved with Genetic Operators. Stud. Inform. Control 2012, 21, 137–146. [Google Scholar] [CrossRef]
- Dorigo, M.; Birattari, M. Ant Colony Optimization; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar]
- Yang, X.S. Firefly Algorithms for Multimodal Optimization. In Stochastic Algorithms: Foundations and Applications; Watanabe, O., Zeugmann, T., Eds.; Springer: Berlin/Heidelberg, Germany, 2009; pp. 169–178. [Google Scholar]
- Strumberger, I.; Bacanin, N.; Tuba, M. Enhanced Firefly Algorithm for Constrained Numerical Optimization, IEEE Congress on Evolutionary Computation. In Proceedings of the IEEE International Congress on Evolutionary Computation (CEC 2017), San Sebastián, Spain, 5–8 June 2017; pp. 2120–2127. [Google Scholar]
- Gandomi, A.H.; Yang, X.S.; Alavi, A.H. Cuckoo search algorithm: A metaheuristic approach to solve structural optimisation problems. Eng. Comput. 2013, 29, 17–35. [Google Scholar] [CrossRef]
- Yang, X.S.; Hossein Gandomi, A. Bat algorithm: A novel approach for global engineering optimisation. Eng. Comput. 2012, 29, 464–483. [Google Scholar] [CrossRef] [Green Version]
- Mirjalili, S.; Lewis, A. The whale optimisation algorithm. Adv. Eng. Softw. 2016, 95, 51–67. [Google Scholar] [CrossRef]
- Wang, G.G.; Deb, S.; dos S. Coelho, L. Elephant Herding Optimization. In Proceedings of the 2015 3rd International Symposium on Computational and Business Intelligence (ISCBI), Bali, Indonesia, 7–9 December 2015; pp. 1–5. [Google Scholar]
- Strumberger, I.; Bacanin, N.; Tuba, M. Hybridized Elephant Herding Optimization Algorithm for Constrained Optimization. In Hybrid Intelligent Systems; Abraham, A., Muhuri, P.K., Muda, A.K., Gandhi, N., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 158–166. [Google Scholar]
- Nayak, M.; Das, S.; Bhanja, U.; Senapati, M.R. Elephant herding optimisation technique based neural network for cancer prediction. Inform. Med. Unlocked 2020, 21, 10045. [Google Scholar] [CrossRef]
- Mirjalili, S.; Mirjalili, S.M.; Lewis, A. Grey wolf optimizer. Adv. Eng. Softw. 2014, 69, 46–61. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.S. Flower pollination algorithm for global optimisation. In Proceedings of the International Conference on Unconventional Computing and Natural Computation, Orléans, France, 3–7 September 2012; pp. 240–249. [Google Scholar]
- Preethi, S.; Aishwarya, P. Combining Wavelet Texture Features and Deep Neural Network for Tumor Detection and Segmentation Over MRI. J. Intell. Syst. 2019, 28, 571–588. [Google Scholar] [CrossRef]
- Sayed, G.I.; Hassanien, A.E. Moth-flame swarm optimisation with neutrosophic sets for automatic mitosis detection in breast cancer histology images. Appl. Intell. 2017, 47, 397–408. [Google Scholar] [CrossRef]
- Bueno, G.; García-Rojo, M.; Déniz, O.; Vidal, J.; Galeotti, J.; Relea, F. A fully automated approach to prostate biopsy segmentation based on level-set and mean filtering. J. Pathol. Inform. 2011, 2, 5. [Google Scholar] [CrossRef] [PubMed]
- Stephan, C.; Cammann, H.; Semjonow, A.; Diamandis, E.P.; Wymenga, L.F.; Lein, M.; Sinha, P.; A Loening, S.; Jung, K. Multicenter Evaluation of an Artificial Neural Network to Increase the Prostate Cancer Detection Rate and Reduce Unnecessary Biopsies. Clin. Chem. 2002, 48, 1279–1287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- James, J.Q.; Li, V.O. A Social Spider Algorithm for global optimisation. Appl. Soft Comput. 2015, 30, 614–627. [Google Scholar]
- Breast Cancer Histopathological Database (BreakHis). Available online: https://web.inf.ufpr.br/vri/databases/breast-cancer-histopathological-database-breakhis/ (accessed on 10 November 2021).
- Mirjalili, S. Evolutionary Algorithms and Neural Networks. In Studies in Computational Intelligence; Springer: Berlin/Heidelberg, Germany, 2019. [Google Scholar] [CrossRef] [Green Version]
- Poli, R.; James, K.; Tim, B. Particle swarm optimisation. Swarm Intell. 2007, 1, 33–57. [Google Scholar] [CrossRef]
- Arora, S.; Singh, S. The Firefly Optimization Algorithm: Convergence Analysis and Parameter Selection. Int. J. Comput. Appl. 2013, 69, 48–52. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balaji, P.; Chidambaram, K. Cancer Diagnosis of Microscopic Biopsy Images Using a Social Spider Optimisation-Tuned Neural Network. Diagnostics 2022, 12, 11. https://doi.org/10.3390/diagnostics12010011
Balaji P, Chidambaram K. Cancer Diagnosis of Microscopic Biopsy Images Using a Social Spider Optimisation-Tuned Neural Network. Diagnostics. 2022; 12(1):11. https://doi.org/10.3390/diagnostics12010011
Chicago/Turabian StyleBalaji, Prasanalakshmi, and Kumarappan Chidambaram. 2022. "Cancer Diagnosis of Microscopic Biopsy Images Using a Social Spider Optimisation-Tuned Neural Network" Diagnostics 12, no. 1: 11. https://doi.org/10.3390/diagnostics12010011
APA StyleBalaji, P., & Chidambaram, K. (2022). Cancer Diagnosis of Microscopic Biopsy Images Using a Social Spider Optimisation-Tuned Neural Network. Diagnostics, 12(1), 11. https://doi.org/10.3390/diagnostics12010011






