Residual-Shuffle Network with Spatial Pyramid Pooling Module for COVID-19 Screening
Abstract
:1. Introduction
2. Convolutional Neural Networks for COVID-19 Detection
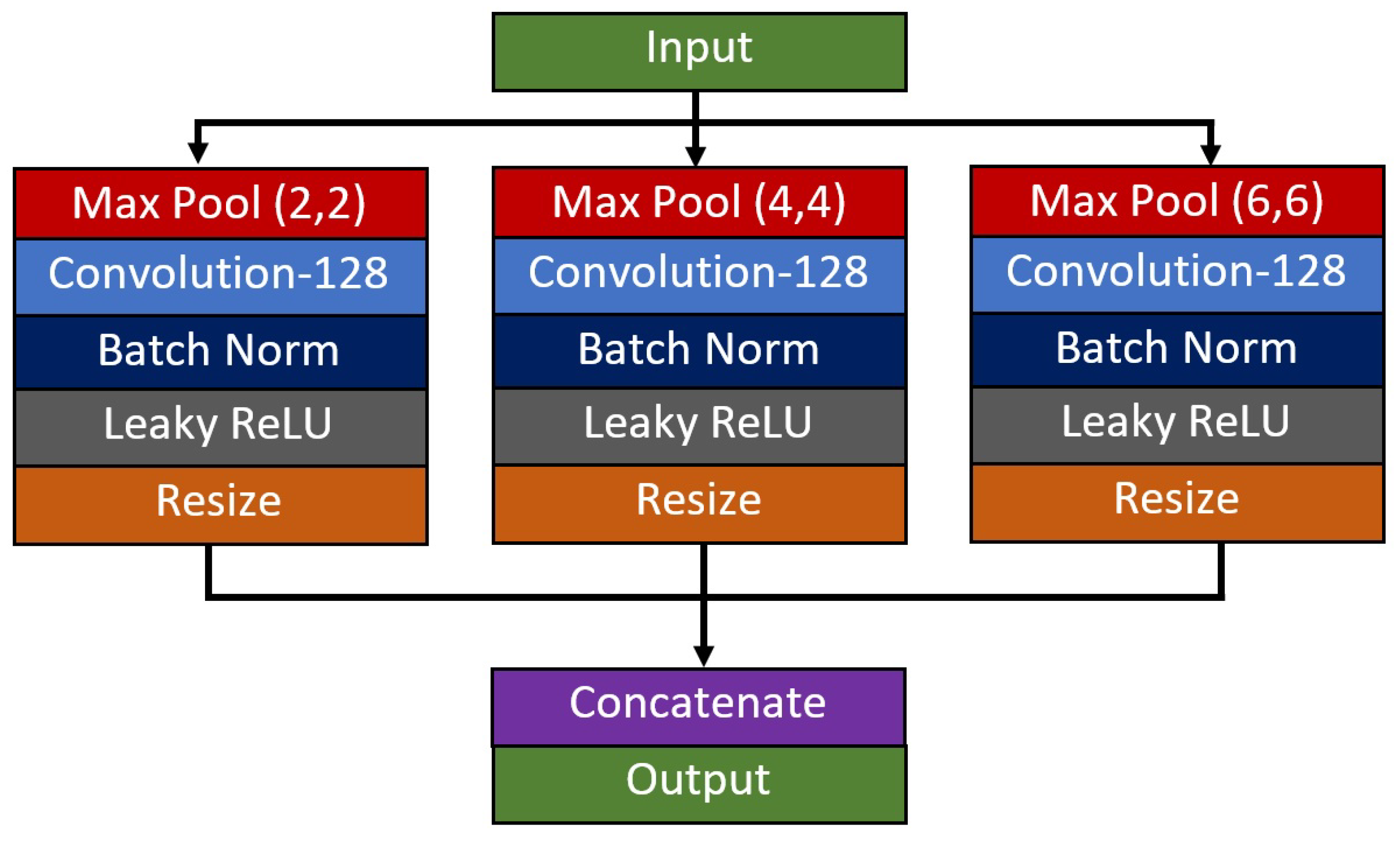
3. Residual-Shuffle Network with Spatial Pyramid Pooling Module
4. Experiments and Discussion
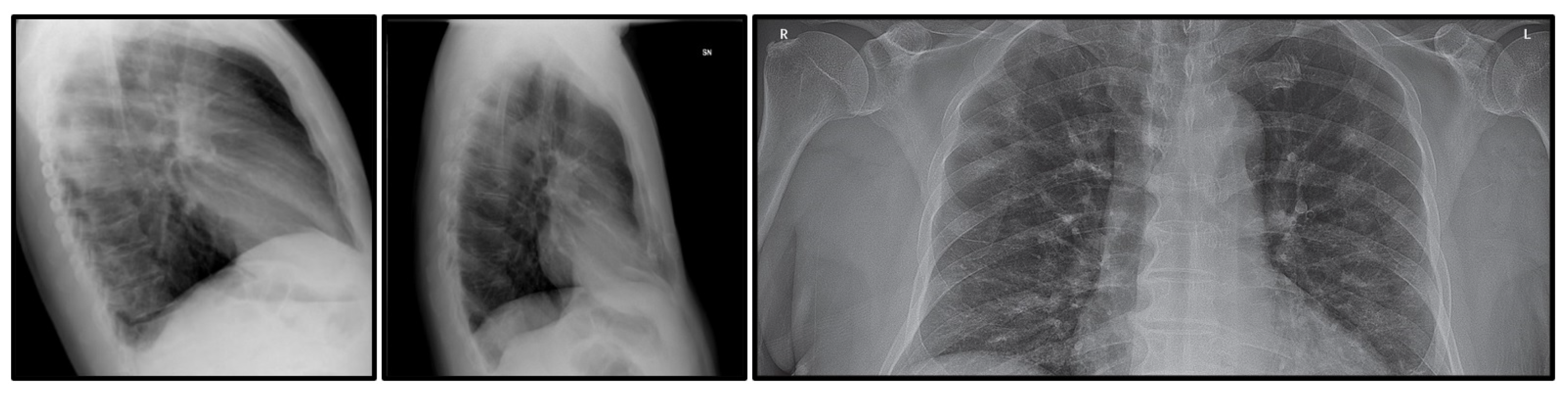
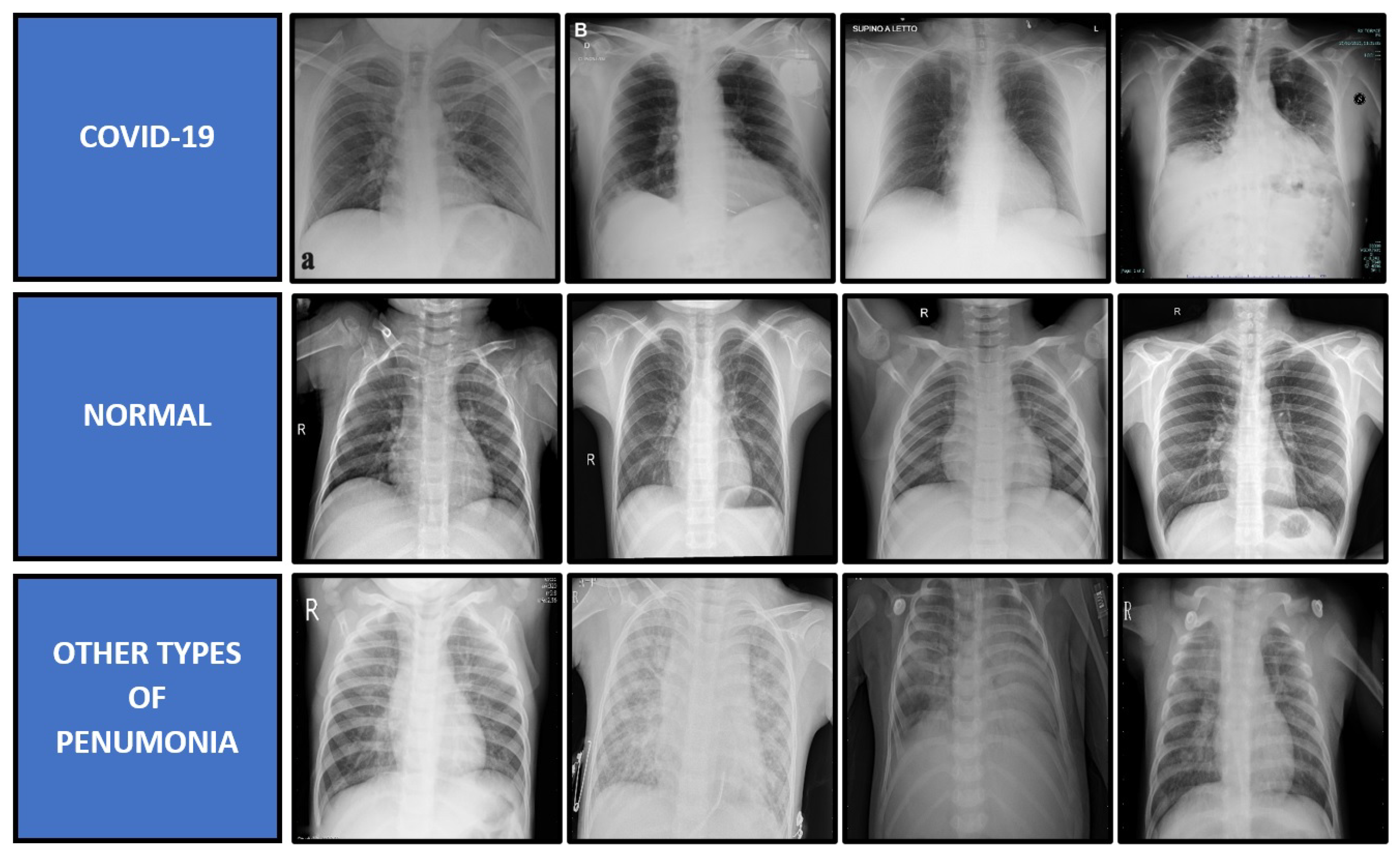
4.1. Dataset
4.2. Evaluation Metrics
4.3. Experimental Setup
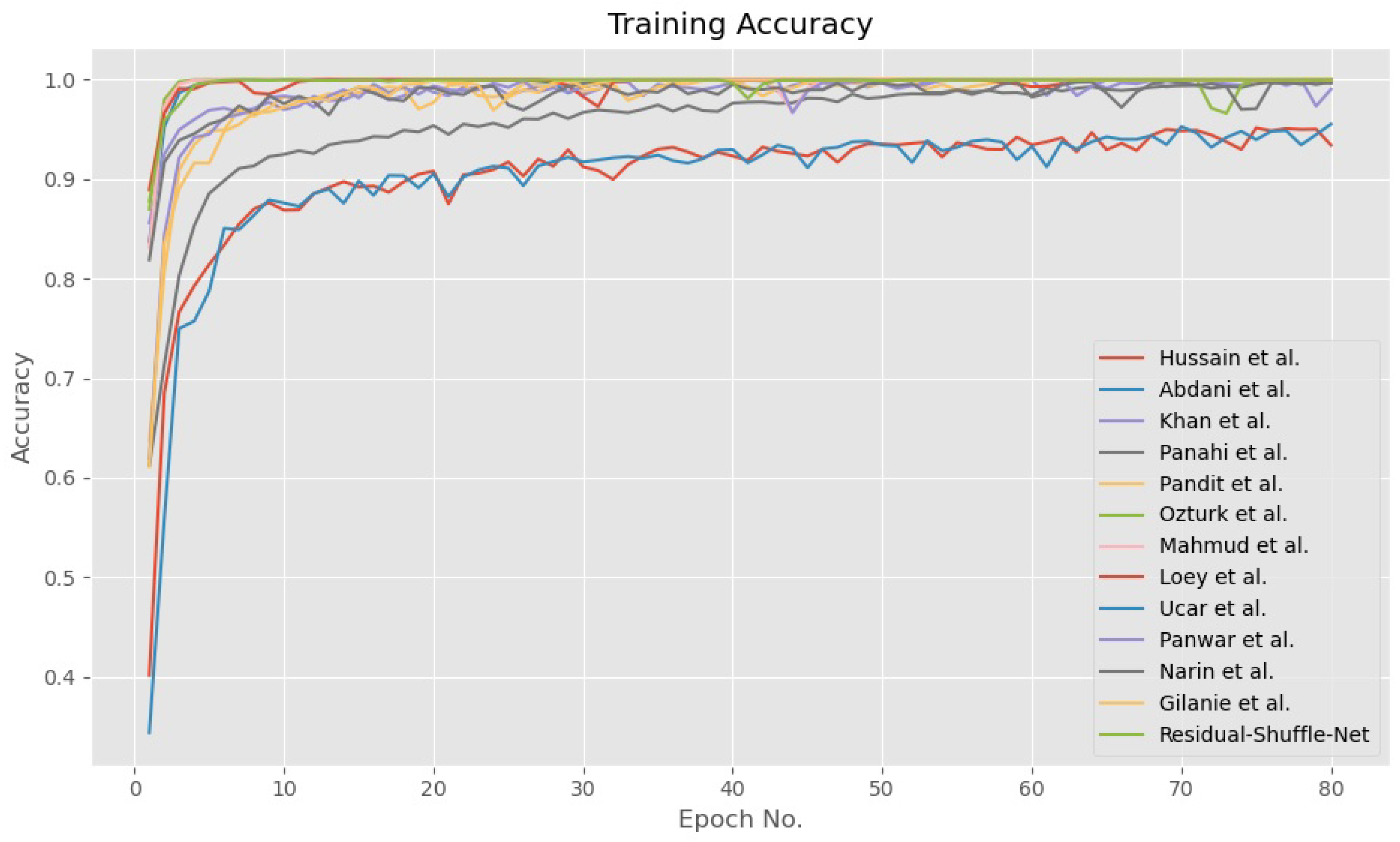
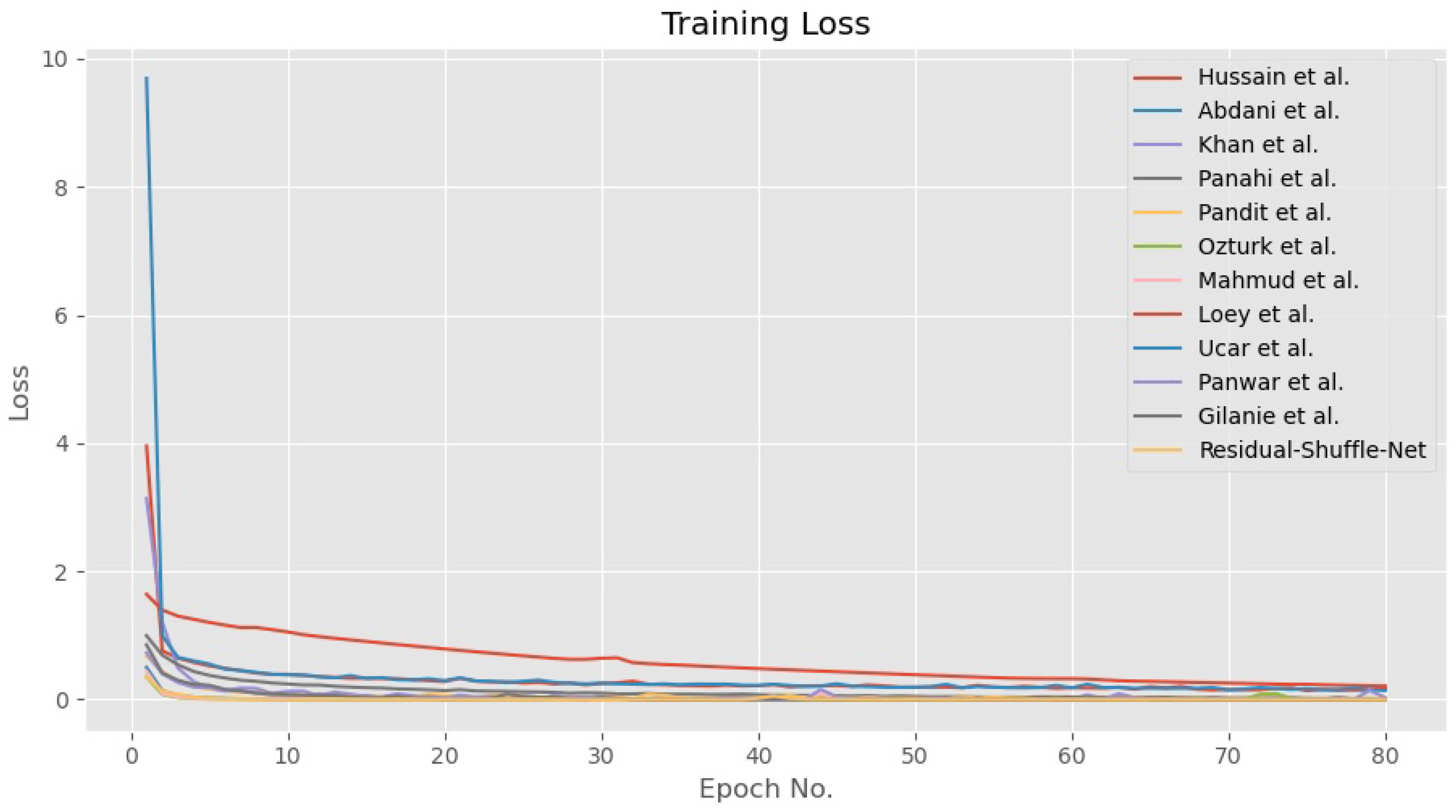
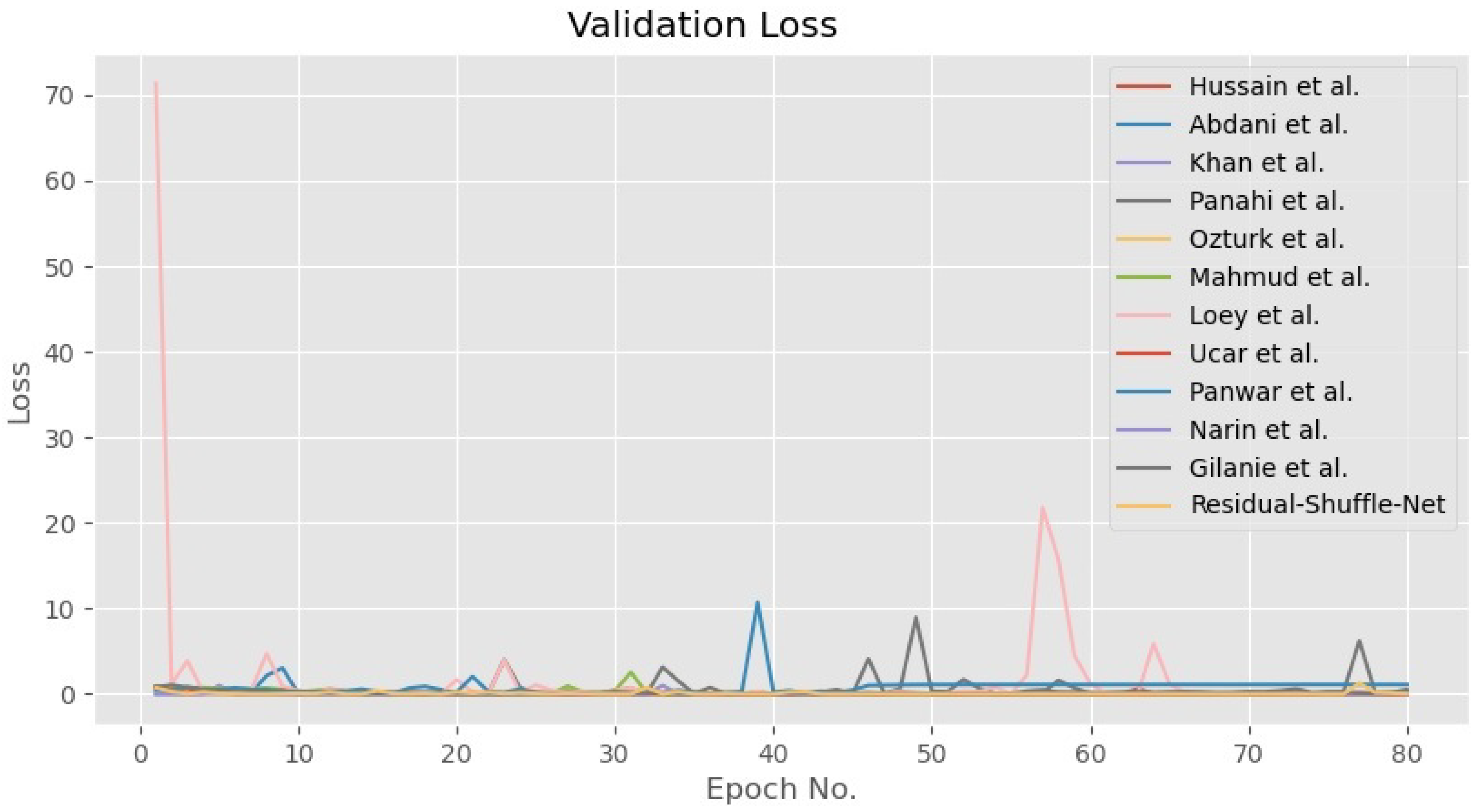
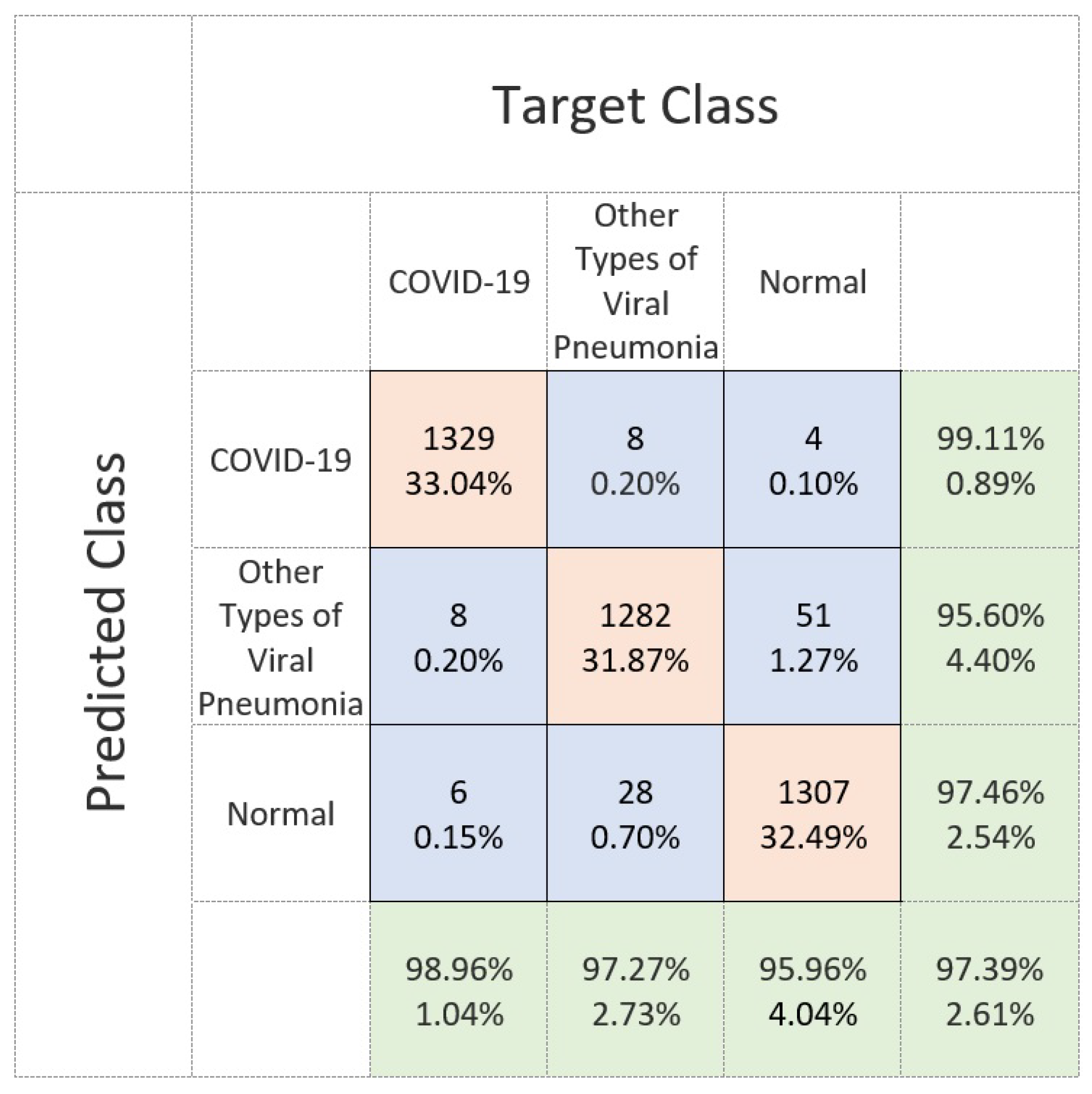
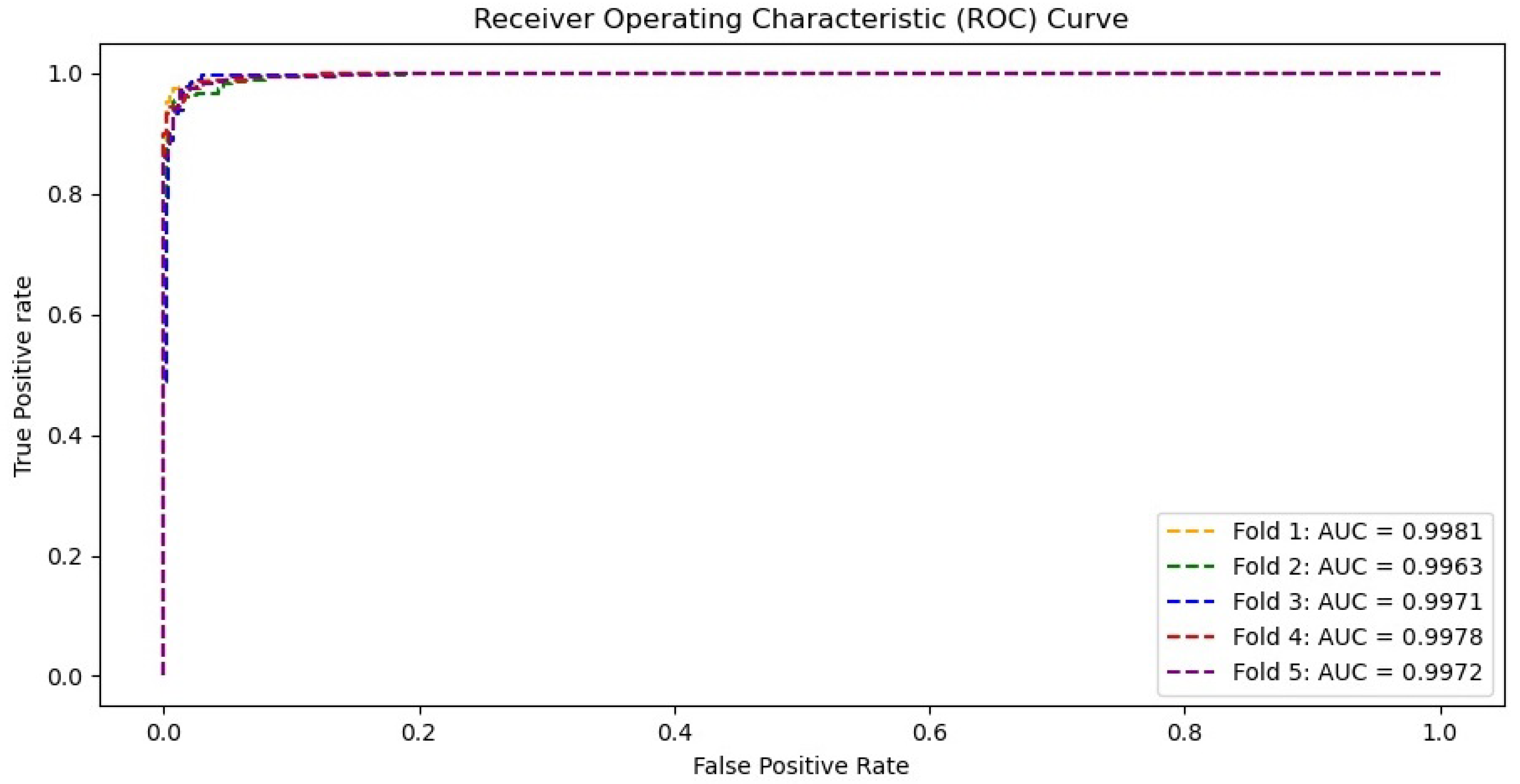
4.4. Discussion on the Residual-Shuffle-Net and Its Benchmarked Models Performance
5. Limitations
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
Abbreviations
| SPP | Spatial Pyramid Pooling |
| CNN | Convolutional Neural Networks |
| COVID-19 | Coronavirus Disease 2019 |
| RT-PCR | Reverse Transcription Polymerase Chain Reaction |
| CT | Computed Tomography |
| GAN | Generative Adversarial Network |
| Residual-Shuffle-Net | Residual-Shuffle Network |
| Leaky ReLU | Leaky Rectified Linear Unit |
| BIMCV | Medical Imaging Databank of the Valencia Region |
| RSNA | Radiological Society of North America |
| DICOM | Digital Imaging and Communications in Medicine |
| PNG | Portable Network Graphics |
| ROC | Receiver Operating Characteristic |
References
- Lancet, T. COVID-19: The worst may be yet to come. Lancet 2020, 396, 71. [Google Scholar] [CrossRef]
- da Silva, J.C.; Félix, V.B.; Leão, S.A.B.F.; Trindade Filho, E.M.; Scorza, F.A. New Brazilian Variant Of The Sars-Cov-2 (P1) of Covid-19 in Alagoas State. Braz. J. Infect. Dis. 2021, 25, 101588. [Google Scholar] [CrossRef]
- Vallee, A.; Chan-Hew-Wai, A.; Bonan, B.; Lesprit, P.; Parquin, F.; Catherinot, É.; Choucair, J.; Billard, D.; Amiel-Taieb, C.; Camps, È.; et al. Oxford—AstraZeneca COVID-19 vaccine: Need of a reasoned and effective vaccine campaign. Public Health 2021, 196, 135–137. [Google Scholar] [CrossRef] [PubMed]
- McGrath, J.; Kenny, C.; Smyth, H.; McGinty, T.; Sheehan, G.; Gaine, S.; McCullagh, B.; MacMahon, P.; Egan, J.; Cotter, A. A multidisciplinary evaluation of suspected, non-confirmed cases of COVID-19 including chest CT, as compared to World Health Organization recommendations. Clin. Radiol. 2021, 76, 384–390. [Google Scholar] [CrossRef] [PubMed]
- Bruzzone, B.; De Pace, V.; Caligiuri, P.; Ricucci, V.; Guarona, G.; Pennati, B.M.; Boccotti, S.; Orsi, A.; Domnich, A.; Da Rin, G.; et al. Comparative diagnostic performance of rapid antigen detection tests for COVID-19 in a hospital setting. Int. J. Infect. Dis. 2021, 107, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Bouassa, R.S.M.; Veyer, D.; Péré, H.; Bélec, L. Analytical performances of the point-of-care SIENNA™ COVID-19 Antigen Rapid Test for the detection of SARS-CoV-2 nucleocapsid protein in nasopharyngeal swabs: A prospective evaluation during the COVID-19 s wave in France. Int. J. Infect. Dis. 2021, 106, 8–12. [Google Scholar] [CrossRef]
- Book COVID—19 Drive—Thru, Clinic and Home Screening Test Services Online. 2021. Available online: https://www.doctoroncall.com.my/medicine/coronavirus-covid-19-test-kit (accessed on 21 June 2021).
- Abdani, S.R.; Zulkifley, M.A.; Zulkifley, N.H. A Lightweight Deep Learning Model for Covid-19 Detection. In Proceedings of the IEEE Symposium on Industrial Electronics & Applications (ISIEA), Shah Alam, Malaysia, 17–18 July 2020. [Google Scholar]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (covid-19) using X-ray images and deep convolutional neural networks. arXiv 2020, arXiv:2003.10849. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. CoRR 2015. Available online: http://xxx.lanl.gov/abs/1512.03385 (accessed on 6 June 2021).
- He, K.; Zhang, X.; Ren, S.; Sun, J. Spatial Pyramid Pooling in Deep Convolutional Networks for Visual Recognition. In Proceedings of the Computer Vision—ECCV, Zurich, Switzerland, 5–12 September 2014; Fleet, D., Pajdla, T., Schiele, B., Tuytelaars, T., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 346–361. [Google Scholar]
- Abdani, S.R.; Zulkifley, M.A. DenseNet with Spatial Pyramid Pooling for Industrial Oil Palm Plantation Detection. In Proceedings of the 2019 International Conference on Mechatronics, Robotics and Systems Engineering, Bali, Indonesia, 4–6 December 2019. [Google Scholar]
- Zulkifley, M.A.; Trigoni, N. Multiple-Model Fully Convolutional Neural Networks for Single Object Tracking on Thermal Infrared Video. IEEE Access 2018, 6, 42790–42799. [Google Scholar] [CrossRef]
- Sumbul, G.; Demir, B. A Novel Multi-Attention Driven System for Multi-Label Remote Sensing Image Classification. In Proceedings of the IGARSS 2019—2019 IEEE International Geoscience and Remote Sensing Symposium, Yokohama, Japan, 28 July–2 August 2019; pp. 5726–5729. [Google Scholar] [CrossRef] [Green Version]
- Shao, G.; Chen, Y.; Wei, Y. Deep Fusion for Radar Jamming Signal Classification Based on CNN. IEEE Access 2020, 8, 117236–117244. [Google Scholar] [CrossRef]
- Zulkifley, M.A.; Abdani, S.R.; Zulkifley, N.H. Pterygium-Net: A deep learning approach to pterygium detection and localization. Multimed. Tools Appl. 2019, 78, 34563–34584. [Google Scholar] [CrossRef]
- Lal, H.; Nguyen, T.; Li, H.; Abbasi, A.A.; Lone, K.J.; Zhao, Z.; Zaib, M.; Chen, A.; Duong, T.Q. Machine-learning classification of texture features of portable chest X-ray accurately classifies COVID-19 lung infection. Biomed. Eng. Online 2020, 19, 88. [Google Scholar]
- Sverzellati, N.; Ryerson, C.J.; Milanese, G.; Renzoni, E.A.; Volpi, A.; Spagnolo, P.; Bonella, F.; Comelli, I.; Affanni, P.; Veronesi, L.; et al. Chest X-ray or CT for COVID-19 pneumonia? Comparative study in a simulated triage setting. Eur. Respir. J. 2021. [Google Scholar] [CrossRef]
- Romanov, A.; Bach, M.; Yang, S.; Franzeck, F.C.; Sommer, G.; Anastasopoulos, C.; Bremerich, J.; Stieltjes, B.; Weikert, T.; Sauter, A.W. Automated CT Lung Density Analysis of Viral Pneumonia and Healthy Lungs Using Deep Learning-Based Segmentation, Histograms and HU Thresholds. Diagnostics 2021, 11, 738. [Google Scholar] [CrossRef]
- Pham, T.D. Classification of COVID-19 chest X-rays with deep learning: New models or fine tuning? Health Inf. Sci. Syst. 2021, 9, 1–11. [Google Scholar] [CrossRef]
- Pandit, M.K.; Banday, S.A.; Naaz, R.; Chishti, M.A. Automatic detection of COVID-19 from chest radiographs using deep learning. Radiography 2021, 27, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Panwar, H.; Gupta, P.; Siddiqui, M.K.; Morales-Menendez, R.; Singh, V. Application of deep learning for fast detection of COVID-19 in X-rays using nCOVnet. Chaos Solitons Fractals 2020, 138, 109944. [Google Scholar] [CrossRef] [PubMed]
- Kikkisetti, S.; Zhu, J.; Shen, B.; Li, H.; Duong, T.Q. Deep-learning convolutional neural networks with transfer learning accurately classify COVID-19 lung infection on portable chest radiographs. PeerJ 2020, 8, e10309. [Google Scholar] [CrossRef] [PubMed]
- Apostolopoulos, I.D.; Mpesiana, T.A. Covid-19: Automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys. Eng. Sci. Med. 2020, 43, 635–640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition; Technical Report; University of Oxford: Oxford, UK, 2014. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L. MobileNetV2: Inverted Residuals and Linear Bottlenecks. In Proceedings of the 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–22 June 2018; pp. 4510–4520. [Google Scholar]
- Szegedy, C.; Wei, L.; Yangqing, J.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar] [CrossRef] [Green Version]
- Chollet, F. Xception: Deep Learning with Depthwise Separable Convolutions. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 1800–1807. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception-v4, inception-resnet and the impact of residual connections on learning. In Proceedings of the AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; Volume 31. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826. [Google Scholar]
- Loey, M.; Smarandache, F.; Khalifa, M.E.N. Within the Lack of Chest COVID-19 X-ray Dataset: A Novel Detection Model Based on GAN and Deep Transfer Learning. Symmetry 2020, 12, 651. [Google Scholar] [CrossRef] [Green Version]
- Dewi, C.; Chen, R.C.; Liu, Y.T.; Jiang, X.; Hartomo, K.D. Yolo V4 for Advanced Traffic Sign Recognition With Synthetic Training Data Generated by Various GAN. IEEE Access 2021, 9, 97228–97242. [Google Scholar] [CrossRef]
- Zulkifley, M.A.; Abdani, S.R.; Zulkifley, N.H. COVID-19 Screening Using a Lightweight Convolutional Neural Network with Generative Adversarial Network Data Augmentation. Symmetry 2020, 12, 1530. [Google Scholar] [CrossRef]
- Ucar, F.; Korkmaz, D. COVIDiagnosis-Net: Deep Bayes-SqueezeNet based diagnosis of the coronavirus disease 2019 (COVID-19) from X-ray images. Med. Hypotheses 2020, 140, 109761. [Google Scholar] [CrossRef] [PubMed]
- Iandola, F.N.; Moskewicz, M.W.; Ashraf, K.; Han, S.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50× fewer parameters and <1 MB model size. CoRR 2016. Available online: http://xxx.lanl.gov/abs/1602.07360 (accessed on 6 June 2021).
- Muad, A.M.; Zaki, S.K.M.; Jasim, S.A. Optimizing Hopfield Neural Network for Super-Resolution Mapping. J. Kejuruter. 2020, 32, 91–97. [Google Scholar] [CrossRef]
- Khan, A.I.; Shah, J.L.; Bhat, M.M. CoroNet: A deep neural network for detection and diagnosis of COVID-19 from chest X-ray images. Comput. Methods Programs Biomed. 2020, 196, 105581. [Google Scholar] [CrossRef]
- Panahi, A.H.; Rafiei, A.; Rezaee, A. FCOD: Fast COVID-19 Detector based on deep learning techniques. Inform. Med. Unlocked 2021, 22, 100506. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, T.; Rahman, M.A.; Fattah, S.A. CovXNet: A multi-dilation convolutional neural network for automatic COVID-19 and other pneumonia detection from chest X-ray images with transferable multi-receptive feature optimization. Comput. Biol. Med. 2020, 122, 103869. [Google Scholar] [CrossRef]
- Gilanie, G.; Bajwa, U.I.; Waraich, M.M.; Asghar, M.; Kousar, R.; Kashif, A.; Aslam, R.S.; Qasim, M.M.; Rafique, H. Coronavirus (COVID-19) detection from chest radiology images using convolutional neural networks. Biomed. Signal Process. Control 2021, 66, 102490. [Google Scholar] [CrossRef]
- Hussain, E.; Hasan, M.; Rahman, M.A.; Lee, I.; Tamanna, T.; Parvez, M.Z. CoroDet: A deep learning based classification for COVID-19 detection using chest X-ray images. Chaos Solitons Fractals 2021, 142, 110495. [Google Scholar] [CrossRef]
- Zhu, J.; Shen, B.; Abbasi, A.; Hoshmand-Kochi, M.; Li, H.; Duong, T.Q. Deep transfer learning artificial intelligence accurately stages COVID-19 lung disease severity on portable chest radiographs. PLoS ONE 2020, 15, e0236621. [Google Scholar] [CrossRef]
- Wong, A.; Lin, Z.Q.; Wang, L.; Chung, A.G.; Shen, B.; Abbasi, A.; Hoshmand-Kochi, M.; Duong, T.Q. Towards computer-aided severity assessment via deep neural networks for geographic and opacity extent scoring of SARS-CoV-2 chest X-rays. Sci. Rep. 2021, 11, 9315. [Google Scholar] [CrossRef] [PubMed]
- Ma, N.; Zhang, X.; Zheng, H.T.; Sun, J. Shufflenet V2: Practical guidelines for efficient cnn architecture design. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 116–131. [Google Scholar]
- Abdani, S.R.; Zulkifley, M.A.; Zulkifley, N.H. Group and Shuffle Convolutional Neural Networks with Pyramid Pooling Module for Automated Pterygium Segmentation. Diagnostics 2021, 11, 1104. [Google Scholar] [CrossRef]
- Vaya, M.d.l.I.; Saborit, J.M.; Montell, J.A.; Pertusa, A.; Bustos, A.; Cazorla, M.; Galant, J.; Barber, X.; Orozco-Beltrán, D.; Garcia, F.; et al. BIMCV COVID-19+: A large annotated dataset of RX and CT images from COVID-19 patients. arXiv 2020, arXiv:2006.01174. [Google Scholar]
- Wang, X.; Peng, Y.; Lu, L.; Lu, Z.; Bagheri, M.; Summers, R.M. ChestX-ray8: Hospital-scale chest X-ray database and benchmarks on weakly-supervised classification and localization of common thorax diseases. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2097–2106. [Google Scholar]
- Ozturk, T.; Talo, M.; Yildirim, E.A.; Baloglu, U.B.; Yildirim, O.; Acharya, U.R. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 2020, 121, 103792. [Google Scholar] [CrossRef] [PubMed]

| No. | Layer | Output Size | Filter Size | Kernel Size | Stride |
|---|---|---|---|---|---|
| 1 | Convolution | 256 × 256 | 8 | 3 × 3 | 1 × 1 |
| 2 | Max Pooling | 128 × 128 | - | 2 × 2 | 2 × 2 |
| 3 | Convolution | 128 × 128 | 16 | 3 × 3 | 1 × 1 |
| 4 | Max Pooling | 64 × 64 | - | 2 × 2 | 2 × 2 |
| 5 | Residual-Shuffle (1) | 64 × 64 | 32-16-32 | 3 × 3 − 1 × 1 − 3 × 3 | 1 × 1 |
| 6 | Max Pooling | 32 × 32 | - | 2 × 2 | 2 × 2 |
| 7 | Residual-Shuffle (2) | 32 × 32 | 64-32-64 | 3 × 3 − 1 × 1 − 3 × 3 | 1 × 1 |
| 8 | Max Pooling | 16 × 16 | - | 2 × 2 | 2 × 2 |
| 9 | Residual-Shuffle (3) | 16 × 16 | 128-64-128 | 3 × 3 − 1 × 1 − 3 × 3 | 1 × 1 |
| 10 | Max Pooling | 8 × 8 | - | 2 × 2 | 2 × 2 |
| 11 | Residual-Shuffle (4) | 8 × 8 | 256-128-256 | 3 × 3 − 1 × 1 − 3 × 3 | 1 × 1 |
| 12 | Spatial Pyramid Pooling | 8 × 8 | 128-128-128 | 2 × 2 − 4 × 4 − 6 × 6 | 1 × 1 |
| 13 | Global Average Pooling | 1 × 1 | - | 8 × 8 | 1 × 1 |
| 14 | Dense + SoftMax | - | 3 | 1 × 1 | 1 × 1 |
| Criteria | Hyper-Parameter Setting |
|---|---|
| Batch size | 64 |
| Training epoch | 80 |
| Backpropagation method | Adam optimizer |
| Input image size | 256 × 256 pixels |
| Optimizer learning rate | 0.0001 |
| Optimizer momentums | = 0.9, = 0.999 |
| Loss function | categorical cross-entropy |
| Labeling format | One-hot encoded |
| Method | Total Parameters | Trainable Parameters | |||||
|---|---|---|---|---|---|---|---|
| Hussain et al. [41] | 0.78695 | 0.78695 | 0.89347 | 0.81998 | 0.77785 | 3,798,083 | 3,798,083 |
| Narin et al. [9] | 0.83765 | 0.76155 | 0.88078 | 0.8376 | 0.72718 | 23,567,299 | 23,514,179 |
| Ozturk et al. [48] | 0.84629 | 0.84629 | 0.92318 | 0.92572 | 0.81911 | 1,167,363 | 1,164,143 |
| Ucar et al. [34] | 0.92841 | 0.92841 | 0.96420 | 0.93130 | 0.92866 | 1,078,211 | 1,078,211 |
| Mahmud et al. [39] | 0.93686 | 0.93686 | 0.96843 | 0.94025 | 0.93609 | 1,338,291 | 1,164,143 |
| Loey et al. [31] | 0.94058 | 0.94058 | 0.97029 | 0.94387 | 0.94067 | 11,192,003 | 11,182,275 |
| Panahi et al. [38] | 0.94406 | 0.94406 | 0.97203 | 0.94674 | 0.94385 | 85,321 | 83,849 |
| Pandit et al. [21] | 0.94858 | 0.94858 | 0.97429 | 0.94859 | 0.94834 | 33,609,539 | 33,609,539 |
| Panwar et al. [22]. | 0.95700 | 0.95700 | 0.978500 | 0.95746 | 0.95692 | 14,747,715 | 14,747,715 |
| Khan et al. [37] | 0.96247 | 0.96247 | 0.98123 | 0.96334 | 0.96247 | 88,054,139 | 87,958,763 |
| Abdani et al. [8] | 0.96395 | 0.96395 | 0.98197 | 0.96454 | 0.96388 | 1,862,331 | 859,883 |
| Gilanie et al. [40] | 0.96868 | 0.96868 | 0.98434 | 0.96884 | 0.96863 | 2,339,267 | 2,339,267 |
| Residual-Shuffle-Net | 0.97390 | 0.97390 | 0.98695 | 0.97403 | 0.97387 | 2,090,491 | 2,087,275 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zulkifley, M.A.; Abdani, S.R.; Zulkifley, N.H.; Shahrimin, M.I. Residual-Shuffle Network with Spatial Pyramid Pooling Module for COVID-19 Screening. Diagnostics 2021, 11, 1497. https://doi.org/10.3390/diagnostics11081497
Zulkifley MA, Abdani SR, Zulkifley NH, Shahrimin MI. Residual-Shuffle Network with Spatial Pyramid Pooling Module for COVID-19 Screening. Diagnostics. 2021; 11(8):1497. https://doi.org/10.3390/diagnostics11081497
Chicago/Turabian StyleZulkifley, Mohd Asyraf, Siti Raihanah Abdani, Nuraisyah Hani Zulkifley, and Mohamad Ibrani Shahrimin. 2021. "Residual-Shuffle Network with Spatial Pyramid Pooling Module for COVID-19 Screening" Diagnostics 11, no. 8: 1497. https://doi.org/10.3390/diagnostics11081497
APA StyleZulkifley, M. A., Abdani, S. R., Zulkifley, N. H., & Shahrimin, M. I. (2021). Residual-Shuffle Network with Spatial Pyramid Pooling Module for COVID-19 Screening. Diagnostics, 11(8), 1497. https://doi.org/10.3390/diagnostics11081497







