Passive Microwave Radiometry for the Diagnosis of Coronavirus Disease 2019 Lung Complications in Kyrgyzstan
Abstract
1. Introduction
2. Materials and Methods
Statistical Approach
3. Results
3.1. Clinical Results
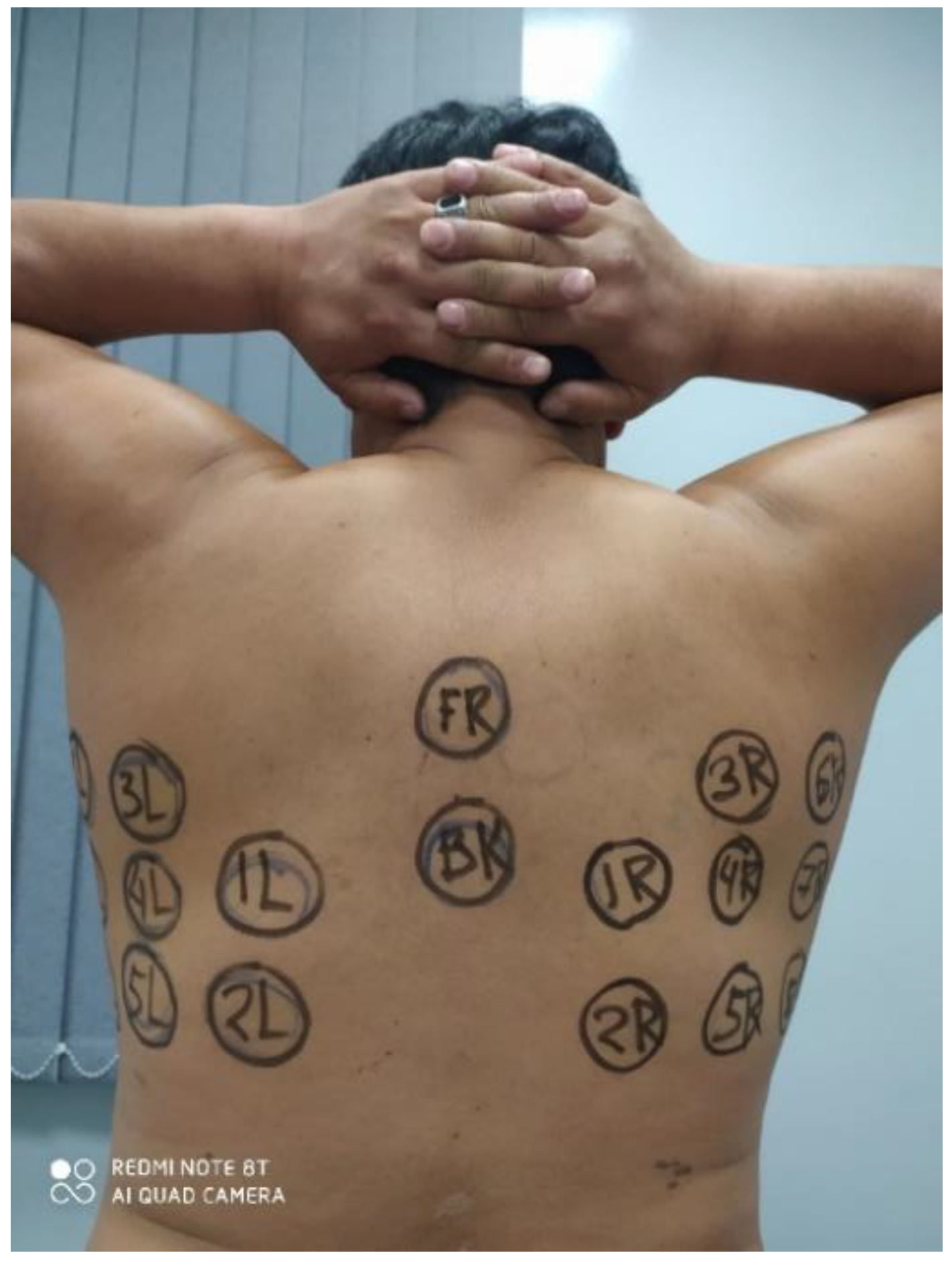
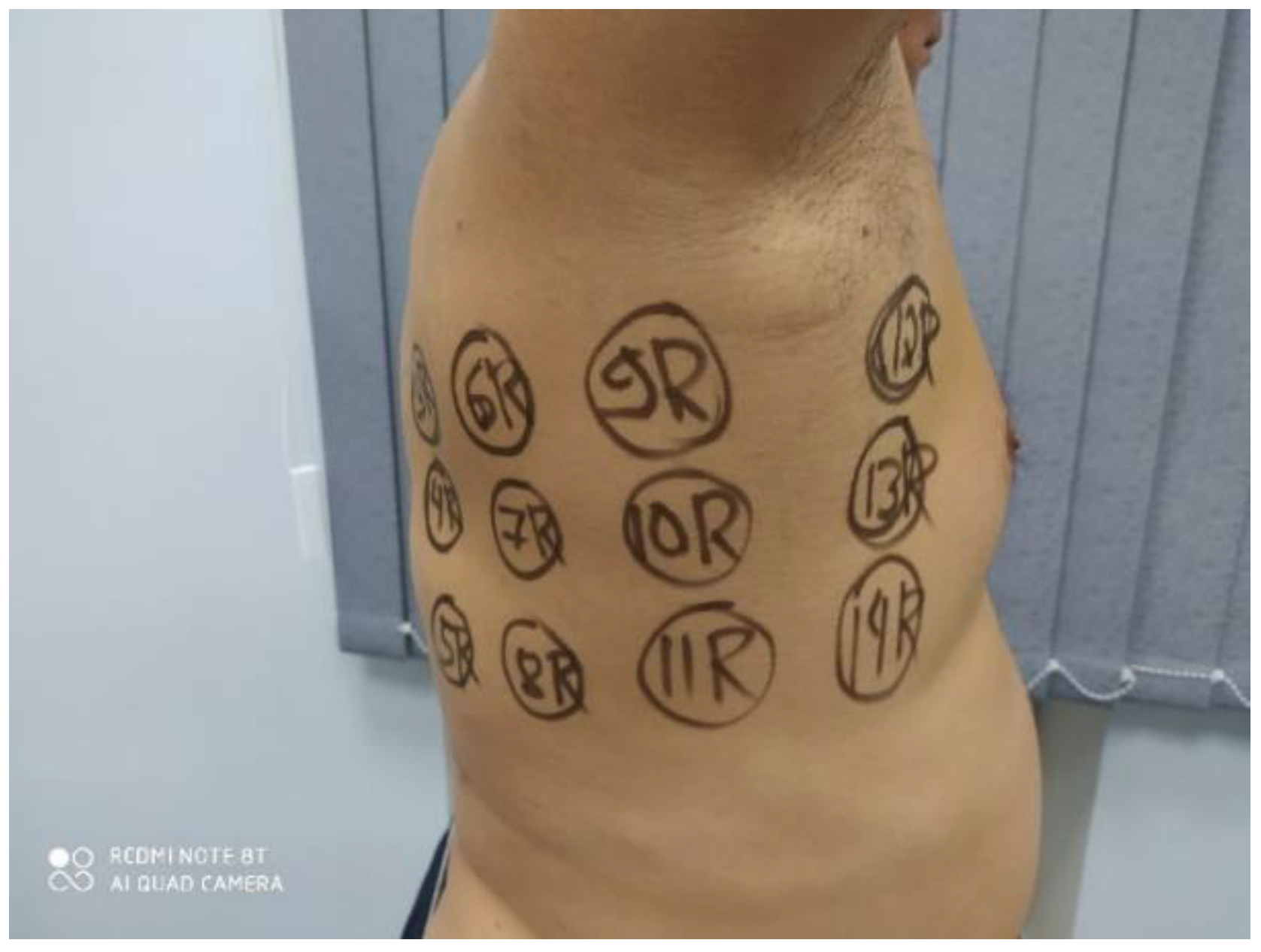
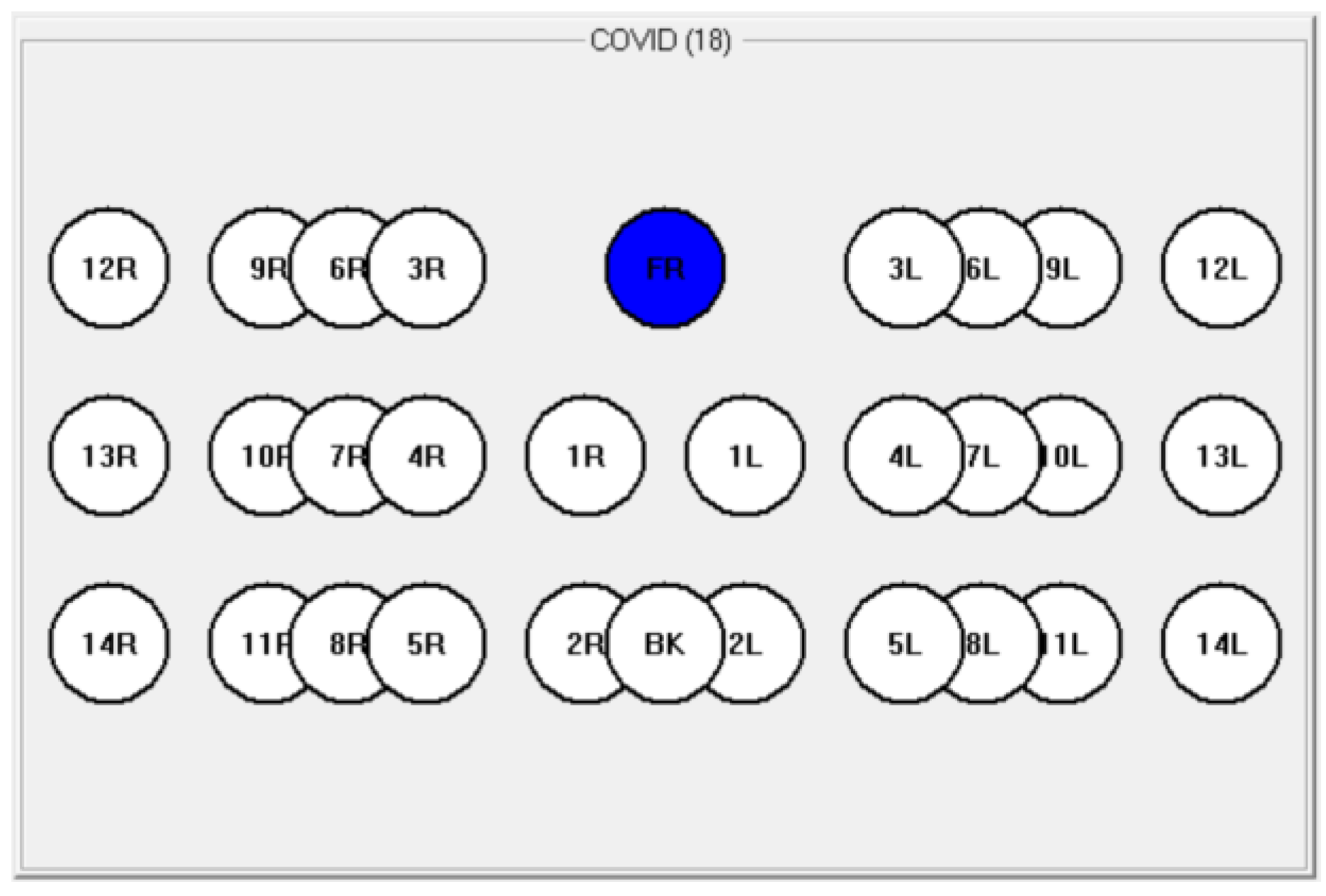
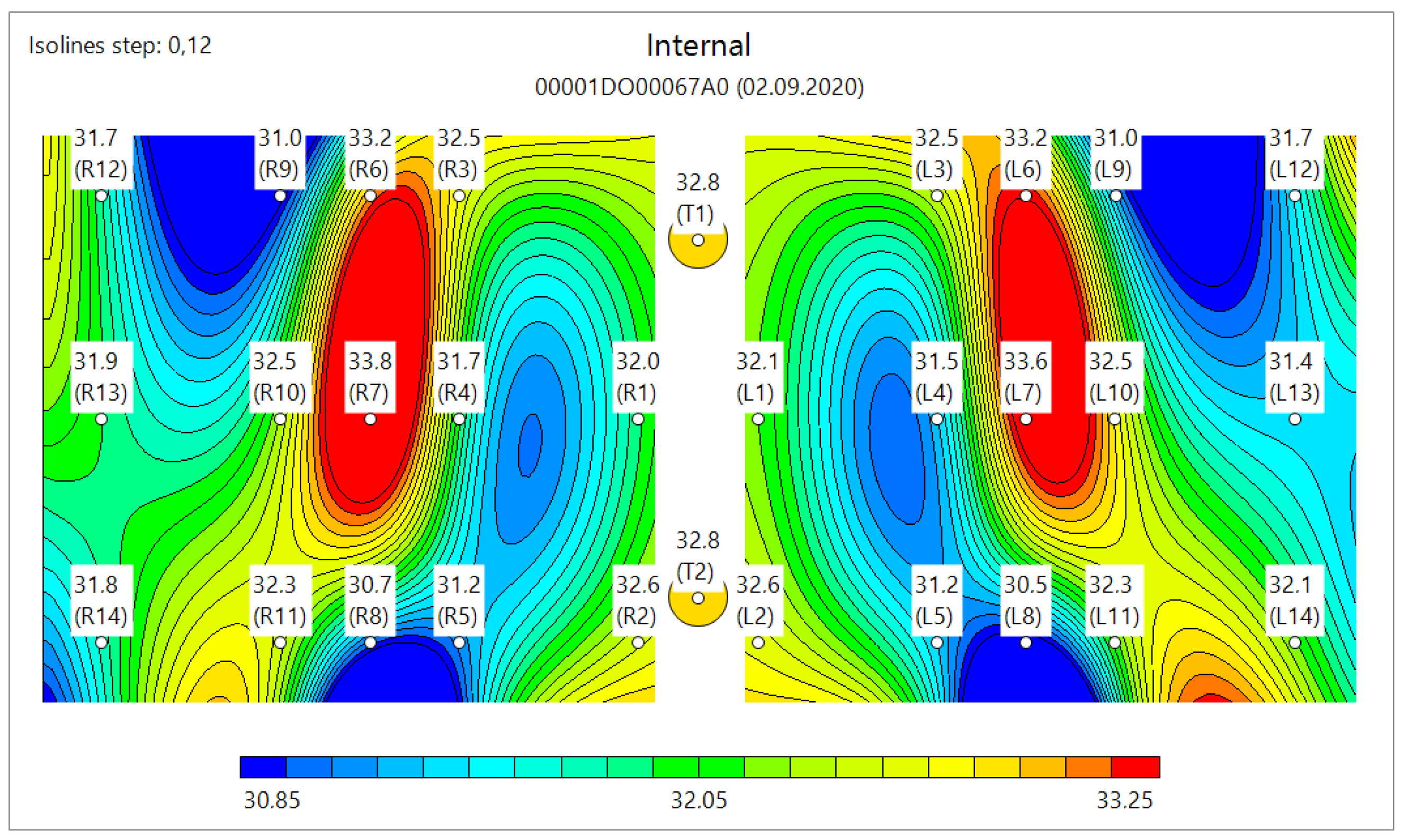
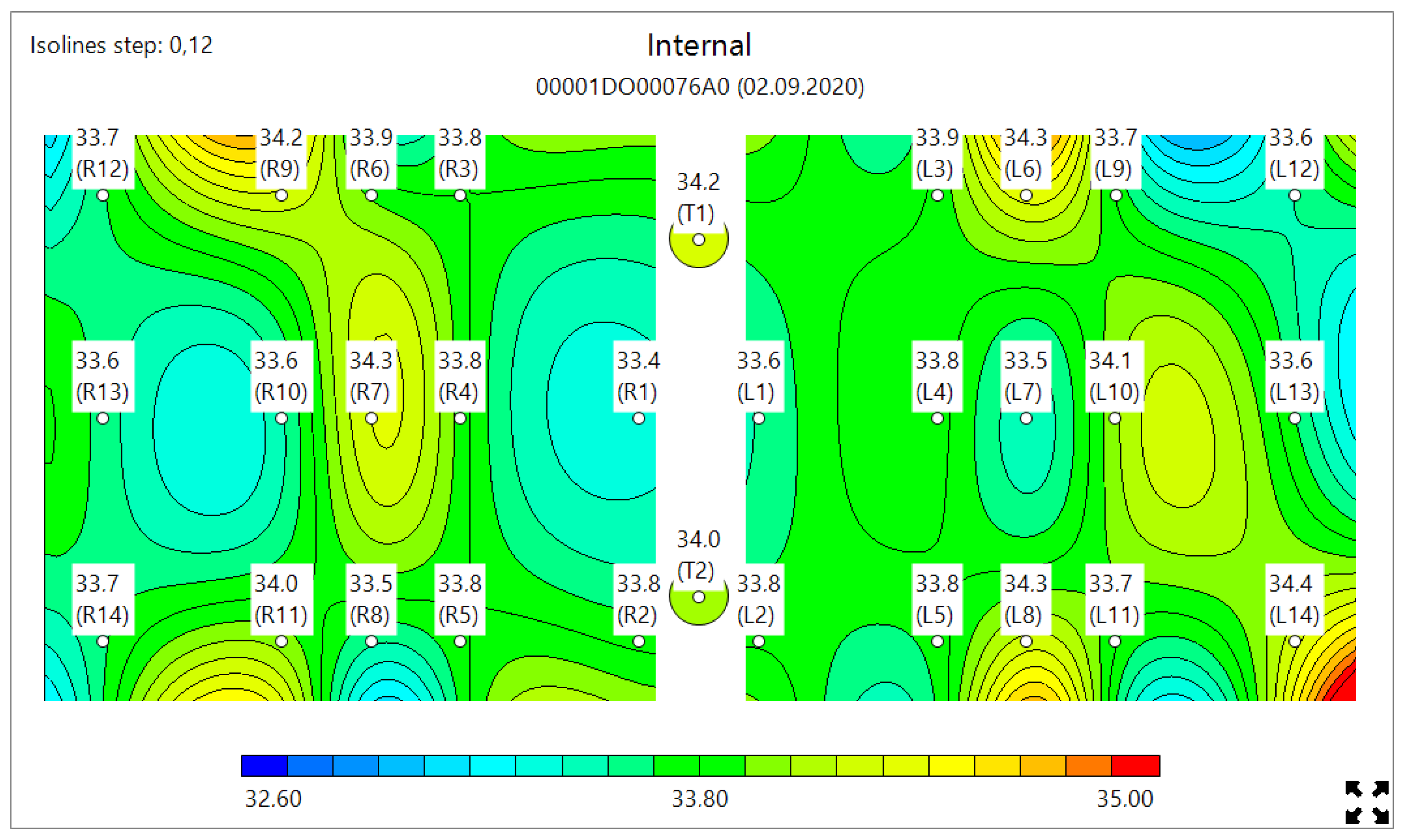
3.2. Clinical Images
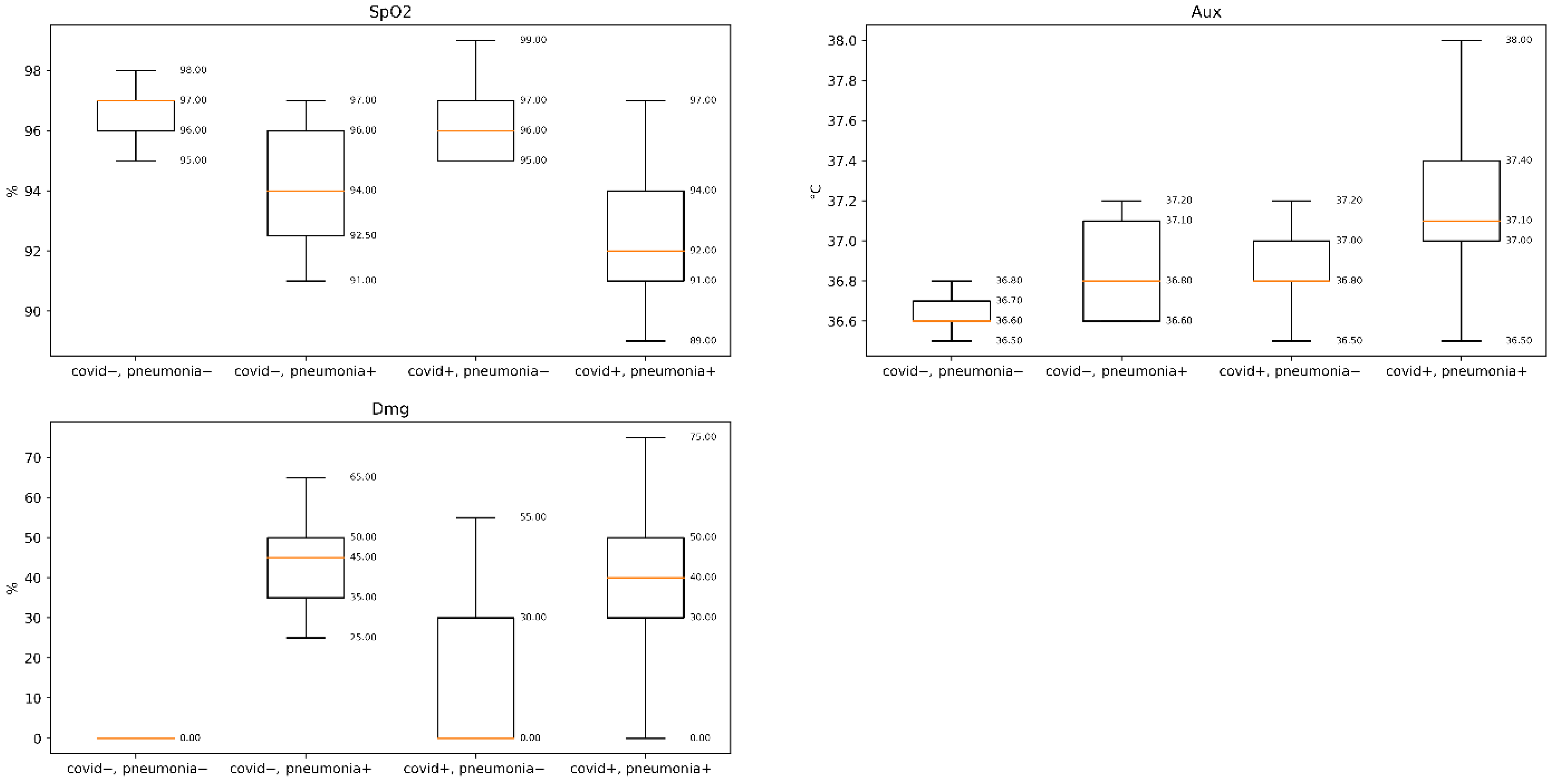
3.3. Statistical Results
- The difference in symmetrical points should be less than 0.5 °C in healthy patients and be of greater value in patients with pneumonia;
- The difference between internal and skin temperatures should be uniform in healthy patients but might show local abnormalities in patients with pneumonia;
- The difference between the hottest and coldest points should be less than 2 °C in healthy patients.
- Asymmetry (AS) expresses the difference between symmetrical positions, and was calculated as follows:
- Median asymmetry (#1) shows the overall disbalance between the left and right sides; the standard deviation of asymmetry (#5) aggregates the local irregularities, while the maximum asymmetry highlights the asymmetry at a single point (#9).
- Asymmetry inverse (ASIN) (#2) reveals the condition when the internal and skin temperatures show asymmetry with significantly different magnitudes or signs.
- Spread (SP) expresses the difference between the internal and skin temperatures.
- SP median (#3) shows the overall change, which is caused by the systematic changes in the metabolism or thermal properties; SP std (#7) shows the non-uniformity of the internal-skin difference; SP max (#11) highlights the anomaly at a single point.
- Relative increase (RI) aimed to reveal the irregular points of increase or decrease. It is calculated as the value of the difference between the internal value and the median of all internal values for that patient
- Internal median (#13) shows a shift in the baseline level of the internal temperature.
- Internal percentile interval (#14) (5–95%) is aimed at measuring the spread between the hottest and coldest measured points. The same applies for the skin (#15, #16).
3.4. Deep Neural Network Results
4. Discussion
- Individual data points are hardly informative, due to inaccuracies in measurements and noise-induced randomness.
- The mean value has limited informativeness due to individual variations in metabolism, conductivity of the tissues, and changes in the ambient temperature.
- An increase or a decrease in a point relative to its neighbors might be informative (so-called thermal heterogeneity).
- An increase or a decrease in a point relative to the symmetric point on the body might be informative (so-called thermal asymmetry).
- An increase or a decrease in the microwave temperature value relative to the infrared temperature value might be informative (so-called thermal convergence), especially compared to its neighbors.
- Covid+, pneumonia+, MWR+, accident and emergency hospitalization; usually, the patients are already hospitalized.
- Covid− or RT-PCR test not available, pneumonia+, MWR+ hospitalization; usually the patients are already hospitalized.
- Covid− or RT-PCR test not available, pneumonia− (or CT test not available,), MWR+; consultancy with a specialist, repeat or take PCR test; repeat, or take CT test and MWR examination.
- Covid+, pneumonia−, MWR−, repeat MWR test; most likely it is asymptomatic COVID-19, and no further action is required.
- Covid−, pneumonia+, MWR−, repeat PCR and MWR tests; usually, the patients are already hospitalized.
- Covid−, pneumonia−, MWR−; no further action is required.
- Nursery homes;
- Ships;
- Remote locations (highlands, islands, deserts);
- Boarder security;
- Detention centers.
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kant, S.K.; Kumar, M.A.; Amos, L. Comprehensive update on current outbreak of novel coronavirus infection (2019-nCoV). Ann. Transl. Med. 2020, 8, 393. [Google Scholar]
- Liu, K.; Chen, Y.; Lin, R.; Han, K. Clinical feature of COVID-19 in elderly patients: A comparison with young and middle-aged patients. J. Infect. 2020, 80, e14. [Google Scholar] [CrossRef] [PubMed]
- Lake Mary, A. What we know so far: COVID-19 current clinical knowledge and research. Clin. Med. 2020, 20, 124. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Wang, Y.; Zhu, T.; Xia, L. LCT features of coronavirus disease 2019 (COVID-19) pneumonia in 62 patients in Wuhan, China. AJR. Am. J. Roentgenol. 2020, 1–8, 1287. [Google Scholar] [CrossRef] [PubMed]
- Chung, M.; Bernheim, A.; Mei, X.; Zhang, N.; Cui, J.; Jacobi, A.; Li, K.; Li, S.; Shan, H.; Xu, W.; et al. CT imaging features of 2019 novel coronavirus (2019–, nCoV). Radiol. Febr. 2020, 295, 202–207. [Google Scholar] [CrossRef]
- ACR Recommendations for the Use of Chest Radiography and Computed Tomography (CT) for Suspected COVID-19 Infection|American College of Radiology. 2020. Available online: http://www.acr.org/Advocacy-and-Economics/ACR-Position-Statements/Recommendations-for-Chest-Radiography-and-CT-for-Suspected-COVID19-Infection (accessed on 12 December 2020).
- Buyun, X.; Yangbo, X.; Jiahao, P.; Zheng, Z.; Tang, W.; Sun, Y.; Xu, C.; Peng, F. Chest CT for detecting COVID-19: A systematic review and meta-analysis of diagnostic accuracy. Eur. Radiol. 2020, 30, 5720–5727. [Google Scholar] [CrossRef]
- Watson, J.; Whiting, P.F.; Brush, J.E. Interpreting a covid-19 test result. BMJ 2020, 369, m1808. [Google Scholar] [CrossRef]
- Chinese National Health Commission. Chinese Clinical Guidance for COVID-19 Pneumonia Diagnosis and Treatment, 7th ed.; American College of Cardiology: Washington, DC, USA, 2020. [Google Scholar]
- Hope, M.D.; Raptis, C.A.; Henry, T.S. Chest computed tomography for detection of coronavirus Disease 2019 (COVID-19): Don’t rush the science. Ann. Internal Med. 2020, 173, 147. [Google Scholar] [CrossRef] [PubMed]
- Raptis, C.A.; Hammer, M.M.; Short, R.G.; Hope, M.D.; Kligerman, S.J.; Shah, A.; Bhalla, S.; Henry, T.S.; Jeudy, J.; Filev, P.D.; et al. Chest CT and coronavirus disease (COVID-19): A critical review of the literature to date. Am. J. Roentgenol. 2020, 1–4, 8. [Google Scholar] [CrossRef]
- Hope, M.D.; Raptis, C.A.; Shah, A.; Hammer, M.M.; Henry, T.S. Six signatories. A role for CT in COVID-19? What data really tell us so far. Lancet 2020, 395, 1189. [Google Scholar] [CrossRef]
- Fang, Y.; Zhang, H.; Xie, J.; Lin, M.; Ying, L.; Pang, P.; Ji, W. Sensitivity of chest CT for COVID-19: Comparison to RT-PCR. Radiology 2020, 296, E115. [Google Scholar] [CrossRef] [PubMed]
- Youxin, W.; Haifeng, H.; Wenrui, W.; Wei, W. Combination of CT and RT-PCR in the screening or diagnosis of COVID-19. J. Glob. Health 2020, 10, 010347. [Google Scholar]
- Rubin, G.D.; Ryerson, C.J.; Haramati, L.B.; Goldin, J.; Humnert, M.; Mazzone, P.J.; Wells, A.U.; Richeldi, L.; Prokop, M.; Inoue, Y.; et al. The Role of Chest Imaging in Patient Management during the COVID-19 Pandemic: A Multinational Consensus Statement from the Fleischner Society. Radiology 2020, 296, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Simpson, S.; Kay, F.U.; Abbara, S.; Bhalla, S.; Chung, J.H.; Chung, M.; Henry, T.S.; Kanne, J.P.; Kligerman, S.; Ko, J.P.; et al. Radiological Society of North America expert consensus statement on reporting chest CT findings related to COVID-19. Endorsed by the Society of Thoracic Radiology, the American College of Radiology, and RSNA. J. Thorac. Imaging 2020, 35, 219. [Google Scholar] [CrossRef] [PubMed]
- Arevalo-Rodriguez, I.; Buitrago-Garcia, D.; Simancas-Racines, D.; Zambrano-Achig, P.; del Campo, R.; Ciapponi, A.; Sued, O.; Martinez-García, L.; Rutjes, A.W.; Low, N.; et al. False-negative results of initial RT-PCR assays for COVID-19: A systematic review. medRXIV 2020, 15, e0242958. [Google Scholar]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 2020, 323, 1843. [Google Scholar] [CrossRef] [PubMed]
- Goryanin, I.; Karbainov, S.; Shevelev, O.; Tarakanov, A.; Redpath, K.; Vesnin, S.; Ivanov, Y. Passive microwave radiometry in biomedical studies. Drug Discov. Today 2020, 25, 757. [Google Scholar] [CrossRef] [PubMed]
- Iskander, M.F.; Durney, C.H., Sr.; Grange, T.; Smith, C.S. Radiometric technique for measuring changes in lung water. IEEE Trans. Microw. Theory Tech. 1984, 32, 554–556. [Google Scholar] [CrossRef]
- Iskander, M.F.; Durney, C.H. Microwave Methods of measuring Changes in Lung water. J. Microw. Power Electromagn. Energy 1983, 18, 265. [Google Scholar] [CrossRef]
- Leroy, Y.; Bocquet, B.; Mamouni, A. Non-invasive microwave radiometry thermometry. Physiol. Meas. 1998, 19, 127. [Google Scholar] [CrossRef]
- Ginzburg, L.I.; Glagolev, N.A. Radiothermometry in the diagnosis of lung diseases. Electr. Industry USSR 1987, 1, 27–58. (In Russian) [Google Scholar]
- World Health Organization. Global Surveillance for COVID-19 Disease Caused by Human Infection with Novel Coronavirus (COVID-19): Interim Guidance; WHO: Geneva, Switzerland, 2020. [Google Scholar]
- Tarakanov, A.; Vesnin, S.; Efremov, V.; Roberts, N.S. An introduction to kernel and nearest-neighbor nonparametric regression. Am. Stat. 1992, 46, 175–185. [Google Scholar]
- Tarakanov, A.V.; Tarakanov, A.A.; Vesnin, S.; Efremov, V.V.; Roberts, N.; Goryanin, I. Influence of Ambient Temperature on Recording of Skin and Deep Tissue Temperature in Region of Lumbar Spine. Eur. J. Transl. Clin. Med. 2020, 7, 21–26. [Google Scholar] [CrossRef]
- Available online: https://en.wikipedia.org/wiki/Tukey’s_range_test (accessed on 5 October 2020).
- Available online: https://pythonhealthcare.org/2018/04/13/55-statistics-multi-comparison-with-tukeys-test-and-the-holm-bonferroni-method/ (accessed on 12 May 2019).
- Galazis, C.; Vesnin, S.; Goryanin, I. Application of artificial intelligence in microwave radiometry (MWR). Bioinformatics 2019, 3, 112–122. [Google Scholar]
- Levshinskii, V.; Galazis, C.; Ovchinnikov, L.; Vesnin, S.; Losev, A.; Goryanin, I. Application of Data Mining and Machine Learning in Microwave Radiometry (MWR). In International Joint Conference on Biomedical Engineering Systems and Technologies BIOSTEC 2019—Biomedical Engineering Systems and Technologies; Roque, A., Tomczyk, A., De Maria, E., Putze, F., Moucek, R., Fred, A., Gamboa, H., Eds.; Springer: Cham, Switzerland, 2020; Volume 1211. [Google Scholar]
- Ai, T.; Yang, Z.; Hou, H.; Zhan, C.; Chen, C.; Tao, Q.; Sun, Z.; Xia, L.; Lv, W. Correlation of chest CT and RT-PCR testing in coronavirus Disease 2019 (COVID-19) in China: A report of 1014 cases. Radiology 2020, 296, E32–E40. [Google Scholar] [CrossRef] [PubMed]
- Eng, J.; Bluemke, D.A. Imaging publications in the COVID-19 pandemic: Applying new research results to clinical practice. Radiology 2020, 297, E228. [Google Scholar] [CrossRef]
- De Smet, K.; De Smet, D.; Ryckaert, T.; Laridon, E.; Heremans, B.; Vandenbulcke, E.; Demedts, I.; Bouckaert, B.; Martens, G.A.; Gryspeerdt, S. Diagnostic Performance of Chest CT for SARS-CoV-2 Infection in Individuals with or without COVID-19 Symptoms. Radiology 2021, 298, E30–E37. [Google Scholar] [CrossRef]
- Zampeli, E.; Raftakis, I.; Michelongona, A.; Nikolaou, C.; Elezoglou, A.; Antoniadis, C.; Toutouzas, K.; Sfikakis, P.P. AB1250 Measurement of joint temperature by microwave radiometry as a novel and simple method for the detection of synovial inflammation: A pilot study. Ann. Rheum. Dis. 2013, 71 (Suppl. 3), 709.5. [Google Scholar] [CrossRef]
- Toutouzas, K.; Synetos, A.; Drakopoulou, M.; Tsiamis, E.; Karanasos, A.; Michelongona, A.; Grassos, H.; Stefanadi, E.; Siores, E.; Stefanadis, C. An experimental study with microwave thermography for the assessment of atherosclerotic vascular disease. Am. Heart Assoc. 2009, 120, S1113. [Google Scholar]
- Zamechnik, T.; Losev, A.; Petrenko, A. Guided Classifier in the Diagnosis of Breast Cancer According to Microwave Radiothermometry. Math. Phys. Comput. Simul. 2019, 22, 52–66. [Google Scholar] [CrossRef]
- Cheboksarov, D.V.; Butrov, A.V.; Shevelev, O.A.; Amcheslavsky, V.G.; Pulina, N.N.; Buntina, M.A.; Sokolov, I.M. Diagnostic opportunities of noninvasive brain thermomonitoring. Anesteziol. Reanimatol 2015, 60, 66. [Google Scholar] [PubMed]
- Vesnin, S.G.; Sedankin, M.K.; Ovchinnikov, L.M.; Gudkov, A.G.; Leushin, V.Y.; Sidorov, I.A.; Goryanin, I.I. Portable microwave radiometer for wearable devices. Sens. Actuat. A Phys. 2021, 318, 112506. [Google Scholar]
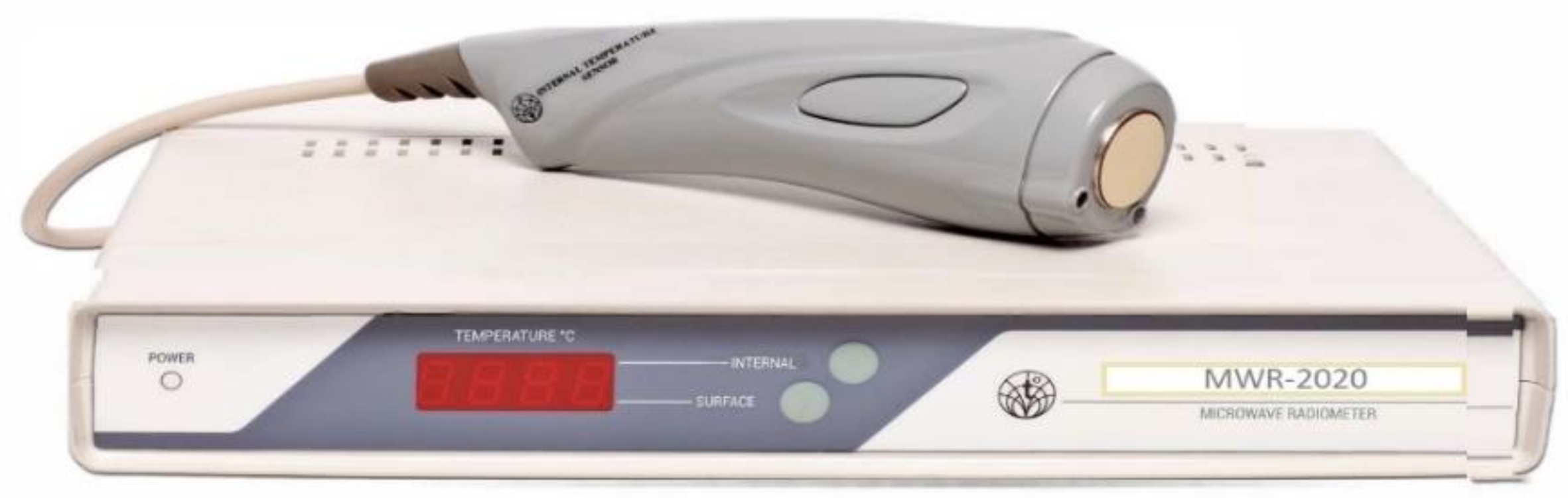

| Key Features (MWR2020) | |
|---|---|
| Temperature detection depth, cm | 3–7 |
| Accuracy of internal temperature measurements (microwave range), °C | 0.2 |
| Measurement time, s | 8 |
| Antenna diameter, mm | 39 |
| Accuracy of skin temperature measurements (infrared range), °C | 0.2 |
| Weight, kg | 2.5 |
| Power, W | 20 |
| Group | Covid− | Covid+ |
|---|---|---|
| Pneumonia− | 77 | 9 |
| Pneumonia+ | 19 | 103 |
| Clothes on | Clothes off | |||||
|---|---|---|---|---|---|---|
| Aggregated Metrics | f-Value | p-Value | Reject Null-Hyp (p < 0.05) | f-Value | p-Value | Reject Null-Hyp (p < 0.05) |
| 1. AS median | 0.34 | 0.79 | No | 3.21 | 0.03 | Yes |
| 2. ASIN median | 0.05 | 0.98 | No | 3.08 | 0.03 | Yes |
| 3. SP median | 3.65 | 0.02 | Yes | 12.35 | 4.18 × 107 | Yes |
| 4. RI median | 0.81 | 0.49 | No | 2.90 | 0.04 | Yes |
| 5. AS std | 0.30 | 0.82 | No | 0.05 | 0.98 | No |
| 6. ASIN std | 0.58 | 0.62 | No | 1.32 | 0.27 | No |
| 7. SP std | 0.30 | 0.83 | No | 2.73 | 0.05 | Yes |
| 8. RI std | 0.32 | 0.81 | No | 0.11 | 0.96 | No |
| 9. AS max | 0.24 | 0.86 | No | 0.05 | 0.99 | No |
| 10. ASIN max | 0.67 | 0.57 | No | 1.05 | 0.37 | No |
| 11. SP max | 0.08 | 0.97 | No | 5.67 | 1.1 × 103 | Yes |
| 12. RI max | 0.71 | 0.55 | No | 1.12 | 0.34 | No |
| 13. Int median | 2.54 | 0.06 | No | 2.88 | 0.039 | Yes |
| 14. Sk median | 7.26 | 2.2 × 104 | Yes | 6.09 | 6.7 × 104 | Yes |
| 15. Int interval | 3.16 | 0.03 | Yes | 0.22 | 0.87 | No |
| 16. Sk interval | 0.93 | 0.42 | No | 0.36 | 0.77 | No |
| Criteria | Significant Pairs (Clothes on) | Significant Pairs (Clothes off) |
|---|---|---|
| SP median | Covid− pneumonia−/covid+ pneumonia+ (delta = +1.1, p = 0.0075) | Covid− pneumonia−/covid+ pneumonia+ (delta = +1.48, p < 0.001) |
| SP std | Covid− pneumonia−/covid+ pneumonia+ (delta = +0.40, p = 0.025) | |
| SP max | Covid− pneumonia−/covid+ pneumonia+ (delta = +2.72, p < 0.001) | |
| Int median | Covid− pneumonia−/covid+ pneumonia− (delta = +1.31, p = 0.043) | |
| Sk median | Covid− pneumonia−/covid+ pneumonia+ (delta = −1.00, p = 0.001) Covid− pneumonia+/covid+ pneumonia+ (delta = −0.85, p = 0.008) | Covid− pneumonia−/covid+ pneumonia+ (delta = −0.64, p = 0.008) Covid− pneumonia+/covid+ pneumonia− (delta = +3.12, p = 0.006) |
| Experiment | Sensitivity | Specificity |
|---|---|---|
| Raw temperatures | 71.05% | 57.52% |
| Raw temperatures and metadata | 79.85% | 48.37% |
| Metrics | 50.99% | 77.29% |
| Raw temperatures and metrics | 71.05% | 74.35% |
| Raw temperatures, metadata, and metrics | 70.95% | 48.18% |
| Experiment | Sensitivity | Specificity |
|---|---|---|
| Raw temperatures and metrics | 71.05% | 74.35% |
| Raw temperatures, metrics, and clothes flag | 76.19% | 76.47% |
| Raw temperatures and metrics (ensemble clothes on/off) | 76.19% | 47.06% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Osmonov, B.; Ovchinnikov, L.; Galazis, C.; Emilov, B.; Karaibragimov, M.; Seitov, M.; Vesnin, S.; Losev, A.; Levshinskii, V.; Popov, I.; et al. Passive Microwave Radiometry for the Diagnosis of Coronavirus Disease 2019 Lung Complications in Kyrgyzstan. Diagnostics 2021, 11, 259. https://doi.org/10.3390/diagnostics11020259
Osmonov B, Ovchinnikov L, Galazis C, Emilov B, Karaibragimov M, Seitov M, Vesnin S, Losev A, Levshinskii V, Popov I, et al. Passive Microwave Radiometry for the Diagnosis of Coronavirus Disease 2019 Lung Complications in Kyrgyzstan. Diagnostics. 2021; 11(2):259. https://doi.org/10.3390/diagnostics11020259
Chicago/Turabian StyleOsmonov, Batyr, Lev Ovchinnikov, Christopher Galazis, Berik Emilov, Mustafa Karaibragimov, Meder Seitov, Sergey Vesnin, Alexander Losev, Vladislav Levshinskii, Illarion Popov, and et al. 2021. "Passive Microwave Radiometry for the Diagnosis of Coronavirus Disease 2019 Lung Complications in Kyrgyzstan" Diagnostics 11, no. 2: 259. https://doi.org/10.3390/diagnostics11020259
APA StyleOsmonov, B., Ovchinnikov, L., Galazis, C., Emilov, B., Karaibragimov, M., Seitov, M., Vesnin, S., Losev, A., Levshinskii, V., Popov, I., Mustafin, C., Kasymbekov, T., & Goryanin, I. (2021). Passive Microwave Radiometry for the Diagnosis of Coronavirus Disease 2019 Lung Complications in Kyrgyzstan. Diagnostics, 11(2), 259. https://doi.org/10.3390/diagnostics11020259






