Abstract
Despite the recent implementation of immunotherapy as a single treatment or in combination with chemotherapy for first-line treatment of advanced non-small cell lung cancer (NSCLC), many patients do not benefit from this regimen due to primary treatment resistance or toxicity. Consequently, there is an urgent need to develop efficient biomarkers that can select patients who will benefit from immunotherapy thereby providing the appropriate treatment and avoiding toxicity. One of the biomarkers recently described for the stratification of NSCLC patients undergoing immunotherapy are mutations in STK11/LKB1, which are often associated with a lack of response to immunotherapy in some patients. Therefore, the purpose of this review is to describe the different cellular mechanisms associated with STK11/LKB1 mutations, which may explain the lack of response to immunotherapy. Moreover the review addresses the co-occurrence of additional mutations that may influence the response to immunotherapy and the current clinical studies that have further explored STK11/LKB1 as a predictive biomarker. Additionally this work includes the opportunities and limitations to look for the STK11/LKB1 status in the therapeutic strategy for NSCLC patients.
1. Introduction
Over the past few years, the mortality due to non-small cell lung cancer (NSCLC) decreased significantly, mainly due to early diagnosis, but also to the development of new therapeutic strategies, including targeted therapies [1]. Additionally, the introduction of immunotherapy alone or in combination with chemotherapy has had a dramatic impact on the prognosis of patients by providing a substantial improvement in overall survival [2]. In this context, many clinical trials have evaluated immunotherapy in the first- and second-line settings with the aim to develop a more efficient treatment strategy with less toxicity [3,4,5,6,7,8,9,10,11].
In line with the tremendous clinical progress, the understanding of the efficacy of immunotherapy in NSCLC and its pathophysiology has unraveled new cellular mechanisms associated with its response to treatment and to intrinsic resistance [12,13,14,15]. Moreover, bioinformatic analyses are becoming increasingly sophisticated allowing the analysis and integration of complex clinical and biological data to further understand the biology of cancer, notably of lung carcinoma [16,17,18,19]. Unfortunately, and despite recent progress, the development and clinical validation of novel robust biomarkers that predict response, resistance and/or toxicity to the treatment in routine clinical care remain major challenges in thoracic oncology.
Many research studies and clinical trials have evaluated the potential of different prognostic and predictive biomarkers in thoracic oncology [20,21,22,23,24,25,26,27,28,29,30,31]. Importantly, an increasing number of biomarkers have been developed and evaluated specifically to determine the response to immunotherapy in NSCLC, including a few developed specifically to assess toxicity [27,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49]. These biomarkers demonstrated caveats that limit their implementation [27,34,36,38,40].
Recently, different studies have associated the presence of STK11 mutations with a lack of response to immunotherapy in NSCLC [50,51,52,53,54]. Additionally, some preclinical and translational studies have shed further light on the biological role of STK11 mutations leading to primary resistance to immunotherapy [55,56,57]. Nevertheless, the implementation of STK11 mutations as a routine biomarker in NSCLC, in addition to the current mandatory or recommended therapeutic targets (EGFR, ALK, ROS1, BRAF, NTRK, PDL-1, RET, MET and HER2), remains controversial and is not performed in daily practice [58].
Therefore, this review aims to highlight the current research of STK11 mutations in late stage NSCLC, the considerations for its potential implementation in routine clinical care, and finally the current limitations of using the STK11 mutational status in decision making of the global therapeutic strategy in thoracic oncology.
2. The Double-Edged Sword of STK11 in Cancer Cell Metabolism
The STK11 gene is located on the short arm of chromosome 19 (19p13.3) and its nine exons codes for liver kinase B1 (LKB1), a protein kinase and master metabolic sensor that acts as an energy gauge to sustain cancer cell survival [59,60,61]. Upon nutrient deprivation, as occurs often in the center of large tumors, LKB1 orchestrates cell metabolism by reducing ATP-consuming processes while simultaneously stimulating ATP-generation. Mechanistically, LKB1 is the upstream serine/threonine kinase for the AMP-activated protein kinase (AMPK). Activation of AMPK redirects metabolism towards decreased fatty acid (FA) synthesis, increased glycolysis, and FA oxidation to replenish ATP stores [60,62,63,64,65,66]. Consequently, LKB1 activates several downstream kinases of the AMPK family by direct phosphorylation in the T-loop domain [60]. In particular, the activation of AMPK upon energetic stress has been intensively analyzed in various diseases, including cancer to induce a metabolic switch from anabolism towards catabolism to regulate energy homeostasis and cell survival (Figure 1).
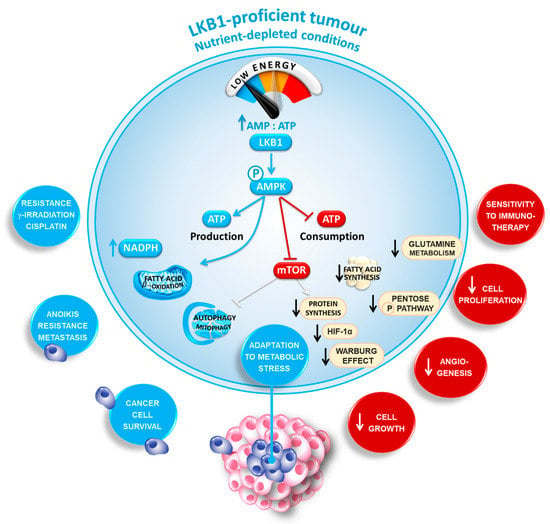
Figure 1.
The double-edged sword of liver kinase B1 (LKB1) in cancer cell metabolism. LKB1 is a master metabolic sensor that acts as an energy gauge to sustain cancer cell survival. Upon nutrient deprivation within the center of large tumors, LKB1 reprograms cell metabolism by slowing down all ATP-consuming processes while simultaneously stimulating ATP-generating processes. By organizing this overall stress response, LKB1 may adapt cancer cell growth under conditions of energy shortage.
Besides its “classical” role as a metabolic checkpoint inhibitor, AMPK directly phosphorylates and inhibits the mammalian target of rapamycin (mTOR), shutting down protein synthesis, the cellular process that consumes the most ATP [67]. Inhibition of mTOR downregulates hypoxia-inducible factor 1α (HIF-1α), thereby negatively affecting angiogenesis and anaerobic glycolysis (the “Warburg effect”) [68]. Advances in molecular profiling have identified mutations or amplifications of specific genes involved in the mTOR pathway (e.g., PIK3CA, PTEN, STK11 and RICTOR) as the most common mechanism leading to mTOR hyperactivation [69]. Noteworthy, LKB1, AMPK and mTORC1 are recruited onto the lysosomal surface upon glucose deprivation where they inhibit mTOR [70]. This activation of the LKB1/AMPK/mTOR signaling pathway on lysosomes commits the cells to basically cannibalize themselves via the autophagy-lysosomal pathway [71,72]. Indeed, suppression of the activity of mTORC1 releases the inhibitory phosphorylation on Unc-51-Like Autophagy Activating Kinase 1 (ULK1), a kinase essential for initiation of autophagy [73]. As a safeguard, AMPK also directly phosphorylates and activates ULK1 and the proautophagy lipid kinase VPS34 [74,75]. Finally, inactivation of mTORC1 releases the inhibitory phosphorylation of TFEB and TFE3 [76]. Simultaneously AMPK directly phosphorylates and activates the transcription factors FOXO and PGC-1α (peroxisome proliferator-activated receptor-gamma coactivator 1α) [77,78]. TFEB, TFE3, PGC-1α and FOXO translocate to the nucleus, where they drive the expression of genes involved in autophagy, mitochondrial processes and lysosomes. Autophagy may then help the cell degrade dysfunctional mitochondria (mitophagy) to fuel the nutrients required for oxidative metabolism [79].
It is also critical to highlight that LKB1/AMPK may safeguard against oxidative stress by inhibiting NADPH-consuming FA synthesis and increasing NADPH-producing FA oxidation [80]. Meanwhile, activated AMPK also phosphorylates and activates the transcription factor NRF2 [81]. NRF2 then activates the transcription of antioxidant genes involved in the production of NADPH. The high NADPH levels, together with autophagy, protect the LKB1-proficient cancers from oxidative stress and ROS-inducing chemotherapies (cisplatin, paclitaxel and doxorubicin) [59]. Consequently, NRF2 activation has been associated with more aggressive lung cancer and decreased survival in patients [82]. Interestingly, the activation of the LKB1/autophagy pathway enables circulating tumor cells to resist anoikis [83]. Accordingly, cells lacking LKB1 undergo apoptosis under metabolic stress as they are unable to respond to a deficiency in energy and restore homeostasis [64].
By orchestrating this overall stress response, LKB1 may help cancer cells to continuously “fine-tune” their growth rate in response to fluctuations in energy in their environment. As such, these dual pro- and antitumoral roles of LKB1 implicitly suggest that LKB1 is not always functioning as a tumor suppressor, as was initially thought and may link LKB1 signaling to tumor progression. In addition to AMPK two other AMPK-related kinases, the salt-inducible kinases SIK1 and SIK3, emerge as predominant downstream targets of LKB1 in NSCLC [84,85]. It seems of great interest to look for the impact of these different pathways on the potential efficiency of the immune checkpoint inhibitors (ICIs), notably those targeting PD-L1/PD1 [2,12,13,14]. Hence, many developments have granted a clearer perceptive on the different factors that reduce an antitumor immune response, leading to the discovery of several agents that work on immune costimulatory and inhibitory checkpoint pathways. So, the best examples of the advanced checkpoint molecules that mediate tumor-induced immune suppression are PD-1 and PD-L1 [12,13,14]. Whether the SIK-dependent pathway plays a role in the resistance to the ICIs however, remains to be determined.
More recently, a connection between STK11 expression and the stimulator of interferon genes (STING) pathway was also highlighted [55,86]. STING is a cytosolic protein activated by the presence of free double-stranded DNA (dsDNA) in the cytoplasm, due to pathogen infection or neoplastic transformation. Aberrant cytoplasmic dsDNA activates STING oligomerization on the endoplasmic reticulum–Golgi membrane. STING then recruits and activates TBK1 (tank-binding kinase 1), which phosphorylates the transcription factor IRF3 to induce the production of type-I interferons and other chemokines, and finally the T cell recruitment. In cancer, STING may play a crucial role as one of the initial steps needed for immune evasion. Interestingly, Kitajima et al. described the downregulation of STING in KRAS/STK11 comutated cancer cells [87]. STK11 inactivation dysregulates serine metabolism, leading to increased levels of S-adenosyl methionine (SAM). SAM is a substrate for multiple epigenetic silencing enzymes, such as DNMT1 and EZH2, that are both directly involved in the methylation of the STING promoter, causing its down-modulation and repression [87] (Figure 2).
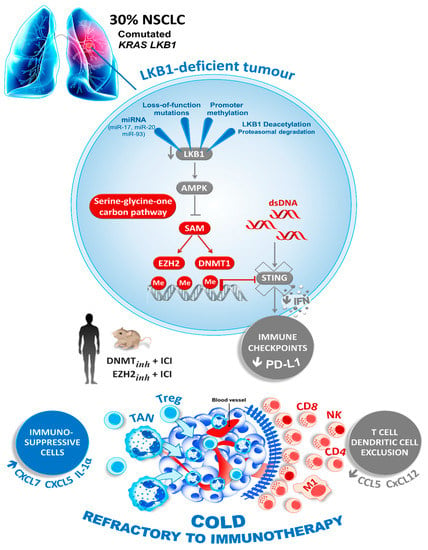
Figure 2.
Loss of LKB1 drives the tumor immune escape. The loss of LKB1, the second most commonly altered tumor suppressor in NSCLC, promotes the production of SAM, a substrate for the DNA and histone methyltransferases DNMT1, EZH2 and other epigenetic silencing enzymes. This downregulates the expression of STING, impairing dsDNA sensing, and thereby the expression of immune checkpoint regulating proteins like PD-L1 and T cell chemokines. Therefore, the LKB1-deficient tumors are characterized by a so-called “cold” immunosuppressive tumor microenvironment (blue), which shows the striking infiltration of immunosuppressive cells (TAN; tumor-associated neutrophil; Treg, T lymphocyte regulator, blue) and the exclusion of inflamed immune cells (CD8 T cells, NK, CD4 T cells and M1; Macrophage type 1, red). Epigenetic therapies that reactivate LKB1 or the STING pathway may boost an anticancer immune response in LKB1-deficient cancers with the resistance to immune-checkpoint blockade (ICI, immune checkpoint inhibitor).
Furthermore, the regulation of the expression of STK11 and its role in cancer cell proliferation remain very complex [88]. Recent studies for example showed that asparagine and aspartate could regulate AMPK-mediated p53 activation by physically binding to LKB1 and modulating LKB1 activity. It seems that p53 can regulate asparagine metabolism to control cell survival by generating an auto-amplification loop via asparagine-aspartate-mediated LKB1-AMPK signaling [88].
3. STK11 and Associated Genomic Alterations in Lung Cancer
STK11 mutations are present in many different tumor types but with varying frequencies [89]. They are more frequently observed in NSCLC, notably in lung adenocarcinoma [89,90,91]. However, reports of the frequency of mutations in lung cancer vary in the literature, and it is also different depending on the patient ethnicity [91,92,93,94,95]. STK11 mutations are less frequent in Asian (4–7%) than in Caucasian patients (16%) in NSCLC analyzed by the TCGA project [96]. They were also significantly higher among Afro-American patients [97]. Interestingly, in one series only 77/1385 (6%) NSCLC patients harbored an STK11 mutation according to next generation sequencing (NGS) analysis while the AACR genie database (v8.1 public) reported STK11 mutations in 1495/14,303 (10.5%) of samples [92]. In addition to lung adenocarcinoma, some pulmonary large-cell neuroendocrine carcinomas can also harbor STK11 mutations [98,99,100]. The most frequently comutated genes are highlighted in Table 1.

Table 1.
Most frequently comutated genes with STK11 in non-small cell lung cancer according to AACR Genie Database (v8.1-public).
Importantly, these STK11 mutations have to be analyzed in association with other comutations of interest, notably in KRAS, KEAP1, TP53 and SMARCA4 [92,101,102]. Arbour et al. showed that among 330 patients with late stage KRAS-mutant NSCLC, the most frequent comutations were found in TP53 (42%), STK11 (29%) and KEAP1/NFE2L2 (27%) [101]. Furthermore, in 62 patients with STK11 mutated tumors analyzed by Bange et al., 18 had an STK11 mutation alone, while 19, 18 and 7 patients had STK11/KRAS, STK11/TP53 and STK11/KRAS/TP53 comutations, respectively [92]. In contrast, a recent study analyzing 69 patients showed that the mutations in STK11 were more frequently observed in the KRAS wild type population than in the KRAS mutated tumors [103]. Additionally, SMARCA4 mutations were associated with STK11 mutations in 39% of all cases [104]. SMARCA4 mutations can be classified into class-I with SMARCA4 truncating mutations, some fusions and homozygous deletions, and into class 2 with SMARCA4 missense mutations or variants of unknown significance [104]. Importantly, STK11 mutations are mainly associated with SMARCA4 class-I mutations [104]. Moreover, the loss of BRG1 expression, which can be detected by immunohistochemistry with an anti-BRG1 antibody, is associated with SMARCA4 class I mutations, and was found to be predominantly detected in adenocarcinomas with co-occurring mutations in KRAS, STK11, TP53 and KEAP1 [105]. Other mutations or gene activations, such as NFE2L2 mutations are also often associated with STK11 mutations [82,106]. It was recently demonstrated that NRF2 activation acts as a critical oncogenic driver, and can promote aggressive lung adenocarcinoma by cooperating with STK11 loss and KRAS activation [82]. Additionally, patients with NRF2-activated non-squamous or squamous tumors have a poor prognosis and limited response to anti-PD-L1 treatment [82,107].
4. STK11 as a Prognostic Biomarker in Lung Cancer
The presence of STK11 mutations in association with additional mutations in KRAS, KEAP1 and SMARCA4 has been reported as an independent negative prognostic factor for overall survival in NSCLC patients [102]. Different studies reported clusters of KRAS-mutant lung adenocarcinomas bearing STK11 mutations, TP53 mutations or CDKN2A/B inactivation [108,109]. In additional studies, patients with a KRAS/STK11 comutation had a poorer overall survival than KRAS mutated patients without a STK11 mutation or with KRAS/TP53 comutations [50,92,110]. La Fleur et al. showed that patients with an isolated KRAS mutation had an overall survival similar to those of the wild-type group, whereas patients with co-occurring mutations in either TP53 or STK11 had a worse overall survival in comparison to the wild-type group and the KRAS mutated only group [110]. Conversely other studies showed that the tumors with comutated KRAS and STK11 did not have a worse prognosis than KRAS mutated/STK11 wild-type tumors [101]. Moreover, the analysis of patients’ survival with early stage lung adenocarcinoma in the TCGA PanCancer data set also highlighted poorer overall survival in patients with STK11 mutations. Additionally, a recent study showed that patients with STK11 mut/KRAS wt tumors had a worse prognosis compared to patients with STK11 and KRAS double mutated tumors [103].
5. STK11 as a Predictive Biomarker for the Therapy Response in Lung Cancer
While many studies have highlighted the prognostic value of STK11 mutations, it remains unclear if STK11 mutations are also predictive of response to immunotherapy. In some series of NSCLC with STK11 mutations treated with first-line systemic therapy, a comutation with KRAS was associated with significantly worse progression-free survival and overall survival [92]. In contrast, comutation of STK11 with TP53 conferred a better prognosis in these patients [92].
Additionally, another study showed a higher prolonged progression-free survival in anti-PD-1-treated patients harboring TP53-mut/STK11-EGFR-wt tumors than in patients with TP53-wt/STK11-mut [111]. A few initial studies demonstrated that mutated STK11 tumors, notably with TP53, KRAS and KEAP1 mutations, showed primary resistance to ICIs [15,53,57]. Using whole-exome sequencing to examine NSCLC patients treated with PD-1 plus CTLA-4 blockade, a study demonstrated that a couple of patients with a STK11 mutation had primary resistance to this therapeutic combination [6]. Most of the studies demonstrated a low level of expression of PD-L1 in patients with a STK11 mutation, which could explain the resistance to immunotherapy [112,113]. In the study of Lamberti et al., the PD-L1-negative group, compared with the PD-L1-high group, had a higher number of mutations in STK11 (19% versus 6%) [113]. In contrast, other studies reported that lung cancer patients with STK11 mutated tumors could respond to ICIs [114,115,116]. However, the response could sometimes be associated with co-occurring TP53 mutations [86,116]. Additionally, a retrospective analysis of the Keynote-042 trial, which evaluated pembrolizumab vs. platinum-based chemotherapy in NSCLC in the first-line setting, revealed comparable response rates between STK11 mutated and STK11 wild-type tumors [5]. In line, an analysis of 2276 NSCLC patients treated in first-line with ICIs demonstrated that the presence of comutations in STK11 and KEAP1 were prognostic rather than predictive [117]. Additionally, a recent study showed that despite the presence of a STK11 mutation, the presence of SMARCA4 mutations could have a positive predictive value for ICI responsiveness, highlighting the controversial role of STK11 as a predictive biomarker [102].
Interestingly, a genomic mutation signature (GMS) obtained with eight selected genes (TP53, KRAS, STK11, EGFR, PTPRD, KMT2C, SMAD4 and HGF) was able to predict response of NSCLC patients to immunotherapy, most notably to anti-PD-1 therapy [118]. The GMS was independent of TMB and PD-L1 expression and predicted response across different clinico-pathological features and combining PD-L1 expression with the GMS improved prediction of response [118].
In addition to the evaluation of STK11 mutations, another study examined the expression of LKB1 using immunohistochemistry. LKB1 expression in more than 50% of tumor cells was defined as LKB1 high and results correlated to the efficacy of pembrolizumab monotherapy in untreated patients with advanced NSCLC [119]. In this study, the median progression free survival and overall survival of patients was numerically shorter in the cohort with low LKB1 expression, although the results were not statistically significant [119]. However, only a few patients were included in this study. Most importantly, this study did not assess STK11 mutations and thus the relationship between expression and mutational status of STK11 remains unknown [119]. It would have been of interest to see how the expression of STK11 differed in the context of the presence of other comutations like KEAP1 or KRAS and to analyze the tumor microenvironment, as the composition of different immune cells might be different among different KRAS mutations [120].
Despite the association of STK11 mutations with response to immunotherapy, some studies have also evaluated STK11 mutations in populations that were treated with chemotherapy. In a French cohort of patients receiving chemotherapy in first-line, 25/302 (8%) NSCLC had a STK11 mutation [93]. No statistical difference was observed between the STK11 status and clinico-pathological variables. Overall survival was shorter for STK11 mutated patients in a univariate analysis. However, the STK11 status did not have an impact on overall survival in a multivariate analysis and progression free survival was not significantly different between the populations [93]. Interestingly one patient with both STK11 and NFE2L2 mutations had a good response to platinum-based chemotherapy [106]. In contrast, a recent study from Jeong et al. demonstrated that deletion of KEAP1 conferred chemoresistance in preclinical models of lung adenocarcinoma and that patients with late stage NSCLC with KEAP1/NFE2L2/CUL3 mutations had a shorter time to treatment failure and overall survival when treated with front-line platinum doublet chemotherapy [121]. However, the impact of KRAS and STK11 mutations on these results were not estimated [121].
6. Potential Treatments Targeting STK11 Mutations in Lung Cancer
To overcome STK11 mediated resistance to immunotherapy, novel compounds are being developed to target this distinct group of lung cancers [2]. Among them, the recently emerging KRAS inhibitors that currently target preferentially KRAS G12C are actively being investigated [122,123,124]. Importantly, considering the impact of different comutations together with KRAS, an important source of heterogeneity in KRAS mutated lung cancer, current drug development programs need to take this complexity into account and design studies that also consider the comutations [125]. However, the impact of a KRAS inhibition on a STK11 comutated tumor remains unknown and different alternative approaches might be necessary to better target these tumors [89,126]. For example, metformin, phenformin with or without sapanisertib, everolimus, tunicamycin, brefeldin A or 2-deoxyglucose, which target different metabolic pathways that can be altered by STK11 mutations have been actively studied [89]. Additionally, a recent study demonstrated that ERK inhibitors can be effective in STK11 mutated tumors in vitro, and which was proposed as a novel therapeutic strategy [127]. Interestingly, STK11 loss induced an increase in energetic/redox stress, which is tolerated, in part, through co-occurring KEAP1/NRF2-dependent metabolic adaptations, thus enhancing glutamine dependence, which rendered those tumors sensitive to glutaminase inhibitors [128]. This was further supported by new research using a combination of CRISPR-Cas9-based genetic screening and metabolomic analyses, which further confirmed that KEAP1 or NRF2-mutant cancers are dependent on increased glutaminolysis, which can be therapeutically exploited through the pharmacological inhibition of glutaminase [129].
7. Assessment of the STK11 Status in Lung Cancer
Currently the detection of STK11 mutations is mainly performed using NGS with gene panels of different sizes (from a few genes to large panels of 500 genes) and both tissue and liquid biopsies have been used as testing material [116,130,131]. In the context of STK11 mutations, it is particularly important to carefully consider the most effective panel size for an NGS panel due to the particular importance of comutations in this context [105]. Some genes of interest, such as KEAP1 and/or SMARCA4 but also other genes that are often comutated with STK11 are absent in some small panels [132,133,134,135,136]. Additionally, STK11 itself may be absent with most of the rapid sequencing approaches, highlighting the importance of the careful selection of the appropriate gene panels [135].
Therefore, it might be more suitable to integrate larger gene panels for NGS testing when the STK11 status is of interest to carefully assess all important comutations and not just the currently mandatory genes for baseline assessment in NSCLC.
8. Integrating the STK11 Mutation Status in the Treatment of Lung Cancer
Identifying clinical or molecular factors that predict the benefit of checkpoint inhibitors in advanced NSCLC remains crucial for the selection of appropriate therapies for each patient. Currently, the expression of PD-L1 on tumor cells remains the principal biomarker to predict the efficacy of checkpoint inhibitors on NSCLC and other tumors. Although there is a linear relationship between the level of the benefit of checkpoint inhibitors and the level of tumor PD-L1 expression in NSCLC, a response of tumors to immunotherapy has still been observed in those with low or undetectable PD-L1 expression [36,116,137]. In contrast, tumors may be unresponsive to immunotherapy even when expressing high levels of PD-L1, highlighting the limitations of PD-L1 for therapy selection [36,138]. While the value of the tumor mutational burden (TMB) has been extensively studied for the prediction of response to immunotherapy, which led recently to its highly debated approval by the FDA, TMB is still a controversial biomarker [21,34].
So far, the assessment of KRAS mutations has been considered to be of interest before chemotherapy and/or ICI treatment, but recent studies demonstrated that the assessment of KRAS alone might be of limited value [139]. Moreover, many studies have found a strong association between mutations in KRAS and STK11 [140].
Considering the challenges of using biomarkers for the selection of immunotherapy for patients with NSCLC, it may be of interest to integrate the STK11 status for late stage NSCLC in patients eligible for treatment with first-line ICIs alone or in association with chemotherapy. In this regard, it was proposed that other genes, such as KEAP1 should now be evaluated and also associated [141]. Consequently, one could theoretically argue that patients with tumors showing an expression of PD-L1 in more than 50% of tumor cells, without other druggable mutations in EGFR, ALK, ROS1, BRAF, NTRK, RET, HER2 and MET, but having an STK11 mutation, should not be treated with ICIs, and should potentially only receive chemotherapy [57]. However, it is very important in this context to highlight the limitations of using STK11 mutations for patient stratification as discussed above. Most importantly, it remains unclear if STK11 can serve as a predictive biomarker that can guide treatment selection and prospective evaluation is still missing. Consequently, immunotherapy should not currently be withheld from patients with STK11 mutated tumors. However, in the case of rapid tumor progression under treatment or in the case of a non-tumoral response, consideration of a STK11 mutation may allow rapid adaptation of therapy for these patients [105]. Additionally, STK11 expression could also be considered [142,143]. However, multiplexing different antibodies of interest for diagnosis and/or assessment of predictive markers may be needed to reserve material for further genomic studies [144,145]. For first-line immunotherapy in NSCLC, future developments should at the same time integrate not only several genomic alterations of interest, including STK11 mutation, but also the assessment of the expression of different proteins and cytokines in the same sample, knowing the impact of different associated mutations in the tumor environment [56,116,120].
9. Conclusions
The in vitro research and clinical trials that integrate evaluation of genomic alterations for treatment responsiveness and comparison of efficacy of different molecular strategies have opened up new approaches to lung cancer precision medicine. However, there is an urgent need to integrate simultaneously many biomarkers to propose the best therapeutic strategy for lung cancer patients [58]. These different biomarkers constitute an increasing number of genomic alterations, identified not only in tissue and cytological samples, but also in liquid biopsies [116,146,147,148]. Recent research has expanded dramatically our understanding of the role of STK11 mutations in mediating resistance to anticancer immunotherapy in NSCLC and have revealed novel therapeutic approaches both in vitro and in vivo. However, these recent studies also highlighted the complexity of the biology of NSCLC and the importance of also assessing co-occurring mutations, which influence the response to therapy. Therefore, it seems essential to evaluate the status of combinations of different genes such as STK11, TP53, KRAS, KEAP1 and SMARCA4, to cite a few, and the different classes of genomic alterations present on these different genes. These analyses will certainly extend our understanding of resistance to immunotherapy and will improve the selection of appropriate therapies for personalized medicine of patients with NSCLC.
Author Contributions
Conceptualization, P.H.; writing—original draft preparation, B.M., S.H., P.H.; visualization, B.M.; All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors thank Brahimi-Horn for editing this manuscript, the Conseil Départemental des Alpes Maritimes, the Ligue Départementale de Lutte contre le Cancer des Alpes Maritimes, the French Government (Agence Nationale de Recherche, ANR) through the ‘Investments for the future’ LABEX SIGNALIFE [ANR-11-LABX-0028-01]); “Association pour la Recherche contre le Cancer” (ARC CANC’AIR GENExposomics project), Cancerôpole PACA; Région Sud, Institut National de la Santé et de la Recherche Médicale (INSERM); and Institut National du Cancer Plan Cancer (PHRC; STALKLUNG01 13-APN-01).
Conflicts of Interest
The authors have no conflict of interest to report.
References
- Howlader, N.; Forjaz, G.; Mooradian, M.J.; Meza, R.; Kong, C.Y.; Cronin, K.A.; Mariotto, A.B.; Lowy, D.R.; Feuer, E.J. The Effect of Advances in Lung-Cancer Treatment on Population Mortality. New Engl. J. Med. 2020, 383, 640–649. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.-Y.; Yang, J.C.-H.; Yang, P.-C. Precision Management of Advanced Non-Small Cell Lung Cancer. Annu. Rev. Med. 2020, 71, 117–136. [Google Scholar] [CrossRef] [PubMed]
- Borghaei, H.; Paz-Ares, L.; Horn, L.; Spigel, D.R.; Steins, M.; Ready, N.E.; Chow, L.Q.; Vokes, E.E.; Felip, E.; Holgado, E.; et al. Nivolumab versus Docetaxel in Advanced Nonsquamous Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2015, 373, 1627–1639. [Google Scholar] [CrossRef] [PubMed]
- Brahmer, J.; Reckamp, K.L.; Baas, P.; Crinò, L.; Eberhardt, W.E.; Poddubskaya, E.; Antonia, S.; Pluzanski, A.; Vokes, E.E.; Holgado, E.; et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2015, 373, 123–135. [Google Scholar] [CrossRef]
- Cho, C.B.; Gilberto, L.; Kowalski, D.M.; Kasahara, K.; Wu, Y.-L.; Castro, G., Jr.; Turna, H.Z.; Cristescu, R.; Aurora-Garg, D.; Loboda, A.; et al. CT084—Relationship between STK11 and KEAP1 Mutational Status and Efficacy in KEYNOTE-042: Pembrolizumab Monotherapy Versus Platinum-Based Chemotherapy as Fist-Line Therapy for PD-L1-Positive Advanced NSCLC. AACR 2020. Available online: https://www.abstractsonline.com/pp8/#!/9045/presentation/10785 (accessed on 3 January 2021).
- Hellmann, M.D.; Nathanson, T.; Liu, C.; Sauter, J.L.; Rekhtman, N.; Chang, E.; Callahan, M.K.; Chaft, J.E.; Voss, M.H.; Tenet, M.; et al. Genomic Features of Response to Combination Immunotherapy in Patients with Advanced Non-Small-Cell Lung Cancer. Cancer Cell 2018, 33, 843–852. [Google Scholar] [CrossRef]
- Herbst, R.S.; Baas, P.; Kim, D.-W.; Felip, E.; Pérez-Gracia, J.L.; Han, J.-Y.; Molina, J.; Kim, J.-H.; Arvis, C.D.; Ahn, M.-J.; et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): A randomised controlled trial. Lancet 2016, 387, 1540–1550. [Google Scholar] [CrossRef]
- Passiglia, F.; Cappuzzo, F.; Alabiso, O.; Bettini, A.C.; Bidoli, P.; Chiari, R.; Defferrari, C.; Delmonte, A.; Finocchiaro, G.; Francini, G.; et al. Efficacy of nivolumab in pre-treated non-small-cell lung cancer patients harbouring KRAS mutations. Br. J. Cancer 2019, 120, 57–62. [Google Scholar] [CrossRef]
- Reck, M.; Rodríguez-Abreu, D.; O’Brien, M.; Rao, S.; Hotta, K.; Vandormael, K.; Riccio, A.; Yang, J.; Pietanza, M.C.; Brahmer, J.R.; et al. Updated Analysis of KEYNOTE-024: Pembrolizumab Versus Platinum-Based Chemotherapy for Advanced Non-Small-Cell Lung Cancer With PD-L1 Tumor Proportion Score of 50% or Greater. J. Clin. Oncol. 2019, 37, 537–546. [Google Scholar] [CrossRef]
- Reuss, J.E.; Anagnostou, V.; Cottrell, T.R.; Smith, K.N.; Verde, F.; Zahurak, M.; Lanis, M.; Murray, J.C.; Chan, H.Y.; McCarthy, C.; et al. Neoadjuvant nivolumab plus ipilimumab in resectable non-small cell lung cancer. J. Immunother. Cancer 2020, 8, e001282. [Google Scholar] [CrossRef]
- Rittmeyer, A.; Barlesi, F.; Waterkamp, D.; Park, K.; Ciardiello, F.; Von Pawel, J.; Gadgeel, S.M.; Hida, T.; Kowalski, D.M.; Dols, M.C.; et al. Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): A phase 3, open-label, multicentre randomised controlled trial. Lancet 2017, 389, 255–265. [Google Scholar] [CrossRef]
- Bodor, J.N.; Boumber, Y.; Borghaei, H. Biomarkers for immune checkpoint inhibition in non-small cell lung cancer (NSCLC). Cancer 2020, 126, 260–270. [Google Scholar] [CrossRef] [PubMed]
- Russo, A.; McCusker, M.G.; Scilla, K.A.; Arensmeyer, K.E.; Mehra, R.; Adamo, V.; Rolfo, C. Immunotherapy in Lung Cancer: From a Minor God to the Olympus. Adv. Exp. Med. Biol. 2020, 1244, 69–92. [Google Scholar] [CrossRef] [PubMed]
- Sanmamed, M.F.; Eguren-Santamaria, I.; Schalper, K.A. Overview of Lung Cancer Immunotherapy. Cancer J. 2020, 26, 473–484. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, S.; Zhou, Q. The Resistance Mechanisms of Lung Cancer Immunotherapy. Front. Oncol. 2020, 10, 568059. [Google Scholar] [CrossRef]
- Palmisano, A.; Krushkal, J.; Li, M.-C.; Fang, J.; Sonkin, D.; Wright, G.; Yee, L.; Zhao, Y.; McShane, L.M. Bioinformatics Tools and Resources for Cancer Immunotherapy Study. Methods Mol. Biol. 2020, 2055, 649–678. [Google Scholar] [CrossRef]
- Prokop, J.W.; May, T.; Strong, K.; Bilinovich, S.M.; Bupp, C.; Rajasekaran, S.; Worthey, E.A.; Lazar, J. Genome sequencing in the clinic: The past, present, and future of genomic medicine. Physiol. Genom. 2018, 50, 563–579. [Google Scholar] [CrossRef]
- Rappoport, N.; Shamir, R. Multi-omic and multi-view clustering algorithms: Review and cancer benchmark. Nucleic Acids Res. 2018, 46, 10546–10562. [Google Scholar] [CrossRef]
- Yohe, S.; Thyagarajan, B. Review of Clinical Next-Generation Sequencing. Arch. Pathol. Lab. Med. 2017, 141, 1544–1557. [Google Scholar] [CrossRef]
- Barnes, T.A.; Amir, E. HYPE or HOPE: The prognostic value of infiltrating immune cells in cancer. Br. J. Cancer 2018, 118, e5. [Google Scholar] [CrossRef]
- Berland, L.; Heeke, S.; Humbert, O.; Macocco, A.; Long-Mira, E.; Lassalle, S.; Lespinet-Fabre, V.; Lalvée, S.; Bordone, O.; Cohen, C.; et al. Current views on tumor mutational burden in patients with non-small cell lung cancer treated by immune checkpoint inhibitors. J. Thorac. Dis. 2019, 11, S71–S80. [Google Scholar] [CrossRef]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of Circulating Tumor DNA in Early- and Late-Stage Human Malignancies. Sci. Transl. Med. 2014, 6, 224ra24. [Google Scholar] [CrossRef] [PubMed]
- Camidge, D.R.; Doebele, R.C.; Kerr, K.M. Comparing and contrasting predictive biomarkers for immunotherapy and targeted therapy of NSCLC. Nat. Rev. Clin. Oncol. 2019, 16, 341–355. [Google Scholar] [CrossRef] [PubMed]
- Danaher, P.; Warren, S.; Lu, R.; Samayoa, J.; Sullivan, A.; Pekker, I.; Wallden, B.; Marincola, F.M.; Cesano, A. Pan-cancer adaptive immune resistance as defined by the Tumor Inflammation Signature (TIS): Results from The Cancer Genome Atlas (TCGA). J. Immunother. Cancer 2018, 6, 63. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.A.; Patel, V. The role of PD-L1 expression as a predictive biomarker: An analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J. Immunother. Cancer 2019, 7, 278. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, S.B.; Narayan, A.; Kole, A.J.; Decker, R.H.; Teysir, J.; Carriero, N.J.; Lee, A.; Nemati, R.; Nath, S.K.; Mane, S.M.; et al. Early Assessment of Lung Cancer Immunotherapy Response via Circulating Tumor DNA. Clin. Cancer Res. 2018, 24, 1872–1880. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P.; Heeke, S.; Alix-Panabières, C.; Pantel, K. Liquid biopsy in the era of immuno-oncology: Is it ready for prime-time use for cancer patients? Ann. Oncol. 2019, 30, 1448–1459. [Google Scholar] [CrossRef]
- Ni, L.; Lu, J. Interferon gamma in cancer immunotherapy. Cancer Med. 2018, 7, 4509–4516. [Google Scholar] [CrossRef]
- Schalper, K.A.; Brown, J.; Carvajal-Hausdorf, D.; McLaughlin, J.; Velcheti, V.; Syrigos, K.N.; Herbst, R.S.; Rimm, D.L. Objective Measurement and Clinical Significance of TILs in Non-Small Cell Lung Cancer. J. Natl. Cancer Inst. 2015, 107, 435. [Google Scholar] [CrossRef]
- Silva, E.M.; Mariano, V.S.; Pastrez, P.R.A.; Pinto, M.C.; Castro, A.G.; Syrjanen, K.J.; Longatto-Filho, A. High systemic IL-6 is associated with worse prognosis in patients with non-small cell lung cancer. PLoS ONE 2017, 12, e0181125. [Google Scholar] [CrossRef]
- Sun, R.; Limkin, E.J.; Vakalopoulou, M.; Dercle, L.; Champiat, S.; Han, S.R.; Verlingue, L.; Brandao, D.; Lancia, A.; Ammari, S.; et al. A radiomics approach to assess tumour-infiltrating CD8 cells and response to anti-PD-1 or anti-PD-L1 immunotherapy: An imaging biomarker, retrospective multicohort study. Lancet Oncol. 2018, 19, 1180–1191. [Google Scholar] [CrossRef]
- Chen, G.; Huang, A.C.; Zhang, W.; Zhang, G.; Wu, M.; Xu, W.; Yu, Z.; Yang, J.; Wang, B.; Sun, H.; et al. Exosomal PD-L1 contributes to immunosuppression and is associated with anti-PD-1 response. Nature 2018, 560, 382–386. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Jiang, C.C.; Jin, L.; Zhang, X.D. Regulation of PD-L1: A novel role of pro-survival signalling in cancer. Ann. Oncol. 2016, 27, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Heeke, S.; Hofman, P. Tumor mutational burden assessment as a predictive biomarker for immunotherapy in lung cancer patients: Getting ready for prime-time or not? Transl. Lung Cancer Res. 2018, 7, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Higgs, B.W.; Morehouse, C.A.; Streicher, K.; Brohawn, P.; Pilataxi, F.; Gupta, A.; Ranade, K. Interferon Gamma Messenger RNA Signature in Tumor Biopsies Predicts Outcomes in Patients with Non-Small Cell Lung Carcinoma or Urothelial Cancer Treated with Durvalumab. Clin. Cancer Res. 2018, 24, 3857–3866. [Google Scholar] [CrossRef] [PubMed]
- Ilie, M.; Hofman, V.; Dietel, M.; Soria, J.-C.; Hofman, P. Assessment of the PD-L1 status by immunohistochemistry: Challenges and perspectives for therapeutic strategies in lung cancer patients. Virchows Arch. 2016, 468, 511–525. [Google Scholar] [CrossRef]
- Liu, L.; Ruiz, J.; O’Neill, S.S.; Grant, S.C.; Petty, W.J.; Yang, M.; Chen, K.; Topaloglu, U.; Pasche, B.; Zhang, W. Favorable outcome of patients with lung adenocarcinoma harboring POLE mutations and expressing high PD-L1. Mol. Cancer 2018, 17, 81. [Google Scholar] [CrossRef]
- Möller, M.; Turzer, S.; Schütte, W.; Seliger, B.; Riemann, D. Blood Immune Cell Biomarkers in Patient with Lung Cancer Undergoing Treatment with Checkpoint Blockade. J. Immunother. 2020, 43, 57–66. [Google Scholar] [CrossRef]
- Peng, W.; Chen, J.Q.; Liu, C.; Malu, S.; Creasy, C.; Tetzlaff, M.T.; Xu, C.; McKenzie, J.A.; Zhang, C.; Liang, X.; et al. Loss of PTEN Promotes Resistance to T Cell-Mediated Immunotherapy. Cancer Discov. 2016, 6, 202–216. [Google Scholar] [CrossRef]
- Prelaj, A.; Tay, R.; Ferrara, R.; Chaput, N.; Besse, B.; Califano, R. Predictive biomarkers of response for immune checkpoint inhibitors in non-small-cell lung cancer. Eur. J. Cancer 2019, 106, 144–159. [Google Scholar] [CrossRef]
- Reuben, A.; Zhang, J.; Chiou, S.-H.; Gittelman, R.M.; Li, J.; Lee, W.-C.; Fujimoto, J.; Behrens, C.; Liu, X.; Wang, F.; et al. Comprehensive T cell repertoire characterization of non-small cell lung cancer. Nat. Commun. 2020, 11, 603. [Google Scholar] [CrossRef]
- Rizvi, H.; Sanchez-Vega, F.; La, K.; Chatila, W.; Jonsson, P.; Halpenny, D.; Plodkowski, A.; Long, N.; Sauter, J.L.; Rekhtman, N.; et al. Molecular Determinants of Response to Anti-Programmed Cell Death (PD)-1 and An-ti-Programmed Death-Ligand 1 (PD-L1) Blockade in Patients with Non-Small-Cell Lung Cancer Profiled with Targeted Next-Generation Sequencing. J. Clin. Oncol. 2018, 36, 633–641. [Google Scholar] [CrossRef] [PubMed]
- Samstein, R.M.; Lee, C.-H.; Shoushtari, A.N.; Hellmann, M.D.; Shen, R.; Janjigian, Y.Y.; Barron, D.A.; Zehir, A.; Jordan, E.J.; Omuro, A.; et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 2019, 51, 202–206. [Google Scholar] [CrossRef] [PubMed]
- Singal, G.; Miller, P.G.; Agarwala, V.; Li, G.; Kaushik, G.; Backenroth, D.; Gossai, A.; Frampton, G.M.; Torres, A.Z.; Lehnert, E.M.; et al. Association of Patient Characteristics and Tumor Genomics with Clinical Outcomes Among Patients with Non-Small Cell Lung Cancer Using a Clinicogenomic Database. JAMA 2019, 321, 1391–1399. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Cheng, G.; Xu, C.; Wang, W.; Shao, Y.; Zhang, Y. Clinicopathological characteristics of POLE mutation in patients with non-small-cell lung cancer. Lung Cancer 2018, 118, 57–61. [Google Scholar] [CrossRef]
- Spranger, S.; Gajewski, T.F. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat. Rev. Cancer 2018, 18, 139–147. [Google Scholar] [CrossRef]
- Subudhi, S.K.; Aparicio, A.; Gao, J.; Zurita, A.J.; Araujo, J.C.; Logothetis, C.J.; Tahir, S.A.; Korivi, B.R.; Slack, R.S.; Vence, L.; et al. Clonal expansion of CD8 T cells in the systemic circulation precedes development of ipilimumab-induced toxicities. Proc. Natl. Acad. Sci. USA 2016, 113, 11919–11924. [Google Scholar] [CrossRef]
- Wang, F.; Zhao, Q.; Wang, Y.-N.; Jin, Y.; He, M.-M.; Liu, Z.-X.; Xu, R.-H. Evaluation of POLE and POLD1 Mutations as Biomarkers for Immunotherapy Outcomes Across Multiple Cancer Types. JAMA Oncol. 2019, 5, 1504–1506. [Google Scholar] [CrossRef]
- Zer, A.; Sung, M.R.; Walia, P.; Khoja, L.; Maganti, M.; Labbe, C.; Shepherd, F.A.; Bradbury, P.A.; Feld, R.; Liu, G.; et al. Correlation of Neutrophil to Lymphocyte Ratio and Absolute Neutrophil Count with Outcomes with PD-1 Axis Inhibitors in Patients with Advanced Non-Small-Cell Lung Cancer. Clin. Lung Cancer 2018, 19, 426–434. [Google Scholar] [CrossRef]
- Aredo, J.V.; Padda, S.K.; Kunder, C.A.; Han, S.S.; Neal, J.W.; Shrager, J.B.; Wakelee, H.A. Impact of KRAS mutation subtype and concurrent pathogenic mutations on non-small cell lung cancer outcomes. Lung Cancer 2019, 133, 144–150. [Google Scholar] [CrossRef]
- Kwack, W.G.; Shin, S.Y.; Lee, S.H. Primary Resistance to Immune Checkpoint Blockade in an STK11/TP53/KRAS-Mutant Lung Adenocarcinoma with High PD-L1 Expression. Oncol. Targets Ther. 2020, 13, 8901–8905. [Google Scholar] [CrossRef]
- Shire, N.J.; Klein, A.B.; Golozar, A.; Collins, J.M.; Fraeman, K.H.; Nordstrom, B.L.; McEwen, R.; Hembrough, T.; Rizvi, N.A. STK11 (LKB1) mutations in metastatic NSCLC: Prognostic value in the real world. PLoS ONE 2020, 15, e0238358. [Google Scholar] [CrossRef] [PubMed]
- Skoulidis, F.; Goldberg, M.E.; Greenawalt, D.M.; Hellmann, M.D.; Awad, M.M.; Gainor, J.F.; Schrock, A.B.; Hartmaier, R.J.; Trabucco, S.E.; Gay, L.; et al. STK11/LKB1 Mutations and PD-1 Inhibitor Resistance in KRAS-Mutant Lung Adenocarcinoma. Cancer Discov. 2018, 8, 822–835. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Guo, J.; Shang, X.; Wang, Z. Less immune cell infiltration and worse prognosis after immunotherapy for patients with lung adenocarcinoma who harbored STK11 mutation. Int. Immunopharmacol. 2020, 84, 106574. [Google Scholar] [CrossRef] [PubMed]
- Della Corte, C.M.; Byers, L.A. Evading the STING: LKB1 Loss Leads to STING Silencing and Immune Escape in KRAS-Mutant Lung Cancers. Cancer Discov. 2019, 9, 16–18. [Google Scholar] [CrossRef]
- Koyama, S.; Akbay, E.A.; Li, Y.Y.; Aref, A.R.; Skoulidis, F.; Herter-Sprie, G.S.; Buczkowski, K.A.; Liu, Y.; Awad, M.M.; Denning, W.L.; et al. STK11/LKB1 Deficiency Promotes Neutrophil Recruitment and Proinflammatory Cytokine Production to Suppress T-cell Activity in the Lung Tumor Microenvironment. Cancer Res. 2016, 76, 999–1008. [Google Scholar] [CrossRef]
- Skoulidis, F.; Heymach, J.V. Co-occurring genomic alterations in non-small-cell lung cancer biology and therapy. Nat. Rev. Cancer 2019, 19, 495–509. [Google Scholar] [CrossRef]
- Mosele, F.; Remon, J.; Mateo, J.; Westphalen, C.; Barlesi, F.; Lolkema, M.; Normanno, N.; Scarpa, A.; Robson, M.; Meric-Bernstam, F.; et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: A report from the ESMO Precision Medicine Working Group. Ann. Oncol. 2020, 31, 1491–1505. [Google Scholar] [CrossRef]
- Ciccarese, F.; Zulato, E.; Indraccolo, S. LKB1/AMPK Pathway and Drug Response in Cancer: A Therapeutic Perspective. Oxidative Med. Cell. Longev. 2019, 2019, 1–16. [Google Scholar] [CrossRef]
- Kullmann, L.; Krahn, M.P. Controlling the master—upstream regulation of the tumor suppressor LKB1. Oncogene 2018, 37, 3045–3057. [Google Scholar] [CrossRef]
- Li, T.-T.; Zhu, H.-B. LKB1 and cancer: The dual role of metabolic regulation. Biomed. Pharmacother. 2020, 132, 110872. [Google Scholar] [CrossRef]
- Bonanno, S.; Zulato, E.; Pavan, A.; Attili, I.; Pasello, G.; Conte, P.; Indraccolo, S. LKB1 and Tumor Metabolism: The Interplay of Immune and Angiogenic Microenvironment in Lung Cancer. Int. J. Mol. Sci. 2019, 20, 1874. [Google Scholar] [CrossRef] [PubMed]
- Hardie, D.G. AMP-activated protein kinase: An energy sensor that regulates all aspects of cell function. Genes Dev. 2011, 25, 1895–1908. [Google Scholar] [CrossRef] [PubMed]
- Shaw, R.J.; Kosmatka, M.; Bardeesy, N.; Hurley, R.L.; Witters, L.A.; Depinho, R.A.; Cantley, L.C. The tumor suppressor LKB1 kinase directly activates AMP-activated kinase and regulates apoptosis in response to energy stress. Proc. Natl. Acad. Sci. USA 2004, 101, 3329–3335. [Google Scholar] [CrossRef] [PubMed]
- Shackelford, D.B.; Shaw, R.J. The LKB1–AMPK pathway: Metabolism and growth control in tumour suppression. Nat. Rev. Cancer 2009, 9, 563–575. [Google Scholar] [CrossRef]
- Woods, A.; Johnstone, S.R.; Dickerson, K.; Leiper, F.C.; Fryer, L.G.; Neumann, D.; Schlattner, U.; Wallimann, T.; Carlson, M.; Carling, D. LKB1 Is the Upstream Kinase in the AMP-Activated Protein Kinase Cascade. Curr. Biol. 2003, 13, 2004–2008. [Google Scholar] [CrossRef]
- Xiang, H.; Zhang, J.; Lin, C.; Zhang, L.; Liu, B.; Ouyang, L. Targeting autophagy-related protein kinases for potential therapeutic purpose. Acta Pharm. Sin. B 2020, 10, 569–581. [Google Scholar] [CrossRef]
- Faubert, B.; Boily, G.; Izreig, S.; Griss, T.; Samborska, B.; Dong, Z.; Dupuy, F.; Chambers, C.; Fuerth, B.J.; Viollet, B.; et al. AMPK is a negative regulator of the Warburg effect and suppresses tumor growth in vivo. Cell Metab. 2013, 17, 113–124. [Google Scholar] [CrossRef]
- Krencz, I.; Sebestyen, A.; Khoor, A. mTOR in Lung Neoplasms. Pathol. Oncol. Res. 2020, 26, 35–48. [Google Scholar] [CrossRef]
- Zhang, C.-S.; Hawley, S.A.; Zong, Y.; Li, M.; Wang, Z.; Gray, A.; Ma, T.; Cui, J.; Feng, J.-W.; Zhu, M.; et al. Fructose-1,6-bisphosphate and aldolase mediate glucose sensing by AMPK. Nature 2017, 548, 112–116. [Google Scholar] [CrossRef]
- Chen, X.; Mao, R.; Su, W.; Yang, X.; Geng, Q.; Guo, C.; Wang, Z.; Wang, J.; Kresty, L.A.; Beer, D.G.; et al. Circular RNA circHIPK3 modulates autophagy via MIR124-3p-STAT3-PRKAA/AMPKα signaling in STK11 mutant lung cancer. Autophagy 2020, 16, 659–671. [Google Scholar] [CrossRef]
- Saftig, P.; Puertollano, R. How Lysosomes Sense, Integrate, and Cope with Stress. Trends Biochem. Sci. 2020, 46, 97–112. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kundu, M.; Viollet, B.; Guan, K.-L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 2011, 13, 132–141. [Google Scholar] [CrossRef] [PubMed]
- Herzig, S.; Shaw, R.J. AMPK: Guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell Biol. 2017, 19, 121–135. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Kim, Y.C.; Fang, C.; Russell, R.C.; Kim, J.H.; Fan, W.; Liu, R.; Zhong, Q.; Guan, K.-L. Differential Regulation of Distinct Vps34 Complexes by AMPK in Nutrient Stress and Autophagy. Cell 2013, 152, 290–303. [Google Scholar] [CrossRef]
- Martina, J.A.; Diab, H.I.; Lishu, L.; Jeong-A, L.; Patange, S.; Raben, N.; Puertollano, R. The Nutrient-Responsive Transcription Factor TFE3 Promotes Autophagy, Lysosomal Biogenesis, and Clearance of Cellular Debris. Sci. Signal. 2014, 7, ra9. [Google Scholar] [CrossRef]
- Eichner, L.J.; Brun, S.N.; Herzig, S.; Young, N.P.; Curtis, S.D.; Shackelford, D.B.; Shokhirev, M.N.; Leblanc, M.; Vera, L.I.; Hutchins, A.; et al. Genetic Analysis Reveals AMPK Is Required to Support Tumor Growth in Murine Kras-Dependent Lung Cancer Models. Cell Metab. 2019, 29, 285–302.e7. [Google Scholar] [CrossRef]
- Mihaylova, M.M.; Shaw, R.J. The AMPK signalling pathway coordinates cell growth, autophagy and metabolism. Nat. Cell Biol. 2011, 13, 1016–1023. [Google Scholar] [CrossRef]
- Belaid, A.; Ndiaye, P.D.; Filippakis, H.; Roux, J.; Röttinger, É.; Graba, Y.; Brest, P.; Hofman, P.; Mograbi, B. Autophagy: Moving Benchside Promises to Patient Bedsides. Curr. Cancer Drug Targets 2015, 15, 684–702. [Google Scholar] [CrossRef]
- Jeon, S.-M.; Chandel, N.S.; Hay, N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature 2012, 485, 661–665. [Google Scholar] [CrossRef]
- Joo, M.S.; Kim, W.D.; Lee, K.Y.; Kim, J.H.; Koo, J.H.; Kim, S.G. AMPK facilitates nuclear accumulation of Nrf2 by phos-phorylating at serine 550. Mol. Cell. Biol. 2016, 36, 1931–1942. [Google Scholar] [CrossRef]
- Singh, A.; Daemen, A.; Nickles, D.; Jeon, S.-M.; Foreman, O.; Sudini, K.; Gnad, F.; Lajoie, S.; Gour, N.; Mitzner, W.; et al. NRF2 activation promotes aggressive lung cancer and associates with poor clinical outcomes. Clin. Cancer Res. 2020. [Google Scholar] [CrossRef] [PubMed]
- Trapp, E.K.; Majunke, L.; Zill, B.; Sommer, H.; Andergassen, U.; Koch, J.; Harbeck, N.; Mahner, S.; Friedl, T.W.P.; Janni, W.; et al. LKB1 pro-oncogenic activity triggers cell survival in circulating tumor cells. Mol. Oncol. 2017, 11, 1508–1526. [Google Scholar] [CrossRef]
- Hollstein, P.E.; Eichner, L.J.; Brun, S.N.; Kamireddy, A.; Svensson, R.U.; Vera, L.I.; Ross, D.S.; Rymoff, T.; Hutchins, A.; Galvez, H.; et al. The AMPK-Related Kinases SIK1 and SIK3 Mediate Key Tumor-Suppressive Effects of LKB1 in NSCLC. Cancer Discov. 2019, 9, 1606–1627. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.W.; Brady, J.J.; Tsai, M.; Li, C.; Winters, I.P.; Tang, R.; Andrejka, L.; Ma, R.K.; Kunder, C.; Chu, P.; et al. An LKB1–SIK Axis Suppresses Lung Tumor Growth and Controls Differentiation. Cancer Discov. 2019, 9, 1590–1605. [Google Scholar] [CrossRef] [PubMed]
- Della Corte, C.M.; Sen, T.; Gay, C.M.; Ramkumar, K.; Diao, L.; Cardnell, R.J.; Rodriguez, B.L.; Stewart, C.A.; Papadimitrakopoulou, V.A.; Gibson, L.; et al. STING Pathway Expression Identifies NSCLC with an Immune-Responsive Phenotype. J. Thorac. Oncol. 2020, 15, 777–791. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, S.; Ivanova, E.; Guo, S.; Yoshida, R.; Campisi, M.; Sundararaman, S.K.; Tange, S.; Mitsuishi, Y.; Thai, T.C.; Masuda, S.; et al. Suppression of STING Associated with LKB1 Loss in KRAS-Driven Lung Cancer. Cancer Discov. 2019, 9, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Yao, P.; Li, L.; Ji, F.; Zhao, S.; Xu, C.; Lan, X.; Jiang, P. p53-mediated control of aspartate-asparagine homeostasis dictates LKB1 activity and modulates cell survival. Nat. Commun. 2020, 11, 1755. [Google Scholar] [CrossRef]
- Laderian, B.; Mundi, P.; Fojo, T.; Bates, S.E. Emerging Therapeutic Implications of STK11 Mutation: Case Series. Oncologist 2020, 25, 733–737. [Google Scholar] [CrossRef]
- Gill, R.K.; Yang, S.-H.; Meerzaman, D.; Mechanic, L.E.; Bowman, E.D.; Jeon, H.-S.; Chowdhuri, S.R.; Shakoori, A.; Dracheva, T.; Hong, K.-M.; et al. Frequent homozygous deletion of the LKB1/STK11 gene in non-small cell lung cancer. Oncogene 2011, 30, 3784–3791. [Google Scholar] [CrossRef]
- Scheffler, M.; Ihle, M.A.; Hein, R.; Merkelbach-Bruse, S.; Scheel, A.H.; Siemanowski, J.; Brägelmann, J.; Kron, A.; Abedpour, N.; Ueckeroth, F.; et al. K-ras Mutation Subtypes in NSCLC and Associated Co-occuring Mutations in Other Oncogenic Pathways. J. Thorac. Oncol. 2019, 14, 606–616. [Google Scholar] [CrossRef]
- Bange, E.; Marmarelis, M.E.; Hwang, W.-T.; Yang, Y.-X.; Thompson, J.C.; Rosenbaum, J.; Bauml, J.M.; Ciunci, C.; Alley, E.W.; Cohen, R.B.; et al. Impact of KRAS and TP53 Co-Mutations on Outcomes After First-Line Systemic Therapy Among Patients with STK11-Mutated Advanced Non-Small-Cell Lung Cancer. JCO Precis. Oncol. 2019, 3. [Google Scholar] [CrossRef] [PubMed]
- Facchinetti, F.; Bluthgen, M.V.; Tergemina-Clain, G.; Faivre, L.; Pignon, J.-P.; Planchard, D.; Remon, J.; Soria, J.-C.; Lacroix, L.; Besse, B. LKB1/STK11 mutations in non-small cell lung cancer patients: Descriptive analysis and prognostic value. Lung Cancer 2017, 112, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Izumi, M.; Suzumura, T.; Ogawa, K.; Matsumoto, Y.; Sawa, K.; Yoshimoto, N.; Tani, Y.; Watanabe, T.; Kaneda, H.; Mitsuoka, S.; et al. Differences in molecular epidemiology of lung cancer among ethnicities (Asian vs. Caucasian). J. Thorac. Dis. 2020, 12, 3776–3784. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Cespedes, M.; Parrella, P.; Esteller, M.; Nomoto, S.; Trink, B.; Engles, J.M.; Westra, W.H.; Herman, J.G.; Sidransky, D. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res. 2002, 62, 3659–3662. [Google Scholar] [PubMed]
- Shen, H.; Zhu, M.; Wang, C. Precision oncology of lung cancer: Genetic and genomic differences in Chinese population. NPJ Precis. Oncol. 2019, 3, 14. [Google Scholar] [CrossRef] [PubMed]
- Arauz, R.F.; Byun, J.S.; Tandon, M.; Sinha, S.; Kuhn, S.; Taylor, S.; Zingone, A.; Mitchell, K.A.; Pine, S.R.; Gardner, K.; et al. Whole-Exome Profiling of NSCLC Among African Americans. J. Thorac. Oncol. 2020, 15, 1880–1892. [Google Scholar] [CrossRef]
- Derks, J.L.; Leblay, N.; Lantuejoul, S.; Dingemans, A.-M.C.; Speel, E.-J.M.; Fernandez-Cuesta, L. New Insights into the Molecular Characteristics of Pulmonary Carcinoids and Large Cell Neuroendocrine Carcinomas, and the Impact on Their Clinical Management. J. Thorac. Oncol. 2018, 13, 752–766. [Google Scholar] [CrossRef]
- Derks, J.; Leblay, N.; Thunnissen, E.; van Suylen, R.J.; den Bakker, M.; Groen, H.J.M.; Smit, E.F.; Damhuis, R.; van den Broek, E.C.; Charbrier, A.; et al. Molecular Subtypes of Pulmonary Large-cell Neuroendocrine Carcinoma Predict Chemotherapy Treatment Outcome. Clin. Cancer Res. 2018, 24, 33–42. [Google Scholar] [CrossRef]
- George, J.; Walter, V.; Peifer, M.; Alexandrov, L.B.; Seidel, D.; Leenders, F.; Maas, L.; Müller, C.; Dahmen, I.; Delhomme, T.M.; et al. Integrative genomic profiling of large-cell neuroendocrine carcinomas reveals distinct subtypes of high-grade neuroendocrine lung tumors. Nat. Commun. 2018, 9, 1048. [Google Scholar] [CrossRef]
- Arbour, K.C.; Jordan, E.; Kim, H.R.; Dienstag, J.; Yu, H.A.; Sanchez-Vega, F.; Lito, P.; Berger, M.; Solit, D.B.; Hellmann, M.; et al. Effects of Co-occurring Genomic Alterations on Outcomes in Patients with KRAS-Mutant Non-Small Cell Lung Cancer. Clin. Cancer Res. 2018, 24, 334–340. [Google Scholar] [CrossRef]
- Schoenfeld, A.J.; Rizvi, H.; Bandlamudi, C.; Sauter, J.L.; Travis, W.D.; Rekhtman, N.; Plodkowski, A.J.; Perez-Johnston, R.; Sawan, P.; Beras, A.; et al. Clinical and molecular correlates of PD-L1 expression in patients with lung adenocarcinomas. Ann. Oncol. 2020, 31, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Gibert, J.; Clavé, S.; Hardy-Werbin, M.; Taus, Á.; Rocha, P.; Longarón, R.; Piquer, G.; Chaib, I.; Carcereny, E.; Morán, T.; et al. Concomitant genomic alterations in KRAS mutant advanced lung adenocarcinoma. Lung Cancer 2020, 140, 42–45. [Google Scholar] [CrossRef] [PubMed]
- Chakravarty, D.; Gao, J.; Phillips, S.M.; Kundra, R.; Zhang, H.; Wang, J.; Rudolph, J.E.; Yaeger, R.; Soumerai, T.; Nissan, M.H.; et al. OncoKB: A Precision Oncology Knowledge Base. JCO Precis. Oncol. 2017, 2017, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Dagogo-Jack, I.; Schrock, A.B.; Kem, M.; Jessop, N.; Lee, J.; Ali, S.M.; Ross, J.S.; Lennerz, J.K.; Shaw, A.T.; Mino-Kenudson, M. Clinicopathologic Characteristics of BRG1-Deficient NSCLC. J. Thorac. Oncol. 2020, 15, 766–776. [Google Scholar] [CrossRef] [PubMed]
- Frank, R.; Scheffler, M.; Merkelbach-Bruse, S.; Ihle, M.A.; Kron, A.; Rauer, M.; Ueckeroth, F.; Koenig, K.; Michels, S.; Fischer, R.; et al. Clinical and Pathological Characteristics of KEAP1- and NFE2L2-Mutated Non-Small Cell Lung Carcinoma (NSCLC). Clin. Cancer Res. 2018, 24, 3087–3096. [Google Scholar] [CrossRef]
- Taguchi, K.; Yamamoto, M. The KEAP1-NRF2 System in Cancer. Front. Oncol. 2017, 7, 85. [Google Scholar] [CrossRef]
- Amanam, I.; Mambetsariev, I.; Gupta, R.; Achuthan, S.; Wang, Y.; Pharaon, R.; Massarelli, E.; Koczywas, M.; Reckamp, K.; Salgia, R. Role of immunotherapy and co-mutations on KRAS-mutant non- small cell lung cancer survival. J. Thorac. Dis. 2020, 12, 5086–5095. [Google Scholar] [CrossRef]
- Skoulidis, F.; Byers, L.A.; Diao, L.; Papadimitrakopoulou, V.A.; Tong, P.; Izzo, J.G.; Behrens, C.; Kadara, H.; Parra, E.R.; Canales, J.R.; et al. Co-occurring Genomic Alterations Define Major Subsets of KRAS-Mutant Lung Adenocarcinoma with Distinct Biology, Immune Profiles, and Therapeutic Vulnerabilities. Cancer Discov. 2015, 5, 860–877. [Google Scholar] [CrossRef]
- La Fleur, L.; Falk-Sörqvist, E.; Smeds, P.; Berglund, A.; Sundström, M.; Mattsson, J.S.; Brandén, E.; Koyi, H.; Isaksson, J.; Brunnström, H.; et al. Mutation patterns in a population-based non-small cell lung cancer cohort and prognostic impact of concomitant mutations in KRAS and TP53 or STK11. Lung Cancer. 2019, 130, 50–58. [Google Scholar] [CrossRef]
- Biton, J.; Mansuet-Lupo, A.; Pécuchet, N.; Alifano, M.; Ouakrim, H.; Arrondeau, J.; Boudou-Rouquette, P.; Goldwasser, F.; Leroy, K.; Goc, J.; et al. TP53, STK11, and EGFR Mutations Predict Tumor Immune Profile and the Response to Anti-PD-1 in Lung Adenocarcinoma. Clin. Cancer Res. 2018, 24, 5710–5723. [Google Scholar] [CrossRef]
- Kadara, H.; Choi, M.; Zhang, J.; Parra, E.R.; Rodriguez-Canales, J.; Gaffney, S.G.; Zhao, Z.; Behrens, C.; Fujimoto, J.; Chow, C.; et al. Whole-exome sequencing and immune profiling of early-stage lung adenocarcinoma with fully annotated clinical follow-up. Ann. Oncol. 2017, 28, 75–82. [Google Scholar] [PubMed]
- Lamberti, G.; Spurr, L.F.; Li, Y.; Ricciuti, B.; Recondo, G.; Umeton, R.; Nishino, M.; Sholl, L.M.; Meyerson, M.L.; Cherniack, A.D.; et al. Clinicopathological and genomic correlates of programmed cell death ligand 1 (PD-L1) expression in nonsquamous non-small-cell lung cancer. Ann. Oncol. 2020, 31, 807–814. [Google Scholar] [CrossRef] [PubMed]
- Domingues, I.; Cedres, S.; Callejo, A.; Vivancos, A.; Martínez-Marti, A.; Felip, E.; Perez, S. Long duration of immunotherapy in a STK11 mutated/KRAS wild-type non-small cell lung cancer patient. Pulmonology 2020, 26, 49–50. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Yu, M.; Zhou, L.; Jiang, L.; Huang, M. Durable response to combination radiotherapy and immunotherapy in EP-resistant lung large-cell neuroendocrine carcinoma with B2M and STK11 mutations: A case report. Immunotherapy 2020, 12, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Nadal, E.; Heeke, S.; Benzaquen, J.; Vilariño, N.; Navarro, A.; Azuara, D.; Varela, M.; Otto, J.; Baixeras, N.; Shahbazian, D.; et al. Two Cases of Large Stage Lung Adenocarcinoma with Radiological Compete Responses to Nivolumab Treatment Harboring a STK11/LKB1 Mutation. JCO Precis. Oncol. 2020, 4, 1239–1245. [Google Scholar] [CrossRef]
- Papillon-Cavanagh, S.; Doshi, P.; Dobrin, R.; Szustakowski, J.; Walsh, A.M. STK11 and KEAP1 mutations as prognostic biomarkers in an observational real-world lung adenocarcinoma cohort. ESMO Open 2020, 5, e000706. [Google Scholar] [PubMed]
- Bai, X.; Wu, D.H.; Ma, S.C.; Wang, J.; Tang, X.R.; Kang, S.; Fu, Q.J.; Cao, C.H.; Luo, H.S.; Chen, Y.H.; et al. Development and validation of a genomic mutation signature to predict response to PD-1 inhibitors in non-squamous NSCLC: A multicohort study. J. Immunother. Cancer 2020, 8, e000381. [Google Scholar] [CrossRef]
- Hasegawa, T.; Yanagitani, N.; Ninomiya, H.; Sakamoto, H.; Tozuka, T.; Yoshida, H.; Amino, Y.; Uematsu, S.; Yoshizawa, T.; Ariyasu, R.; et al. Association Between the Efficacy of Pembrolizumab and Low STK11/LKB1 Expression in High-PD-L1-expressing Non-small-cell Lung Cancer. In Vivo 2020, 34, 2997–3003. [Google Scholar] [CrossRef]
- Falk, A.T.; Yazbeck, N.; Guibert, N.; Chamorey, E.; Paquet, A.; Ribeyre, L.; Bence, C.; Zahaf, K.; Leroy, S.; Marquette, C.-H.; et al. Effect of mutant variants of the KRAS gene on PD-L1 expression and on the immune microenvironment and association with clinical outcome in lung adenocarcinoma patients. Lung Cancer 2018, 121, 70–75. [Google Scholar] [CrossRef]
- Jeong, Y.; Hellyer, J.A.; Stehr, H.; Hoang, N.T.; Niu, X.; Das, M.; Padda, S.K.; Ramchandran, K.; Neal, J.W.; Wakelee, H.A.; et al. Role of KEAP1/NFE2L2 Mutations in the Chemotherapeutic Response of Patients with Non-Small Cell Lung Cancer. Clin. Cancer Res. 2020, 26, 274–281. [Google Scholar] [CrossRef]
- Ghimessy, A.; Radeczky, P.; Laszlo, V.; Hegedus, B.; Renyi-Vamos, F.; Fillinger, J.; Klepetko, W.; Lang, C.; Dome, B.; Megyesfalvi, Z. Current therapy of KRAS-mutant lung cancer. Cancer Metastasis Rev. 2020, 39, 1159–1177. [Google Scholar] [CrossRef] [PubMed]
- Guibert, N.; Ilie, M.; Long, E.; Hofman, V.; Bouhlel, L.; Brest, P.; Mograbi, B.; Marquette, C.H.; Didier, A.; Mazieres, J.; et al. KRAS Mutations in Lung Adenocarcinoma: Molecular and Epidemiological Characteristics, Methods for Detection, and Therapeutic Strategy Perspectives. Curr. Mol. Med. 2015, 15, 418–432. [Google Scholar] [CrossRef] [PubMed]
- Hallin, J.; Engstrom, L.D.; Hargis, L.; Calinisan, A.; Aranda, R.; Briere, D.M.; Sudhakar, N.; Bowcut, V.; Baer, B.R.; Ballard, J.A.; et al. The KRAS(G12C) Inhibitor MRTX849 Provides Insight toward Therapeutic Susceptibility of KRAS-Mutant Cancers in Mouse Models and Patients. Cancer Discov. 2020, 10, 54–71. [Google Scholar] [CrossRef] [PubMed]
- Schabath, M.B.; Welsh, E.A.; Fulp, W.J.; Chen, L.; Teer, J.K.; Thompson, Z.J.; Engel, B.E.; Xie, M.; Berglund, A.E.; Creelan, B.C.; et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene 2016, 35, 3209–3216. [Google Scholar] [CrossRef] [PubMed]
- Momcilovic, M.; Shackelford, D.B. Targeting LKB1 in cancer—Exposing and exploiting vulnerabilities. Br. J. Cancer 2015, 113, 574–584. [Google Scholar] [CrossRef]
- Caiola, E.; Iezzi, A.; Tomanelli, M.; Bonaldi, E.; Scagliotti, A.; Colombo, M.; Guffanti, F.; Micotti, E.; Garassino, M.C.; Minoli, L.; et al. LKB1 Deficiency Renders NSCLC Cells Sensitive to ERK Inhibitors. J. Thorac. Oncol. 2020, 15, 360–370. [Google Scholar] [CrossRef]
- Galan-Cobo, A.; Sitthideatphaiboon, P.; Qu, X.; Poteete, A.; Pisegna, M.A.; Tong, P.; Chen, P.-H.; Boroughs, L.K.; Rodriguez, M.L.M.; Zhang, W.; et al. LKB1 and KEAP1/NRF2 Pathways Cooperatively Promote Metabolic Reprogramming with Enhanced Glutamine Dependence in KRAS-Mutant Lung Adenocarcinoma. Cancer Res. 2019, 79, 3251–3267. [Google Scholar] [CrossRef]
- Romero, R.; Sayin, V.I.; Davidson, S.M.; Bauer, M.R.; Singh, S.X.; Leboeuf, S.E.; Karakousi, T.R.; Ellis, D.C.; Bhutkar, A.; Sánchez-Rivera, F.J.; et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat. Med. 2017, 23, 1362–1368. [Google Scholar] [CrossRef]
- Aggarwal, C.; Thompson, J.C.; Black, T.A.; Katz, S.I.; Fan, R.; Yee, S.S.; Chien, A.L.; Evans, T.L.; Bauml, J.M.; Alley, E.W.; et al. Clinical Implications of Plasma-Based Genotyping with the Delivery of Personalized Therapy in Metastatic Non-Small Cell Lung Cancer. JAMA Oncol. 2019, 5, 173–180. [Google Scholar] [CrossRef]
- Kruglyak, K.M.; Lin, E.; Ong, F.S. Next-Generation Sequencing and Applications to the Diagnosis and Treatment of Lung Cancer. Adv. Exp. Med. Biol. 2016, 890, 123–136. [Google Scholar] [CrossRef]
- Heeke, S.; Hofman, V.; Long-Mira, E.; Lespinet, V.; Lalvée, S.; Bordone, O.; Ribeyre, C.; Tanga, V.; Benzaquen, J.; Leroy, S.; et al. Use of the Ion PGM and the GeneReader NGS Systems in Daily Routine Practice for Advanced Lung Adenocarcinoma Patients: A Practical Point of View Reporting a Comparative Study and Assessment of 90 Patients. Cancers 2018, 10, 88. [Google Scholar] [CrossRef] [PubMed]
- DiBardino, D.M.; Rawson, D.W.; Saqi, A.; Heymann, J.J.; Pagan, C.A.; Bulman, W.A. Next-generation sequencing of non-small cell lung cancer using a customized, targeted sequencing panel: Emphasis on small biopsy and cytology. CytoJournal 2017, 14, 7. [Google Scholar] [CrossRef] [PubMed]
- Malapelle, U.; Pisapia, P.; Rocco, D.; Smeraglio, R.; Di Spirito, M.; Bellevicine, C.; Troncone, G. Next generation sequencing techniques in liquid biopsy: Focus on non-small cell lung cancer patients. Transl. Lung Cancer Res. 2016, 5, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Sholl, L.M. Molecular diagnostics of lung cancer in the clinic. Transl. Lung Cancer Res. 2017, 6, 560–569. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.C.; Yee, S.S.; Troxel, A.B.; Savitch, S.L.; Fan, R.; Balli, D.; Lieberman, D.B.; Morrissette, J.D.; Evans, T.L.; Bauml, J.M.; et al. Detection of Therapeutically Targetable Driver and Resistance Mutations in Lung Cancer Patients by Next-Generation Sequencing of Cell-Free Circulating Tumor DNA. Clin. Cancer Res. 2016, 22, 5772–5782. [Google Scholar] [CrossRef] [PubMed]
- Kalbasi, A.; Ribas, A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat. Rev. Immunol. 2020, 20, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Lamberti, G.; Sisi, M.; Andrini, E.; Palladini, A.; Giunchi, F.; Lollini, P.-L.; Ardizzoni, A.; Gelsomino, F. The Mechanisms of PD-L1 Regulation in Non-Small-Cell Lung Cancer (NSCLC): Which Are the Involved Players? Cancers 2020, 12, 3129. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.K.; Man, J.; Lord, S.; Cooper, W.; Links, M.; Gebski, V.; Herbst, R.S.; Gralla, R.J.; Mok, T.; Yang, J.C. Clinical and Molecular Characteristics Associated with Survival Among Patients Treated with Checkpoint Inhibitors for Advanced Non-Small Cell Lung Carcinoma: A Systematic Review and Meta-analysis. JAMA Oncol. 2018, 4, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Legras, A.; Barritault, M.; Tallet, A.; Fabre, E.; Guyard, A.; Rance, B.; Digan, W.; Pecuchet, N.; Giroux-Leprieur, E.; Julie, C.; et al. Validity of Targeted Next-Generation Sequencing in Routine Care for Identifying Clinically Relevant Molecular Profiles in Non-Small-Cell Lung Cancer: Results of a 2-Year Experience on 1343 Samples. J. Mol. Diagn. 2018, 20, 550–564. [Google Scholar] [CrossRef]
- Marinelli, D.; Mazzotta, M.; Scalera, S.; Terrenato, I.; Sperati, F.; D’Ambrosio, L.; Pallocca, M.; Corleone, G.; Krasniqi, E.; Pizzuti, L.; et al. KEAP1-driven co-mutations in lung adenocarcinoma unresponsive to immunotherapy despite high tumor mutational burden. Ann. Oncol. 2020, 31, 1746–1754. [Google Scholar] [CrossRef]
- Mitchell, K.G.; Parra, E.R.; Zhang, J.; Nelson, D.B.; Corsini, E.M.; Villalobos, P.; Moran, C.A.; Skoulidis, F.; Wistuba, I.I.; Fujimoto, J.; et al. LKB1/STK11 Expression in Lung Adenocarcinoma and Associations with Patterns of Recurrence. Ann. Thorac. Surg. 2020, 110, 1131–1138. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.H.; Zhao, F.J.; Mo, H.Y.; Jia, R.R.; Tang, J.; Zhao, X.H.; Wei, J.L.; Huo, R.R.; Li, Q.Q.; You, X.M. Association between LKB1 expression and prognosis of patients with solid tumours: An updated systematic review and meta-analysis. BMJ Open 2019, 9, e027185. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P.; Badoual, C.; Henderson, F.; Berland, L.; Hamila, M.; Long-Mira, E.; Lassalle, S.; Roussel, H.; Hofman, V.; Tartour, E.; et al. Multiplexed Immunohistochemistry for Molecular and Immune Profiling in Lung Cancer-Just About Ready for Prime-Time? Cancers 2019, 11, 283. [Google Scholar] [CrossRef] [PubMed]
- Ilie, M.; Beaulande, M.; Hamila, M.; Erb, G.; Hofman, V.; Hofman, P. Automated chromogenic multiplexed immunohisto-chemistry assay for diagnosis and predictive biomarker testing in non-small cell lung cancer. Lung Cancer 2018, 124, 90–94. [Google Scholar] [CrossRef]
- Diaz, L.A., Jr.; Bardelli, A. Liquid biopsies: Genotyping circulating tumor DNA. J. Clin. Oncol. 2014, 32, 579–586. [Google Scholar] [CrossRef]
- Jiang, J.; Adams, H.-P.; Yao, L.; Yaung, S.; Lal, P.; Balasubramanyam, A.; Fuhlbrück, F.; Tikoo, N.; Lovejoy, A.F.; Froehler, S.; et al. Concordance of Genomic Alterations by Next-Generation Sequencing in Tumor Tissue versus Cell-Free DNA in Stage I–IV Non-Small Cell Lung Cancer. J. Mol. Diagn. 2020, 22, 228–235. [Google Scholar] [CrossRef]
- Remon, J.; Lacroix, L.; Jovelet, C.; Caramella, C.; Howarth, K.; Plagnol, V.; Rosenfeld, N.; Morris, C.; Mezquita, L.; Pannet, C.; et al. Real-World Utility of an Amplicon-Based Next-Generation Sequencing Liquid Biopsy for Broad Molecular Profiling in Patients with Advanced Non-Small-Cell Lung Cancer. JCO Precis. Oncol. 2019, 3, 1–14. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).