Influence of Biopsy Technique on Molecular Genetic Tumor Characterization in Non-Small Cell Lung Cancer—The Prospective, Randomized, Single-Blinded, Multicenter PROFILER Study Protocol
Abstract
1. Background
2. Methods/Design
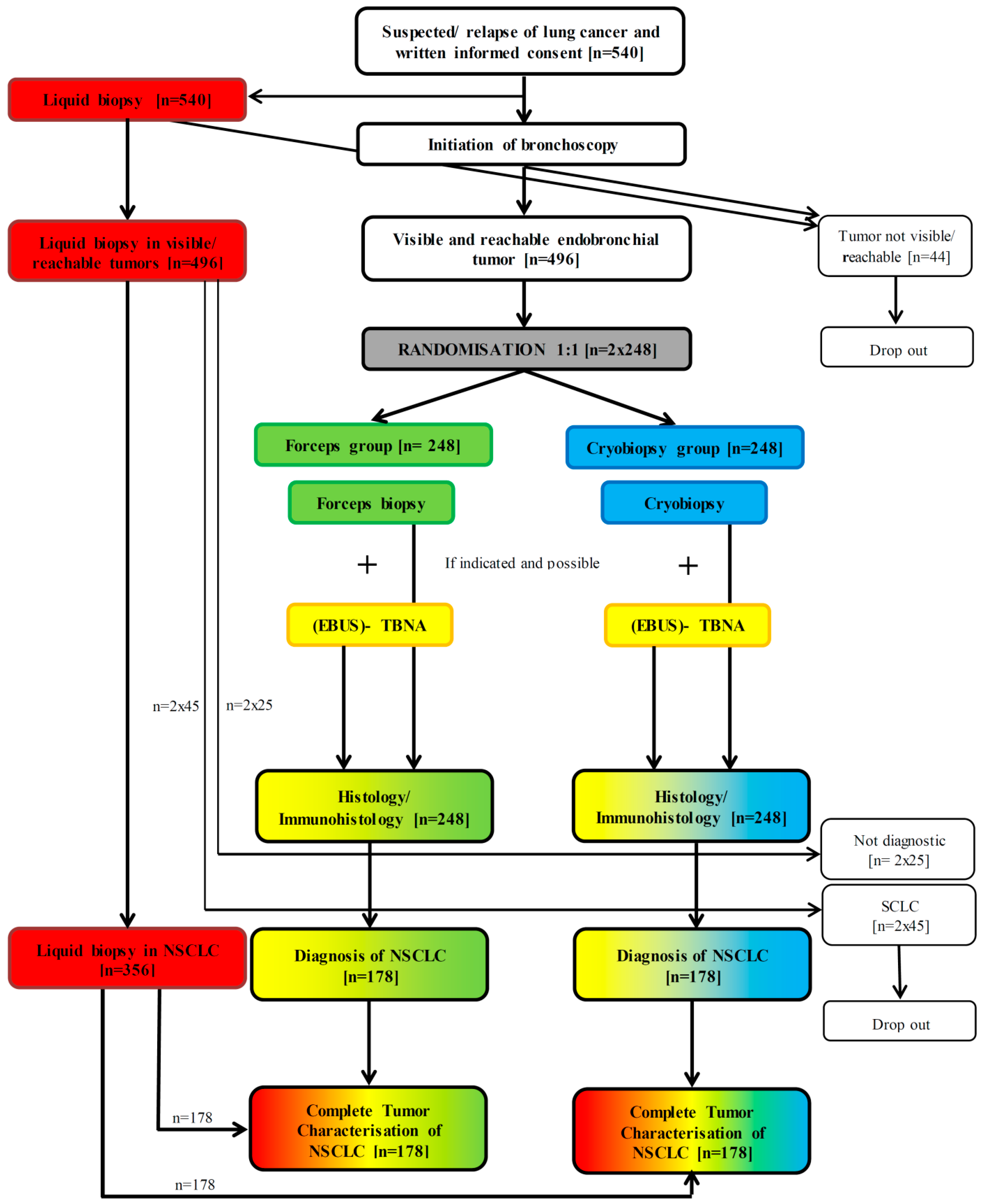
2.1. Trial Design
2.2. Patient Selection
2.3. Study Objectives
2.4. Pre-Procedural Training and Standardization
2.5. Procedural Protocol
2.5.1. Patient Enrollment
2.5.2. Randomization
2.5.3. Liquid Biopsy Procedure
2.5.4. Bronchoscopy
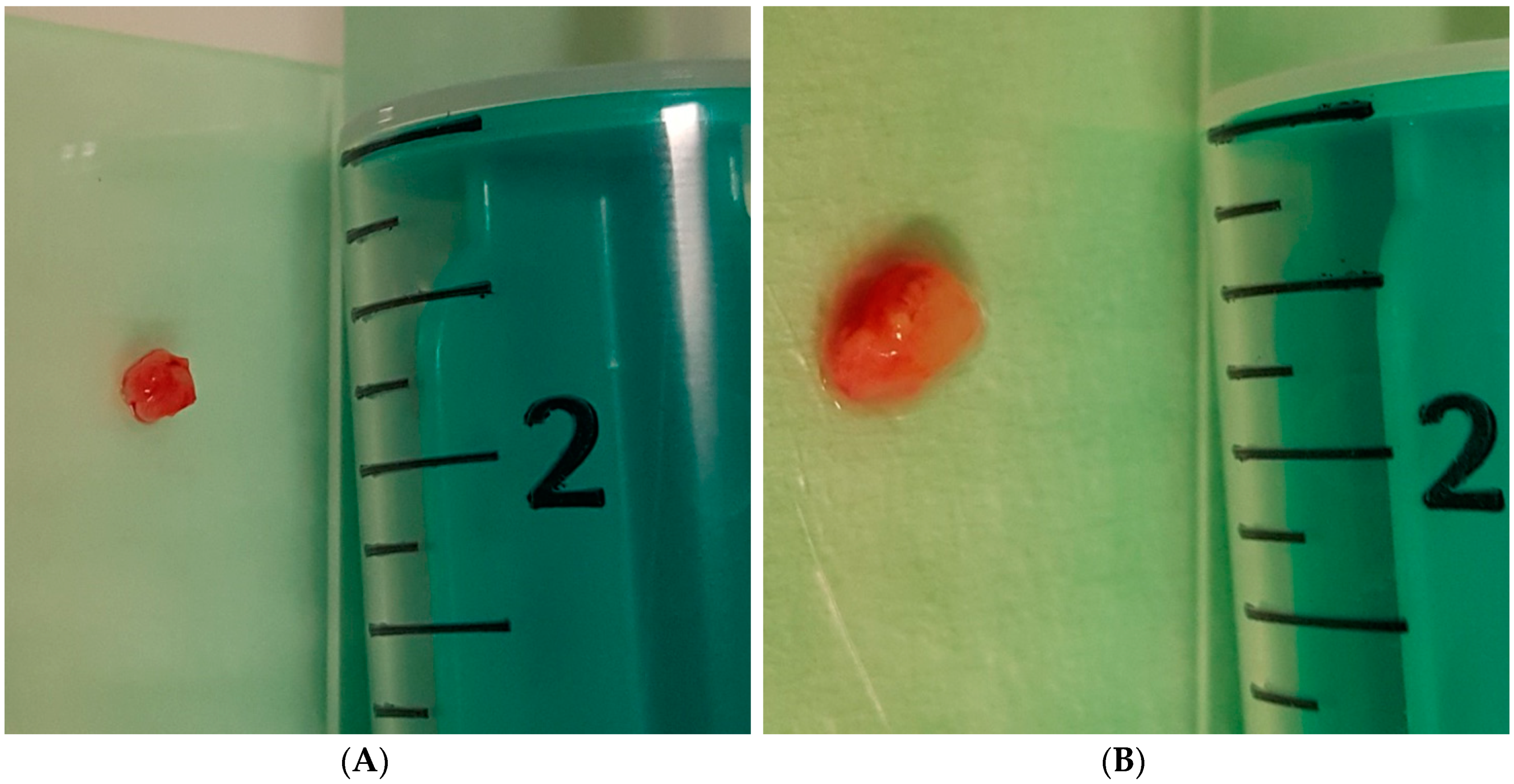
2.5.5. Forceps and Cryobiopsy Procedure
2.5.6. TBNA Procedure
2.6. Pathological Analysis
2.6.1. Baseline Pathological Evaluation
2.6.2. Molecular Genetic Evaluation
2.7. Safety Data
2.7.1. Bleeding
2.7.2. Other side Effects
2.8. Statistical Methodology
2.8.1. Sample Size Estimate
2.8.2. Randomization
2.8.3. Blinding
2.8.4. Primary Endpoint Analysis
2.8.5. Secondary Endpoint Analysis
2.9. Trial Oversight and Ethics Approval
3. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Availability of Data and Materials
References
- Siegel, R.; Ma, J.; Zou, Z.; Jemal, A. Cancer statistics, 2014. CA A Cancer J. Clin. 2014, 64, 9–29. [Google Scholar] [CrossRef] [PubMed]
- Cheng, T.Y.; Cramb, S.M.; Baade, P.D.; Youlden, D.R.; Nwogu, C.; Reid, M.E. The International Epidemiology of Lung Cancer: Latest Trends, Disparities, and Tumor Characteristics. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2016, 11, 1653–1671. [Google Scholar] [CrossRef]
- Ferlay, J.; Steliarova-Foucher, E.; Lortet-Tieulent, J.; Rosso, S.; Coebergh, J.W.; Comber, H.; Forman, D.; Bray, F. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries in 2012. Eur. J. Cancer (Oxf. Engl. 1990) 2013, 49, 1374–1403. [Google Scholar] [CrossRef]
- Miller, K.D.; Nogueira, L.; Mariotto, A.B.; Rowland, J.H.; Yabroff, K.R.; Alfano, C.M.; Jemal, A.; Kramer, J.L.; Siegel, R.L. Cancer treatment and survivorship statistics, 2019. CA A Cancer J. Clin. 2019. [Google Scholar] [CrossRef]
- Meza, R.; Meernik, C.; Jeon, J.; Cote, M.L. Lung cancer incidence trends by gender, race and histology in the United States, 1973–2010. PLoS ONE 2015, 10, e0121323. [Google Scholar] [CrossRef]
- Jemal, A.; Siegel, R.; Ward, E.; Hao, Y.; Xu, J.; Thun, M.J. Cancer statistics, 2009. CA A Cancer J. Clin. 2009, 59, 225–249. [Google Scholar] [CrossRef]
- Morgensztern, D.; Ng, S.H.; Gao, F.; Govindan, R. Trends in stage distribution for patients with non-small cell lung cancer: A National Cancer Database survey. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2010, 5, 29–33. [Google Scholar] [CrossRef]
- Goldstraw, P.; Chansky, K.; Crowley, J.; Rami-Porta, R.; Asamura, H.; Eberhardt, W.E.; Nicholson, A.G.; Groome, P.; Mitchell, A.; Bolejack, V. The IASLC Lung Cancer Staging Project: Proposals for Revision of the TNM Stage Groupings in the Forthcoming (Eighth) Edition of the TNM Classification for Lung Cancer. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2016, 11, 39–51. [Google Scholar] [CrossRef]
- Aisner, D.L.; Marshall, C.B. Molecular pathology of non-small cell lung cancer: A practical guide. Am. J. Clin. Pathol. 2012, 138, 332–346. [Google Scholar] [CrossRef]
- Lindeman, N.I.; Cagle, P.T.; Aisner, D.L.; Arcila, M.E.; Beasley, M.B.; Bernicker, E.H.; Colasacco, C.; Dacic, S.; Hirsch, F.R.; Kerr, K.; et al. Updated Molecular Testing Guideline for the Selection of Lung Cancer Patients for Treatment With Targeted Tyrosine Kinase Inhibitors: Guideline From the College of American Pathologists, the International Association for the Study of Lung Cancer, and the Association for Molecular Pathology. Arch. Pathol. Lab. Med. 2018, 142, 321–346. [Google Scholar] [CrossRef]
- Yang, J.C.; Wu, Y.L.; Schuler, M.; Sebastian, M.; Popat, S.; Yamamoto, N.; Zhou, C.; Hu, C.P.; O’Byrne, K.; Feng, J.; et al. Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): Analysis of overall survival data from two randomised, phase 3 trials. Lancet Oncol. 2015, 16, 141–151. [Google Scholar] [CrossRef]
- Zhou, C.; Wu, Y.L.; Chen, G.; Feng, J.; Liu, X.Q.; Wang, C.; Zhang, S.; Wang, J.; Zhou, S.; Ren, S.; et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): A multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011, 12, 735–742. [Google Scholar] [CrossRef]
- Rosell, R.; Carcereny, E.; Gervais, R.; Vergnenegre, A.; Massuti, B.; Felip, E.; Palmero, R.; Garcia-Gomez, R.; Pallares, C.; Sanchez, J.M.; et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): A multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012, 13, 239–246. [Google Scholar] [CrossRef]
- Wu, Y.L.; Zhou, C.; Liam, C.K.; Wu, G.; Liu, X.; Zhong, Z.; Lu, S.; Cheng, Y.; Han, B.; Chen, L.; et al. First-line erlotinib versus gemcitabine/cisplatin in patients with advanced EGFR mutation-positive non-small-cell lung cancer: Analyses from the phase III, randomized, open-label, ENSURE study. Ann. Oncol. Off. J. Eur. Soc. Med Oncol. 2015, 26, 1883–1889. [Google Scholar] [CrossRef]
- Fukuoka, M.; Wu, Y.L.; Thongprasert, S.; Sunpaweravong, P.; Leong, S.S.; Sriuranpong, V.; Chao, T.Y.; Nakagawa, K.; Chu, D.T.; Saijo, N.; et al. Biomarker analyses and final overall survival results from a phase III, randomized, open-label, first-line study of gefitinib versus carboplatin/paclitaxel in clinically selected patients with advanced non-small-cell lung cancer in Asia (IPASS). J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2011, 29, 2866–2874. [Google Scholar] [CrossRef]
- Sequist, L.V.; Yang, J.C.; Yamamoto, N.; O’Byrne, K.; Hirsh, V.; Mok, T.; Geater, S.L.; Orlov, S.; Tsai, C.M.; Boyer, M.; et al. Phase III study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2013, 31, 3327–3334. [Google Scholar] [CrossRef]
- Yang, J.C.; Hirsh, V.; Schuler, M.; Yamamoto, N.; O’Byrne, K.J.; Mok, T.S.; Zazulina, V.; Shahidi, M.; Lungershausen, J.; Massey, D.; et al. Symptom control and quality of life in LUX-Lung 3: A phase III study of afatinib or cisplatin/pemetrexed in patients with advanced lung adenocarcinoma with EGFR mutations. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2013, 31, 3342–3350. [Google Scholar] [CrossRef]
- Mok, T.S.; Cheng, Y.; Zhou, X.; Lee, K.H.; Nakagawa, K.; Niho, S.; Lee, M.; Linke, R.; Rosell, R.; Corral, J.; et al. Improvement in Overall Survival in a Randomized Study That Compared Dacomitinib With Gefitinib in Patients With Advanced Non-Small-Cell Lung Cancer and EGFR-Activating Mutations. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2018, 36, 2244–2250. [Google Scholar] [CrossRef]
- Zhou, C.; Wu, Y.L.; Chen, G.; Feng, J.; Liu, X.Q.; Wang, C.; Zhang, S.; Wang, J.; Zhou, S.; Ren, S.; et al. Final overall survival results from a randomised, phase III study of erlotinib versus chemotherapy as first-line treatment of EGFR mutation-positive advanced non-small-cell lung cancer (OPTIMAL, CTONG-0802). Ann. Oncol. Off. J. Eur. Soc. Med Oncol. 2015, 26, 1877–1883. [Google Scholar] [CrossRef] [PubMed]
- Mok, T.S.; Wu, Y.L.; Thongprasert, S.; Yang, C.H.; Chu, D.T.; Saijo, N.; Sunpaweravong, P.; Han, B.; Margono, B.; Ichinose, Y.; et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N. Engl. J. Med. 2009, 361, 947–957. [Google Scholar] [CrossRef] [PubMed]
- Maemondo, M.; Inoue, A.; Kobayashi, K.; Sugawara, S.; Oizumi, S.; Isobe, H.; Gemma, A.; Harada, M.; Yoshizawa, H.; Kinoshita, I.; et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N. Engl. J. Med. 2010, 362, 2380–2388. [Google Scholar] [CrossRef]
- Yoneda, K.; Imanishi, N.; Ichiki, Y.; Tanaka, F. A liquid biopsy in primary lung cancer. Surg. Today 2019, 49, 1–14. [Google Scholar] [CrossRef]
- Castello, A.; Russo, C.; Grizzi, F.; Qehajaj, D.; Lopci, E. Prognostic Impact of Intratumoral Heterogeneity Based on Fractal Geometry Analysis in Operated NSCLC Patients. Mol. Imaging Biol. Mib Off. Publ. Acad. Mol. Imaging 2018. [Google Scholar] [CrossRef]
- Chen, Z.Y.; Zhong, W.Z.; Zhang, X.C.; Su, J.; Yang, X.N.; Chen, Z.H.; Yang, J.J.; Zhou, Q.; Yan, H.H.; An, S.J.; et al. EGFR mutation heterogeneity and the mixed response to EGFR tyrosine kinase inhibitors of lung adenocarcinomas. Oncologist 2012, 17, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Chen, Z.; Xu, C.; Zhang, X.; Yan, H.; Su, J.; Yang, J.; Xie, Z.; Guo, W.; Li, F.; et al. Intratumoral heterogeneity of EGFR-activating mutations in advanced NSCLC patients at the single-cell level. BMC Cancer 2019, 19, 369. [Google Scholar] [CrossRef] [PubMed]
- Jamal-Hanjani, M.; Wilson, G.A.; McGranahan, N.; Birkbak, N.J.; Watkins, T.B.K.; Veeriah, S.; Shafi, S.; Johnson, D.H.; Mitter, R.; Rosenthal, R.; et al. Tracking the Evolution of Non-Small-Cell Lung Cancer. N. Engl. J. Med. 2017, 376, 2109–2121. [Google Scholar] [CrossRef]
- Jia, Q.; Wu, W.; Wang, Y.; Alexander, P.B.; Sun, C.; Gong, Z.; Cheng, J.N.; Sun, H.; Guan, Y.; Xia, X.; et al. Local mutational diversity drives intratumoral immune heterogeneity in non-small cell lung cancer. Nat. Commun. 2018, 9, 5361. [Google Scholar] [CrossRef]
- Kwon, D.; Koh, J.; Kim, S.; Go, H.; Kim, Y.A.; Keam, B.; Kim, T.M.; Kim, D.W.; Jeon, Y.K.; Chung, D.H. MET exon 14 skipping mutation in triple-negative pulmonary adenocarcinomas and pleomorphic carcinomas: An analysis of intratumoral MET status heterogeneity and clinicopathological characteristics. Lung Cancer (Amst. Neth.) 2017, 106, 131–137. [Google Scholar] [CrossRef]
- Mansuet-Lupo, A.; Zouiti, F.; Alifano, M.; Tallet, A.; Charpentier, M.C.; Ducruit, V.; Devez, F.; Lemaitre, F.; Laurent-Puig, P.; Damotte, D.; et al. Intratumoral distribution of EGFR mutations and copy number in metastatic lung cancer, what impact on the initial molecular diagnosis? J. Transl. Med. 2014, 12, 131. [Google Scholar] [CrossRef][Green Version]
- Nakamura, S.; Hayashi, K.; Imaoka, Y.; Kitamura, Y.; Akazawa, Y.; Tabata, K.; Groen, R.; Tsuchiya, T.; Yamasaki, N.; Nagayasu, T.; et al. Intratumoral heterogeneity of programmed cell death ligand-1 expression is common in lung cancer. PLoS ONE 2017, 12, e0186192. [Google Scholar] [CrossRef]
- Remon, J.; Majem, M. EGFR mutation heterogeneity and mixed response to EGFR tyrosine kinase inhibitors of non small cell lung cancer: A clue to overcoming resistance. Transl. Lung Cancer Res. 2013, 2, 445–448. [Google Scholar] [CrossRef]
- Soucheray, M.; Capelletti, M.; Pulido, I.; Kuang, Y.; Paweletz, C.P.; Becker, J.H.; Kikuchi, E.; Xu, C.; Patel, T.B.; Al-Shahrour, F.; et al. Intratumoral Heterogeneity in EGFR-Mutant NSCLC Results in Divergent Resistance Mechanisms in Response to EGFR Tyrosine Kinase Inhibition. Cancer Res. 2015, 75, 4372–4383. [Google Scholar] [CrossRef]
- Taniguchi, K.; Okami, J.; Kodama, K.; Higashiyama, M.; Kato, K. Intratumor heterogeneity of epidermal growth factor receptor mutations in lung cancer and its correlation to the response to gefitinib. Cancer Sci. 2008, 99, 929–935. [Google Scholar] [CrossRef]
- Zhang, L.L.; Kan, M.; Zhang, M.M.; Yu, S.S.; Xie, H.J.; Gu, Z.H.; Wang, H.N.; Zhao, S.X.; Zhou, G.B.; Song, H.D.; et al. Multiregion sequencing reveals the intratumor heterogeneity of driver mutations in TP53-driven non-small cell lung cancer. Int. J. Cancer 2017, 140, 103–108. [Google Scholar] [CrossRef]
- Leighl, N.B.; Rekhtman, N.; Biermann, W.A.; Huang, J.; Mino-Kenudson, M.; Ramalingam, S.S.; West, H.; Whitlock, S.; Somerfield, M.R. Molecular testing for selection of patients with lung cancer for epidermal growth factor receptor and anaplastic lymphoma kinase tyrosine kinase inhibitors: American Society of Clinical Oncology endorsement of the College of American Pathologists/International Association for the study of lung cancer/association for molecular pathology guideline. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2014, 32, 3673–3679. [Google Scholar] [CrossRef]
- Lindeman, N.I.; Cagle, P.T.; Beasley, M.B.; Chitale, D.A.; Dacic, S.; Giaccone, G.; Jenkins, R.B.; Kwiatkowski, D.J.; Saldivar, J.S.; Squire, J.; et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: Guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2013, 8, 823–859. [Google Scholar] [CrossRef]
- Kalemkerian, G.P.; Narula, N.; Kennedy, E.B.; Biermann, W.A.; Donington, J.; Leighl, N.B.; Lew, M.; Pantelas, J.; Ramalingam, S.S.; Reck, M.; et al. Molecular Testing Guideline for the Selection of Patients With Lung Cancer for Treatment With Targeted Tyrosine Kinase Inhibitors: American Society of Clinical Oncology Endorsement of the College of American Pathologists/International Association for the Study of Lung Cancer/Association for Molecular Pathology Clinical Practice Guideline Update. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 2018, 36, 911–919. [Google Scholar] [CrossRef]
- Liu, Y.; Dong, Z.; Jiang, T.; Hou, L.; Wu, F.; Gao, G.; He, Y.; Zhao, J.; Li, X.; Zhao, C.; et al. Heterogeneity of PD-L1 Expression Among the Different Histological Components and Metastatic Lymph Nodes in Patients With Resected Lung Adenosquamous Carcinoma. Clin. Lung Cancer 2018, 19, e421–e430. [Google Scholar] [CrossRef]
- Hetzel, J.; Eberhardt, R.; Herth, F.J.; Petermann, C.; Reichle, G.; Freitag, L.; Dobbertin, I.; Franke, K.J.; Stanzel, F.; Beyer, T.; et al. Cryobiopsy increases the diagnostic yield of endobronchial biopsy: A multicentre trial. Eur. Respir. J. 2012, 39, 685–690. [Google Scholar] [CrossRef] [PubMed]
- Haentschel, M.; Boeckeler, M.; Ehab, A.; Wagner, R.; Spengler, W.; Steger, V.; Boesmueller, H.; Horger, M.; Lewis, R.A.; Fend, F.; et al. Cryobiopsy increases the EGFR detection rate in non-small cell lung cancer. Lung Cancer (Amst. Neth.) 2020, 141, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Du Rand, I.A.; Barber, P.V.; Goldring, J.; Lewis, R.A.; Mandal, S.; Munavvar, M.; Rintoul, R.C.; Shah, P.L.; Singh, S.; Slade, M.G.; et al. British Thoracic Society guideline for advanced diagnostic and therapeutic flexible bronchoscopy in adults. Thorax 2011, 66 (Suppl. 3), iii1–iii21. [Google Scholar] [CrossRef]
- Du Rand, I.A.; Blaikley, J.; Booton, R.; Chaudhuri, N.; Gupta, V.; Khalid, S.; Mandal, S.; Martin, J.; Mills, J.; Navani, N.; et al. British Thoracic Society guideline for diagnostic flexible bronchoscopy in adults: Accredited by NICE. Thorax 2013, 68 (Suppl. 1), i1–i44. [Google Scholar] [CrossRef] [PubMed]
- Hetzel, J.; Hetzel, M.; Hasel, C.; Moeller, P.; Babiak, A. Old meets modern: The use of traditional cryoprobes in the age of molecular biology. Respir. Int. Rev. Thorac. Dis. 2008, 76, 193–197. [Google Scholar] [CrossRef] [PubMed]
- Colella, S.; Haentschel, M.; Shah, P.; Poletti, V.; Hetzel, J. Transbronchial Lung Cryobiopsy in Interstitial Lung Diseases: Best Practice. Respir. Int. Rev. Thorac. Dis. 2018, 95, 383–391. [Google Scholar] [CrossRef]
- Wahidi, M.M.; Herth, F.; Yasufuku, K.; Shepherd, R.W.; Yarmus, L.; Chawla, M.; Lamb, C.; Casey, K.R.; Patel, S.; Silvestri, G.A.; et al. Technical Aspects of Endobronchial Ultrasound-Guided Transbronchial Needle Aspiration: CHEST Guideline and Expert Panel Report. Chest 2016, 149, 816–835. [Google Scholar] [CrossRef]
- van der Heijden, E.H.; Casal, R.F.; Trisolini, R.; Steinfort, D.P.; Hwangbo, B.; Nakajima, T.; Guldhammer-Skov, B.; Rossi, G.; Ferretti, M.; Herth, F.F.; et al. Guideline for the acquisition and preparation of conventional and endobronchial ultrasound-guided transbronchial needle aspiration specimens for the diagnosis and molecular testing of patients with known or suspected lung cancer. Respir. Int. Rev. Thorac. Dis. 2014, 88, 500–517. [Google Scholar] [CrossRef]
- Buttner, R.; Wolf, J.; Kron, A. [The national Network Genomic Medicine (nNGM): Model for innovative diagnostics and therapy of lung cancer within a public healthcare system]. Der Pathol. 2019, 40, 276–280. [Google Scholar] [CrossRef]
- National Network Genomic Medicine (nNGM). Centralized Testing—Decentral Treatment. Available online: https://www.nngm.de/en/ (accessed on 25 June 2020).
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 2004, 350, 2129–2139. [Google Scholar] [CrossRef]
- Paez, J.G.; Janne, P.A.; Lee, J.C.; Tracy, S.; Greulich, H.; Gabriel, S.; Herman, P.; Kaye, F.J.; Lindeman, N.; Boggon, T.J.; et al. EGFR mutations in lung cancer: Correlation with clinical response to gefitinib therapy. Science (N. Y.) 2004, 304, 1497–1500. [Google Scholar] [CrossRef]
- Kobayashi, T.; Koizumi, T.; Agatsuma, T.; Yasuo, M.; Tsushima, K.; Kubo, K.; Eda, S.; Kuraishi, H.; Koyama, S.; Hachiya, T.; et al. A phase II trial of erlotinib in patients with EGFR wild-type advanced non-small-cell lung cancer. Cancer Chemother. Pharmacol. 2012, 69, 1241–1246. [Google Scholar] [CrossRef]
- Tsunoda, A.; Morikawa, K.; Inoue, T.; Miyazawa, T.; Hoshikawa, M.; Takagi, M.; Mineshita, M. A prospective observational study to assess PD-L1 expression in small biopsy samples for non-small-cell lung cancer. BMC Cancer 2019, 19, 546. [Google Scholar] [CrossRef]
- Naito, J.; Toyoda, T.; Nakajima, T.; Fujiwara, T.; Iwasawa, S.; Suzuki, H.; Takiguchi, Y.; Yoshino, I. A Repeated Biopsy by EBUS-TBNA Contributed to the Selection of an Appropriate Therapeutic Regimen for a Lung Cancer Patient. J. Bronchol. Interv. Pulmonol. 2019, 26, 129–131. [Google Scholar] [CrossRef]
- Wisniewski, P.; Glogowski, M.; Olszewski, W. Assessment of diagnostic value of cytological examination in lung carcinoma in own material obtained by EBUS and EUS methods. Pol. J. Pathol. Off. J. Pol. Soc. Pathol. 2018, 69, 278–284. [Google Scholar] [CrossRef] [PubMed]
- Raad, S.; Hanna, N.; Jalal, S.; Bendaly, E.; Zhang, C.; Nuguru, S.; Oueini, H.; Diab, K. Endobronchial Ultrasound-Guided Transbronchial Needle Aspiration Use for Subclassification and Genotyping of Lung Non-Small-Cell Carcinoma. South. Med. J. 2018, 111, 484–488. [Google Scholar] [CrossRef]
- Goag, E.K.; Lee, J.M.; Chung, K.S.; Kim, S.Y.; Leem, A.Y.; Song, J.H.; Jung, J.Y.; Park, M.S.; Chang, Y.S.; Kim, Y.S.; et al. Usefulness of Bronchoscopic Rebiopsy of Non-Small Cell Lung Cancer with Acquired Resistance to Epidermal Growth Factor Receptor-Tyrosine Kinase Inhibitor. J. Cancer 2018, 9, 1113–1120. [Google Scholar] [CrossRef] [PubMed]
- Stoy, S.P.; Segal, J.P.; Mueller, J.; Furtado, L.V.; Vokes, E.E.; Patel, J.D.; Murgu, S. Feasibility of Endobronchial Ultrasound-guided Transbronchial Needle Aspiration Cytology Specimens for Next Generation Sequencing in Non-small-cell Lung Cancer. Clin. Lung Cancer 2018, 19, 230–238.e232. [Google Scholar] [CrossRef]
- Bonifazi, M.; Tramacere, I.; Zuccatosta, L.; Mei, F.; Sediari, M.; Paonessa, M.C.; Gasparini, S. Conventional versus Ultrasound-Guided Transbronchial Needle Aspiration for the Diagnosis of Hilar/Mediastinal Lymph Adenopathies: A Randomized Controlled Trial. Respir. Int. Rev. Thorac. Dis. 2017, 94, 216–223. [Google Scholar] [CrossRef]
- Kirita, K.; Izumo, T.; Matsumoto, Y.; Hiraishi, Y.; Tsuchida, T. Bronchoscopic Re-biopsy for Mutational Analysis of Non-small Cell Lung Cancer. Lung 2016, 194, 371–378. [Google Scholar] [CrossRef]
- Izumo, T.; Matsumoto, Y.; Chavez, C.; Tsuchida, T. Re-biopsy by endobronchial ultrasound procedures for mutation analysis of non-small cell lung cancer after EGFR tyrosine kinase inhibitor treatment. BMC Pulm. Med. 2016, 16, 106. [Google Scholar] [CrossRef]
- Rooper, L.M.; Nikolskaia, O.; Carter, J.; Ning, Y.; Lin, M.T.; Maleki, Z. A single EBUS-TBNA procedure can support a large panel of immunohistochemical stains, specific diagnostic subtyping, and multiple gene analyses in the majority of non-small cell lung cancer cases. Hum. Pathol. 2016, 51, 139–145. [Google Scholar] [CrossRef]
- Casadio, C.; Guarize, J.; Donghi, S.; Di Tonno, C.; Fumagalli, C.; Vacirca, D.; Dell’Orto, P.; De Marinis, F.; Spaggiari, L.; Viale, G.; et al. Molecular Testing for Targeted Therapy in Advanced Non-Small Cell Lung Cancer: Suitability of Endobronchial Ultrasound Transbronchial Needle Aspiration. Am. J. Clin. Pathol. 2015, 144, 629–634. [Google Scholar] [CrossRef]
- Oki, M.; Saka, H.; Ando, M.; Kitagawa, C.; Kogure, Y.; Seki, Y. Endoscopic ultrasound-guided fine needle aspiration and endobronchial ultrasound-guided transbronchial needle aspiration: Are two better than one in mediastinal staging of non-small cell lung cancer? J. Thorac. Cardiovasc. Surg. 2014, 148, 1169–1177. [Google Scholar] [CrossRef][Green Version]
- Navani, N.; Brown, J.M.; Nankivell, M.; Woolhouse, I.; Harrison, R.N.; Jeebun, V.; Munavvar, M.; Ng, B.J.; Rassl, D.M.; Falzon, M.; et al. Suitability of endobronchial ultrasound-guided transbronchial needle aspiration specimens for subtyping and genotyping of non-small cell lung cancer: A multicenter study of 774 patients. Am. J. Respir. Crit. Care Med. 2012, 185, 1316–1322. [Google Scholar] [CrossRef]
- Schmid-Bindert, G.; Wang, Y.; Jiang, H.; Sun, H.; Henzler, T.; Wang, H.; Pilz, L.R.; Ren, S.; Zhou, C. EBUS-TBNA provides highest RNA yield for multiple biomarker testing from routinely obtained small biopsies in non-small cell lung cancer patients—A comparative study of three different minimal invasive sampling methods. PLoS ONE 2013, 8, e77948. [Google Scholar] [CrossRef]
- Navani, N.; Nankivell, M.; Lawrence, D.R.; Lock, S.; Makker, H.; Baldwin, D.R.; Stephens, R.J.; Parmar, M.K.; Spiro, S.G.; Morris, S.; et al. Lung cancer diagnosis and staging with endobronchial ultrasound-guided transbronchial needle aspiration compared with conventional approaches: An open-label, pragmatic, randomised controlled trial. Lancet. Respir. Med. 2015, 3, 282–289. [Google Scholar] [CrossRef]
- Cheng, Y.; Wang, Y.; Zhao, J.; Liu, Y.; Gao, H.; Ma, K.; Zhang, S.; Xin, H.; Liu, J.; Han, C.; et al. Real-world EGFR testing in patients with stage IIIB/IV non-small-cell lung cancer in North China: A multicenter, non-interventional study. Thorac. Cancer 2018, 9, 1461–1469. [Google Scholar] [CrossRef]
- Hong, M.H.; Kim, H.R.; Ahn, B.C.; Heo, S.J.; Kim, J.H.; Cho, B.C. Real-World Analysis of the Efficacy of Rebiopsy and EGFR Mutation Test of Tissue and Plasma Samples in Drug-Resistant Non-Small Cell Lung Cancer. Yonsei Med. J. 2019, 60, 525–534. [Google Scholar] [CrossRef] [PubMed]
- Komiya, K.; Nakashima, C.; Nakamura, T.; Hirakawa, H.; Abe, T.; Ogusu, S.; Takahashi, K.; Takeda, Y.; Egashira, Y.; Kimura, S.; et al. Current Status and Problems of T790M Detection, a Molecular Biomarker of Acquired Resistance to EGFR Tyrosine Kinase Inhibitors, with Liquid Biopsy and Re-biopsy. Anticancer Res. 2018, 38, 3559–3566. [Google Scholar] [CrossRef]
- Kim, T.O.; Oh, I.J.; Kho, B.G.; Park, H.Y.; Chang, J.S.; Park, C.K.; Shin, H.J.; Lim, J.H.; Kwon, Y.S.; Kim, Y.I.; et al. Feasibility of re-biopsy and EGFR mutation analysis in patients with non-small cell lung cancer. Thorac. Cancer 2018, 9, 856–864. [Google Scholar] [CrossRef]
- Chougule, A.; Basak, S. Epidermal growth factor receptor T790M testing in progressed lung cancer: A review of sensitive methods for analysis of tissue and liquid biopsy samples. Indian J. Cancer 2017, 54, S45–S54. [Google Scholar] [CrossRef]
- Zanwar, S.; Noronha, V.; Joshi, A.; Patil, V.M.; Chougule, A.; Kumar, R.; More, S.; Goud, S.; Janu, A.; Mahajan, A.; et al. Repeat biopsy in epidermal growth factor receptor mutation-positive nonsmall cell lung cancer: Feasibility, limitations, and clinical utility in Indian patients. Indian J. Cancer 2017, 54, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Clery, E.; Pisapia, P.; Feliciano, S.; Vigliar, E.; Marano, A.; De Luca, C.; Malapelle, U.; Troncone, G.; Bellevicine, C. There is still a role for cytology in the ‘liquid biopsy’ era. A lesson from a TKI-treated patient showing adenocarcinoma to squamous cell carcinoma transition during disease progression. J. Clin. Pathol. 2017, 70, 798–802. [Google Scholar] [CrossRef] [PubMed]
- Ehab, A.; Khairy El-Badrawy, M.; Abdelhamed Moawad, A.; El-Dosouky Abo-Shehata, M. Cryobiopsy versus forceps biopsy in endobronchial lesions, diagnostic yield and safety. Adv. Respir. Med. 2017, 85, 301–306. [Google Scholar] [CrossRef] [PubMed][Green Version]
| (A) Inclusion Criteria |
| Informed consent to the study and the study-specific procedures prior to any study intervention |
| Age ≥ 18 years |
| Patients with known NSCLC and suspected relapse after therapy |
| Patients with known NSCLC and suspected relapse after therapy |
| Bronchoscopically visible tumor |
| (B) Exclusion Criteria |
| Pre-existing malignancy other than NSCLC |
Contraindication for bronchoscopy follow the international guidelines [41,42], daily clinical practice and local regulations, and exclude:
|
| Category | Intervention for Bleeding Control |
|---|---|
| No | Self-limiting bleeding without need for any intervention for bleeding control |
| Mild | Self-limiting bleeding, manageable with suction alone and without the need for any specific intervention |
| Moderate | Non self-limiting bleeding with need for suction plus additional intervention (alone or in combination) including application of ice-cold saline or vasoconstrictors, or transient balloon tamponade, leading to a termination of bleeding |
| Severe | Non self-limiting bleeding with need for suction, plus any additional intervention (alone or in combination) and need for prolonged observation, stay in the hospital or intensive care therapy |
| Persistent/Fatal | Any persisting health impairment or death as a direct consequence of bleeding |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Haentschel, M.; Boeckeler, M.; Bonzheim, I.; Schimmele, F.; Spengler, W.; Stanzel, F.; Petermann, C.; Darwiche, K.; Hagmeyer, L.; Buettner, R.; et al. Influence of Biopsy Technique on Molecular Genetic Tumor Characterization in Non-Small Cell Lung Cancer—The Prospective, Randomized, Single-Blinded, Multicenter PROFILER Study Protocol. Diagnostics 2020, 10, 459. https://doi.org/10.3390/diagnostics10070459
Haentschel M, Boeckeler M, Bonzheim I, Schimmele F, Spengler W, Stanzel F, Petermann C, Darwiche K, Hagmeyer L, Buettner R, et al. Influence of Biopsy Technique on Molecular Genetic Tumor Characterization in Non-Small Cell Lung Cancer—The Prospective, Randomized, Single-Blinded, Multicenter PROFILER Study Protocol. Diagnostics. 2020; 10(7):459. https://doi.org/10.3390/diagnostics10070459
Chicago/Turabian StyleHaentschel, Maik, Michael Boeckeler, Irina Bonzheim, Florian Schimmele, Werner Spengler, Franz Stanzel, Christoph Petermann, Kaid Darwiche, Lars Hagmeyer, Reinhard Buettner, and et al. 2020. "Influence of Biopsy Technique on Molecular Genetic Tumor Characterization in Non-Small Cell Lung Cancer—The Prospective, Randomized, Single-Blinded, Multicenter PROFILER Study Protocol" Diagnostics 10, no. 7: 459. https://doi.org/10.3390/diagnostics10070459
APA StyleHaentschel, M., Boeckeler, M., Bonzheim, I., Schimmele, F., Spengler, W., Stanzel, F., Petermann, C., Darwiche, K., Hagmeyer, L., Buettner, R., Tiemann, M., Schildhaus, H.-U., Muche, R., Boesmueller, H., Everinghoff, F., Mueller, R., Atique, B., Lewis, R. A., Zender, L., ... Hetzel, J. (2020). Influence of Biopsy Technique on Molecular Genetic Tumor Characterization in Non-Small Cell Lung Cancer—The Prospective, Randomized, Single-Blinded, Multicenter PROFILER Study Protocol. Diagnostics, 10(7), 459. https://doi.org/10.3390/diagnostics10070459







