COVID-19 Serological Tests: How Well Do They Actually Perform?
Abstract
1. Background
2. COVID-19 Diagnostic Tests
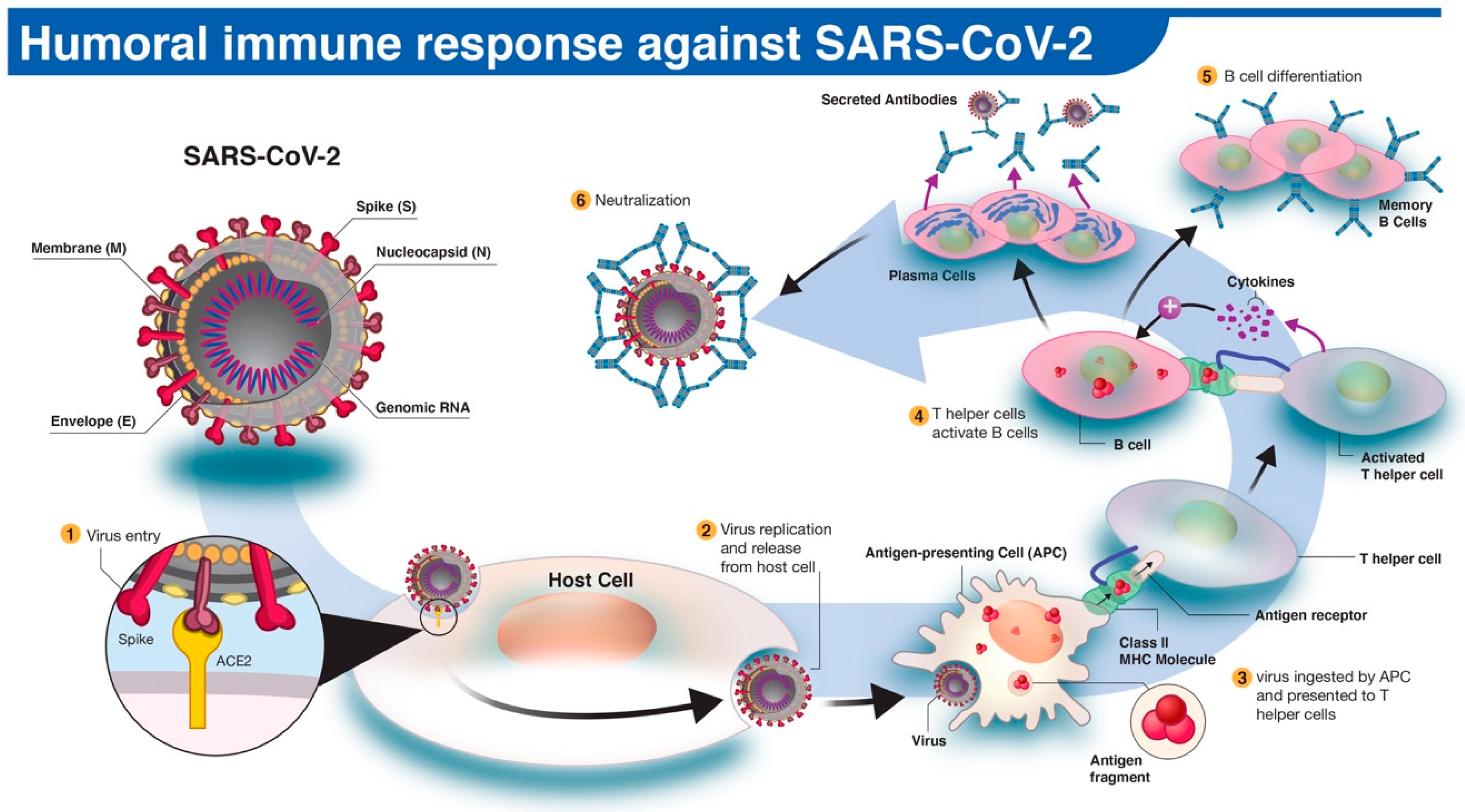
3. Humoral Immune Response to SARS-COV-2
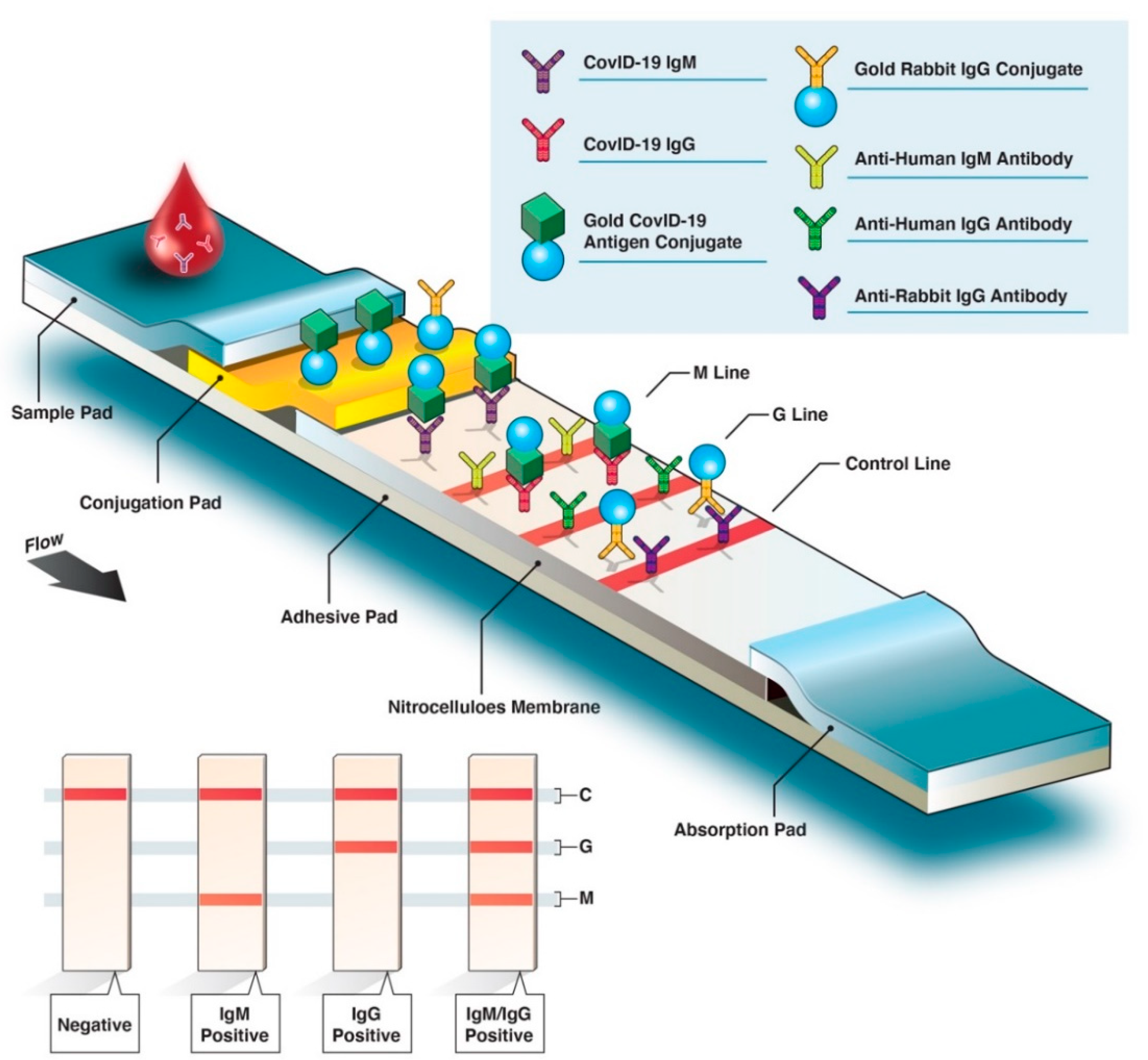
4. Serological Antibody Test
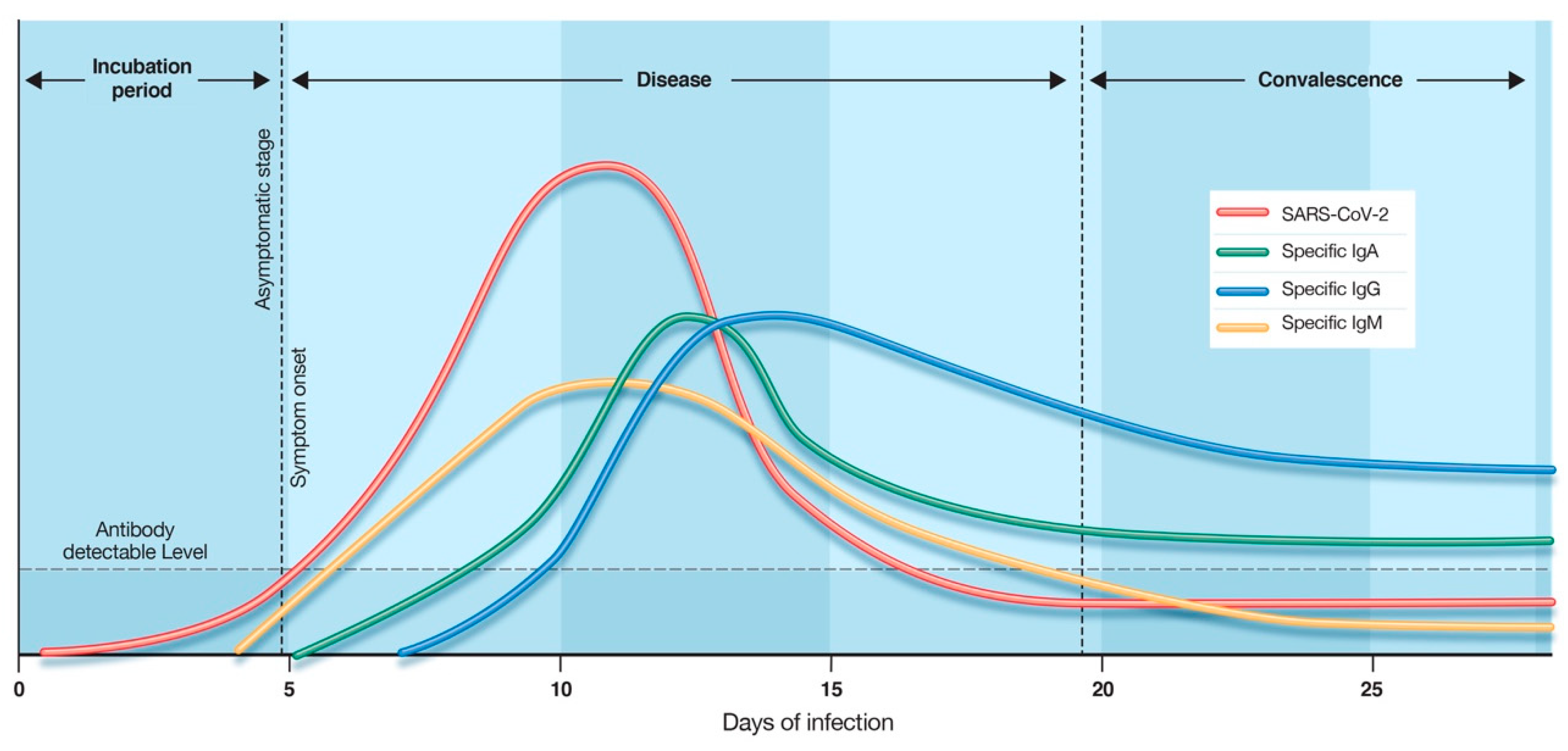
5. Time Kinetics of Antibody Response in COVID-19
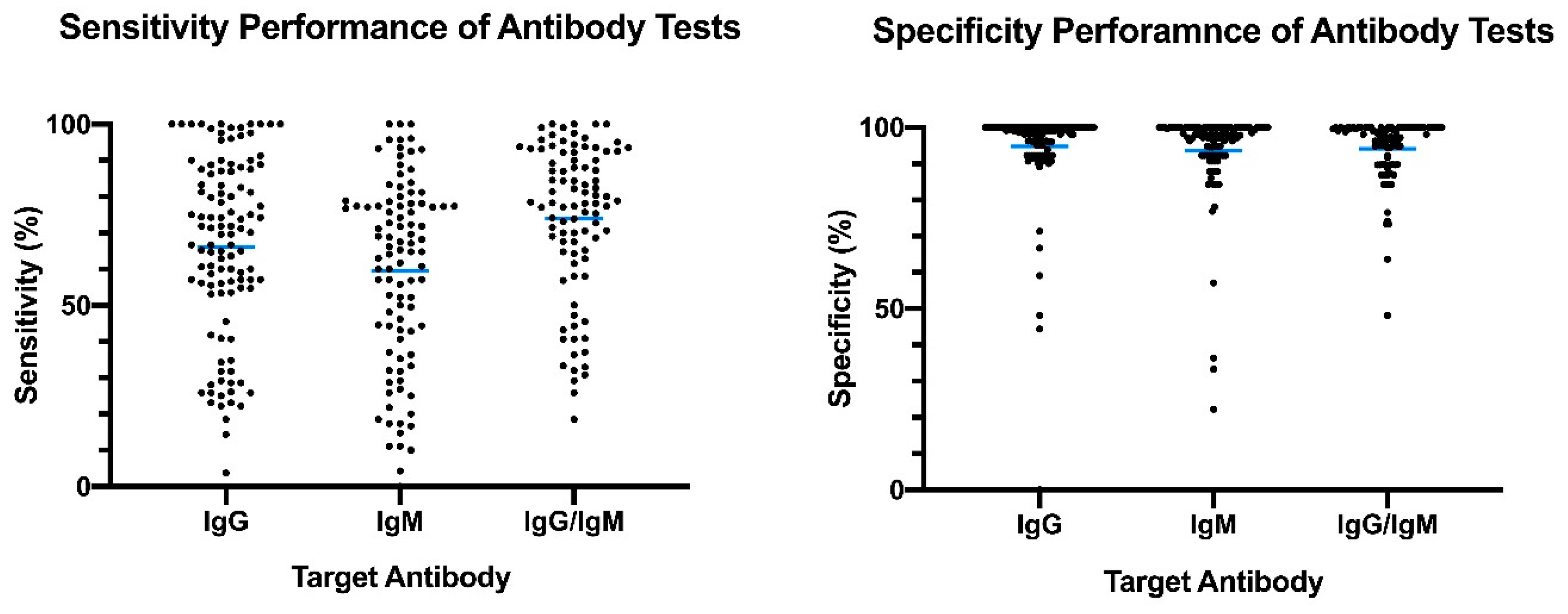
6. Serological Test Performance
7. Seroprevalence of SARS-COV-2-Specific Antibodies
8. Concluding Remarks
- Which serological tests can identify SARS-CoV-2-neutralizing antibodies?
- Is there cross-reactivity between neutralizing antibodies and other coronaviruses?
- Which SARS-CoV-2 antigens are optimal for the detection of neutralizing antibodies?
- What is the correlation between SARS-CoV-2-specific antibodies and protective immunity status?
- How long does protective immunity last in recovered patients? Are individuals susceptible to reinfection with SARS-CoV-2?
- Is humoral antibody response the best indicator for protective immunity, or are there other immune-cell-based mechanisms?
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| COVID-19 | Coronavirus Disease 2019 |
| SARS | Severe Acute Respiratory Syndrome |
| SARS-CoV | Severe Acute Respiratory Syndrome Coronavirus |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome Coronavirus 2 |
| MERS | Middle East Respiratory Syndrome |
| BAL | Bronchoalveolar Lavage |
| EUA | Emergency Use Authorization |
| RT-PCR | Reverse Transcriptase Polymerase Chain Reaction |
| S | SARS-CoV-2 Spike Protein |
| E | SARS-CoV-2 Envelope Protein |
| M | SARS-CoV-2 Membrane Protein |
| NP | SARS-CoV-2 Nucleocapsid Protein |
| RBD | Receptor-Binding Domain |
| ELISA | Enzyme-Linked Immunosorbent Assay |
| RDT | Rapid Diagnostic Test |
| CLIA | Chemiluminescence Immunoassay |
| LFIA | Lateral Flow Immunoassay |
| PoC | Point of Care |
| DABA | Double-Antigen-Bridging Assay |
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- World Health Organization COVID-19 Situation Report 127. Available online: https://www.who.int/docs/default-source/coronaviruse/situation-reports (accessed on 27 May 2020).
- Wang, Y.; Zhang, D.; Du, G.; Du, R.; Zhao, J.; Jin, Y.; Fu, S.; Gao, L.; Cheng, Z.; Lu, Q.; et al. Remdesivir in adults with severe COVID-19: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet 2020, 395, 1569–1578. [Google Scholar] [CrossRef]
- Grein, J.; Ohmagari, N.; Shin, D.; Diaz, G.; Asperges, E.; Castagna, A.; Feldt, T.; Green, G.; Green, M.L.; Lescure, F.X.; et al. Compassionate Use of Remdesivir for Patients with Severe Covid-19. N. Engl. J. Med. 2020, 382, 2327–2336. [Google Scholar] [CrossRef]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef]
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.Y.; Chen, L.; Wang, M. Presumed Asymptomatic Carrier Transmission of COVID-19. JAMA 2020. [Google Scholar] [CrossRef]
- Winter, A.K.; Hegde, S.T. The important role of serology for COVID-19 control. Lancet Infect. Dis 2020. [Google Scholar] [CrossRef]
- Janeway, C.A. Immunobiology, 5th edition: The Immune System in Health and Disease; Garland Publishing: New York, NY, USA, 2001. [Google Scholar]
- Azkur, A.K.; Akdis, M.; Azkur, D.; Sokolowska, M.; van de Veen, W.; Bruggen, M.C.; O’Mahony, L.; Gao, Y.; Nadeau, K.; Akdis, C.A. Immune response to SARS-CoV-2 and mechanisms of immunopathological changes in COVID-19. Allergy 2020. [Google Scholar] [CrossRef]
- Yu, H.Q.; Sun, B.Q.; Fang, Z.F.; Zhao, J.C.; Liu, X.Y.; Li, Y.M.; Sun, X.Z.; Liang, H.F.; Zhong, B.; Huang, Z.F.; et al. Distinct features of SARS-CoV-2-specific IgA response in COVID-19 patients. Eur. Respir. J. 2020. [Google Scholar] [CrossRef]
- Parks, J.M.; Smith, J.C. How to Discover Antiviral Drugs Quickly. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cong, Y.; Ulasli, M.; Schepers, H.; Mauthe, M.; V’Kovski, P.; Kriegenburg, F.; Thiel, V.; de Haan, C.A.M.; Reggiori, F. Nucleocapsid Protein Recruitment to Replication-Transcription Complexes Plays a Crucial Role in Coronaviral Life Cycle. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [PubMed]
- Ni, L.; Ye, F.; Cheng, M.L.; Feng, Y.; Deng, Y.Q.; Zhao, H.; Wei, P.; Ge, J.; Gou, M.; Li, X.; et al. Detection of SARS-CoV-2-Specific Humoral and Cellular Immunity in COVID-19 Convalescent Individuals. Immunity 2020. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Hillyer, C.; Du, L. Neutralizing Antibodies against SARS-CoV-2 and Other Human Coronaviruses: (Trends in Immunology 41, 355-359; 2020). Trends Immunol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Vashist, S.K. In Vitro Diagnostic Assays for COVID-19: Recent Advances and Emerging Trends. Diagnostics 2020, 10, 202. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Chang, L.; Wang, L. Laboratory testing of SARS-CoV, MERS-CoV, and SARS-CoV-2 (2019-nCoV): Current status, challenges, and countermeasures. Rev. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- To, K.K.; Tsang, O.T.; Leung, W.S.; Tam, A.R.; Wu, T.C.; Lung, D.C.; Yip, C.C.; Cai, J.P.; Chan, J.M.; Chik, T.S.; et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet Infect. Dis. 2020, 20, 565–574. [Google Scholar] [CrossRef]
- Wolfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Muller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020. [Google Scholar] [CrossRef]
- Guo, L.; Ren, L.; Yang, S.; Xiao, M.; Chang, D.; Yang, F.; Dela Cruz, C.S.; Wang, Y.; Wu, C.; Xiao, Y.; et al. Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19). Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Tan, W.L.Y.; Zhang, J. Viral Kinetics and Antibody Responses in Patients with COVID-19. medRxiv (preprint) 2020. [Google Scholar] [CrossRef]
- Mandrekar, J.N. Simple statistical measures for diagnostic accuracy assessment. J. Thorac. Oncol. 2010, 5, 763–764. [Google Scholar] [CrossRef]
- U.S. Food & Drug Administratin. Independent Evaluations of COVID-19 Serological Tests. Available online: https://open.fda.gov/apis/device/covid19serology/ (accessed on 6 June 2020).
- FINDDx-SARS-COV-2 DIAGNOSTICS: PERFORMANCE DATA. Available online: https://www.finddx.org/covid-19/dx-data/ (accessed on 20 May 2020).
- Ria Lassaunière, A.F.; Harboe, Z.B.; Nielsen, A.C.Y.; Fomsgaard, A.; Krogfelt, K.A.; Charlotte S. Jørgensen, C.S. Evaluation of nine commercial SARS-CoV-2 immunoassays. medRxiv (preprint) 2020. [Google Scholar] [CrossRef]
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; et al. Evaluation of Nucleocapsid and Spike Protein-Based Enzyme-Linked Immunosorbent Assays for Detecting Antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58. [Google Scholar] [CrossRef] [PubMed]
- Lou, B. Serology characteristics of SARS-CoV-2 infection since the exposure and post symptoms onset. medRxiv (preprint) 2020. [Google Scholar] [CrossRef]
- Zhang, J.; Li, N.; Liu, Y.; Ye, R.; Qin, X.; Zheng, R. Serological detection of 2019-nCoV respond to the epidemic: A useful complement to nucleic acid testing. medRxiv (preprint) 2020. [Google Scholar] [CrossRef]
- Bryan, A.; Pepper, G.; Wener, M.H.; Fink, S.L.; Morishima, C.; Chaudhary, A.; Jerome, K.R.; Mathias, P.C.; Greninger, A.L. Performance Characteristics of the Abbott Architect SARS-CoV-2 IgG Assay and Seroprevalence in Boise, Idaho. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, M.C.; Freedberg, K.A.; Hyle, E.P.; Paltiel, A.D. Waiting for Certainty on Covid-19 Antibody Tests-At What Cost? N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Sood, N.; Simon, P.; Ebner, P.; Eichner, D.; Reynolds, J.; Bendavid, E.; Bhattacharya, J. Seroprevalence of SARS-CoV-2-Specific Antibodies Among Adults in Los Angeles County, California, on April 10-11, 2020. JAMA 2020. [Google Scholar] [CrossRef]
- New York State Governor Website. Available online: https://www.governor.ny.gov/news/amid-ongoing-covid-19-pandemic-governor-cuomo-announces-results-completed-antibody-testing (accessed on 2 May 2020).
- Popovich, N.; Sanger-Katz, M. The World Is Still Far From Herd Immunity for Coronavirus. Available online: https://www.nytimes.com/interactive/2020/05/28/upshot/coronavirus-herd-immunity.html (accessed on 28 June 2020).
- Cohen, J.; Kupferschmidt, K. Countries test tactics in ‘war’ against COVID-19. Science 2020, 367, 1287–1288. [Google Scholar] [CrossRef]

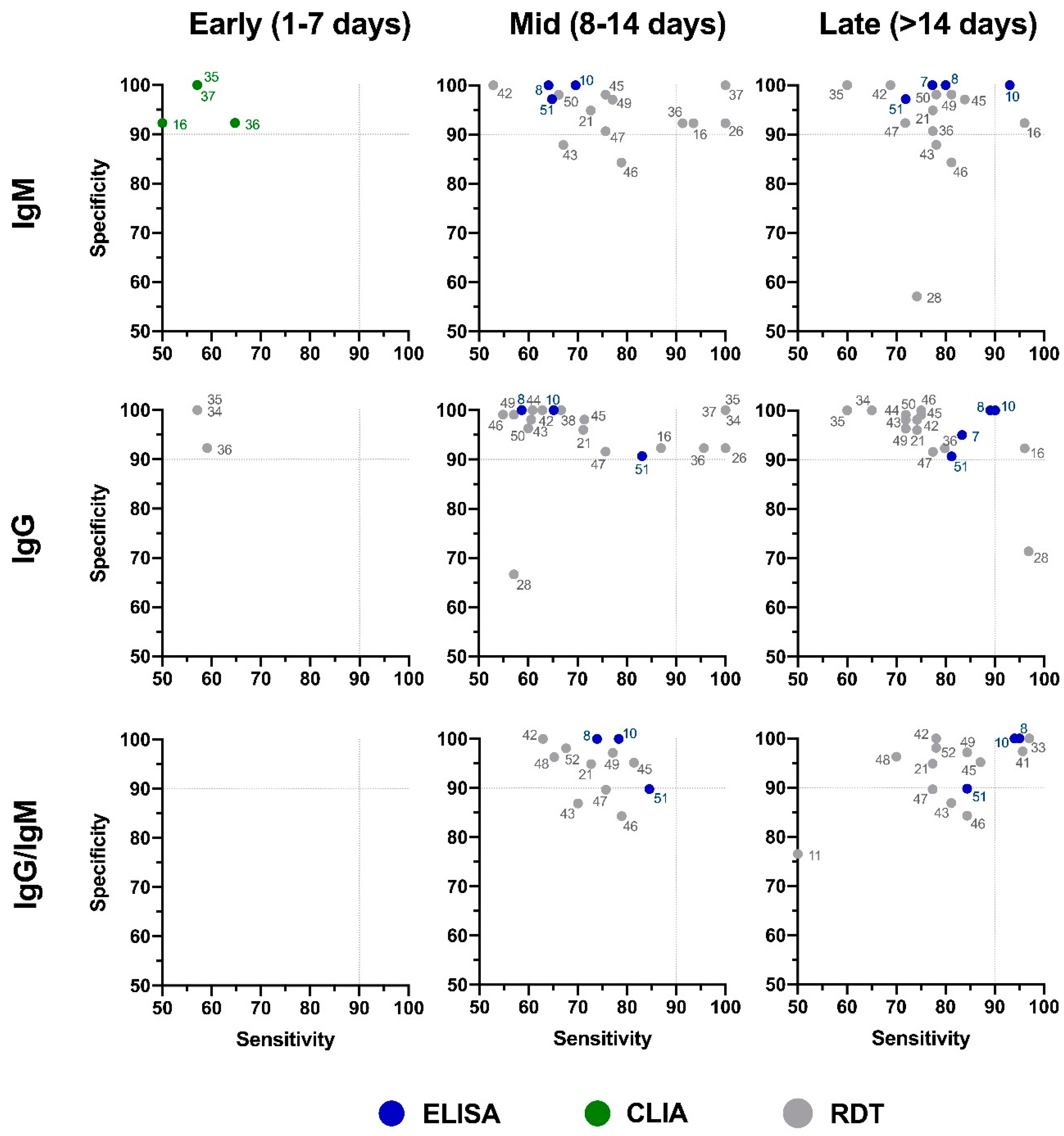
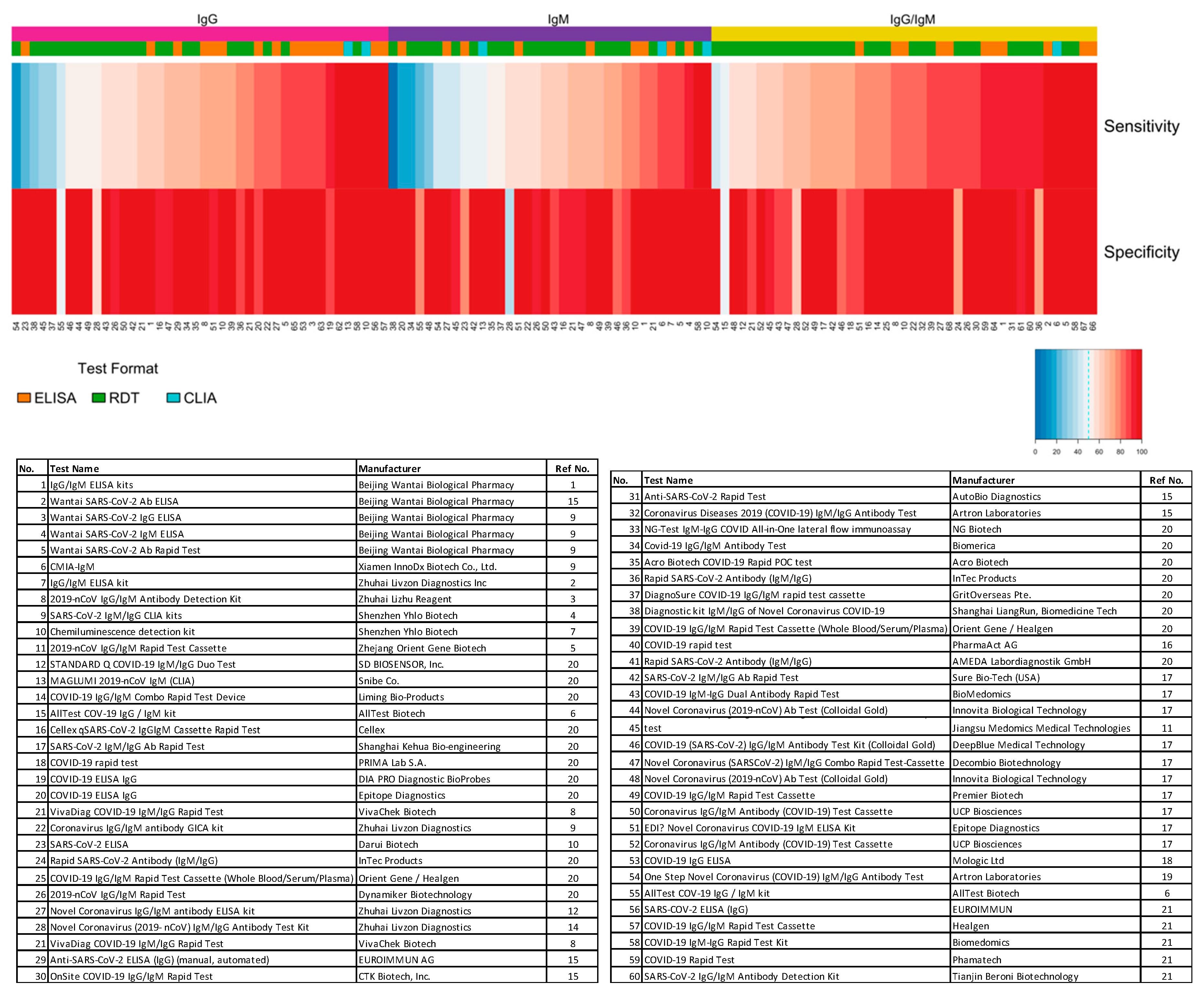
| No. | Test Name [ref] | Manufacturer | Format | Target Antibody | Disease Stage | Sensitivity (%) | Specificity (%) | Antigen |
|---|---|---|---|---|---|---|---|---|
| 2 | Wantai SARS-CoV-2 Ab ELISA [26] | Beijing Wantai Biological Pharmacy | ELISA | IgG/IgM/IgA | Overall | 95.4 | 100 | RBD |
| 8 | 2019-nCoV IgG/IgM Antibody Detection Kit [27] | Zhuhai Lizhu Reagent | ELISA | IgG/IgM | Late (>14 days) | 95 | 100 | NP |
| 6 | CMIA-Ab [28] | Xiamen InnoDx Biotech Co., Ltd. | CLIA | IgG/IgM | Overall | 96.2 | 99.3 | RBD |
| 10 | Chemiluminescen. detection kit [29] | Shenzhen Yhlo Biotech | CLIA | IgM IgG | Overall | 100 100 | 98.5 99 | n/a |
| 13 | MAGLUMI 2019-nCoV IgG [25] | Snibe Co. | CLIA | IgG | Overall | 98.8 | 95.1 | n/a |
| 61 | Architect SARS-CoV-2 IgG Assay [30] | Abbott | CLIA | IgG | Late (>14 days) | 97.2 | 100 | NP |
| 5 | Wantai SARS-CoV-2 Ab Rapid Test [28] | Beijing Wantai Biological Pharmacy | RDT | IgG/IgM/IgA | Overall | 97.5 | 95.2 | RBD |
| 33 | NG-Test IgM-IgG COVID All-in-One Lateral Flow Immunoassay [25] | NG Biotech | RDT | IgG/IgM | Late (>14 days) | 97 | 100 | n/a |
| 34 | Covid-19 IgG/IgM Antibody Test [25] | Biomerica | RDT | IgG | Mid. (8–14 days) | 100 | 100 | n/a |
| 35 | Acro Biotech COVID-19 Rapid POC Test [25] | Acro Biotech | RDT | IgG | Mid. (8–14 days) | 100 | 100 | n/a |
| 37 | DiagnoSure COVID-19 IgG/IgM Rapid Test Cassette [25] | GritOverseas Pte. | RDT | IgG IgM | Mid. (8–14 days) | 100 100 | 100 100 | n/a |
| 41 | Rapid SARS-CoV-2 Antibody (IgM/IgG) [25] | AMEDA Labordiagnostik GmbH | RDT | IgG/IgM | Late (>14 days) | 95.7 | 97.4 | n/a |
| 57 | COVID-19 IgG/IgM Rapid Test Cassette [24] | Healgen | RDT | IgM IgG | Overall | 100 96.7 | 100 97.5 | n/a |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghaffari, A.; Meurant, R.; Ardakani, A. COVID-19 Serological Tests: How Well Do They Actually Perform? Diagnostics 2020, 10, 453. https://doi.org/10.3390/diagnostics10070453
Ghaffari A, Meurant R, Ardakani A. COVID-19 Serological Tests: How Well Do They Actually Perform? Diagnostics. 2020; 10(7):453. https://doi.org/10.3390/diagnostics10070453
Chicago/Turabian StyleGhaffari, Abdi, Robyn Meurant, and Ali Ardakani. 2020. "COVID-19 Serological Tests: How Well Do They Actually Perform?" Diagnostics 10, no. 7: 453. https://doi.org/10.3390/diagnostics10070453
APA StyleGhaffari, A., Meurant, R., & Ardakani, A. (2020). COVID-19 Serological Tests: How Well Do They Actually Perform? Diagnostics, 10(7), 453. https://doi.org/10.3390/diagnostics10070453






