Uncovering Potential Roles of Differentially Expressed Genes, Upstream Regulators, and Canonical Pathways in Endometriosis Using an In Silico Genomics Approach
Abstract
:1. Introduction
Etiology
2. Materials and Methods
2.1. Data Retrieval and Sample Description
2.2. Gene Expression Analysis
2.3. Gene Ontology, Pathway, and Upstream Regulators Analysis
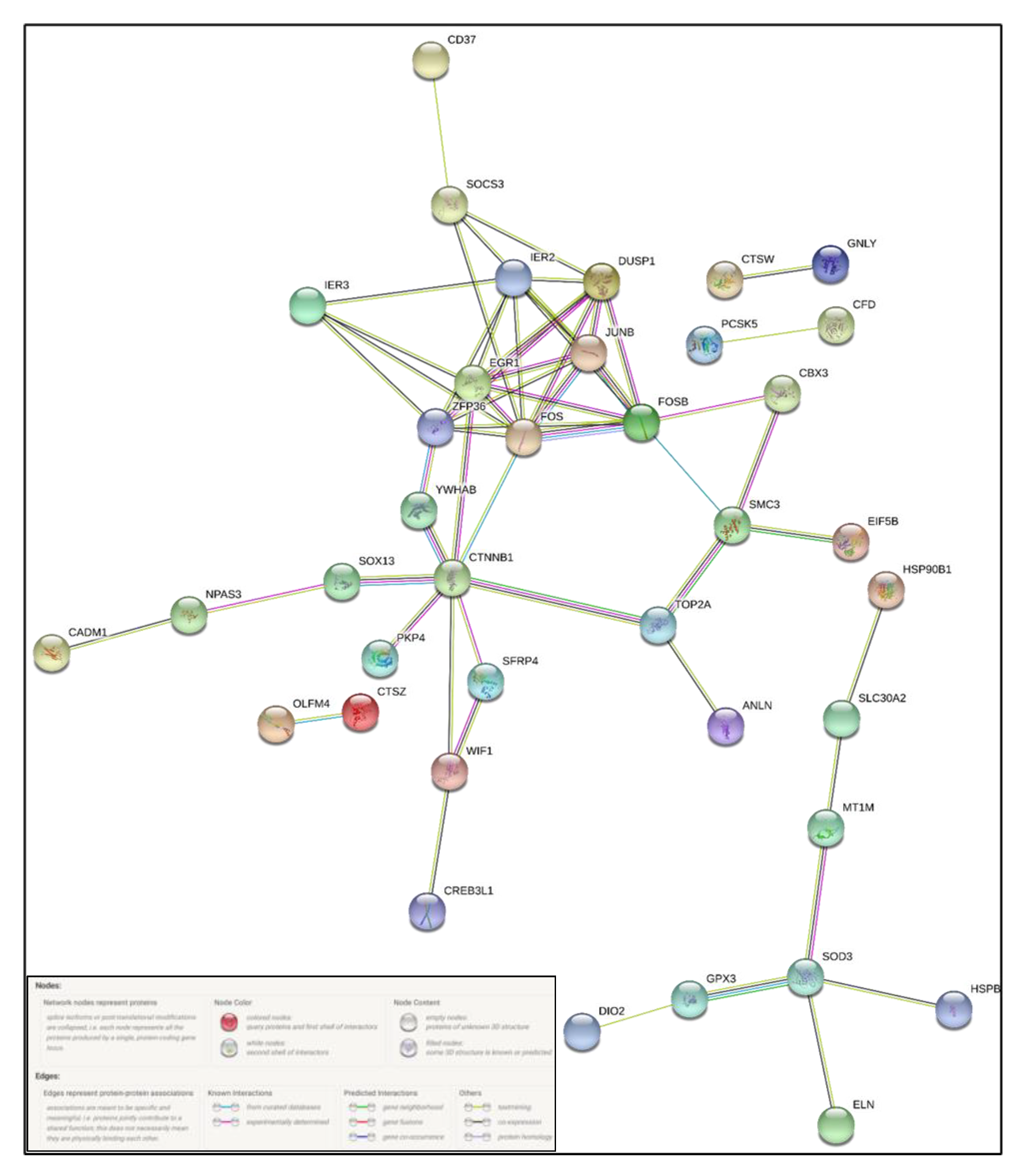
2.4. Protein–Protein Interaction Analysis
3. Results
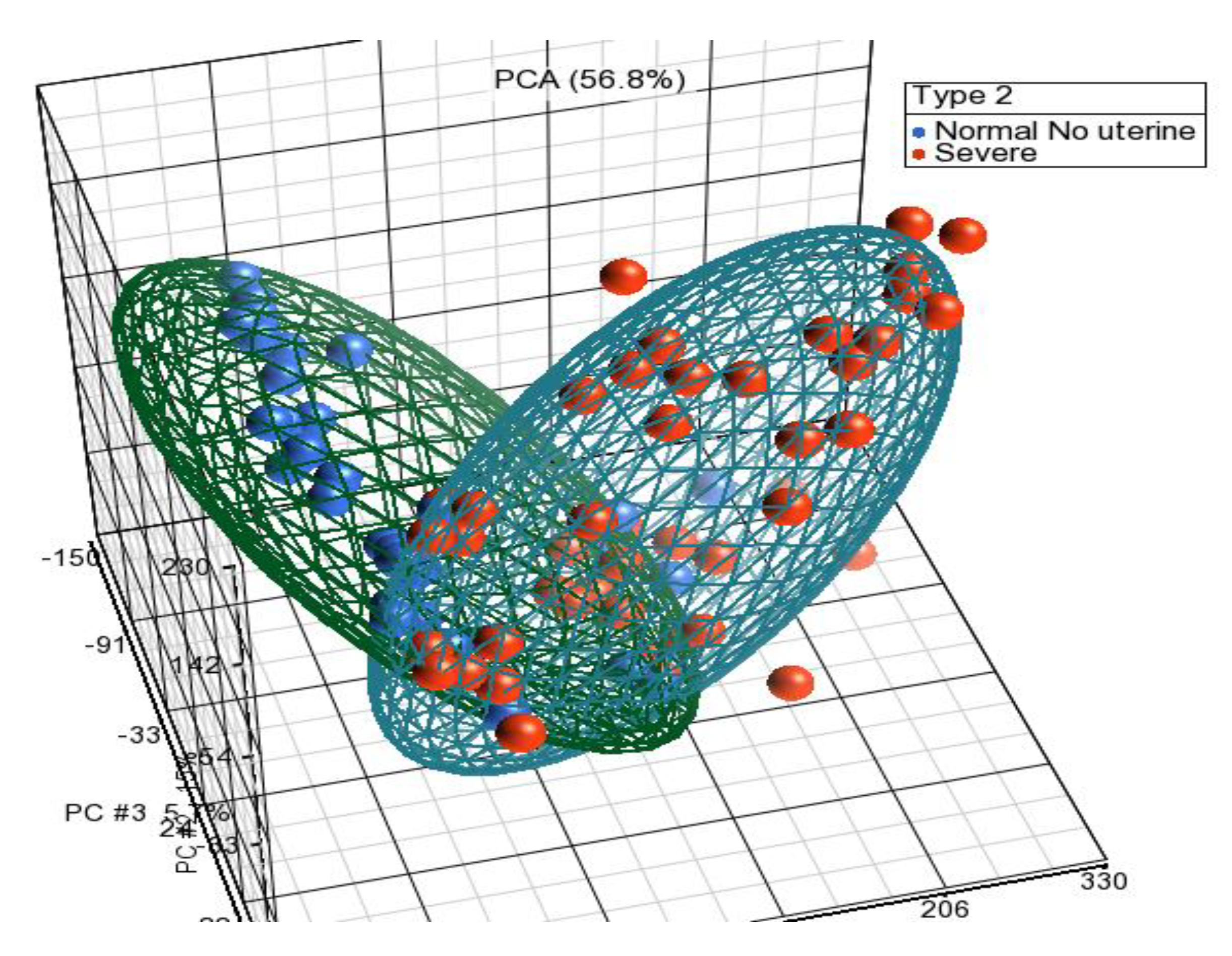
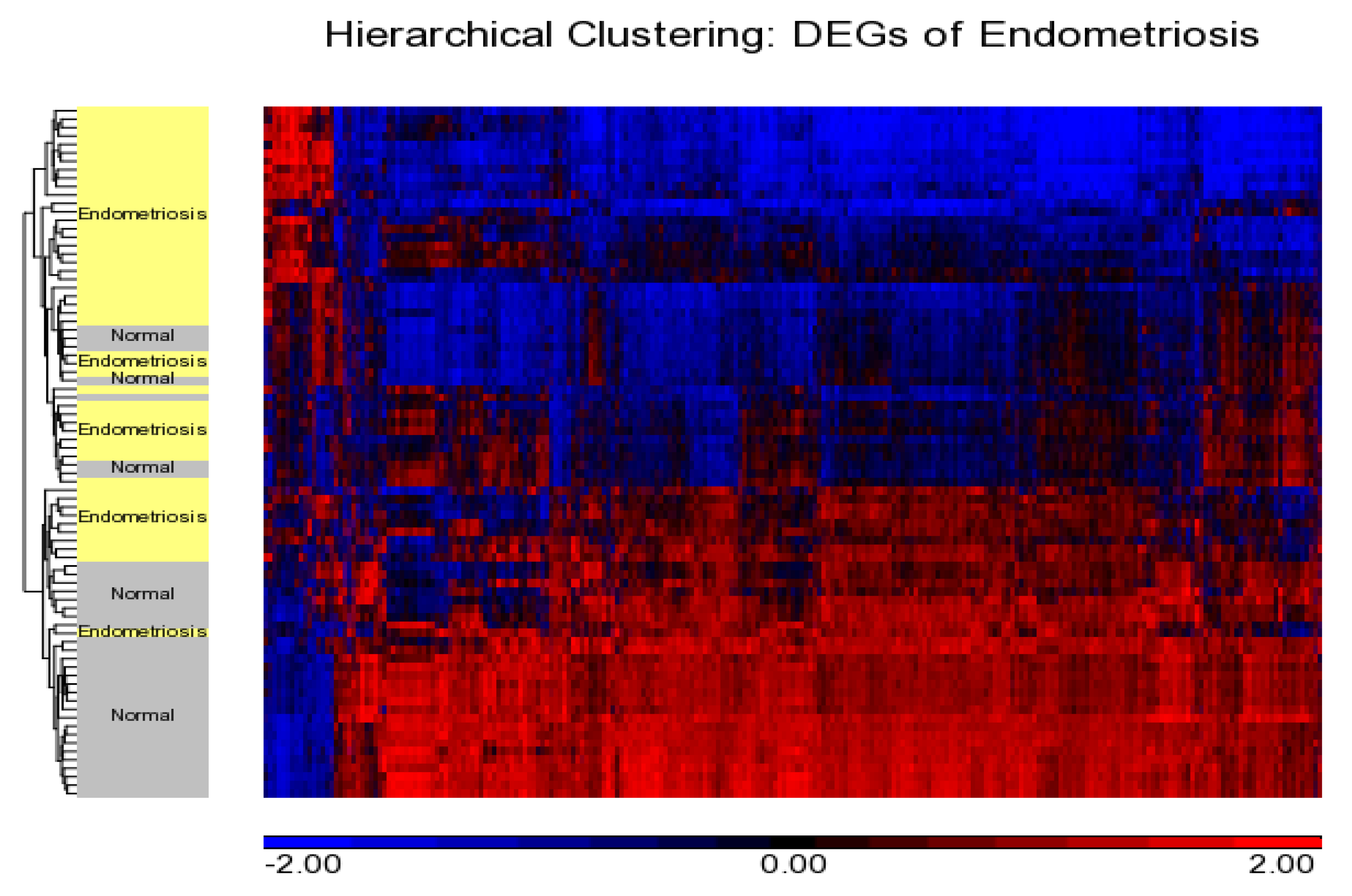
Differentially Expressed Genes from Meta-Analysis
4. Discussion
4.1. Molecular Etiology of Endometriosis
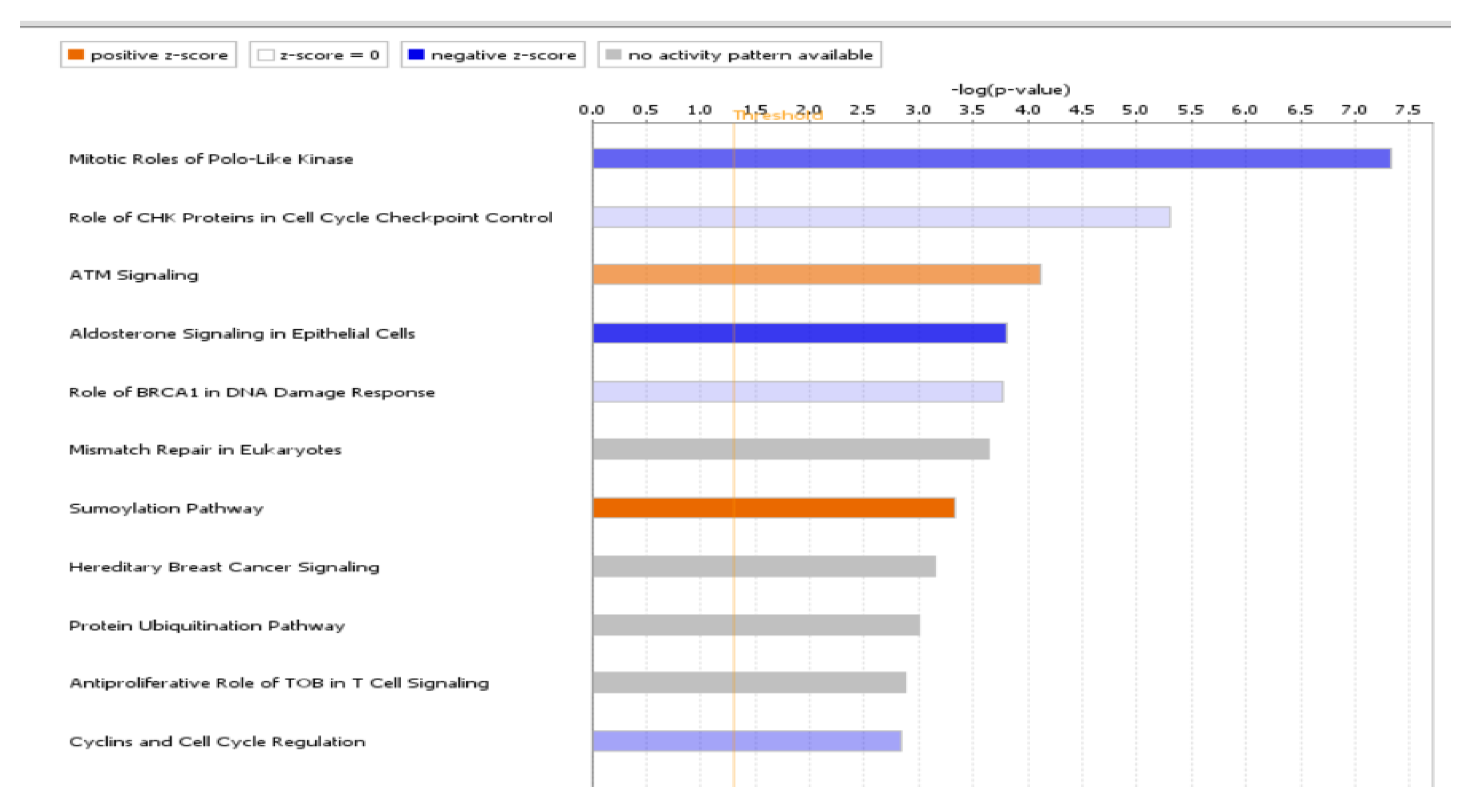
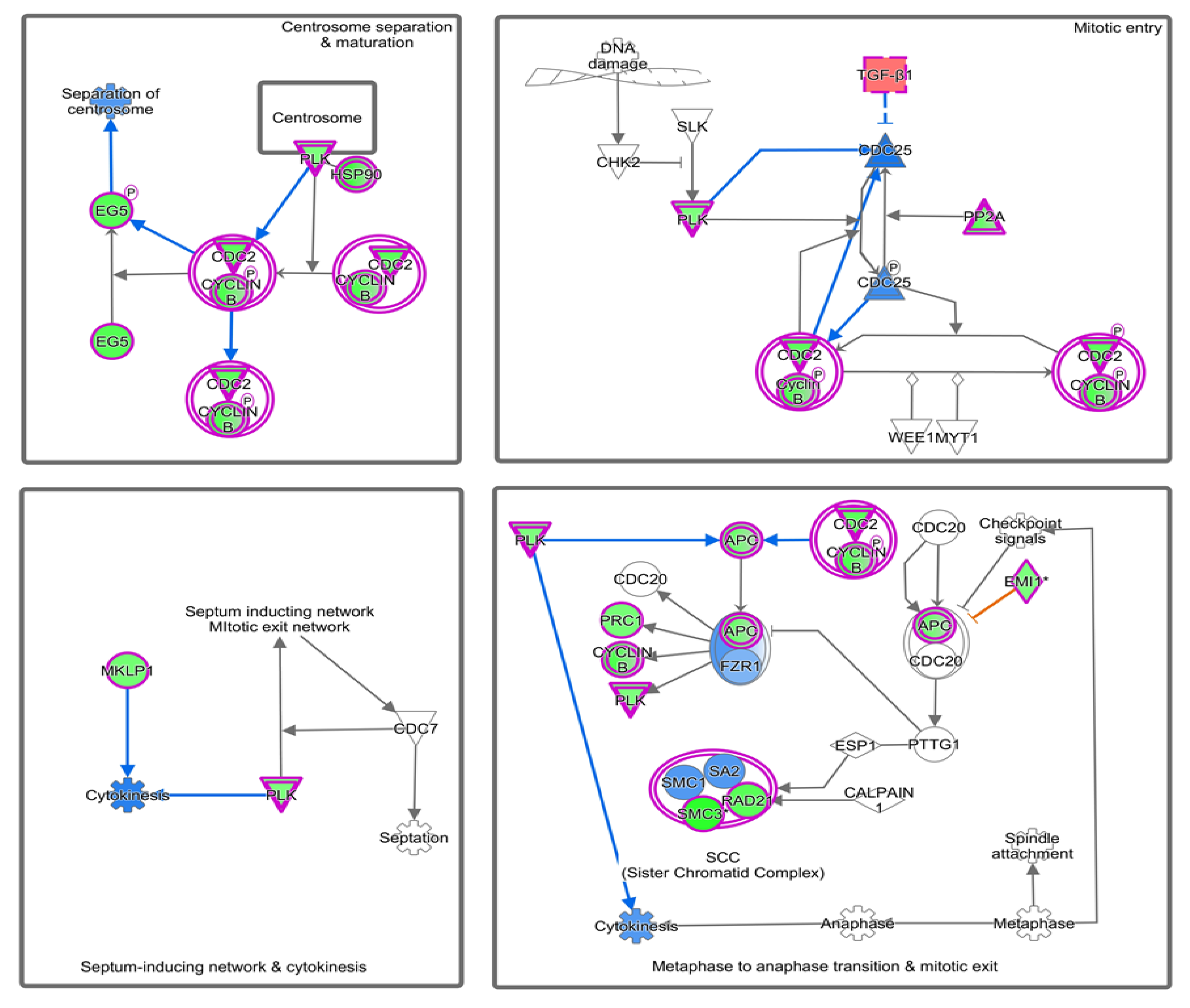
4.2. Canonical Pathways Involved in Endometriosis
4.3. Future Directions
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Cho, Y.J.; Lee, S.H.; Park, J.W.; Han, M.; Park, M.J.; Han, S.J. Dysfunctional signaling underlying endometriosis: Current state of knowledge. J. Mol. Endocrinol. 2018, 60, R97–R113. [Google Scholar] [CrossRef] [Green Version]
- Tani, H.; Sato, Y.; Ueda, M.; Miyazaki, Y.; Suginami, K.; Horie, A.; Konishi, I.; Shinomura, T. Role of Versican in the Pathogenesis of Peritoneal Endometriosis. J. Clin. Endocrinol. Metab. 2016, 101, 4349–4356. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alimi, Y.; Iwanaga, J.; Loukas, M.; Tubbs, R.S. The Clinical Anatomy of Endometriosis: A Review. Cureus 2018, 10, e3361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yeung, P., Jr. The laparoscopic management of endometriosis in patients with pelvic pain. Obstet. Gynecol. Clin. N. Am. 2014, 41, 371–383. [Google Scholar] [CrossRef] [PubMed]
- Rouzi, A.A.; Sahly, N.; Kafy, S.; Sawan, D.; Abduljabbar, H. Prevalence of endometriosis at a university hospital in Jeddah, Saudi Arabia. Clin. Exp. Obstet. Gynecol. 2015, 42, 785–786. [Google Scholar]
- Zheng, Q.M.; Mao, H.I.; Zhao, Y.J.; Zhao, J.; Wei, X.; Liu, P.S. Risk of endometrial polyps in women with endometriosis: A meta-analysis. Reprod. Biol. Endocrinol. 2015, 13, 103. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.S.; Kim, T.H.; Chung, H.H.; Song, Y.S. Risk and prognosis of ovarian cancer in women with endometriosis: A meta-analysis. Br. J. Cancer 2014, 110, 1878–1890. [Google Scholar] [CrossRef]
- Vaughan, S.; Coward, J.I.; Bast, R.C.; Berchuck, A.; Berek, J.S.; Brenton, J.D.; Coukos, G.; Crum, C.C.; Drapkin, R.; Etemadmoghadam, D.; et al. Rethinking ovarian cancer: Recommendations for improving outcomes. Nat. Rev. Cancer 2011, 11, 719–725. [Google Scholar] [CrossRef] [Green Version]
- Hughes, C.; Foster, W.; Agarwal, S. The impact of endometriosis across the lifespan of women: Foreseeable research and therapeutic prospects. Biomed. Res. Int. 2015, 2015, 158490. [Google Scholar] [CrossRef]
- Sorlie, T.; Tibshirani, R.; Parker, J.; Hastie, T.; Marron, J.S.; Nobel, A.; Deng, S.; Johnsen, H.; Pesich, R.; Geisler, S.; et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 2003, 100, 8418–8423. [Google Scholar] [CrossRef] [Green Version]
- Hedenfalk, I.; Duggan, D.; Chen, Y.; Radmacher, M.; Bittner, M.; Simon, R.; Meltzer, P.; Gusterson, B.; Esteller, M.; Kallioniemi, O.P.; et al. Gene-expression profiles in hereditary breast cancer. N. Engl. J. Med. 2001, 344, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.C.; Wooten, E.C.; Tsimelzon, A.; Hilsenbeck, S.G.; Gutierrez, M.C.; Elledge, R.; Mohsin, S.; Osborne, C.K.; Chamness, G.C.; Allred, D.C.; et al. Gene expression profiling for the prediction of therapeutic response to docetaxel in patients with breast cancer. Lancet 2003, 362, 362–369. [Google Scholar] [CrossRef]
- Borghese, B.; Zondervan, K.T.; Abrao, M.S.; Chapron, C.; Vaiman, D. Recent insights on the genetics and epigenetics of endometriosis. Clin. Genet. 2017, 91, 254–264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ponnampalam, A.P.; Weston, G.C.; Trajstman, A.C.; Susil, B.; Rogers, P.A.W. Molecular classification of human endometrial cycle stages by transcriptional profiling. Mol. Hum. Reprod. 2004, 10, 879–893. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sherwin, R.; Catalano, R.; Sharkey, A. Large-scale gene expression studies of the endometrium: What have we learnt? Reproduction 2006, 132, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Miller, L.M.; Johnson, N.P. EPHect—The Endometriosis Phenome (and Biobanking) Harmonisation Project—May be very helpful for clinicians and the women they are treating. F1000Res 2017, 6, 14. [Google Scholar] [CrossRef]
- Parazzini, F.; Vercellini, P.; Pelucchi, C. Endometriosis: Epidemiology, and Etiological Factors. In Endometriosis; Linda, C.G., Johannes, L.H.E., David, L.H., Eds.; Wiley-Blackwell: Oxford, UK, 2012; pp. 19–26. [Google Scholar] [CrossRef]
- Porpora, M.G.; Resta, S.; Fuggetta, E.; Storelli, P.; Megiorni, F.; Manganaro, L.; De Felip, E. Role of environmental organochlorinated pollutants in the development of endometriosis. Clin. Exp. Obstet. Gynecol. 2013, 40, 565–567. [Google Scholar]
- Mathur, S.; Peress, M.R.; Williamson, H.O.; Youmans, C.D.; Maney, S.A.; Garvin, A.J.; Rust, P.F.; Fudenberg, H.H. Autoimmunity to endometrium and ovary in endometriosis. Clin. Exp. Immunol. 1982, 50, 259–266. [Google Scholar] [CrossRef] [Green Version]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- Attar, E.; Bulun, S.E. Aromatase inhibitors: The next generation of therapeutics for endometriosis? Fertil. Steril. 2006, 85, 1307–1318. [Google Scholar] [CrossRef]
- Sharma, G.; Hu, C.; Staquicini, D.I.; Brigman, J.L.; Liu, M.; Mauvais-Jarvis, F.; Pasqualini, R.; Arap, W.; Arterburn, J.B.; Hathaway, H.J. Preclinical efficacy of the GPER-selective agonist G-1 in mouse models of obesity and diabetes. Sci. Transl. Med. 2020, 12. [Google Scholar] [CrossRef] [PubMed]
- Marques, C.; Silva, T.S.; Dias, M.F. Clear cell carcinoma arising from abdominal wall endometriosis - Brief report and review of the literature. Gynecol. Oncol. Rep. 2017, 20, 78–80. [Google Scholar] [CrossRef] [PubMed]
- Yıldırım, D.; Tatar, C.; Doğan, O.; Hut, A.; Dönmez, T.; Akıncı, M.; Toptaş, M.; Bayık, R.N. Post-cesarean scar endometriosis. Turk. J. Obstet. Gynecol. 2018, 15, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Sosa-Durán, E.E.; Aboharp-Hasan, Z.; Mendoza-Morales, R.C.; García-Rodríguez, F.M.; Jiménez-Villanueva, X.; Peñavera-Hernández, J.R. Clear cell adenocarcinoma arising from abdominal wall endometriosis. Cirugía Cir. 2016, 84, 245–249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Milde-Langosch, K. The Fos family of transcription factors and their role in tumourigenesis. Eur. J. Cancer 2005, 41, 2449–2461. [Google Scholar] [CrossRef] [PubMed]
- Nemos, C.; Delage-Mourroux, R.; Jouvenot, M.; Adami, P. Onset of direct 17-beta estradiol effects on proliferation and c-fos expression during oncogenesis of endometrial glandular epithelial cells. Exp. Cell Res. 2004, 296, 109–122. [Google Scholar] [CrossRef]
- Mahner, S.; Baasch, C.; Schwarz, J.; Hein, S.; Wölber, L.; Jänicke, F.; Milde-Langosch, K. C-Fos expression is a molecular predictor of progression and survival in epithelial ovarian carcinoma. Br. J. Cancer 2008, 99, 1269–1275. [Google Scholar] [CrossRef]
- Krämer, A.; Green, J.; Pollard, J., Jr.; Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 2014, 30, 523–530. [Google Scholar] [CrossRef]
- Gilmore, T.D.; Gerondakis, S. The c-Rel Transcription Factor in Development and Disease. Genes Cancer 2011, 2, 695–711. [Google Scholar] [CrossRef] [Green Version]
- Pallares, J.; Martínez-Guitarte, J.L.; Dolcet, X.; Llobet, D.; Rue, M.; Palacios, J.; Prat, J.; Matias-Guiu, X. Abnormalities in the NF-κB family and related proteins in endometrial carcinoma. J. Pathol. 2004, 204, 569–577. [Google Scholar] [CrossRef]
- Lecarpentier, Y.; Schussler, O.; Hébert, J.-L.; Vallée, A. Multiple Targets of the Canonical WNT/β-Catenin Signaling in Cancers. Front. Oncol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Morin, P.J. beta-catenin signaling and cancer. Bioessays News Rev. Mol. Cell. Dev. Biol. 1999, 21, 1021–1030. [Google Scholar] [CrossRef]
- Chen, Y.; Song, W. Wnt/catenin β1/microRNA 183 predicts recurrence and prognosis of patients with colorectal cancer. Oncol Lett. 2018, 15, 4451–4456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsuzaki, S.; Darcha, C. Involvement of the Wnt/beta-Catenin Signaling Pathway in the Cellular and Molecular Mechanisms of Fibrosis in Endometriosis. PLoS ONE 2013, 8, e76808. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.-M.; Yang, W.-X. Epithelial-to-mesenchymal transition in the development of endometriosis. Oncotarget 2017, 8, 41679–41689. [Google Scholar] [CrossRef] [Green Version]
- Kischel, P.; Waltregny, D.; Dumont, B.; Turtoi, A.; Greffe, Y.; Kirsch, S.; De Pauw, E.; Castronovo, V. Versican overexpression in human breast cancer lesions: Known and new isoforms for stromal tumor targeting. Int. J. Cancer 2010, 126, 640–650. [Google Scholar] [CrossRef]
- Du, W.W.; Yang, W.; Yee, A.J. Roles of versican in cancer biology—Tumorigenesis, progression and metastasis. Histol. Histopathol. 2013, 28, 701–713. [Google Scholar] [CrossRef]
- Andersson-Sjöland, A.; Hallgren, O.; Rolandsson, S.; Weitoft, M.; Tykesson, E.; Larsson-Callerfelt, A.K.; Rydell-Törmänen, K.; Bjermer, L.; Malmström, A.; Karlsson, J.C.; et al. Versican in inflammation and tissue remodeling: The impact on lung disorders. Glycobiology 2015, 25, 243–251. [Google Scholar] [CrossRef] [Green Version]
- Miyazaki, Y.; Horie, A.; Tani, H.; Ueda, M.; Okunomiya, A.; Suginami, K.; Kondoh, E.; Baba, T.; Konishi, I.; Shinomura, T.; et al. Versican V1 in human endometrial epithelial cells promotes BeWo spheroid adhesion in vitro. Reproduction 2019, 157, 53. [Google Scholar] [CrossRef] [Green Version]
- Kodama, J.; Kusumoto, T.; Seki, N.; Matsuo, T.; Ojima, Y.; Nakamura, K.; Hongo, A.; Hiramatsu, Y. Prognostic significance of stromal versican expression in human endometrial cancer. Ann. Oncol. 2007, 18, 269–274. [Google Scholar] [CrossRef]
- AlKusayer, G.M.; Pon, J.R.; Peng, B.; Klausen, C.; Lisonkova, S.; Kinloch, M.; Yong, P.; Muhammad, E.M.S.; Leung, P.C.K.; Bedaiwy, M.A. HOXB4 Immunoreactivity in Endometrial Tissues From Women With or Without Endometriosis. Reprod. Sci. 2017. [Google Scholar] [CrossRef]
- Ozkan, Z.S.; Cilgin, H.; Simsek, M.; Cobanoglu, B.; Ilhan, N. Investigation of apelin expression in endometriosis. J. Reprod. Infertil. 2013, 14, 50–55. [Google Scholar]
- Luo, Q.; Ning, W.; Wu, Y.; Zhu, X.; Jin, F.; Sheng, J.; Huang, H. Altered expression of interleukin-18 in the ectopic and eutopic endometrium of women with endometriosis. J. Reprod. Immunol. 2006, 72, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.K.; Bulmer, J.N.; Searle, R.F. Immunohistochemical characterization of proliferation, oestrogen receptor and progesterone receptor expression in endometriosis: Comparison of eutopic and ectopic endometrium with normal cycling endometrium. Hum. Reprod. 1995, 10, 3272–3279. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Yang, X.L.; Sun, X.; Peng, C.; Li, X.; Wang, M.; Zhou, Y.F. [Expression of integrin beta3 and osteopontin in endometrium of patients with adenomyosis]. Zhonghua Fu Chan Ke Za Zhi 2009, 44, 354–358. [Google Scholar] [PubMed]
- Joshi, N.R.; Miyadahira, E.H.; Afshar, Y.; Jeong, J.W.; Young, S.L.; Lessey, B.A.; Serafini, P.C.; Fazleabas, A.T. Progesterone Resistance in Endometriosis Is Modulated by the Altered Expression of MicroRNA-29c and FKBP4. J. Clin. Endocrinol. Metab. 2017, 102, 141–149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hapangama, D.K.; Raju, R.S.; Valentijn, A.J.; Barraclough, D.; Hart, A.; Turner, M.A.; Platt-Higgins, A.; Barraclough, R.; Rudland, P.S. Aberrant expression of metastasis-inducing proteins in ectopic and matched eutopic endometrium of women with endometriosis: Implications for the pathogenesis of endometriosis. Hum. Reprod. 2012, 27, 394–407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bulun, S.E.; Yang, S.; Fang, Z.; Gurates, B.; Tamura, M.; Zhou, J.; Sebastian, S. Role of aromatase in endometrial disease. J. Steroid Biochem. Mol. Biol. 2001, 79, 19–25. [Google Scholar] [CrossRef]
- Ebert, A.D.; Bartley, J.; David, M. Aromatase inhibitors and cyclooxygenase-2 (COX-2) inhibitors in endometriosis: New questions—Old answers? Eur. J. Obstet. Gynecol. Reprod. Biol. 2005, 122, 144–150. [Google Scholar] [CrossRef]
- Simmen, F.A.; Su, Y.; Xiao, R.; Zeng, Z.; Simmen, R.C.M. The Krüppel-like factor 9 (KLF9) network in HEC-1-A endometrial carcinoma cells suggests the carcinogenic potential of dys-regulated KLF9 expression. Reprod. Biol. Endocrinol. 2008, 6, 41. [Google Scholar] [CrossRef] [Green Version]
- Pabona, J.M.; Simmen, F.A.; Nikiforov, M.A.; Zhuang, D.; Shankar, K.; Velarde, M.C.; Zelenko, Z.; Giudice, L.C.; Simmen, R.C. Kruppel-like factor 9 and progesterone receptor coregulation of decidualizing endometrial stromal cells: Implications for the pathogenesis of endometriosis. J. Clin. Endocrinol. Metab. 2012, 97, E376–E392. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heard, M.E.; Simmons, C.D.; Simmen, F.A.; Simmen, R.C. Kruppel-like factor 9 deficiency in uterine endometrial cells promotes ectopic lesion establishment associated with activated notch and hedgehog signaling in a mouse model of endometriosis. Endocrinology 2014, 155, 1532–1546. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Signorile, P.G.; Baldi, A. Looking for an effective and non-invasive diagnostic test for endometriosis: Where are we? Ann. Transl. Med. 2018, 17, S106. [Google Scholar] [CrossRef] [PubMed]
- Bockaj, M.; Fung, B.; Tsoulis, M.; Foster, W.G.; Soleymani, L. Method for Electrochemical Detection of Brain Derived Neurotrophic Factor (BDNF) in Plasma. Anal. Chem. 2018, 90, 8561–8566. [Google Scholar] [CrossRef] [PubMed]
- Takai, N.; Miyazaki, T.; Fujisawa, K.; Nasu, K.; Hamanaka, R.; Miyakawa, I. Polo-like kinase (PLK) expression in endometrial carcinoma. Cancer Lett. 2001, 169, 41–49. [Google Scholar] [CrossRef]
- Takai, N.; Miyazaki, T.; Fujisawa, K.; Nasu, K.; Hamanaka, R.; Miyakawa, I. Expression of polo-like kinase in ovarian cancer is associated with histological grade and clinical stage. Cancer Lett. 2001, 164, 41–49. [Google Scholar] [CrossRef]
- de Cárcer, G. The Mitotic Cancer Target Polo-Like Kinase 1: Oncogene or Tumor Suppressor? Genes 2019, 10, 208. [Google Scholar] [CrossRef] [Green Version]
- Fu, Z.; Wen, D. The Emerging Role of Polo-Like Kinase 1 in Epithelial-Mesenchymal Transition and Tumor Metastasis. Cancers (Basel) 2017, 9, 131. [Google Scholar] [CrossRef] [Green Version]
- Tang, L.; Wang, T.-T.; Wu, Y.-T.; Zhou, C.-Y.; Huang, H.-F. High expression levels of cyclin B1 and Polo-like kinase 1 in ectopic endometrial cells associated with abnormal cell cycle regulation of endometriosis. Fertil. Steril. 2009, 91, 979–987. [Google Scholar] [CrossRef]
- Gutteridge, R.E.A.; Ndiaye, M.A.; Liu, X.; Ahmad, N. Plk1 Inhibitors in Cancer Therapy: From Laboratory to Clinics. Mol. Cancer Ther. 2016, 15, 1427–1435. [Google Scholar] [CrossRef] [Green Version]
- Zhou, B.-B.S.; Elledge, S.J. The DNA damage response: Putting checkpoints in perspective. Nature 2000, 408, 433–439. [Google Scholar] [CrossRef] [PubMed]
- Vassileva, V.; Millar, A.; Briollais, L.; Chapman, W.; Bapat, B. Genes involved in DNA repair are mutational targets in endometrial cancers with microsatellite instability. Cancer Res. 2002, 62, 4095–4099. [Google Scholar] [PubMed]
- Itamochi, H.; Nishimura, M.; Oumi, N.; Kato, M.; Oishi, T.; Shimada, M.; Sato, S.; Naniwa, J.; Sato, S.; Kudoh, A.; et al. Checkpoint Kinase Inhibitor AZD7762 Overcomes Cisplatin Resistance in Clear Cell Carcinoma of the Ovary. Int. J. Gynecol. Cancer 2014, 24, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Daniel, J.A.; Pellegrini, M.; Lee, J.-H.; Paull, T.T.; Feigenbaum, L.; Nussenzweig, A. Multiple autophosphorylation sites are dispensable for murine ATM activation in vivo. J. Cell. Biol. 2008, 183, 777–783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, J.; Mun Tho, L.; Xu, N.; A. Gillespie, D. Chapter 3—The ATM–Chk2 and ATR–Chk1 Pathways in DNA Damage Signaling and Cancer. In Advances in Cancer Research; Vande Woude, G.F., Klein, G., Eds.; Academic Press: Cambridge, MA, USA, 2010; Volume 108, pp. 73–112. [Google Scholar]
- Takeuchi, M.; Tanikawa, M.; Nagasaka, K.; Oda, K.; Kawata, Y.; Oki, S.; Agapiti, C.; Sone, K.; Miyagawa, Y.; Hiraike, H.; et al. Anti-Tumor Effect of Inhibition of DNA Damage Response Proteins, ATM and ATR, in Endometrial Cancer Cells. Cancers (Basel) 2019, 11, 1913. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smith, J.R.; Hayman, G.T.; Wang, S.J.; Laulederkind, S.J.F.; Hoffman, M.J.; Kaldunski, M.L.; Tutaj, M.; Thota, J.; Nalabolu, H.S.; Ellanki, S.L.R.; et al. The Year of the Rat: The Rat Genome Database at 20: A multi-species knowledgebase and analysis platform. Nucleic Acids Res. 2020, 48, D731–D742. [Google Scholar] [CrossRef]
- Sabbadin, C.; Andrisani, A.; Ambrosini, G.; Bordin, L.; Donà, G.; Manso, J.; Ceccato, F.; Scaroni, C.; Armanini, D. Aldosterone in Gynecology and Its Involvement on the Risk of Hypertension in Pregnancy. Front. Endocrinol. 2019, 10, 575. [Google Scholar] [CrossRef] [Green Version]
- Letsiou, S.; Peterse, D.P.; Fassbender, A.; Hendriks, M.M.; van den Broek, N.J.; Berger, R.; Dorien, F.O.; Vanhie, A.; Vodolazkaia, A.; Van Langendonckt, A.; et al. Endometriosis is associated with aberrant metabolite profiles in plasma. Fertil. Steril. 2017, 107, 699–706.e696. [Google Scholar] [CrossRef] [Green Version]
| GEO Data Series | Total No. of Cases (Diseased + Normal) | Platform | Sample Description |
|---|---|---|---|
| GSE 7846 | 10 (5 + 5) | GPL570 | Endometriosis (n = 5), Normal (n = 5) |
| GSE 7305 | 20 (10 +10) | GPL570 | Endometriosis (n = 10), Normal (n = 10) |
| GSE 6364 | 37 (21 + 16) | GPL570 | Proliferative (n = 6), Proliferative normal (n = 5); Early-secretory (n = 6), Early-secretory normal (n = 3); Mid-secretory (n = 9), Mid-secretory normal (n = 8) |
| GSE 4888 | 27 (21 + 6) | GPL570 | Proliferative (n = 4), Early-secretory (n = 3), Mid-secretory (n = 8), Late-secretory (n = 6), Ambiguous histology (n = 6) |
| GSE 51981 | 148 (77 + 71) | GPL570 | Severe endometriosis (n = 48), Mild endometriosis (n = 29), Normal without pelvic/uterine pathology (n = 34), Normal with pelvic/uterine pathology (n = 37) |
| GSE 31683 | 10 (6 + 4) | GPL570 | KLF9 silenced (n = 2), PGR silenced (n = 2), Both KLF9 and PGR silenced (n = 2), Normal (n = 4) |
| GSE 25628 | 22 (16 + 6) | GPL571 | Eutopic (n = 9), Ectopic (n = 7), Normal (n = 6) |
| Gene Symbol | Gene Name | p-Value | Fold-Change |
|---|---|---|---|
| FOS | FBJ murine osteosarcoma viral oncogene homolog | 3.02 × 10−13 | 9.94558 |
| EGR1 | early growth response 1 | 7.67 × 10−17 | 7.95823 |
| FOSB | FBJ murine osteosarcoma viral oncogene homolog B | 1.87 × 10−10 | 7.14284 |
| ZFP36 | ZFP36 ring finger protein | 5.23 × 10−14 | 4.01565 |
| JUNB | jun B proto-oncogene | 8.28 × 10−15 | 4.01404 |
| APOD | apolipoprotein D | 1.21 × 10−7 | 3.7376 |
| CST1 | cystatin SN | 2.52 × 10−5 | 3.42216 |
| GPX3 | glutathione peroxidase 3 | 0.0057 | 3.34348 |
| PER1 | period circadian clock 1 | 6.14 × 10−13 | 3.23599 |
| CTSW | cathepsin W | 8.08 × 10−7 | 3.22223 |
| SOCS3 | suppressor of cytokine signaling 3 | 4.99 × 10−5 | 3.11348 |
| CFD | complement factor D (adipsin) | 0.000993 | 2.99141 |
| HSPB6 | heat shock protein, alpha-crystallin-related, B6 | 9.31 × 10−8 | 2.98362 |
| LEFTY1 | left-right determination factor 1 | 0.004712 | 2.89285 |
| PAEP | progestagen-associated endometrial protein | 0.029113 | 2.88223 |
| DUSP1 | dual specificity phosphatase 1 | 1.48 × 10−7 | 2.88219 |
| GNLY | Granulysin | 1.52 × 10−5 | 2.83639 |
| EPHX1 | epoxide hydrolase 1, microsomal (xenobiotic) | 6.69 × 10−10 | 2.78618 |
| MT1M | metallothionein 1M | 0.011648 | 2.76574 |
| CLEC3B///EXOSC7 | C-type lectin domain family 3, member B///exosome component 7 | 7.13 × 10−5 | 2.74002 |
| GNLY | Granulysin | 1.61 × 10−5 | 2.73433 |
| TPSAB1///TPSB2 | tryptase alpha/beta 1///tryptase beta 2 (gene/pseudogene) | 1.20 x10−6 | 2.73381 |
| IER2 | immediate early response 2 | 4.65 × 10−14 | 2.67138 |
| PTPRO | protein tyrosine phosphatase, receptor type, O | 2.70 × 10−14 | 2.66719 |
| ELN | Elastin | 2.26 × 10−7 | 2.6637 |
| IER3 | immediate early response 3 | 5.70 × 10−5 | 2.61981 |
| SOX13 | SRY box 13 | 4.59 × 10−9 | 2.60666 |
| SOD3 | superoxide dismutase 3, extracellular | 2.00 × 10−9 | 2.60457 |
| SLC30A2 | solute carrier family 30 (zinc transporter), member 2 | 0.002566 | 2.59508 |
| AQP3 | aquaporin 3 (Gill blood group) | 0.000604 | 2.58706 |
| HSPB6 | heat shock protein, alpha-crystallin-related, B6 | 2.81 × 10−6 | 2.57421 |
| CD37 | CD37 molecule | 3.56 × 10−8 | 2.48625 |
| IRX3 | iroquois homeobox 3 | 0.00585 | 2.48441 |
| CREB3L1 | cAMP responsive element binding protein 3-like 1 | 1.53 × 10−7 | 2.48023 |
| Gene Symbol | Gene Title | p-Value | Fold-Change |
|---|---|---|---|
| DIO2 | deiodinase, iodothyronine, type II | 6.84 × 10−12 | −5.38638 |
| CPM | carboxypeptidase M | 1.52 × 10−7 | −5.20763 |
| OLFM4 | olfactomedin 4 | 1.22 × 10−6 | −4.77761 |
| PALLD | palladin, cytoskeletal associated protein | 1.20 × 10−13 | −4.47775 |
| BAG5 | BCL2-associated athanogene 5 | 1.62 × 10−8 | −4.38993 |
| TOP2A | topoisomerase (DNA) II alpha | 4.50 × 10−8 | −4.22713 |
| PKP4 | plakophilin 4 | 5.35 × 10−19 | −3.97475 |
| CDC20B | cell division cycle 20B | 1.36 × 10−6 | −3.96043 |
| SNTN | sentan, cilia apical structure protein | 5.43 × 10−12 | −3.95899 |
| SET | SET nuclear proto-oncogene | 5.39 × 10−11 | −3.90488 |
| CRISPLD1 | cysteine-rich secretory protein LCCL domain containing 1 | 5.12 × 10−7 | −3.85982 |
| NPAS3 | neuronal PAS domain protein 3 | 1.57 × 10−6 | −3.83298 |
| CADM1 | cell adhesion molecule 1 | 4.63 × 10−15 | −3.78248 |
| SMC3 | structural maintenance of chromosomes 3 | 2.59 × 10−16 | −3.7407 |
| SFRP4 | secreted frizzled-related protein 4 | 0.000481 | −3.72407 |
| ANK2 | ankyrin 2, neuronal | 2.00 × 10−7 | −3.71876 |
| ANLN | anillin actin binding protein | 4.54 × 10−8 | −3.70838 |
| WIF1 | WNT inhibitory factor 1 | 1.73 × 10−6 | −3.65345 |
| MMP26 | matrix metallopeptidase 26 | 0.002107 | −3.63998 |
| PCSK5 | proprotein convertase subtilisin/kexin type 5 | 7.33 × 10−8 | −3.63558 |
| OXR1 | oxidation resistance 1 | 1.62 × 10−13 | −3.61905 |
| CTNNB1 | catenin (cadherin-associated protein), beta 1 | 6.39 × 10−12 | −3.55057 |
| PCSK5 | proprotein convertase subtilisin/kexin type 5 | 1.69 × 10−8 | −3.53439 |
| EIF5B | eukaryotic translation initiation factor 5B | 1.60 × 10−8 | −3.52153 |
| HSP90B1 | heat shock protein 90kDa beta (Grp94), member 1 | 2.11 × 10−9 | −3.49988 |
| PCYOX1 | prenylcysteine oxidase 1 | 3.06 × 10−11 | −3.47374 |
| MIB1 | mindbomb E3 ubiquitin protein ligase 1 | 1.96 × 10−9 | −3.46544 |
| OSBPL1A | oxysterol binding protein-like 1A | 2.08 × 10−14 | −3.46429 |
| CBX3 | chromobox homolog 3 | 2.18 × 10−12 | −3.45307 |
| TCAF1 | TRPM8 channel-associated factor 1 | 1.87 × 10−13 | −3.42665 |
| KMO | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 2.06 × 10−5 | −3.42531 |
| CTSZ | cathepsin Z | 3.75 × 10−13 | −3.40201 |
| KMO | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 2.13 × 10−5 | −3.38857 |
| PCSK5 | proprotein convertase subtilisin/kexin type 5 | 5.06 × 10−8 | −3.38299 |
| YWHAB | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta | 2.64 × 10−13 | −3.35541 |
| NMT2 | N-myristoyltransferase 2 | 5.60 × 10−12 | -3.34186 |
| CADM1 | cell adhesion molecule 1 | 3.18 × 10−9 | −3.3281 |
| CEP57 | centrosomal protein 57kDa | 3.21 × 10−15 | −3.28628 |
| Ingenuity Canonical Pathways | −log (p-Value) | Ratio | Predicted z-Score | Molecules |
|---|---|---|---|---|
| Mitotic Roles of Polo-Like Kinase | 7.34 | 0.333 | −2.714 (Inhibited) | ANAPC4, CCNB1, CCNB2, CDC23, CDC27, CDK1, FBXO5, HSP90AA1, HSP90AB1, HSP90B1, KIF11, KIF23, PLK2, PPM1L, PPP2R2C, PPP2R5C, PPP2R5E, PRC1, RAD21, SMC3, TGFB1 |
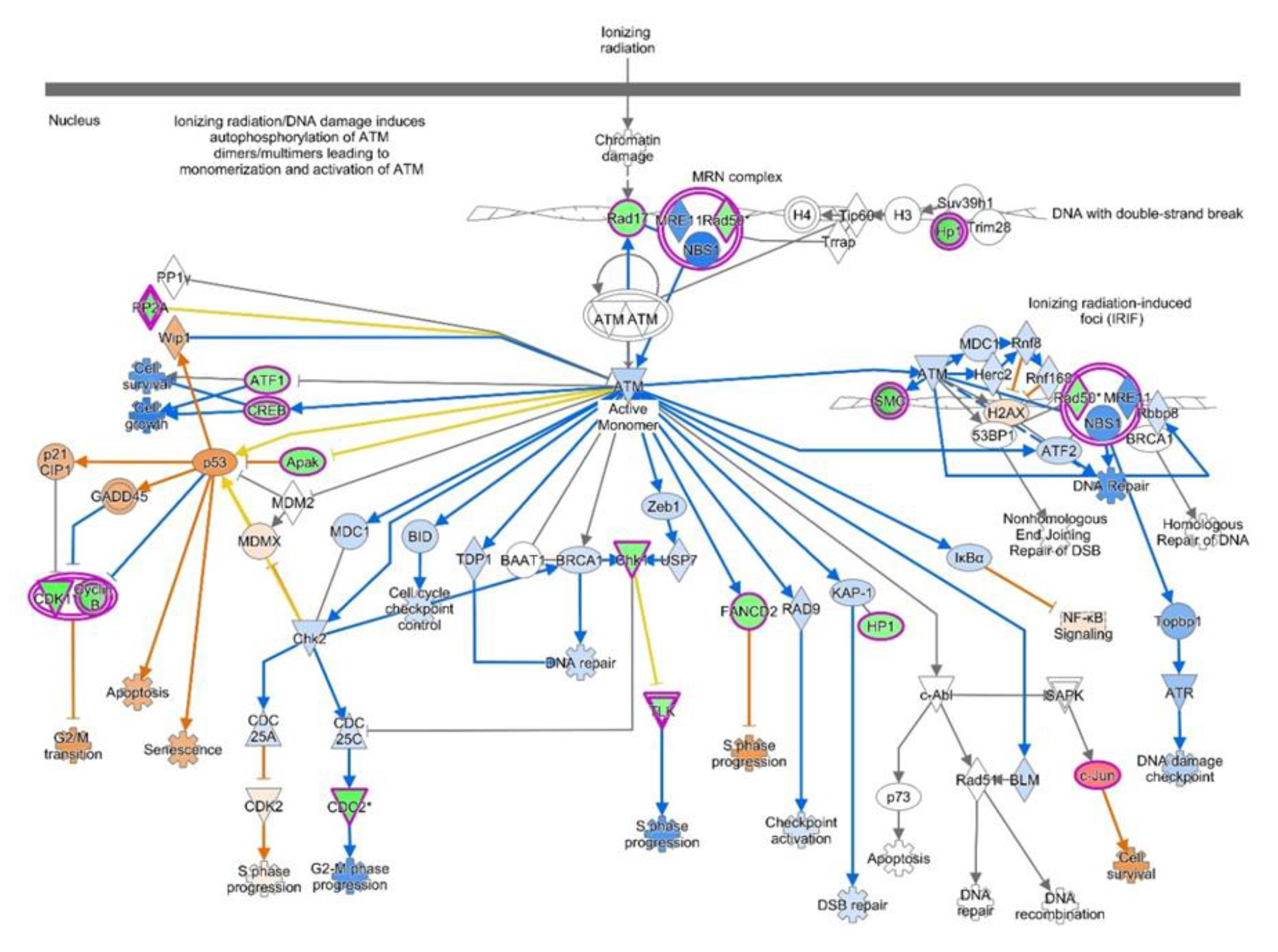
| ATM Signaling | 4.11 | 0.219 | 1.698 (Activated) | ATF1, CBX1, CBX3, CBX5, CCNB1, CCNB2, CDK1, CHEK1, CREB1, FANCD2, JUN, PPM1L, PPP2R2C, PPP2R5C, PPP2R5E, RAD17, RAD50, SMC2, SMC3, TLK1, ZNF420 |
| Aldosterone Signaling in Epithelial Cells | 3.8 | 0.183 | −3.464 (Inhibited) | DNAJA1, DNAJB14, DNAJC10, DNAJC27, DNAJC3, DNAJC9, DUSP1, HSP90AA1, HSP90AB1, HSP90B1, HSPA4, HSPA9, HSPB6, HSPD1, HSPH1, ITPR2, PDIA3, PDPK1, PIK3C2A, PIK3CA, PIK3R3, PIP5K1B, PLCB1, PRKCI, PRKD3, SACS, SCNN1G |
| Role of CHK Proteins in Cell Cycle Checkpoint Control | 5.3 | 0.298 | −0.632 | ATMIN, CDK1, CHEK1, E2F7, E2F8, PCNA, PPM1L, PPP2R2C, PPP2R5C, PPP2R5E, RAD17, RAD50, RFC3, RFC4, RFC5, RPA1, TLK1 |
| SUMOylation Pathway | 3.34 | 0.198 | 2.668 (Activated) | DNMT3A, EP300, FOS, HDAC2, JUN, MYB, PCNA, PIAS1, RFC3, RFC4, RFC5, RHOB, RHOBTB1, RHOQ, RHOT1, RND3, RPA1, SERBP1, SMAD4, UBA2 |
| Role of BRCA1 in DNA Damage Response | 3.77 | 0.225 | −0.707 | ABRAXAS1, ATF1, BRCC3, BRD7, BRIP1, CHEK1, E2F7, E2F8, FANCD2, FANCL, MSH2, MSH6, RAD50, RFC3, RFC4, RFC5, RPA1, SMARCC1 |
| Cyclins and Cell Cycle Regulation | 2.83 | 0.2 | −1.604 | CCNA1, CCNA2, CCNB1, CCNB2, CCNE2, CDK1, CDK6, E2F7, E2F8, HDAC2, PPM1L, PPP2R2C, PPP2R5C, PPP2R5E, SKP2, TGFB1 |
| PI3K/AKT Signaling | 2.7 | 0.171 | −1.091 | CTNNB1, EIF4E, HSP90AA1, HSP90AB1, HSP90B1, INPP5F, ITGB1, JAK1, PDPK1, PIK3CA, PIK3R3, PPM1L, PPP2R2C, PPP2R5C, PPP2R5E, PTEN, RASD1, RHEB, RPS6KB1, SFN, SYNJ2, YWHAB |
| Upstream Regulator | Molecule Type | Predicted Activation z-score | p-Value of Overlap | Target Molecules in Dataset |
|---|---|---|---|---|
| REL | transcription regulator | −4.137 (Inhibited) | 0.00021 | AGA, AGPS, ANLN, APP, ARFGAP3, ATXN1, BCL3, CAMK2D, CCNY, CDC6 |
| CTNNB1 | transcription regulator | −3.208 (Inhibited) | 0.0104 | AKAP13, ALDH1A1, APOD, APP, ARFGAP3, ARMH4, AURKA, CADM1, CALM1, CCL3 |
| PGR | ligand-dependent nuclear receptor | −2.237 (Inhibited) | 0.00051 | ABCG2, ACOX1, AHCYL1, AK3, AKAP13, ATP1B1, ATXN1, BUB1, CA12, CCNB1 |
| VCAN | Proteoglycan | −2.625 (Inhibited) | 0.024 | COMP, CPE, ELN, IFI44L, IFIT1, ITGB1, MYH10, PCSK5, PENK, PLA2G2A |
| ACTL6A | Other | −2.236 (Inhibited) | 0.093 | CCNA2, CCNB1, CCNB2, CCNE2, SFN |
| DUSP1 | phosphatase | −2.155 (Inhibited) | 1 | CMPK2, DUSP1, IER3, IFIT1, IFIT3, JUN, PTEN, ZFP36 |
| HELLS | Enzyme | −2 (Inhibited) | 0.0024 | CCNA2, CCNB1, CDC6, HSPD1, PCNA, SLC44A1 |
| RASSF8 | Other | −2 (Inhibited) | 0.0016 | ENPP5, MRPL30, NEDD9, POSTN |
| TCF4 | transcription regulator | −1.912 | 0.040 | CCNA2, CCNB2, CDK1, CEP55, E2F8, FOS, HMGB2, HMMR, HSP90B1, IFI16 |
| IGF2R | transmembrane receptor | −2 (Inhibited) | 0.0080 | ENPP5, MRPL30, NEDD9, POSTN |
| TGFB1 | growth factor | −1.276 | 0.00087 | ABCG2, ACKR1, ADAM12, ALDH5A1, APP, ARHGAP19, ASPM, ATG12, ATXN1, BCL3 |
| HSF2 | transcription regulator | −1.408 | 0.026 | CCT2, HSBP1, HSPA4, HSPH1, JUN, PSMA5, TCP1 |
| EDN3 | other | 0.816 | 0.0024 | CDH2, CTNNB1, EGR1, FOS, ITGB1, LAMA1 |
| FOS | transcription regulator | 0.917 | 0.0020 | ACOX1, ADAM12, AGPS, ANK3, AQP3, ATP2C1, CADM1, CALU, CAMK2D, CAT |
| RTN4 | other | 1.51 | 0.0041 | APP, CFL1, IMPACT, JUN, JUND, LAP3, MAP2, RHOB, RTN4, YWHAB |
| ZFP36 | transcription regulator | 1.873 | 0.00015 | CCNE2, CDC6, CENPA, CLCN3, CLMP, CTSS, E2F8, FOS, IER3, JUN |
| EPHB1 | kinase | 1.98 | 0.0052 | EGR1, FOS, JUN, JUNB |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mirza, Z.; Abdel-dayem, U.A. Uncovering Potential Roles of Differentially Expressed Genes, Upstream Regulators, and Canonical Pathways in Endometriosis Using an In Silico Genomics Approach. Diagnostics 2020, 10, 416. https://doi.org/10.3390/diagnostics10060416
Mirza Z, Abdel-dayem UA. Uncovering Potential Roles of Differentially Expressed Genes, Upstream Regulators, and Canonical Pathways in Endometriosis Using an In Silico Genomics Approach. Diagnostics. 2020; 10(6):416. https://doi.org/10.3390/diagnostics10060416
Chicago/Turabian StyleMirza, Zeenat, and Umama A. Abdel-dayem. 2020. "Uncovering Potential Roles of Differentially Expressed Genes, Upstream Regulators, and Canonical Pathways in Endometriosis Using an In Silico Genomics Approach" Diagnostics 10, no. 6: 416. https://doi.org/10.3390/diagnostics10060416
APA StyleMirza, Z., & Abdel-dayem, U. A. (2020). Uncovering Potential Roles of Differentially Expressed Genes, Upstream Regulators, and Canonical Pathways in Endometriosis Using an In Silico Genomics Approach. Diagnostics, 10(6), 416. https://doi.org/10.3390/diagnostics10060416





