COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2
Abstract
1. Introduction
2. Isothermal Amplification Technologies (IAT)
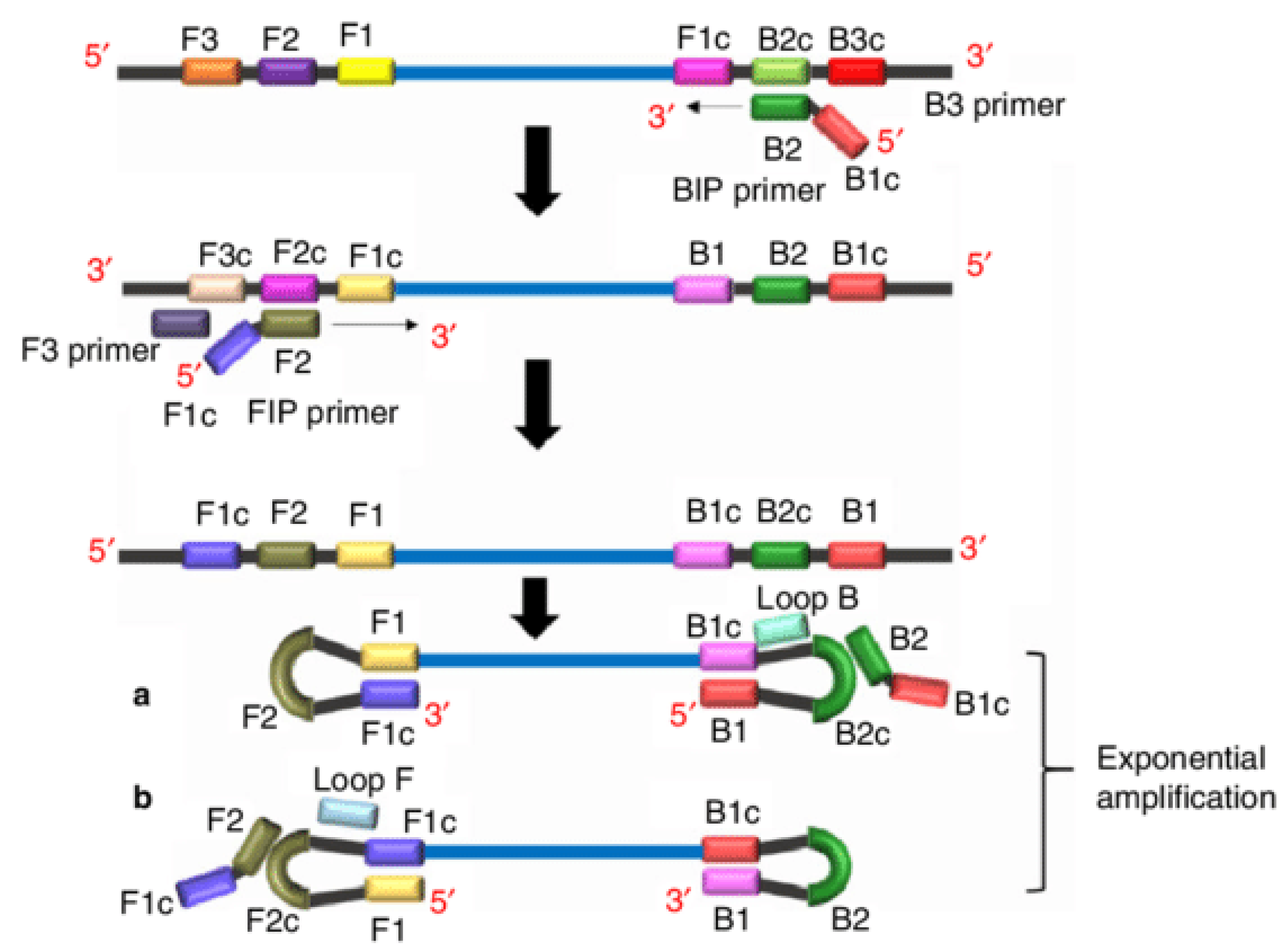
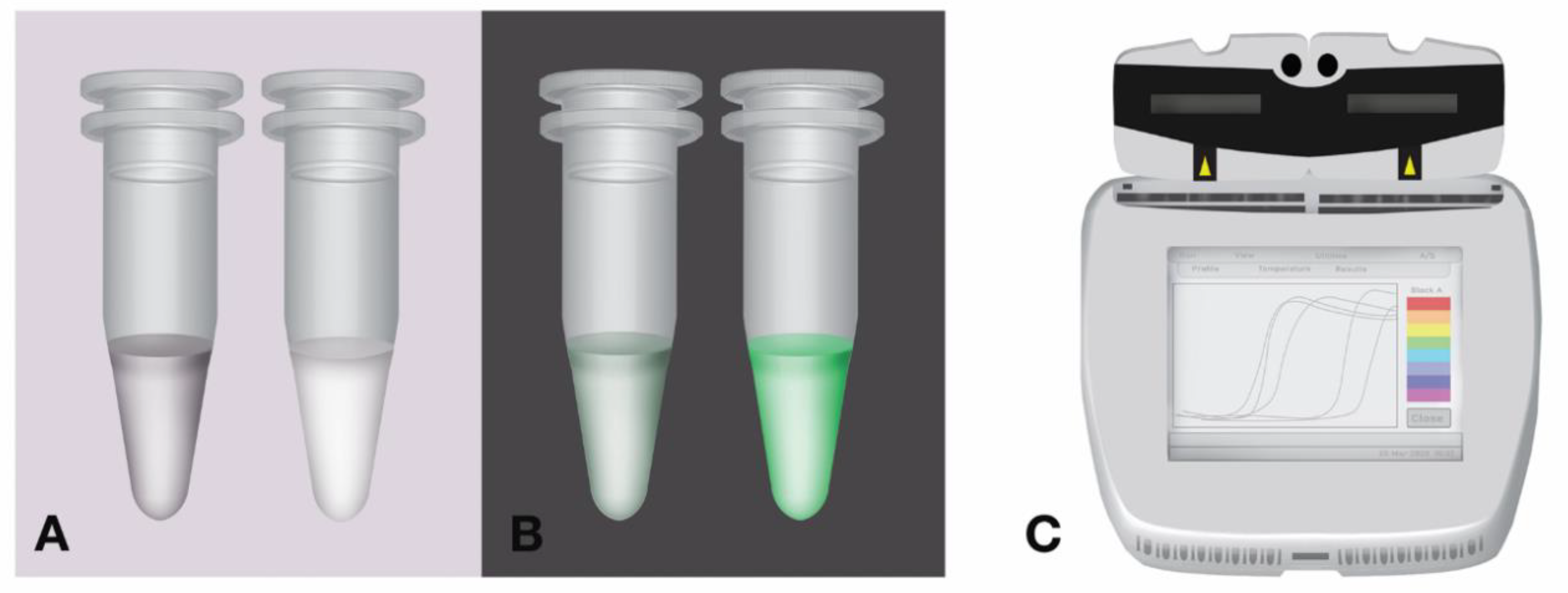
2.1. Loop-Mediated Isothermal Amplification
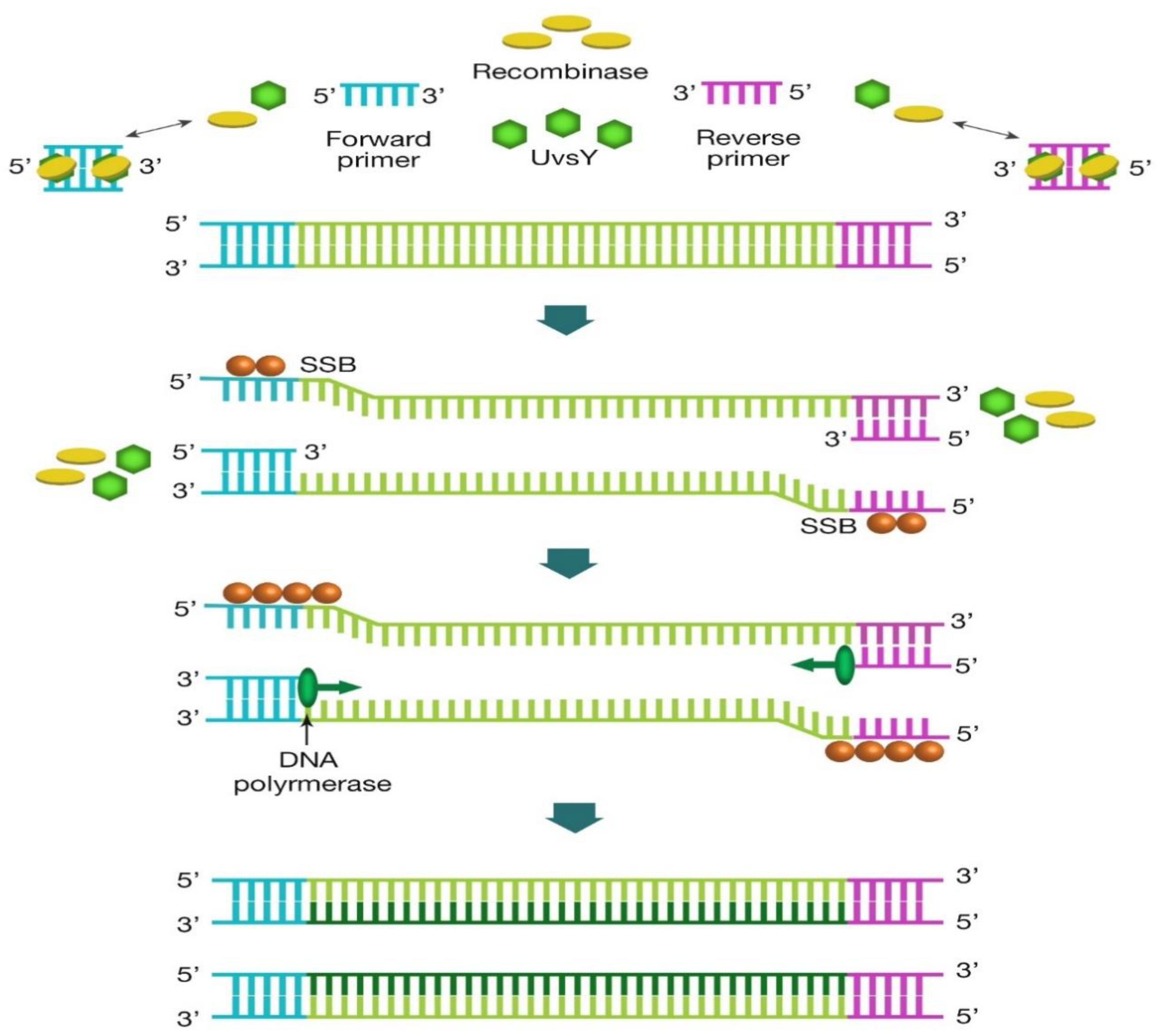
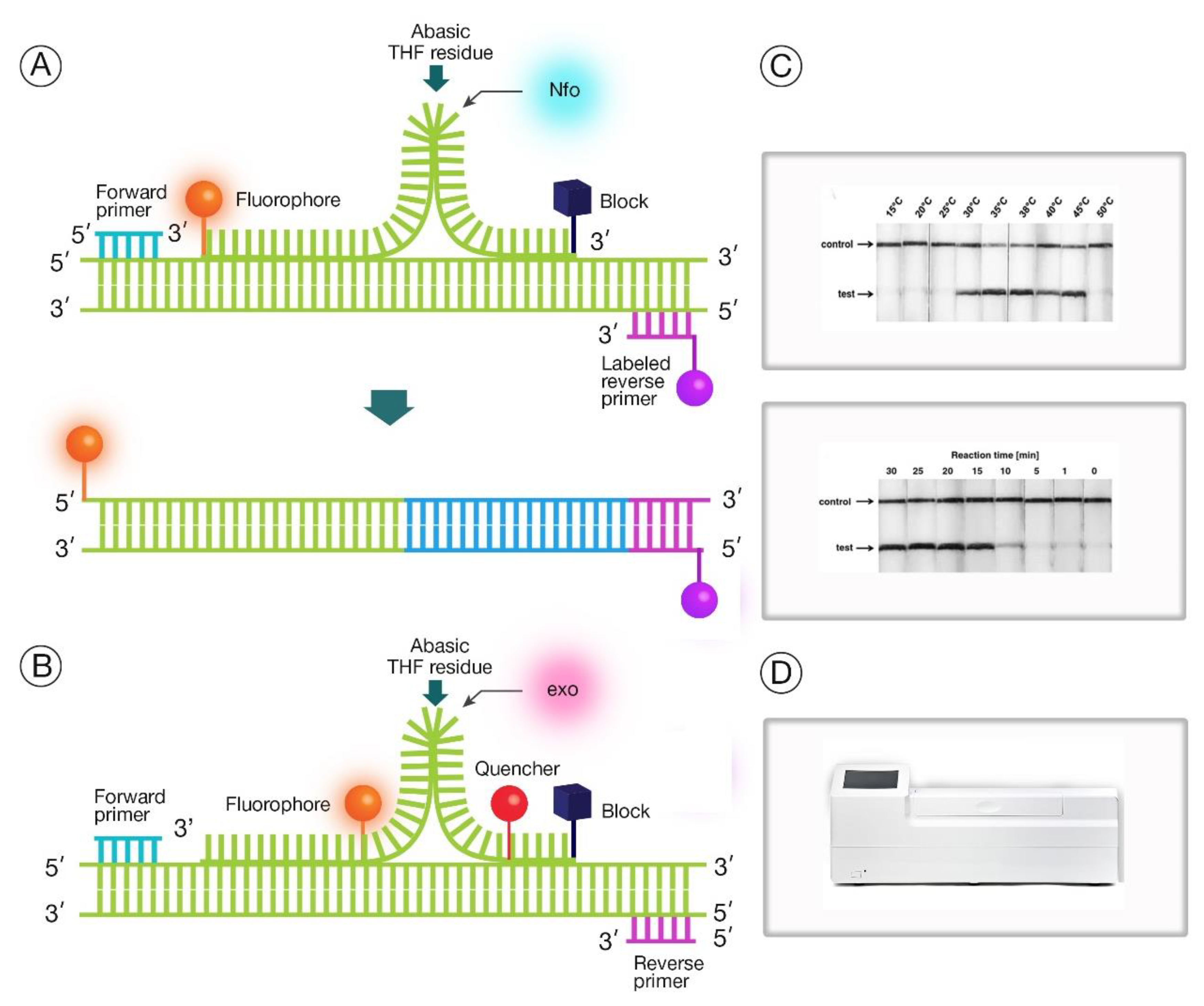
2.2. Recombinase Polymerase Amplification
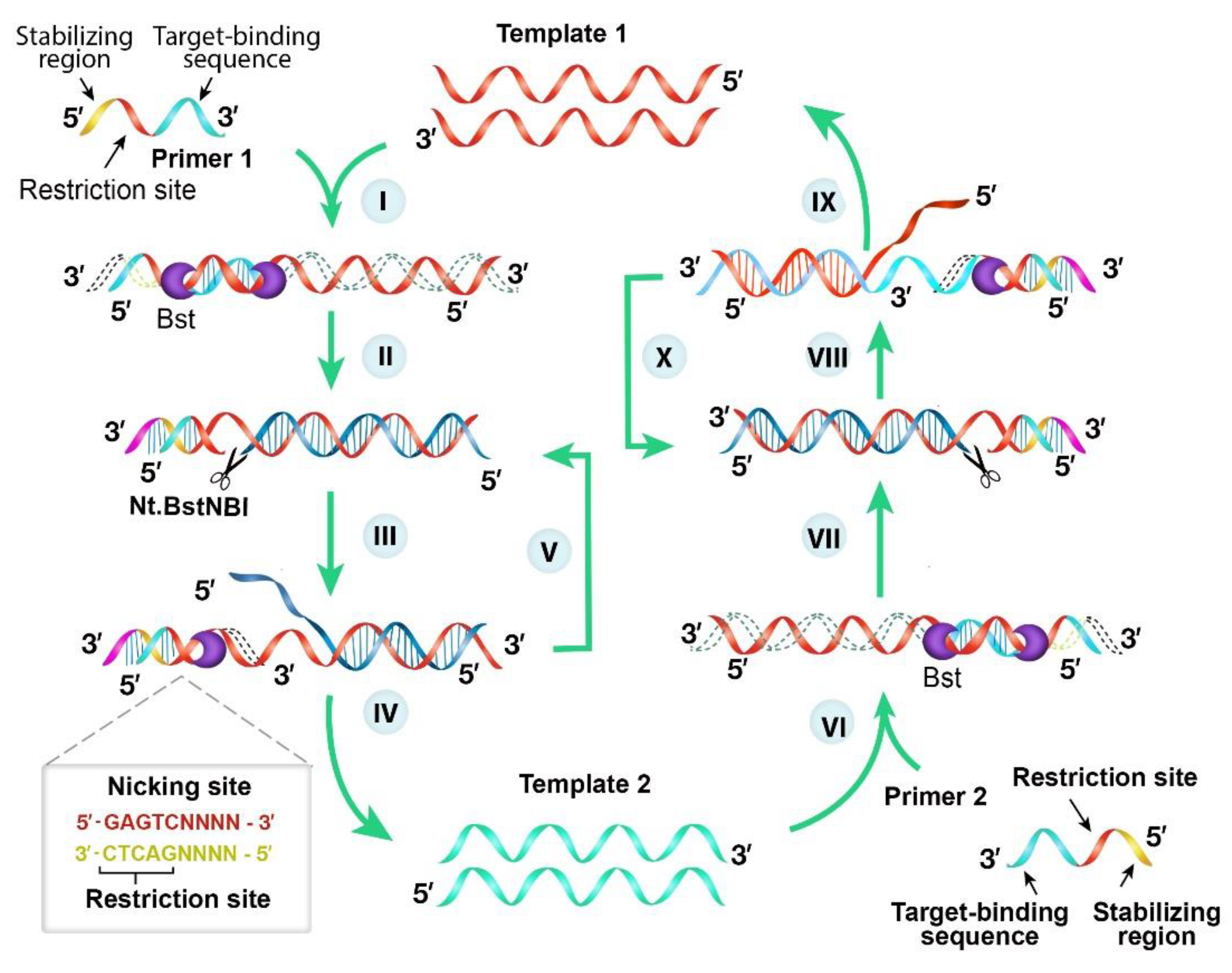
2.3. Nicking Enzyme-Assisted Reaction
3. Future Direction of IAT for COVID-19 Diagnosis
3.1. Safety Measures for Sample Process and IAT Testing
3.2. Development of Multiplex Detection of SARS-CoV-2
3.3. Integrating Sample Preparation, Amplification and Detection to Reduce Cross Contamination
Author Contributions
Funding
Conflicts of Interest
References
- Novel Coronavirus (2019-nCoV), Situation Report 11. Available online: https://www.who.int/docs/defaultsource/coronaviruse/situation-reports/20200131-sitrep-11-ncov.pdf?sfvrsn=de7c0f7_4 (accessed on 13 March 2020).
- Novel Coronavirus (2019-nCoV) Situation Report 48. Available online: https://www.who.int/docs/defaultsource/coronaviruse/situation-reports/20200308-sitrep-48-covid-19.pdf?sfvrsn=16f7ccef_4 (accessed on 13 March 2020).
- The Global Coronavirus Recession Is Beginning. Available online: https://edition.cnn.com/2020/03/16/economy/global-recession-coronavirus/index.html (accessed on 13 March 2020).
- The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team. The epidemiological characteristics of an outbreak of 2019 novel coronavirus disease (COVID-19)—China, 2020. China CDC Wkly. 2020, 2, 1–10. [Google Scholar]
- Munster, V.J.; Koopmans, M.; van Doremalen, N.; van Riel, D.; de Wit, E. A novel coronavirus emerging in China—key questions for impact assessment. N. Engl. J. Med. 2020, 382, 692–694. [Google Scholar] [CrossRef] [PubMed]
- Hoehl, S.; Rabenau, H.; Berger, A.; Kortenbusch, M.; Cinatl, J.; Bojkova, D.; Behrens, P.; Böddinghaus, B.; Götsch, U.; Naujoks, F.; et al. Evidence of SARS-CoV-2 infection in returning travelers from Wuhan, China. N. Engl. J. Med. 2020, 382, 1278–1280. [Google Scholar] [CrossRef] [PubMed]
- Gates, B. Responding to Covid-19—A Once-in-a-Century Pandemic? N. Engl. J. Med. 2020, 382, 1677–1679. [Google Scholar] [CrossRef]
- Wang, W.; Tang, J.; Wei, F. Updated understanding of the outbreak of 2019 novel coronavirus (2019-nCoV) in Wuhan, China. J. Med. Virol. 2020, 92, 441–447. [Google Scholar] [CrossRef]
- Tang, B.; Bragazzi, N.L.; Li, Q.; Tang, S.; Xiao, Y.; Wu, J. An updated estimation of the risk of transmission of the novel coronavirus (2019-nCov). Infect. Dis. Model. 2020, 5, 248–255. [Google Scholar] [CrossRef]
- Cyranoski, D. Did pangolins spread the China coronavirus to people? Nature 2020. [Google Scholar] [CrossRef]
- Banerjee, A.; Kulcsar, K.; Misra, V.; Frieman, M.; Mossman, K. Bats and Coronaviruses. Viruses 2019, 11, 41. [Google Scholar] [CrossRef]
- WHO Recommendations to Reduce Risk of Transmission of Emerging Pathogens from Animals to Humans in Live Animal Markets. Available online: https://www.who.int/health-topics/coronavirus/whorecommendations-to-reduce-risk-of-transmission-of-emerging-pathogens-from-animals-to-humans-inlive-animal-markets (accessed on 13 March 2020).
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Ou, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.; et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Rothe, C.; Schunk, M.; Sothmann, P.; Bretzel, G.; Froeschl, G.; Wallrauch, C.; Zimmer, T.; Thiel, V.; Janke, C.; Guggemos, W.; et al. Transmission of 2019-nCoV infection from an asymptomatic contact in Germany. N. Engl. J. Med. 2020, 382, 970–971. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.M.; Wang, W.; Song, Z.G.; Hu, Y.; Tao, Z.W.; Tian, J.H.; Pei, Y.Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.T.; Leung, K.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study. Lancet 2020. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S.; Schneider, J.; Schmidt, M.L.; et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 2020, 25. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, T.; Andreasen, S.Z.; Wol, A.; Bang, D.D. From Lab on a Chip to Point of Care Devices: The Role of Open Source Microcontrollers. Micromachines 2018, 9, 403. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization (WHO). Outbreak Investigation. Available online: https://www.who.int/hac/techguidance/training/outbreak%20investigation_en.pdf (accessed on 13 March 2020).
- Isere, E.E.; Fatiregun, A.; Ajayi, I.O. An overview of disease surveillance and notification system in Nigeria and the roles of clinicians in disease outbreak prevention and control. Niger. Med. J. 2015, 56, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.; Le Hello, S.; Silva, N.J.-D.; Vaillant, V.; De Valk, H.; Weill, F.-X.; Le Strat, Y. The French human Salmonella surveillance system: Evaluation of timeliness of laboratory reporting and factors associated with delays, 2007 to 2011. Eurosurveillance 2014, 19, 20664. [Google Scholar] [CrossRef]
- Nguyen, T.; Bang, D.D.; Wolff, A. 2019 Novel Coronavirus Disease (COVID-19): Paving the Road for Rapid Detection and Point-of-Care Diagnostics. Micromachines 2020, 11, 306. [Google Scholar] [CrossRef]
- Chen, Y.; Chan, K.H.; Hong, C.; Kang, Y.; Ge, S.; Chen, H.; Wong, E.Y.; Joseph, S.; Patteril, N.G.; Wernery, U.; et al. A highly specific rapid antigen detection assay for onsite diagnosis of MERS. J. Infect. 2016, 73, 82–84. [Google Scholar] [CrossRef]
- Lau, S.K.; Woo, P.C.; Wong, B.H.; Tsoi, H.W.; Woo, G.K.; Poon, R.W.; Chan, K.H.; Wei, W.I.; Peiris, J.S.; Yuen, K.Y. Detection of severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein in sars patients by enzyme-linked immunosorbent assay. J. Clin. Microbiol. 2004, 42, 2884–2889. [Google Scholar] [CrossRef]
- Sastre, P.; Dijkman, R.; Camunas, A.; Ruiz, T.; Jebbink, M.F.; van der Hoek, L.; Vela, C.; Rueda, P. Differentiation between human coronaviruses NL63 and 229E using a novel double-antibody sandwich enzyme-linked immunosorbent assay based on specific monoclonal antibodies. Clin. Vaccine Immunol. 2011, 18, 113–118. [Google Scholar] [CrossRef]
- Liu, I.J.; Chen, P.J.; Yeh, S.H.; Chiang, Y.P.; Huang, L.M.; Chang, M.F.; Chen, S.Y.; Yang, P.C.; Chang, S.C.; Wang, W.K. Immunofluorescence assay for detection of the nucleocapsid antigen of the severe acute respiratory syndrome (SARS)-associated coronavirus in cells derived from throat wash samples of patients with SARS. J. Clin. Microbiol. 2005, 43, 2444–2448. [Google Scholar] [CrossRef] [PubMed]
- Sizun, J.; Arbour, N.; Talbot, P.J. Comparison of immunofluorescence with monoclonal antibodies and RT-PCR for the detection of human coronaviruses 229E and OC43 in cell culture. J. Virol. Methods 1998, 72, 145–152. [Google Scholar] [CrossRef]
- James, A.; Macdonald, J. Recombinase polymerase amplification: Emergence as a critical molecular technology for rapid, low-resource diagnostics. Expert Rev. Mol. Diagn. 2015, 15, 1475–1489. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Wu, S.; Hao, X.; Dong, X.; Mao, L.; Pelechano, V.; Chen, W.H.; Yin, X. Rapid colorimetric detection of COVID-19 coronavirus using a reverse transcriptional loop-mediated isothermal amplification (RT-LAMP) diagnostic platform: iLACO. MedRxiv 2020. [Google Scholar] [CrossRef]
- Lu, R.; Wu, X.; Wan, Z.; Li, Y.; Zuo, L.; Qin, J.; Jin, X.; Zhang, C. Development of a Novel Reverse Transcription Loop-Mediated Isothermal Amplification Method for Rapid Detection of SARS-CoV-2. Virol. Sin. 2020. [Google Scholar] [CrossRef]
- Lamb, L.E.; Bartolone, S.N.; Ward, E.; Chancellor, M.B. Rapid Detection of Novel Coronavirus (COVID-19) by Reverse Transcription-Loop-Mediated Isothermal Amplification. MedRxiv 2020. [Google Scholar] [CrossRef]
- Wua, G.; Zamana, M.H. Low-cost tools for diagnosing and monitoring HIV infection in low-resource settings. Bull. World Health Organ. 2012, 90, 914–920. [Google Scholar] [CrossRef]
- Mori, Y.; Kanda, H.; Notomi, T. Loop-mediated isothermal amplification (LAMP): Recent progress in research and development. J. Infect. Chemother. 2013, 19, 404–411. [Google Scholar] [CrossRef]
- Mori, Y.; Notomi, T. Loop-mediated isothermal amplification (LAMP): A rapid, accurate, and cost-effective diagnostic method for infectious diseases. J. Infect. Chemother. 2009, 15, 62–69. [Google Scholar] [CrossRef]
- De Paz, H.D.; Brotons, P.; Muñoz-Almagro, C. Molecular isothermal techniques for combating infectious diseases: Towards low-cost point-of-care diagnostics. Expert Rev. Mol. Diagn. 2014, 14, 827–843. [Google Scholar] [CrossRef]
- Compton, J. Nucleic acid sequence-based amplification. Nature 1991, 350, 91–92. [Google Scholar] [CrossRef] [PubMed]
- Gill, P.; Ghaemi, A. Nucleic acid isothermal amplification technologies–a review. Nucleos Nucleot Nucleic Acids 2008, 27, 224–243. [Google Scholar] [CrossRef] [PubMed]
- Dean, F.B.; Nelson, J.R.; Giesler, T.L.; Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 2011, 11, 1095–1099. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Vincent, M.; Xu, Y.; Kong, H. Helicase-dependent isothermal DNA amplification. EMBO Rep. 2004, 5, 795–800. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S. Rolling replication of short DNA circles. Proc. Natl. Acad. Sci. USA 1995, 92, 4641–4645. [Google Scholar] [CrossRef]
- Wharam, S.D.; Marsh, P.; Lloyd, J.S.; Ray, T.D.; Mock, G.A.; Assenberg, R.; McPhee, J.E.; Brown, P.; Weston, A.; Cardy, D.L. Specific detection of DNA and RNA targets using a novel isothermal nucleic acid amplification assay based on the formation of a three-way junction structure. Nucleic Acids Res. 2001, 29, e54. [Google Scholar] [CrossRef] [PubMed]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA Detection using recombination proteins. PLoS Biol. 2006, 4, e204. [Google Scholar] [CrossRef]
- Guatelli, J.C.; Whitfield, K.M.; Kwoh, D.Y.; Barringer, K.J.; Richman, D.D.; Gingeras, T.R. Isothermal, in vitro amplification of nucleic acids by a multienzyme reaction modeled after retroviral replication. Proc. Natl. Acad. Sci. USA 1990, 87, 1874–1878. [Google Scholar] [CrossRef]
- Zhang, D.Y.; Brandwein, M.; Hsuih, T.C.; Li, H. Amplification of target-specific, ligation-dependent circular probe. Gene 1998, 211, 77–85. [Google Scholar] [CrossRef]
- Walker, G.T.; Little, M.C.; Nadeau, J.G.; Shank, D.D. Isothermal in vitro amplification of DNA by a restriction enzyme/DNA polymerase system. Proc. Natl. Acad. Sci. USA 1992, 89, 392–396. [Google Scholar] [CrossRef] [PubMed]
- Van Ness, J.; Van Ness, L.K.; Galas, D.J. Isothermal reactions for the amplification of oligonucleotides. Proc. Natl. Acad. Sci. USA 2003, 100, 4504–4509. [Google Scholar] [CrossRef] [PubMed]
- Chotiwan, N.; Brewster, C.D.; Magalhaes, T.; Weger-Lucarelli, J.; Duggal, N.K.; Rückert, C.; Nguyen, C.; Luna, S.M.; Fauver, J.R.; Andre, B.; et al. Rapid and specific detection of Asian- and African-lineage Zika viruses. Sci. Transl. Med. 2017, 9, eaag0538. [Google Scholar] [CrossRef] [PubMed]
- Wong, Y.P.; Othman, S.; Lau, Y.L.; Radu, S.; Chee, H.-Y. Loop-mediated isothermal amplification (LAMP): A versatile technique for detection of microorganisms. J. Appl. Microbiol. 2018, 124, 626–643. [Google Scholar] [CrossRef] [PubMed]
- Tomita, N.; Mori, Y.; Kanda, H.; Notomi, T. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nat. Protoc. 2008, 3, 877–882. [Google Scholar] [CrossRef]
- Crannell, Z.A.; Rohrman, B.; Richards-Kortum, R. Equipment-free incubation of recombinase polymerase amplification reactions using body heat. PLoS ONE 2014, 9, e112146. [Google Scholar] [CrossRef]
- James, A.S.; Todd, S.; Pollak, N.M.; Marsh, G.A.; Macdonald, J. Marsh, and Joanne Macdonald. Ebolavirus diagnosis made simple, comparable and faster than molecular detection methods: Preparing for the future. Virol. J. 2018, 15, 75. [Google Scholar] [CrossRef]
- Qian, C.; Wang, R.; Wu, H.; Ji, F.; Wu, J. Nicking enzyme-assisted amplification (NEAA) technology and its applications: A review. Anal. Chim. Acta 2019, 1050, 1–15. [Google Scholar] [CrossRef]
- Kersting, S.; Rausch, V.; Bier, F.F.; von Nickisch-Rosenegk, M. Rapid detection of Plasmodium falciparum with isothermal recombinase polymerase amplification and lateral flow analysis. Malaria J. 2014, 13, 1–9. [Google Scholar] [CrossRef]
- Kashir, J.; Yaqinuddin, A. Loop mediated isothermal amplification (LAMP) assays as a rapid diagnostic for COVID-19 [published online ahead of print, 2020 Apr 25]. Med. Hypotheses 2020, 141, 109786. [Google Scholar] [CrossRef]
- Tang, Y.W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. The Laboratory Diagnosis of COVID-19 Infection: Current Issues and Challenges. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Blow, J.A.; Dohm, D.J.; Negley, D.L.; Mores, C.N. Virus inactivation by nucleic acid extraction reagents. J. Virol. Methods 2004, 119, 195–198. [Google Scholar] [CrossRef] [PubMed]
- Burton, J.E.; Easterbrook, L.; Pitman, J.; Anderson, D.; Roddy, S.; Bailey, D.; Vipond, R.; Bruce, C.B.; Roberts, A.D. The effect of a non-denaturing detergent and a guanidinium-based inactivation agent on the viability of Ebola virus in mock clinical serum samples. J. Virol. Methods 2017, 250, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Mazur, S.; Ork, B.L.; Postnikova, E.; Hensley, L.E.; Jahrling, P.B.; Johnson, R.; Holbrook, M.R. Inactivation and safety testing of Middle East Respiratory Syndrome Coronavirus. J. Virol. Methods 2015, 223, 13–18. [Google Scholar] [CrossRef]
- Krõlov, K.; Frolova, J.; Tudoran, O.; Suhorutsenko, J.; Lehto, T.; Sibul, H.; Mäger, I.; Laanpere, M.; Tulp, I.; Langel, Ü. Sensitive and rapid detection of Chlamydia trachomatis by recombinase polymerase amplification directly from urine samples. J. Mol. Diagn. 2014, 16, 127–135. [Google Scholar] [CrossRef]
- Cui, J.; Li, F.; Shi, Z.L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2019, 30, 30251–30258. [Google Scholar] [CrossRef]
- Chan, J.F.; Yip, C.C.; To, K.K.; Tang, T.H.; Wong, S.C.; Leung, K.H.; Fung, A.Y.; Ng, A.C.; Zou, Z.; Tsoi, H.W.; et al. Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-polymerase chain reaction assay validated in vitro and with clinical specimens. J. Clin. Microbiol. 2020, 4, 00310–00320. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Ma, B.; Li, J.; Chen, K.; Yu, X.; Sun, C.; Zhang, M. Multiplex Recombinase Polymerase Amplification Assay for the Simultaneous Detection of Three Foodborne Pathogens in Seafood. Foods 2020, 9, 278. [Google Scholar] [CrossRef]
- Mahony, J.; Chong, S.; Bulir, D.; Ruyter, A.; Mwawasi, K.; Waltho, D. Multiplex loop-mediated isothermal amplification (M-LAMP) assay for the detection of influenza A/H1, A/H3 and influenza B can provide a specimen-to-result diagnosis in 40 min with single genome copy sensitivity. J. Clin. Virol. 2013, 58, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Niemz, A.; Ferguson, T.M.; Boyle, D.S. Point-of-care nucleic acid testing for infectious diseases. Trends Biotechnol. 2011, 29, 240–250. [Google Scholar] [CrossRef] [PubMed]
| Isothermal Amplification Technology | Method for Denaturing Nucleic Acids | Estimated Reaction Time | Reference |
|---|---|---|---|
| Nucleic acid sequence-based amplification | Thermal | 90 min | [36] |
| Transcription-mediated amplification | Thermal | 1–2 h | [37] |
| Multiple displacement amplification | Enzymatic | 8–10 h | [38] |
| Loop-mediated amplification | Enzymatic | <1 h | [39] |
| Helicase-dependent amplification | Enzymatic | 2 h | [40] |
| Rolling circle amplification | Enzymatic | 1 h | [41] |
| Signal-mediated amplification | Thermal | 2 h | [42] |
| Recombinase polymerase amplification | Enzymatic | 10 min | [43] |
| Self-sustained sequence replication | Thermal | <1 h | [44] |
| Ramification amplification | Enzymatic | 1 h | [45] |
| Strand displacement amplification | Thermal | 2 h | [46] |
| Nicking enzyme-assisted reaction | Enzymatic | 15–30 min | [47] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
James, A.S.; Alawneh, J.I. COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2. Diagnostics 2020, 10, 399. https://doi.org/10.3390/diagnostics10060399
James AS, Alawneh JI. COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2. Diagnostics. 2020; 10(6):399. https://doi.org/10.3390/diagnostics10060399
Chicago/Turabian StyleJames, Ameh S., and John I. Alawneh. 2020. "COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2" Diagnostics 10, no. 6: 399. https://doi.org/10.3390/diagnostics10060399
APA StyleJames, A. S., & Alawneh, J. I. (2020). COVID-19 Infection Diagnosis: Potential Impact of Isothermal Amplification Technology to Reduce Community Transmission of SARS-CoV-2. Diagnostics, 10(6), 399. https://doi.org/10.3390/diagnostics10060399





